Abstract
Promoter CpG methylation is a fundamental regulatory process of gene expression. TET proteins are active CpG demethylases converting 5-methylcytosine to 5-hydroxymethylcytosine, with loss of 5 hmC as an epigenetic hallmark of cancers, indicating critical roles of TET proteins in epigenetic tumorigenesis. Through analysis of tumor methylomes, we discovered TET1 as a methylated target, and further confirmed its frequent downregulation/methylation in cell lines and primary tumors of multiple carcinomas and lymphomas, including nasopharyngeal, esophageal, gastric, colorectal, renal, breast and cervical carcinomas, as well as non-Hodgkin, Hodgkin and nasal natural killer/T-cell lymphomas, although all three TET family genes are ubiquitously expressed in normal tissues. Ectopic expression of TET1 catalytic domain suppressed colony formation and induced apoptosis of tumor cells of multiple tissue types, supporting its role as a broad bona fide tumor suppressor. Furthermore, TET1 catalytic domain possessed demethylase activity in cancer cells, being able to inhibit the CpG methylation of tumor suppressor gene (TSG) promoters and reactivate their expression, such as SLIT2, ZNF382 and HOXA9. As only infrequent mutations of TET1 have been reported, compared to TET2, epigenetic silencing therefore appears to be the dominant mechanism for TET1 inactivation in cancers, which also forms a feedback loop of CpG methylation during tumorigenesis.
Similar content being viewed by others
Introduction
DNA methylation at the C5 position of cytosine (5-methylcytosine, 5-mC), known as the “fifth base”, is a key epigenetic modification at CpG dinucleotides, playing critical roles in normal development and disease pathogenesis including tumorigenesis1. Regional promoter CpG methylation together with genome-wide hypomethylation, as a fundamental epigenetic hallmark of cancers, lead to the silencing of tumor suppressor genes (TSG) and activation of oncogenes, contributing to cancer initiation and progression. Recently, various whole-genome sequencing studies of virtually all human cancers also demonstrate that the most commonly mutated genes are epigenetic modifiers including CpG methylation machinery components across diverse cancers2,3,4,5, highlighting the direct and crucial involvement of epigenetic programming dysregulation in tumorigenesis.
DNA methylation is a reversible process, through either passive or active demethylation. Passive demethylation has been well-documented owing to reduction in activities or absence of DNA methyltransferases (DNMTs) during DNA replication. The newly identified 5-hydroxymethylcytosine (5 hmC) in mammalian genomic DNA6, as an intermediate of active DNA demethylation, has been recognized as the “sixth base”, which provides us new insight into the regulation of CpG methylation dynamics via active demethylation. 5 hmC is readily expressed in human normal tissues and embryonic stem cells, but becomes greatly decreased in multiple cancer tissues7,8,9. 5 hmC modification is relatively stable, not just as a transient intermediate10, arising as a novel epigenetic hallmark of tumors11.
The ten-eleven translocation (TET) family of DNA hydroxylases, including TET1, TET2, and TET3, mediates the conversion of 5 mC to 5 hmC and final DNA demethylation through sequential oxidation reactions, thus as key executers for establishing 5 hmC pattern and maintaining a hypomethylated genome state12,13. TET1 was firstly identified as a fusion partner of MLL in acute myeloid leukemia (AML)6. Inactive mutations or deletions of TET2 with impaired catalytic activity were frequently detected in hematopoietic malignancies14, along with decreased 5 hmC levels4,15,16, while no somatic TET1 or TET3 mutation was found in myeloid and lymphoid tumors. The biological functions of TET family members or 5 hmC on the reprogramming and development of embryotic stem cells have been extensively studied17,18,19,20,21. Recent reports also demonstrate that TET gene expression are reduced in some solid tumors, associated with 5 hmC depletion and gene downregulation, thus playing critical functional roles in tumor initiation and metastasis22,23,24,25,26. Some mechanisms have been proposed to mediate TET disruption in cancers, including post-transcriptional regulation by miR-2227, post-translational modification by cellular proteolytic system28, and nuclear exclusion of TET proteins29,30. However, a systematic study of the expression and transcriptional regulation of TET members in most human cancers is still needed.
Here, we have studied the expression and transcriptional regulation of TET family genes in a large collection of human normal and tumor samples. We examined the epigenetic and genetic alterations of TET1 through analyzing cancer methylomes previously established by us31 and also online genomics database of common tumors. We discovered frequent promoter methylation of TET1 in a large set of tumor cell lines and primary tumors, and confirmed its tumor suppressive functions and demethylation activity in tumor cells.
Results and Discussion
Epigenomic identification of TET1 as a methylated target in multiple cancers
During our analysis of whole-genome CpG methylation profiles (methylomes) of multiple tumor cell lines and primary tumors31, the promoter of one of the CpG demethylases, TET1, turned out to be a target in multiple methylomes (Fig. 1A). Bioinformatics analysis of the methylome data showed significant positive enrichment of CpG methylation (Cut off = 2) at the TET1 promoter and exon 1 region in multiple tumors, including nasopharyngeal carcinoma (NPC) xenografts (C15, C18) and primary tumor (OCT83), esophageal squamous cell carcinoma (ESCC) cell lines (KYSE140, KYSE510), hepatocellular carcinoma (HCC) cell lines (HuH7, HepG2) and primary tumor (418T), as well as nasal NK/T-cell lymphoma (NKTCL) cell lines (SNK6, NK-YS) and primary tumor (NK1) (Fig. 1A). The TET1 promoter and exon 1 region contain a typical CpG island (Fig. 2A), indicating that CpG methylation most likely regulates its expression in human cells.
(A) Representative methylome data. TET1 gene structure, promoter and exon 1 (NCBI database GRCh37.p13) are shown on the top panel. E1: exon 1. Positive methylation signal peaks identified by MeDIP-chip are shown in pink shadow for: NPC xenografts (C15, C18) and primary tumor (OCT83), ESCC cell lines (KYSE140, KYSE510), HCC cell lines (HuH7, HepG2) and primary tumor (HCC418T), NKTCL cell lines (SNK6, NK-YS) and primary tumor (NK1). (B) Expression of TET family genes (TET1, −2, −3) in human normal adult and fetal tissues by semi-quantitative RT-PCR, with GAPDH as a control. Sk. M., skeleton muscle.
(A) Structure of the TET1 promoter CpG island (CGI). CpG sites are shown as short vertical lines. MSP primer sites and BGS region analyzed are also indicated. (B) TET1 methylation was not detected in not-bisulfited DNA samples, indicating that the MSP system is specific. m4/m8 represents specific MSP primer set of TET1 methylation detection. (C,D) TET1 was frequently silenced and methylated in multiple carcinoma and lymphoma cell lines, detected by semi-quantitative RT-PCR and MSP, but expressed and unmethylated in immortalized but non-transformed normal epithelial cell lines (with names green underlined). M, methylated; U, unmethylated. (E) Abundant expression of TET2 and TET3 in TET1-downregulated tumor cell lines. Ca, carcinoma; NPC, nasopharyngeal carcinoma; ESCC, esophageal squamous cell carcinoma; CRC, colorectal cancer; RCC, renal cancer; NKTCL, nasal NK/T-cell lymphoma.
We thus further examined the expression and methylation profiles of TET1 in multiple cancers. Results showed that, although all three TET genes (TET1, −2, −3) were ubiquitously expressed in a series of human normal adult and fetal tissues (Fig. 1B), only TET1 neither TET2 nor TET3, was frequently downregulated or totally silenced in a variety of tumor cell lines including multiple carcinomas (nasopharyngeal, esophageal, lung, gastric, colon, breast, cervical, renal) and lymphomas (Hodgkin, non-Hodgkin and NKTCL), while TET1 is readily expressed in all immortalized normal epithelial cell lines of different tissue origins (Fig. 2 and Suppl. Fig. S1A).
Methylation-specific PCR (MSP) primers for TET1 was tested for not amplifying any not-bisulfited DNA, confirming the detection specificity of TET1 methylation in our study (Fig. 2B). Then by MSP, we detected TET1 promoter methylation in virtually all downregulated cell lines of nasopharyngeal, esophageal, lung, gastric, colon, breast, cervical and renal carcinomas, as well as Hodgkin (HL), non-Hodgkin (NHL) and NKTCL lymphomas, but not in immortalized normal epithelial cell lines (Fig. 2C,D; Table 1). Moreover, TET1 downregulation and methylation were infrequently detected in hepatocellular (HCC) and prostate cancer cell lines but not in the bladder and melanoma cell lines examined (Suppl. Fig. S1B).
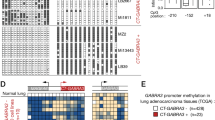
We further studied the detailed methylation profile of TET1 promoter by bisulfite genomic sequencing (BGS). A 384-bp region (+151-bp to +534-bp) spanning TET1 promoter and exon 1, containing 39 CpG sites was analyzed (Fig. 2A). BGS results showed heavily methylated alleles in representative cell lines, including NPC, ESCC, lung, gastric, colon, breast, cervical and renal carcinomas, as well as lymphomas, while barely present in immortalized normal cell lines of nasopharyngeal (NP69, NP460), esophageal (Het-1A), colon (CCD841con) and kidney (HEK293) epithelial cells, consistent with the MSP data (Fig. 3A). Thus, TET1 silencing by promoter CpG methylation is a common event in multiple tumors.
(A) Detection of TET1 methylation in multiple tumor cell lines and normal cell lines by BGS. (B) Treatment with Aza or combined with TSA (A + T) demethylated TET1 promoter in silenced cell lines of multiple tissue types. Expression and methylation changes were detected by semi-quantitative RT-PCR and MSP. (C) BGS analysis of TET1 promoter in cell lines with or without treatment. NPC, nasopharyngeal carcinoma; ESCC, esophageal squamous cell carcinoma; CRC, colorectal cancer; BrCa, breast cancer; RCC, renal cancer; Ca, carcinoma.
We further investigated whether TET1 promoter methylation directly mediates its repression. DNA methyltransferase inhibitor 5-aza-dC (Aza) was used or in combination with histone deacetylase (HDAC) inhibitor to treat tumor cell lines of nasopharyngeal, esophageal, colon, breast and renal, all with methylated and downregulated TET1. After the treatment, restoration of TET1 expression was observed, along with increased unmethylated promoter alleles as detected by MSP (Fig. 3B). Demethylation of the TET1 promoter was confirmed by BGS analysis, which shows dramatically demethylated CpG sites (Fig. 3C), indicating that CpG methylation directly mediates TET1 silencing in tumor cells.
In this study, we demonstrated that epigenetic silencing is a common regulatory mechanism for TET1 inactivation at the transcriptional level in multiple human cancers. Additional alternative mechanisms regulating expression and activities of TET family members have been reported32. For examples, high mobility group AT-hook 2 (HMGA2), a chromatin remodeling factor, suppresses TET1 expression by directly binding to its promoter or indirectly through other components in breast cancer cells24. Polycomb repressive complex 2 (PRC2) mediates Tet1 downregulation through H3K27me3 histone mark deposition33. PARP activity increases TET1 expression levels through maintaining a permissive chromatin state34. miR-22 suppresses TET expression levels in breast cancer cells through directly targeting the 3′-untranslated regions (UTRs) of TET mRNAs27. As direct substrates of calpains (calcium-activated cysteine proteases), TET proteins also undergo calpain-mediated degradation28. Nuclear exclusion of TET1 and TET2 is significantly correlated with loss of 5mC in glioma and colon cancer29,30. Thus, TET expression could be regulated at multiple levels of transcription, post-transcription or post-translation in different cell context, although TET1 silencing through promoter CpG methylation appears to be more common and predominant in multiple tumors.
Frequent silencing of TET1 by promoter methylation in primary tumors
As promoter CpG methylation in tumor cell lines might be derived from cell culture-induced secondary effect, we further examined TET1 methylation and expression in primary tumor samples. We detected frequent TET1 methylation in multiple tumors, including 55% (31/56) of NPC, 55% (30/55) of gastric, 27% (3/11) of colon, 42% (5/12) of hepatocellular, 36% (18/50) of breast and 28% (13/46) of renal tumor samples, as well as 78% of primary Hodgkin and 83% (10/12) of NKTCL lymphoma samples (Fig. 4A, Suppl. Fig. S2, Table 1), but infrequently in primary ESCC, lung, prostate tumors and other non-Hodgkin lymphomas (Suppl. Fig. S2, Table 1). TET1 methylation could even be detected in 50% of 16 nose swab samples from suspected NPC patients (Fig. 4B). In contrast, TET1 methylation was not detected in a panel of human normal adult and fetal tissues except for being barely seen in normal small intestine and colon (Fig. 4C). Further detailed BGS methylation analysis confirmed the presence of methylated promoter alleles in primary tumors but not normal tissues (Fig. 4D). TET1 downregulation was also detected in paired primary tumors of several tissue types (lung, stomach, colon, rectum, breast and kidney) and primary NPC tumors (Fig. 4E). Furthermore, through online GENT and Oncomine database analysis, we found that TET1 mRNA levels were significantly reduced in multiple solid tumors and leukemia, compared with their corresponding normal tissues (Suppl. Fig. S3). These results clearly demonstrate that TET1 silencing by promoter CpG methylation is a common event for multiple tumors of epithelial and lymphoid origins.
TET1 promoter methylation in (A) multiple primary tumors and (B) nose swab samples from NPC patients, detected by MSP. (C) TET1 methylation is barely seen in normal tissues by MSP analysis. (D) Representative BGS analysis of TET1 promoter methylation in primary tumors and normal tissues. Circles, CpG sites analyzed; row of circles, an individual promoter allele that was cloned, randomly selected and sequenced; filled circle, methylated CpG site; open circle, unmethylated site. (E) Levels of TET1 mRNA expression in representative paired tumor (T)/normal (N) tissues, and primary tumor tissues (NPC), measured by semi-quantitative RT-PCR. Ca, carcinoma; NPC, nasopharyngeal carcinoma; CRC, colorectal cancer; RCC, renal cancer; GsCa, gastric cancer; Sk. muscle, skeleton muscle; S. intestine, small intestine.
Several studies have shown that TET genes are readily expressed in normal esophageal, gastric, colon, liver and breast tissues by PCR or immunohistochemistry22,23,25, but decreased in tumor cell lines and primary tumors to varied grades, with TET1 as the most significantly downregulated member. A previous report through analyzing Cancer Genome Atlas TCGA database found that TET1 is downregulated in primary tumors of colorectal, breast and lung since early stage, and associated with patient poor survival23. TET1 is significantly decreased at mRNA and protein levels in gastric primary tumors compared to surgical margins and associated with tumor localization and TNM grades35. DNA methylation and bivalent histone marks at the CpG island 3′-shore mediate TET1 silencing in gastric cancer36. Reduced TET1 expression or 5 hmC level in breast cancer tissues could be biomarkers for breast cancer progression37. TET1 methylation in colorectal cancer tissues, not TET2 and TET338, has been found as an early event in CRC tumorigenesis, thus as a valuable biomarker for metastasis prediction39. Our results are consistent with these previous studies. TET1 methylation appears to be tumor-specific and thus could serve as a potential epigenetic biomarker for cancer detection.
Genetic alteration of TET1 is uncommon in human cancers
As alterations of cancer gene are through either genetic or epigenetic mechanisms, we further investigated possible genetic alterations of TET1 in cancers. Somatically acquired mutations of TET1 in human cancers were analyzed using the COSMIC database. Only <1% of tumor cases (most cases with ≤0.25%) had detectable TET1 mutations (Fig. 5A), consisting of 80% of missense mutations, 10% of nonsense and 10% of synonymous mutations (Fig. 5B), with most of the mutations located in coding regions (Fig. 5C). We also detected hemizygous deletion of TET1 in some tumor cell lines with TET1 silencing and methylation, but not in TET1-expresssing cells (Suppl. Fig. S4A,B). Consistently, TET1 gene deletion was also observed in solid tumors by analyzing DNA copy number alterations using the Oncomine database (Suppl. Fig. S4C). These results demonstrate that TET1 mutation is uncommon in human cancers, although TET1 deletion is indeed present in some tumor samples.
TET1 functions as a tumor suppressor which requires its catalytic activity
The TET1 catalytic domain (CD) (containing the Cys-rich and DSBH regions) remains intact hydroxylase activity in embryonic development and reprogramming6,13, displaying ability to induce 5 hmC formation, demethylation and gene transcription in differentiated cells33. We test whether the catalytic activity of TET1 was required for its possible tumor suppression functions, using TET1-CD and its enzymatic dead mutant (TET1-CD-mut) (Fig. 6A). Ectopic expression of TET1-CD significantly suppressed tumor cell clonogenicity (to ~40–50% of control cells) in colony formation assays of NPC, ESCC, gastric, colon and breast tumor cells, while the TET1-CD-mut lost this ability (Fig. 6B). TUNEL assay showed significantly increased numbers of apoptotic cells in TET1-CD expressing-tumor cells, compared with vector or TET1-CD-mut controls (Fig. 6C). These results demonstrate that TET1 possesses bona fide tumor suppressive functions in tumor cells of multiple types.
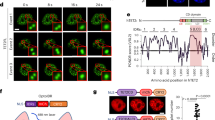
(A) Structure and functional domains of the human TET1 protein, containing a C-terminal CD domain including the Cys-rich and DSBH regions, and a CXXC domain. The positions of three nuclear localization sequences (NLS) are shown. TET1 catalytic domain (TET1-CD) containing the Cys-rich and DSBH regions and TET1 mutant (TET1-CD-mut) with two amino acid substitutions (H1672A; D1674A) in the catalytic domain are also shown. (B) Ectopic expression of TET1-CD inhibited tumor cell growth of multiple tissue types. Representative colony formation assays of TET1-CD- and TET1-CD-mut-expressing tumor cells of nasopharyngeal, esophageal, gastric, colon, and breast cancers are shown. Quantitative analyses of colony numbers are shown as values of mean ± S.D. (lower panel), ***p < 0.001. NPC, nasopharyngeal carcinoma; ESCC, esophageal squamous cell carcinoma; GsCa, gastric cancer; CRC, colorectal cancer; BrCa, breast cancer. (C) Ectopic expression of TET1-CD induced tumor cell apoptosis. TET1-CD, TET1-CD-mut, and vector-expressing NPC tumor cells (HONE1) were analyzed by TUNEL assays. (D) TET1-CD upregulated multiple TSGs expression in tumor cells, as examined by semi-quantitative RT-PCR. (E) TET1-CD upregulated multiple TSGs expression as measured by qRT-PCR in NPC (HNE1) cells. Fold changes of TSGs expression in TET1-CD and TET1-CD-mut-transcfected cells were calculated by normalizing towards vector-expressing cells (set 1.0). GAPDH was used as an internal control. Data are shown as mean ± SD of three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001. (F) Detection of promoters methylation of HOXA9, SLIT2 and ZNF382 genes by MSP in TET1-CD and TET1-CD-mut-expressing tumor cells.
Consistent with our results, several recent studies have shown similar tumor-suppressive functions of TET1 in cancer cells. TET1 inhibits proliferation and invasion of colon23, breast24,25, renal40 and prostate25 cancer cells in vivo and in vitro. TET1 deficiency promotes B-lineage differentiation, leading eventually to B-cell lymphoma41. TET1 suppression as a key event of the RAS programming is required for KRAS-induced cellular transformation26. Thus, loss of function of TET1 is a common event during multiple tumorigenesis of solid tumors or hematologic malignancies.
TET1 induces TSG promoter demethylation in tumor cells
Several studies identified TET1 target genes in mouse ES cells and some tumor cells, using RNA- or ChIP- sequencing or hydroxymethylated DNA immunoprecipitation sequencing (hMeDIP-seq)12,24,26,27,33,42,43,44,45. A series of TET1-targeted genes including TSGs have been identified, such as TIMP25, HOXA9 and HOXA724, and Wnt signaling antagonists DKK3 and DKK423. To further explore the molecular mechanism of TET1 in tumor suppression, we examined some known and potential target TSGs to assess the demethylase activity of TET1 in tumor cells. Mild upregulation of HOXA9, HOXA5, PCDH7, TCF4, MEIS1, SLIT2 and ZNF382 at mRNA levels was observed in TET1-CD-expressing carcinoma cells by semi-quantitative RT-PCR (Fig. 6D) and qRT-PCR (Fig. 6E). Meanwhile, we also detected decreased methylated alleles of HOXA9, SLIT2 and ZNF382 promoters in TET1-CD-expressing tumor cells, but not in TET1-CD-mut-expressing cells, with increased unmethylated promoter alleles observed concurrently, suggesting that TET1 indeed functions as a CpG demethylase to demethylate and reactivate multiple TSGs in tumor cells (Fig. 6F). In addition to HOXA9, we also found that TSGs like SLIT2, ZNF382, PCDH7, TCF4, MEIS1 and HOXA5 as TET1 target genes which could be demethylated and reactivated by TET1 in tumor cells. Other mechanisms besides demethylase activity could also be involved in regulating target genes by TET1, such as recruiting PRC242, PRDM1443, Sin3A co-repressor complex44 and MBD3/NURD complex45. Further studies on TET1-targeted gene regulation in human cancers would help us to understand more of its role in cancer development.
The discovery of TET enzymes, in addition to DNMTs, establishes a fundamental etiologic role of CpG methylation in human cancers. In response to environment carcinogens46,47,48 like chemical carcinogens and tumor viruses, DNMT activities and expression levels are induced and increased in cells, displaying stronger maintenance and de novo methylation capacity, leading to specific gene CpG island hypermethylation. The epigenetic alterations, especially promoter CpG methylation of TSGs, facilitate genome instability, disrupted cellular signaling and even further genetic mutations, thus are crucial to tumor initiation and progression1,49. Remarkably, promoter CpG methylation-mediated silencing of the CpG demethylase TET1 in human cancers, which in turn, further leads to increased 5 mC levels in tumor cells, thus forming a DNA methylation feedback loop mediated by DNMT/CpG methylation and TET1 (Fig. 7).
When normal cells are exposed to carcinogens (chemical carcinogens, tumor viruses, etc), DNA methyltransferases (DNMTs) are induced, upregulated or overactivated, which further generates higher levels of DNA CpG methylation (5 mC). Elevated level of 5 mC on tumor suppressor gene (TSG) promoters lead to TSGs silencing and functional inactivation, ultimately to tumorigenesis. Ten-eleven-translocation (TET) proteins catalyze DNA CpG demethylation through converting 5 mC to 5-hydroxymethylcytosine (5 hmC), maintaining a delicate balance between CpG methylation and demethylation in normal cells. While in premalignant or tumor cells, CpG demethylation by TET would induce TSG promoter demethylation and functional restoration for further tumor suppression. Thus unlike normal cells where TET proteins are abundant, loss of TET1 expression through promoter CpG methylation frequently occurs in tumor cells, which in turn, increases 5 mC levels and promotes TSG inactivation in tumor pathogenesis.
In summary, our study comprehensively examined TET1 expression and methylation status in multiple tumors, and demonstrated that promoter CpG methylation is a predominant mechanism for TET1 inactivation in human cancers. The tumor-specific methylation of TET1 could serve as a valuable, epigenetic non-invasive biomarker. TET1 as a tumor suppressor and CpG demethylase in tumor cells requires its intact catalytic domain, which provides new insight into the epigenetic master role of TET1 in tumor pathogenesis. Our findings enlighten us on the mechanistic elucidation of the importance of CpG methylation in human cancers.
Material and Methods
Cell lines and tissue samples
Human tumor cell lines of multiple tissue types were used50,51,52,53,54,55, including nasopharyngeal (NPC), esophageal squamous cell (ESCC), lung, gastric, colorectal (CRC), hepatocellular (HCC), breast, cervical, renal (RCC), bladder and prostate carcinomas, melanoma, as well as non-Hodgkin (NHL), Hodgkin (HL) and nasal natural killer (NK)/T-cell (NKTCL) lymphomas. Immortalized, non-transformed normal epithelial cell lines were used as “normal” controls. Cell lines were obtained from either American Type Culture Collection or collaborators. When needed, cell lines were treated with 10 μmol/L 5-aza-2′-deoxycytidine (Aza) (Sigma-aldrich, St Louis, MO) for 3 days, without or with further treatment with 100 nmol/l trichostatin A (TSA) (Cayman Chemical Co., Ann Arbor, MI) for additional ~16 h as previously50,53. Normal adult and fetal tissue RNA and DNA samples were purchased commercially (Stratagene, La Jolla, Ca; Millipore-Chemicon, Billerica, Ma). DNA samples of primary carcinomas, nose swab from suspected NPC patients, as well as surgical margin normal tissues, have been described previously31,51,52.
Establishment of tumor methylomes by MeDIP-chip
Methylated DNA immunoprecipitation (MeDIP) coupled with promoter microarray hybridization was performed as previously31. Briefly, immunoprecipitation of methylated DNA was performed using monoclonal antibody against 5-methylcytidine (33D3, Diagenode, Seraing, Belgium) labeled with magnetic beads. Total input and immunoprecipitated DNA were labeled with Cy3 or Cy5, respectively, and hybridized to NimbleGen™ HG18 Meth (385K CGI plus) promoter arrays or HG19 (2.1 M) Deluxe Promoter arrays (Array Star, Inc., MD). Normal epithelial cell lines and normal tissues were used as controls. Bioinformatics analysis of methylome data was performed as previously31.
Semi-quantitative RT-PCR and quantitative real-time PCR (qRT-PCR)
Semi-quantitative RT-PCR and quantitative real-time PCR were performed as described before50,53, with GAPDH as a control for all the samples shown in our previous publications31,51,52. qRT-PCR was carried out according to the manufacturer’s protocol (HT7900 system; applied Biosystems), with SYBR Green master mix (applied Biosystems) used. Primers used are listed in Supplementary Table S1.
Bisulfite treatment of DNA samples and promoter methylation analysis
CpG island (CGI) analysis for TET1 promoter and exon 1 was performed using CpG island Searcher (http://ccnt.hsc.usc.edu/cpgislands2). Bisulfite modification of genomic DNA was carried out as described previously56,57. For MSP analysis, approximately 50 ng of bisulfited DNA for each sample was amplified with methylation- or unmethylation- specific primer set, according to our previous MSP protocol58. Bisulfite-treated DNA was also amplified using a set of BGS primers, then cloned into pCR4-TOPO vector (Invitrogen, Carlsbad, Ca), with 8–10 clones randomly picked and sequenced. MSP and BGS primers used are shown in Supplementary Table S1. Unmethylated gene alleles for these treated samples have been detected in our previous publications, which shows the good quality of these DNA samples31,51,52.
Genetic deletion analysis for TET1
Homozygous deletion of TET1 coding exons 2 and 4 was examined using multiplex genomic DNA PCR, as previously described51. Primer sequences are shown in Supplementary Table S1.
Colony formation assay of tumor cells
Human TET1 catalytic domain (TET1-CD) cDNA and its catalytic domain mutant (TET1- CD-mut) clones (Addgene, Cambridge, MA) were used as templates to generate TET1 constructs with an N-terminal Flag tag, and subcloned into pcDNA3.1 vector (Invitrogen, Carlsbad, Ca). Cells were cultured overnight in a 12-well plate and transfected with empty vector or TET1-CD, TET1-CD-mut-expressing plasmids using Lipofectamine 2000 (Invitrogen, Carlsbad, Ca). Forty-eight hours later, transfectants were replated in triplicate and cultured for 10–15 days in complete medium containing G418. Surviving colonies were stained with crystal violet (0.5% w/v) after methanol fixation, with visible colonies (≥50 cells) counted.
TUNEL assay
Cells cultured on coverslips were fixed with 4% paraformaldehyde, and permeabilized with 0.1% triton X-100. TUNEL (Terminal deoxynucleotidyl transferase dUTP nick end labeling) staining was performed using the In Situ Cell Death Detection Kit (Roche, Mannheim, Germany).
Statistical analysis
Student’s t-tests were performed. All reported p-values were two-sided, and p < 0.05 was considered statistically significant.
Additional Information
How to cite this article: Li, L. et al. Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Sci. Rep. 6, 26591; doi: 10.1038/srep26591 (2016).
Change history
06 October 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692, doi: 10.1016/j.cell.2007.01.029 (2007).
Schubeler, D. Function and information content of DNA methylation. Nature 517, 321–326, doi: 10.1038/nature14192 (2015).
You, J. S. & Jones, P. A. Cancer genetics and epigenetics: two sides of the same coin? Cancer cell 22, 9–20, doi: 10.1016/j.ccr.2012.06.008 (2012).
Hamidi, T., Singh, A. K. & Chen, T. Genetic alterations of DNA methylation machinery in human diseases. Epigenomics 7, 247–265, doi: 10.2217/epi.14.80 (2015).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558, doi: 10.1126/science.1235122 (2013).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935, doi: 10.1126/science.1170116 (2009).
Jin, S. G. et al. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer research 71, 7360–7365, doi: 10.1158/0008-5472.CAN-11-2023 (2011).
Haffner, M. C. et al. Global 5-hydroxymethylcytosine content is significantly reduced in tissue stem/progenitor cell compartments and in human cancers. Oncotarget 2, 627–637 (2011).
Kudo, Y. et al. Loss of 5-hydroxymethylcytosine is accompanied with malignant cellular transformation. Cancer science 103, 670–676, doi: 10.1111/j.1349-7006.2012.02213.x (2012).
Bachman, M. et al. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nature chemistry 6, 1049–1055, doi: 10.1038/nchem.2064 (2014).
Lian, C. G. et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 150, 1135–1146, doi: 10.1016/j.cell.2012.07.033 (2012).
Tan, L. & Shi, Y. G. Tet family proteins and 5-hydroxymethylcytosine in development and disease. Development 139, 1895–1902, doi: 10.1242/dev.070771 (2012).
Guo, J. U., Su, Y., Zhong, C., Ming, G. L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434, doi: 10.1016/j.cell.2011.03.022 (2011).
Delhommeau, F. et al. Mutation in TET2 in myeloid cancers. The New England journal of medicine 360, 2289–2301, doi: 10.1056/NEJMoa0810069 (2009).
Konstandin, N. et al. Genomic 5-hydroxymethylcytosine levels correlate with TET2 mutations and a distinct global gene expression pattern in secondary acute myeloid leukemia. Leukemia 25, 1649–1652, doi: 10.1038/leu.2011.134 (2011).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer cell 20, 11–24, doi: 10.1016/j.ccr.2011.06.001 (2011).
Ito, S. et al. Role of Tet proteins in 5 mC to 5 hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133, doi: 10.1038/nature09303 (2010).
Yamaguchi, S., Shen, L., Liu, Y., Sendler, D. & Zhang, Y. Role of Tet1 in erasure of genomic imprinting. Nature 504, 460–464, doi: 10.1038/nature12805 (2013).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393, doi: 10.1038/nature09934 (2011).
Dawlaty, M. M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175, doi: 10.1016/j.stem.2011.07.010 (2011).
Spruijt, C. G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–1159, doi: 10.1016/j.cell.2013.02.004 (2013).
Yang, H. et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 32, 663–669, doi: 10.1038/onc.2012.67 (2013).
Neri, F. et al. TET1 is a tumour suppressor that inhibits colon cancer growth by derepressing inhibitors of the WNT pathway. Oncogene, doi: 10.1038/onc.2014.356 (2014).
Sun, M. et al. HMGA2/TET1/HOXA9 signaling pathway regulates breast cancer growth and metastasis. Proceedings of the National Academy of Sciences of the United States of America 110, 9920–9925, doi: 10.1073/pnas.1305172110 (2013).
Hsu, C. H. et al. TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell reports 2, 568–579, doi: 10.1016/j.celrep.2012.08.030 (2012).
Wu, B. K. & Brenner, C. Suppression of TET1-dependent DNA demethylation is essential for KRAS-mediated transformation. Cell reports 9, 1827–1840, doi: 10.1016/j.celrep.2014.10.063 (2014).
Song, S. J. et al. MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 154, 311–324, doi: 10.1016/j.cell.2013.06.026 (2013).
Wang, Y. & Zhang, Y. Regulation of TET protein stability by calpains. Cell reports 6, 278–284, doi: 10.1016/j.celrep.2013.12.031 (2014).
Muller, T. et al. Nuclear exclusion of TET1 is associated with loss of 5-hydroxymethylcytosine in IDH1 wild-type gliomas. The American journal of pathology 181, 675–683, doi: 10.1016/j.ajpath.2012.04.017 (2012).
Huang, Y. et al. Loss of nuclear localization of TET2 in colorectal cancer. Clin Epigenetics 8, 9, doi: 10.1186/s13148-016-0176-7 (2016).
Li, L. et al. Characterization of the nasopharyngeal carcinoma methylome identifies aberrant disruption of key signaling pathways and methylated tumor suppressor genes. Epigenomics 7, 155–173, doi: 10.2217/epi.14.79 (2015).
Huang, Y. & Rao, A. Connections between TET proteins and aberrant DNA modification in cancer. Trends Genet 30, 464–474, doi: 10.1016/j.tig.2014.07.005 (2014).
Neri, F. et al. TET1 is controlled by pluripotency-associated factors in ESCs and downmodulated by PRC2 in differentiated cells and tissues. Nucleic acids research, doi: 10.1093/nar/gkv392 (2015).
Ciccarone, F. et al. Poly(ADP-ribosyl)ation is involved in the epigenetic control of TET1 gene transcription. Oncotarget 5, 10356–10367, doi: 10.18632/oncotarget.1905 (2014).
Gambichler, T. et al. Decreased expression of ten-eleven translocation 2 protein is associated with progressive disease and death in patients with mucosis fungoides. Br J Dermatol, doi: 10.1111/bjd.14174 (2015).
Park, J. L. et al. Decrease of 5 hmC in gastric cancers is associated with TET1 silencing due to with DNA methylation and bivalent histone marks at TET1 CpG island 3′-shore. Oncotarget 6, 37647–37662, doi: 10.18632/oncotarget.6069 (2015).
Tsai, K. W. et al. Reduction of global 5-hydroxymethylcytosine is a poor prognostic factor in breast cancer patients, especially for an ER/PR-negative subtype. Breast Cancer Res Treat 153, 219–234, doi: 10.1007/s10549-015-3525-x (2015).
Rawluszko-Wieczorek, A. A. et al. Clinical significance of DNA methylation mRNA levels of TET family members in colorectal cancer. Journal of cancer research and clinical oncology, doi: 10.1007/s00432-014-1901-2 (2015).
Ichimura, N. et al. Aberrant TET1 Methylation Closely Associated with CpG Island Methylator Phenotype in Colorectal Cancer. Cancer Prev Res (Phila) 8, 702–711, doi: 10.1158/1940-6207.CAPR-14-0306 (2015).
Fan, M., He, X. & Xu, X. Restored expression levels of TET1 decrease the proliferation and migration of renal carcinoma cells. Mol Med Rep, doi: 10.3892/mmr.2015.4058 (2015).
Cimmino, L. et al. TET1 is a tumor suppressor of hematopoietic malignancy. Nat Immunol 16, 653–662, doi: 10.1038/ni.3148 (2015).
Neri, F. et al. Genome-wide analysis identifies a functional association of Tet1 and Polycomb repressive complex 2 in mouse embryonic stem cells. Genome biology 14, R91, doi: 10.1186/gb-2013-14-8-r91 (2013).
Okashita, N. et al. PRDM14 promotes active DNA demethylation through the ten-eleven translocation (TET)-mediated base excision repair pathway in embryonic stem cells. Development 141, 269–280, doi: 10.1242/dev.099622 (2014).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348, doi: 10.1038/nature10066 (2011).
Yildirim, O. et al. Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 147, 1498–1510, doi: 10.1016/j.cell.2011.11.054 (2011).
Li, H. P., Leu, Y. W. & Chang, Y. S. Epigenetic changes in virus-associated human cancers. Cell Res 15, 262–271, doi: 10.1038/sj.cr.7290295 (2005).
Tsai, C. N., Tsai, C. L., Tse, K. P., Chang, H. Y. & Chang, Y. S. The Epstein-Barr virus oncogene product, latent membrane protein 1, induces the downregulation of E-cadherin gene expression via activation of DNA methyltransferases. Proceedings of the National Academy of Sciences of the United States of America 99, 10084–10089, doi: 10.1073/pnas.152059399 (2002).
Ghoshal, K. et al. Inhibitors of histone deacetylase and DNA methyltransferase synergistically activate the methylated metallothionein I promoter by activating the transcription factor MTF-1 and forming an open chromatin structure. Mol Cell Biol 22, 8302–8319 (2002).
Baylin, S. B. & Ohm, J. E. Epigenetic gene silencing in cancer - a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 6, 107–116, doi: 10.1038/nrc1799 (2006).
Jin, H. et al. Epigenetic silencing of a Ca(2+)-regulated Ras GTPase-activating protein RASAL defines a new mechanism of Ras activation in human cancers. Proceedings of the National Academy of Sciences of the United States of America 104, 12353–12358, doi: 10.1073/pnas.0700153104 (2007).
Li, L. et al. The human cadherin 11 is a pro-apoptotic tumor suppressor modulating cell stemness through Wnt/beta-catenin signaling and silenced in common carcinomas. Oncogene 31, 3901–3912, doi: 10.1038/onc.2011.541 (2012).
Li, L. et al. Epigenetic identification of receptor tyrosine kinase-like orphan receptor 2 as a functional tumor suppressor inhibiting beta-catenin and AKT signaling but frequently methylated in common carcinomas. Cellular and molecular life sciences : CMLS 71, 2179–2192, doi: 10.1007/s00018-013-1485-z (2014).
Ying, J. et al. Functional epigenetics identifies a protocadherin PCDH10 as a candidate tumor suppressor for nasopharyngeal, esophageal and multiple other carcinomas with frequent methylation. Oncogene 25, 1070–1080, doi: 10.1038/sj.onc.1209154 (2006).
Murray, P. G. et al. Epigenetic silencing of a proapoptotic cell adhesion molecule, the immunoglobulin superfamily member IGSF4, by promoter CpG methylation protects Hodgkin lymphoma cells from apoptosis. The American journal of pathology 177, 1480–1490, doi: 10.2353/ajpath.2010.100052 (2010).
Wang, Z. et al. Epigenetic silencing of the 3p22 tumor suppressor DLEC1 by promoter CpG methylation in non-Hodgkin and Hodgkin lymphomas. J Transl Med 10, 209, doi: 10.1186/1479-5876-10-209 (2012).
Tao, Q. et al. Methylation status of the Epstein-Barr virus major latent promoter C in iatrogenic B cell lymphoproliferative disease. Application of PCR-based analysis. The American journal of pathology 155, 619–625, doi: 10.1016/S0002-9440(10)65157-7 (1999).
Tao, Q. et al. Defective de novo methylation of viral and cellular DNA sequences in ICF syndrome cells. Human molecular genetics 11, 2091–2102 (2002).
Tao, Q. Cancer research in an era when epigenetics is no longer “epi” - challenges and opportunities. Chinese journal of cancer 32, 1–2, doi: 10.5732/cjc.012.10300 (2013).
Acknowledgements
We thank Profs Bert Vogelstein, George Tsao, Gopesh Srivastava, Riccardo Dalla-Favera (DLBCL), Meenhard Herlyn (melanoma), Sun Young Rha, and Catherine Kelly for some cell lines; DSMZ (German Collection of Microorganisms & Cell Cultures) for KYSE cell lines (Shimada et al., Cancer 69, 277–284, 1992). PGM was supported by a Bloodwise Programme Grant. This study was supported by Hong Kong HMRF (#13120082), RGC (TBRS #T12-401/13R), China Natural Science Foundation (#81572327), and special research funds from The Chinese University of Hong Kong.
Author information
Authors and Affiliations
Contributions
Q.T. and L.L. conceived and supervised the study; L.L., C.L., H.M. and Z.D. acquired and analyzed data; P.G., B.L., R.A., A.T.C., T.S.M., W.Y.C. and F.K.C. provided materials and commented the manuscript; L.L. drafted the manuscript; Q.T. and L.L. finalized the manuscript.
Corresponding author
Ethics declarations
Competing interests
TSKM has received honoraria from Boehringer Ingelheim, BioMarin Pharmaceuticals, AstraZeneca, Roche/Genentech, Pfi zer, Eli Lilly, Merck Serono, Merck Sharp & Dohme, Janssen, Clovis Oncology, GlaxoSmithKline, Novartis, SFJ Pharmaceutical, ACEA Biosciences, Vertex Pharmaceuticals, Bristol-Myers Squibb, AVEO & Biodesix, Prime Oncology, and Amgen; advisory board fees from AstraZeneca, Roche/Genentech, Pfi zer, Eli Lilly, Boehringer Ingelheim, Merck Serono, Merck Sharp & Dohme, Janssen, Clovis Oncology, BioMarin, GlaxoSmithKline, Novartis, SFJ Pharmaceutical, ACEA Biosciences, Vertex Pharmaceuticals, AVEO & Biodesix, and Bristol-Myers Squibb; and is a shareholder in Sanomic. ATC received honoraria from consulting or Advisory role at Merck, Taiho Pharmaceutical, Roche, Amgen and received research funding from Boehringer Ingelheim, Bristol-Myers Squibb, Eli Lilly, Pfizer.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, L., Li, C., Mao, H. et al. Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Sci Rep 6, 26591 (2016). https://doi.org/10.1038/srep26591
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26591
This article is cited by
-
The regulation of plasma gelsolin by DNA methylation in ovarian cancer chemo-resistance
Journal of Ovarian Research (2024)
-
TET2 Is Downregulated in Early Esophageal Squamous Cell Carcinoma and Promotes Esophageal Squamous Cell Malignant Behaviors
Digestive Diseases and Sciences (2024)
-
DNA methylation in the genesis, progress and prognosis of head and neck cancer
Holistic Integrative Oncology (2023)
-
Isoform-specific and ubiquitination dependent recruitment of Tet1 to replicating heterochromatin modulates methylcytosine oxidation
Nature Communications (2022)
-
Nasopharyngeal carcinoma: an evolving paradigm
Nature Reviews Clinical Oncology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.