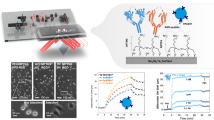
Abstract
While animal experimentations have spearheaded numerous breakthroughs in biomedicine, they also have spawned many logistical concerns in providing toxicity screening for copious new materials. Their prioritization is premised on performing cellular-level screening in vitro. Among the screening assays, secretomic assay with high sensitivity, analytical throughput and simplicity is of prime importance. Here, we build on the over 3-decade-long progress on transistor biosensing and develop the holistic assay platform and procedure called semiconductor electronic label-free assay (SELFA). We demonstrate that SELFA, which incorporates an amplifying nanowire field-effect transistor biosensor, is able to offer superior sensitivity, similar selectivity and shorter turnaround time compared to standard enzyme-linked immunosorbent assay (ELISA). We deploy SELFA secretomics to predict the inflammatory potential of eleven engineered nanomaterials in vitro and validate the results with confocal microscopy in vitro and confirmatory animal experiment in vivo. This work provides a foundation for high-sensitivity label-free assay utility in predictive toxicology.
Similar content being viewed by others
Introduction
With the advent of high-throughput analytical approaches, biomedical scientists have begun to deal with big data sets which rely on innovating and utilizing discovery tools capable of generating rich data in a rapid manner. The same trend is also evident in toxicology which has been facing the challenge of assessing safety profiles of over 90,000 existing and emerging chemicals or materials to humans. Nevertheless, the reality is that such needs are far from satisfied due to both cost and throughput constraints. While assessments have been performed in an observational-based descriptive toxicological fashion using animals, same results can, in our opinion, be obtained more effectively in a predictive manner with target-specific and mechanism-based biological observations1,2,3,4,5,6. This is in agreement with the vision of the National Academy of Sciences (NAS), which advocates a paradigm shift in the approach to toxicological testing in the 21st century, including the development of robust scientific platforms for screening a large number of toxicants at the cellular level1. Central to the implementation of predictive toxicology is the move from predominantly animal screening to in vitro studies where appropriate2,7,8,9. In one embodiment, the data of secretomic assays are utilized to rank material toxicity in vitro and predict their toxicity in vivo, offering great value in planning and prioritizing the expensive and time-consuming animal experiments for validation purpose.
Most biomolecular assays, such as the enzyme-linked immunosorbent assay (ELISA), are labeling-based, thus have limited analytical throughput and/or sensitivity, involve tedious experimental procedures and also necessitate costly infrastructure, reagents and trained personnel. Highly specific biomolecular quantitation can instead be performed with great simplicity via a label-free route, such as the optics-free and enzyme-free field-effect transistor (FET) immunoassay. A FET without a gate electrode designed to detect immunological antigen-antibody binding reactions without labels was patented in 198010 and first unambiguously demonstrated in 198911. The same underlying FET sensor channel has then shrunk to the nanoscale, lain down on an insulator and evolved to become non-planar12. Numerous FET biosensors with nanowire-, nanoribbon-, or nanotube-on-insulator channels have thus emerged13,14,15,16,17 and demonstrated outstanding lower limits of detection (LLOD) when combined with proper electrical biasing18. Nevertheless, their output-signal levels at LLOD are inherently very low, i.e. tens to hundreds of picoamps of drain current change as summarized in Supplementary Data Fig. 1 and are therefore impractical for use outside of the research lab. In addition, most FET immunoassays have not been validated against current analytical standards such as the gold standard ELISA and have not been deployed in quantitating real biological samples in the scenario of interest. As a result, label-free FET immunoassays have not been meaningfully adopted and impacted the biomedical and pharmaceutical industry even after over 3 decades of intense research.
Results
SELFA development and analytical validations
The low output signal level of an FET sensor can be readily boosted in the electrical domain, yet such amplification should occur as close as possible to the sensor in order to minimize the noise induced by parasitic components19. One feasible concept is an integrated nanowire FET sensor-amplifier that can locally amplify the transduced sensing current20,21. In this work, we developed such sensor-amplifier concept into a biomolecular assay platform for the first time, which we call Semiconductor Electronic Label-Free Assay (SELFA). In particular, we explored the viability of the on-the-sensing-spot amplification in immunodetection, developed a calibration strategy and performed analytical validation of the holistic immunoassay.
The amplifying FET sensor in SELFA has a compact T-shape nanowire structure, integrating therein two nanowire FETs – one for sensing and the other for amplification (Fig. 1a). We denote the compound device as a T-nwFET sensor. Resting on an insulator, the T-nwFET sensor embraces the genuine advantage behind the “large surface area-to-volume ratio” preception22 via the sensing nwFET and offers electrical signal amplification in extreme proximity via the connecting amplifying nwFET (see Supplementary Data Fig. 2 for amplification mechanism20,21). We fabricated our T-nwFET sensors on silicon-on-insulator (SOI) substrates using a combination of electron-beam and optical lithography as well as other standard semiconductor fabrication processes (for more details see the ‘Device fabrication’ subsection of the Methods section). The widths of Si nanowires of the fabricated T-nwFET sensors were around 100 nm, as confirmed using scanning electron microscopy (SEM) and shown in Fig. 1b. We further passivated the T-nwFET sensor substrate surface with a 1.5-μm thick SU-8 polymer layer to prevent electrical shorts. The SU-8 in the area immediately above and nearby the sensing Si nanowire was removed via a photolithography step, with the opening becoming part of a microfluidic well for interfacing with reagents and analyte solutions (Fig. 1c,f). Prior to testing in buffer solutions, we verified dry T-nwFETs’ intrinsic amplification function in air (see ‘Electrical measurement’ section of the Methods for more details). In Fig. 1d, the qualitative analysis showed that the T-nwFETs’ drain current IDrain (blue line) did change more rapidly than the corresponding sense current ISen change (red line) upon varying the substrate gate voltage, VSub. We should clarify though the ISen changes due to biosensing as discussed below were not induced by any variation in VSub.
Amplifying T-nanowire FET sensor and Semiconductor Electronic Label-Free Assay (SELFA).
(a) Structure of the label-free amplifying T-nwFET biosensor with integrated sensing and amplifying nwFET. (b) SEM image of a T-nwFET prototype fabricated on an SOI substrate. The Si nanowire width is 100 nm and the source, drain and sense contacts are made of Au-on-Ti. (c) Optical micrograph of an SU-8 microfluidic well exposing the T-nwFET’s sensing nanowire surface. (d) Measured drain and sense currents versus varying substrate gate voltage of a dry T-nwFET in air. (e) Photograph of a ceramic dual inline package embedding an SOI chip integrated with 24 T-nwFET sensors along two microfluidic channels, constituting the optics-free and enzyme-free SELFA. (f) Optical micrograph of an array of T-nwFET sensors along one integrated microfluidic channel.
For subsequent measurement portability, we packaged the SOI chip containing 24 T-nwFET sensors inside a ceramic dual inline package (DIP) via wire bonding (Fig. 1e), allowing concurrent quantitation of multiple analytes as a panel. We selected human inflammatory cytokine interleukin-1 beta (IL-1β) as a representative cell signaling secretome biomarker for device evaluation purposes without losing generality. IL-1β is a well-studied member of the interleukin family of cytokines which has an intermediate molecular weight of 17 kDa and isoelectric point (pI) of around 6.223. More importantly, IL-1β plays an important role in pathological fibrosis in which it is secreted by macrophages receiving toxicants such as engineered nanomaterials (ENMs) and is central to the regulation of inflammatory responses including acute pulmonary inflammation24,25. We immobilized the exposed sensing Si nanowire surface of the T-nwFETs in SELFA package with anti-human IL-1β monoclonal antibody (mAb) capture probes. We also passivated the remaining surface with amine-terminated polyethylene glycol (PEG) to suppress random biofouling or non-specific binding (for more details see ‘Surface modification’ subsection of the Methods section). The resultant optics-free packaged SELFA is readily interfaced with portable readout electronics for in silico data processing, storage and transmission, including the on-site measurement and data recording in Biosafety Level 2 laboratories.
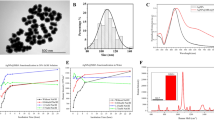
We first examined the LLOD of SELFA versus that of ELISA. To reduce the counter-ion screening effect in FET-based biosensing for SELFA, we prepared 0.01 × phosphate buffered saline (PBS) solutions with different human IL-1β concentrations through serial dilution from a spiked starting concentration of 2,000 pg/mL (C1) to 3.7 × 10−6 pg/mL (C30). We performed quantitative ELISA according to the kit manufacturer’s instructions (for more details see ‘ELISA procedures’ subsection of the Methods section). After sufficient binding and secondary conjugation, we measured the optical density (OD) values at 450 nm using a microplate reader for the aforesaid human IL-1β concentration series as shown in Fig. 2a. We found the ELISA kit’s LLOD to be ~5 pg/mL (between C9 and C10), a value which is close to the manufacturer’s specification and corroborated by the respective bright field images shown in the inset of Fig. 2a. The dynamic range (DR) of ELISA turned out to be about 2 orders of magnitude.
Evaluation of lower limit of detection (LLOD) and cross-reactivity with interferents.
Steady-state calibration curve for (a) ELISA and (b) SELFA. Measured interferent responses for (c) ELISA and (d) SELFA normalized to that of human IL-1β at the same concentration for two interferent concentrations – 10 and 100 pg/mL.
The same human IL-1β samples were used to test the performance of our T-nwFET in SELFA. In brief, we first introduced the pure 0.01 × PBS solution to the T-nwFET sensors and measured the respective IDrain as blank-signal levels (IDrain_bl). This was followed by a 10-minute incubation step, which is substantially shorter than that required in typical ELISA, to allow sufficient IL-1β binding with the mAb probes pre-immobilized onto the sensor’s nanowire surface. With the solutions in place, we measured the steady-state drain current IDrain_ss (and steady-state sense current ISen_ss) at few different VSub while holding the source, drain and sense terminal voltages (VSrc, VDrain and VSen) constant. Finally, we computed the differences between IDrain_ss and IDrain_bl as |ΔIDrain| (equivalent to |ΔGDrain| with constant VDrain)26 and plotted them against the corresponding human IL-1β concentrations (Fig. 2b). We extracted the LLOD of SELFA’s T-nwFET to be ~10 fg/mL, which we define here as the concentration that corresponds to the |ΔIDrain| value equals to 3 times of the IDrain_bl noise27. In addition, we observed the DR of SELFA to be around 4–5 orders of magnitude, which could be useful for quantitating specimens with wider range of biomolecular abundance.
The SELFA paradigm improved upon the LLOD of ELISA by almost 3 orders of magnitude. This was made possible because the attainable output-signal levels in SELFA’s T-nwFET sensors were substantially higher than those in generic nwFET sensors. The typical T-nwFET’s |ΔIDrain| was ~170 nA at ~100 fg/mL of IL-1β (the corresponding drain conductance change |ΔGDrain| is ~85 nS) while that of a typical generic nwFET at ~100 fg/mL of IL-6, another cytokine, was ~90 pA (with |ΔGDrain| of ~450 pS)28. Incidentally, the LLOD of that generic nwFET was ~100 fg/mL as well28 (see also Supplementary Data Fig. 3 for our own generic nwFET data).
Besides LLOD and DR, we evaluated both SELFA and ELISA for any cross reactivity with interferents that might lead to false-positive or inaccurate results. We selected three representative and biologically relevant interfering molecules with respect to the human IL-1β target. The first was mouse IL-1β due to its structural similarity to the target and thus potentially high cross reactivity with the anti-human IL-1β mAb capture probes. The second was human IL-8 in the same interleukin family which molecular weight of 11.1 kDa is similar to human IL-1β. The third molecule was bovine serum albumin (BSA) which is an abundant protein with low cross reactivity.
We prepared 0.01 × PBS solutions of each of the three interferents at two concentrations (i.e., 10 and 100 pg/mL), as well as the human IL-1β calibration standards of the same concentrations. We followed the aforementioned ELISA and SELFA procedures and collected the cross reactivity results as shown in Fig. 2c,d. For ease of comparisons between ELISA and SELFA, we normalized all the measured interferent responses with respect to that of human IL-1β of identical concentrations. Also, it is worth noting that the SELFA responses should be positive for mouse IL-1β and BSA since the pI values of both molecules (4.56 and 4.7, respectively29, signaling-gateway.org) are below the pH value of 0.01 × PBS solution (~7.4); conversely, the SELFA response should be negative for human IL-8 due to its pI value (9.1, signaling-gateway.org).
With 100 pg/mL of interferents (i.e., mouse IL-1β, human IL-8, or BSA), we observed almost equivalent selectivity between ELISA and SELFA, with averaged responses in absolute amplitude being 16.74% and 11.78%, respectively. At interferent concentration of 10 pg/mL, approaching the ELISA’s LLOD, ELISA generated an average response of 62.50% and thus became incapable of differentiating between interferent and target molecules. On the contrary, SELFA still maintained its fidelity yielding an average response of 12.15%.
In summary, SELFA is able to offer a substantially lower analytical LLOD compared to ELISA. This is made possible by the reagent-less and local signal transduction and amplification mechanisms harnessed by the T-nwFET sensors in SELFA. In addition, the label-free SELFA exhibits comparable analytical selectivity versus the labeling-based ELISA, validating the reliability of our SELFA platform.
SELFA implementation for predictive toxicology – A case study in acute lung inflammatory toxicity
We and others have previously developed a predictive toxicological approach for ENMs such as redox-active metal oxides and long aspect ratio (LAR) materials2,30 based on the assessment of pro-inflammatory and pro-fibrogenic cellular responses. In particular, we examined their toxicities via quantification of secretome biomarkers (e.g. IL-1β, other cytokines and growth factors) that is routinely performed using relatively expensive, time-consuming and labor-intensive ELISA assays in 96-well plates31,32,33,34,35. To demonstrate the utility of SELFA secretomics in predictive toxicology, we performed comparative analyses between SELFA and ELISA in quantitating IL-1β released in cultured macrophage receiving some new as well as previously examined ENMs. SELFA quantification of IL-1β release generates in vitro hazard ranking of the examined ENMs that directly indicates their degree of lysosomal damage in cells and acute inflammation in animal lungs. The presented SELFA platform and procedure development, side-by-side comparison and the confirmatory cellular and animal study serve as the first demonstration of its application in a real-life toxicological scenario.
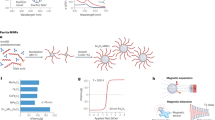
First, we utilized both SELFA and ELISA to quantitate human IL-1β in supernatants from cultured human macrophage cells (THP-1) receiving a library of ENMs (see the ‘Biological sample preparation’ subsection of the Methods section for more details). We obtained 11 well-characterized ENMs from the UC CEIN compositional and combinatorial material library30 for this validation study as tabulated in Fig. 3. In this set of ENMs, the toxicity of #2–#9 were previously assessed while #10–#12 were newly synthesized with unknown toxicity. The morphology of these ENMs was shown in TEM images in Supplementary Data Fig. 4. We also included PBS solution as a negative control as well as the supernatant collected from monosodium urate crystals (MSU) treated THP-1 cell culture as a positive control.
In vitro biomolecular assays for in vivo toxicology prediction.
(a) Various metal oxide and long aspect ratio ENMs were used to treat human macrophage cells (THP-1). PBS and MSU solutions served as the negative and positive control, respectively. (b) SELFA and ELISA quantitated concentration of human IL-1β in supernatants collected from THP-1 cells which received indicated treatment at 40 μg/mL for 24 hours. (c) Correlation between SELFA and ELISA quantitation. Quantitated concentrations of 13 samples (controls included) were linearly fitted, yielding a slope of 0.880 and an intercept of 93.8 pg/mL. R2 = 0.930.
For reliable quantitation of human IL-1β in these biological samples, we developed a novel strategy to calibrate SELFA on a per-sensor basis (for more details see the ‘Sensor calibration’ subsection of the Methods section). In essence, we introduced to each T-nwFET sensor standard solutions whose concentration was below and/or above the anticipated concentration in all supernatant samples to establish the reference calibration points. We were then able to extract the human IL-1β concentrations with confidence by interpolation from the calibration points. The reliable extraction was made possible by correcting for the effects of sensor-to-sensor variations inherited from fabrication and/or probe immobilization (see Supplementary Data Fig. 5 for representative calibration and extraction results). It should be noted that this per-sensor calibration strategy can be simplified to calibrate only a few sensors within a batch so long as the variation among individual sensors is generally minimal.
Next, we deployed and validated the holistic SELFA immunoassay including the hardware, assay procedures and calibration strategy through determination of human IL-1β concentration in the aforementioned supernatant and control samples. We diluted the supernatants by 250 times in 0.01 × PBS solution such that their concentrations were within the DR of T-nwFET sensors. We measured each sample using at least 2 different sensors and under multiple voltage biases to further minimize run-to-run variation like running ELISA in duplicates. The SELFA-quantitated concentrations along with the ELISA values for the supernatants under test (Samples #2–12) and controls (Samples #1 and #13) are shown in Fig. 3b, demonstrating good agreement between the two types of assays. For the purpose of method comparison, we further plotted the SELFA-quantitated concentrations against those obtained by ELISA as shown in Fig. 3c. SELFA demonstrates a linear correlation with ELISA with a R2 > 0.93. From these THP-1 data, we found that ENMs #2, #6, #7, #8 and #10, resulted in a significant increase in IL-1β production compared to PBS treated cells (#1), while ENMs #11–12 led to intermediary elevation. Sample #13, which is MSU positive control, yielded an expected increase in the level of IL-1β.
To further validate the toxicity ranking generated by SELFA, we carried out confocal microscopy study in THP-1 cells, animal experiments and histology assessments. It has been established that IL-1β production in macrophages requires assembly of the NLRP3 inflammasome, which activation is dependent on lysosome damage and the lysosomal enzyme cathepsin B release following the endocytosis of these ENMs36,37,38. Therefore, toward confirming the foregoing IL-1β ranking, we used confocal fluorescence microscopy to discern lysosomal damage by tracking the containment of cathepsin B that could be detected by Magic Red probes in THP-1 cells (see the ‘Confocal microscopy assay’ subsection of the Methods section for more details). As shown in Fig. 4a, PBS-treated cells (Sample #1) revealed a punctate distribution of red fluorescence in the intact lysosome, which is in marked contrast to the diffuse staining pattern observed in MSU-treated cells (positive control Sample #13). Consistent with the IL-1β data (Fig. 3) which identified ENMs (#2, #6, #7, #8 and #10) that triggered strong cytokine release, the Magic Red staining revealed the same set of ENMs that induced obvious cathepsin B release and potent lysosome damage, while ENMs #11–12 incurred intermediary lysosomal damage with a corresponding intermediary level of IL-1β production. The remaining ENMs did not provoke any observable lysosomal damage, as expected from the foregoing SELFA IL-1β cytokine analysis.
In vitro and in vivo experimentation confirming SELFA prediction of lung inflammation potential.
(a) Visualization of cathepsin B localization in THP-1 cells after ENMs treatment. After ENMs treatment, cells were stained with Magic Red (for identifying lysosomal damage and cathepsin B release) and wheat germ agglutinin-Alexa Fluor 488 (for identifying cell membrane), followed by visualization under a confocal 1P/FCS inverted microscope. (b) Anesthetized C57BL/6 mice were exposed to the selected ENMs once via oropharyngeal aspiration and were euthanized after 48 hours. The BAL fluids were collected to determine the neutrophil count. (c) Lung tissues were harvested, fixed and stained with hematoxylin/eosin (H&E). While a small increase in the level of pulmonary inflammation (marked by arrows) was observed for samples #11 and #12 compared to the control (#1), a dramatic acute pulmonary inflammation effect was apparent from #2.
Although we have demonstrated above an excellent correlation between the IL-1β data and confocal visualization, we have yet to show the former’s indication of pulmonary toxicity in animal lungs. In other words, we need to examine how well the SELFA-based cytokine ranking in vitro predicts the degree of acute lung inflammation in vivo. Due to logistic constraints (e.g. number of animals), we decided to validate the induced lung damage for selected ENMs, i.e. #2 (which was assessed before and with predicted high toxicity by SELFA), #11 and #12 (which were new ENMs with predicted intermediary toxicity) in an acute toxicity experiment in vivo. We compared the acute pro-inflammatory effects of introducing 5 μg of both ENM to the animal lungs. We chose this dose based on our experiences and its relevance to occupational exposure level estimated by dosimetry39. After 48 hours post oropharyngeal aspiration, we collected the bronchoalveolar lavage (BAL) fluid and determined the neutrophil count which is an acute response to inflammatory lung injury (see the ‘Animal experiment’ subsection of the Methods section for more details). We found that the neutrophil count in BAL fluid from animal treated with ENM #2 (as-prepared multiwall carbon nanotube) was significantly higher than that with #1 (PBS negative control) (Fig. 4b), which is in agreement with our previous results31. On the other hand, the ENMs #11 and #12 (Co3O4 nanocube and plate, respectively) triggered a weaker response compared to #2 yet the corresponding neutrophil count was still higher than that for #1. We further confirmed the increased potency of ENM #2 via histology of the corresponding lung tissue (Fig. 4c), which showed the typical focal areas of acute inflammation surrounding small airways in the lung (marked by arrows), in contrast to the less severe inflammation in the animal receiving #11 and #12. At the same time, we observed no acute inflammation with the negative control (#1) as expected. Concisely, Fig. 4 constitutes the integrated set of animal validation data that confirms the same degree of ENMs in vivo pulmonary toxicity as predicted by the in vitro toxicological cytokine ranking generated by the novel SELFA platform.
Discussion
Although a FET for detecting immunoreaction has existed for over 3 decades, a practical FET immunoassay with validated operation and calibration procedures that can quantitate biological specimens in the scenario of interest remains lacking. We have proposed and demonstrated in this work a label-free, high-sensitivity biomolecular assay platform, SELFA, based on our amplifying T-nwFET biosensor. We examined its sensitivity and selectivity to a cytokine of interest to predictive toxicology (human IL-1β) and compared the results with an ELISA assay. The results revealed that SELFA has an LLOD ~10 fg/mL, or almost 3 orders of magnitude lower than the conventional ELISA approach and a wider dynamic range spanning across 4–5 orders of magnitude. Moreover, in the presence of various interfering species, SELFA exhibits a high selectivity to human IL-1β comparable to ELISA. We further evaluated SELFA secretomics in predicting the inflammatory toxicity of various ENMs in vitro. Using a novel per-sensor calibration strategy, SELFA is able to quantitate the secreted human IL-1β concentrations in cultured human macrophages receiving different ENMs. The quantitation results agreed well with that of ELISA and corroborated the confocal visualizations of IL-1β related lysosomal damage. Most importantly, the SELFA-generated in vitro toxicity ranking was further substantiated by an acute lung inflammation study on animals instilled with selected ENMs. In sum, the holistic SELFA platform and procedures is able to quantify key biomarkers in real biospecimens and readily deployed for predictive toxicology studies.
Methods
Device fabrication
We fabricated T-nwFET devices on (100)-oriented p-type silicon-on-insulator (SOI) wafers (SOITEC, USA). We patterned the silicon nanowires via e-beam lithography, chrome layer evaporation, photoresist liftoff and subsequent reactive ion etching (RIE) of the top SOI layer not covered by chrome. Then we formed the contact electrodes via photoresist lift-off of an evaporated gold-on-titanium layer followed by rapid thermal annealing at 450 °C for 60 seconds. Next, we spin-coated a 1.5 μm SU-8 layer for surface electrical insulation with openings above and close to the sensing nanowire developed away. The SU-8 layer prevents shorting of metal contacts in the presence of buffer solutions and its hydrophobic surface facilitates biomolecular diffusion to above the sensing nanowire. After immobilizing the sensing surface with antibody capture probes, we packaged the sensor chip into a DIP carrier for portable electrical measurements.
Electrical measurements
We perform all electrical measurements using the Keithley 4200 Semiconductor Characterization System. We applied VDrain = 2 V and VSen = 0.8 V to the drain and sense terminals, respectively and grounded the source terminal. We swept the substrate bias Vsub from 15 V to 30 V between which the T-nwFET device operates in the linear regime. Prior to the introduction of biological samples, we measured the baseline drain current IDrain. In all sensing experiments using SELFA, we took steady-state measurements in 0.01 × PBS solutions to ensure a sufficiently long Debye length.
Surface modification
We functionalized the sensor surface with a 2-step self-assembled monolayer (SAM) layer to allow aldehyde-terminated groups to covalently bond to the antibodies of interest. We first treated the native oxide covering the T-nwFET sensing surface with 2% (v/v) 3-aminopropyltriethoxysilane (APTES) solution (Sigma-Aldrich). Next, we introduced the homobifunctional crosslinker glutaraldehyde (2% v/v) (Sigma-Aldrich) to add an aldehyde moiety to the surface. Subsequently, we coated T-nwFET surface with 10 μg/mL monoclonal anti-human IL-1β antibody in order to covalently attach antibodies to the surface through their amine moiety. Finally, we introduced 1 mg/mL amine-polyethylene glycol (PEG) to block unreacted aldehyde groups and suppress random fouling of the sensor.
ELISA procedures
We measured the IL-1β concentrations in spiked PBS solutions and cultured THP-1 supernatants using the OptEIA ELISA kit (BD Biosciences, CA) according to the manufacturer’s instructions. In brief, we first coated the 96-well plate with monoclonal anti-human IL-1β antibody. After blocking the surface with PBS supplemented with 10% fetal bovine serum to suppress random fouling, we added 50 μL of calibration and biological samples into the 96-well plate for overnight incubation at 4 °C. Next, we conjugated the captured cytokines with secondary polyclonal anti-human IL-1β cross-linked to horseradish peroxidase. In order to minimize non-specific bindings, we performed multiple washing steps per the instructions including 3 washes after sample incubation, 3 washes after secondary antibody conjugation and 7 washes before measurement. We measured the absorbance at 450 nm using a plate reader (SpectroMax M5e, Molecular Devices Corp., Sunnyvale, CA, USA).
Biological sample preparation
We obtained the THP-1 cells from ATCC (Manassas, VA). We cultured the THP-1 cells in RPMI 1640 medium supplemented with 10% fetal bovine serum at 5% CO2 and 37 °C. We seeded aliquots of 5 × 104 THP-1 cells in 0.1 mL complete medium with 1 μg/mL phorbol 12-myristate 13-acetate (PMA) overnight in 96-well plates (Corning, NY, USA). We primed the cells with 10 ng/mL lipopolysaccharide (LPS) to initiate transcriptional activation of the IL-1β promoter. We prepared fresh ENM suspensions (40 μg/mL) in complete RPMI medium and sonicated them before use. After 24 hours post ENMs introduction, we separated the ENMs from the supernatant via a centrifugation process at 15,000 rpm for 10–15 minutes. Such procedure had been confirmed to be effective through the use of ICP-OES elemental analysis and TEM observation34,35. After that, we measured the secreted IL-1β concentrations in resultant culture medium using IL-1β ELISA kit and SELFA.
Sensor calibration
To minimize the impact of sensor-to-sensor variations due to fabrication or surface preparation on the T-nwFET sensing reliability, we proposed and implemented the following per-sensor calibration strategy to quantitate IL-1β concentration in supernatant samples. On each antibody and PEG-modified virgin T-nwFET sensor, we first recorded the baseline IDrain in blank 0.01 × PBS solution as IDrain_bl. Then we measured the IDrain’s in 3 standard spiked solutions that we introduced sequentially in an ascending order of human IL-1β concentration; these concentrations were below those expected of the supernatant samples to be measured. Next we introduced a supernatant sample and noted the corresponding IDrain. After that, we recorded the IDrain of the last standard solution which human IL-1β concentration was higher than the expected supernatant concentration. Note that all foregoing IDrain’s were steady-state values and we subtracted IDrain_bl apiece from them as |ΔIDrain|’s. Finally, we fit the lower and higher concentration |ΔIDrain|’s using a standard Langmuir isotherm based expression and interpolated the supernatant |ΔIDrain| value to extract the corresponding human IL-1β concentration as illustrated in Supplementary Data Fig. 5.
Confocal microscopy assay
We detected cathepsin B in THP-1 cells post-ENM exposure using the Magic Red-cathepsin B substrate. We first treated the THP-1 cells with 40 μg/mL ENMs in complete RPMI 1640 medium for 8 hours. We then washed these cell samples with PBS solution, stained them with Magic Red (ImmunoChemistry Technologies) for 1 hour and fixed them in 4% paraformaldehyde for 20 minutes. After two subsequent washes in PBS solution, we stained the cell membrane with Alexa Fluor 488-conjugated wheat germ agglutinin at room temperature for 1 hour. Finally, we visually examined these cells under a confocal microscope (Leica Confocal SP2 1P/FCS) in the UCLA/CNSI Advanced Light Microscopy/Spectroscopy Shared Facility. We obtained high-magnification images with a 100× objective. We used cells treated with PBS solution and 50 μg/mL of MSU as the negative and positive control, respectively.
Animal experiment
In order to validate the in vitro cytokine data, we performed an animal experiment to assess the acute toxicological responses in the animal lung by oropharyngeal aspiration. We purchased eight week old male C57Bl/6 mice from Charles River Laboratories (Hollister, CA). We housed all mice under standard laboratory conditions that were set up according to an approved protocol. All procedures were conducted in accordance with Division of Laboratory Animal Medicine (DLAM) guidelines and with the approval of the Ethical Committee and Animal Research Committee of UCLA (2004-022-41). We exposed the mice to various ENMs via oropharyngeal aspiration as previously described39. In brief, we anesthetized the mice by intraperitoneal injection of ketamine (100 mg/kg)/xylazine (10 mg/kg) with a total volume of 100 μL. With each anesthetized mouse held in a vertical position, we instilled 50 μL of a suspension with 5 μg of one ENM in water (equivalent to 0.25 mg/kg) at the back of the tongue to allow aspiration in the lung. Control mice received the same volume of PBS solution (negative control). We sacrificed the mice after 48 hours post-exposure and collected the BAL fluid and lung tissues. We used the BAL fluid to perform total and differential neutrophil cell counts. We stained histological sections with hematoxylin/eosin to reveal the level of pulmonary inflammation.
Additional Information
How to cite this article: Mao, Y. et al. Semiconductor Electronic Label-Free Assay for Predictive Toxicology. Sci. Rep. 6, 24982; doi: 10.1038/srep24982 (2016).
References
National Research Council Committee on Toxicity Testing and Assessment of Environmental Agents. Toxicity Testing in the 21st Century: A Vision and a Strategy. National Academies Press. Washington, D.C. (2007).
Meng, H., Xia, T., George, S. & Nel, A. E. A Predictive toxicological paradigm for the safety assessment of nanomaterials. ACS Nano 3, 1620–1627 (2009).
Nel, A., Xia, T., Mädler, L. & Li, N. Toxic potential of materials at the nanolevel. Science 311, 622–627 (2006).
Service, R. F. Can high-speed tests sort out which nanomaterials are safe? Science 321, 1036–1037 (2008).
Pardo-Martin, C. et al. High-throughput in vivo vertebrate screening. Nat Meth 7, 634–636 (2010).
Zhao, F., Meng, H., Yan, L., Wang, B. & Zhao, Y. Nanosurface chemistry and dose govern the bioaccumulation and toxicity of carbon nanotubes, metal nanomaterials and quantum dots in vivo. Sci Bull 60, 3–20 (2015).
Rallo, R. et al. Self-organizing map analysis of toxicity-related cell signaling pathways for metal and metal oxide nanoparticles. Environ Sci Technol 45, 1695–1702 (2011).
Puzyn, T. et al. Using nano-QSAR to predict the cytotoxicity of metal oxide nanoparticles. Nat Nano 6, 175–178 (2011).
Donaldson, K., Borm, P., Castranova, V. & Gulumian, M. The limits of testing particle-mediated oxidative stress in vitro in predicting diverse pathologies; relevance for testing of nanoparticles. Part Fibre Toxicol 6, 13 (2009).
Schenk, J. F. General Electric Company. Field effect transistor for detection of biological reactions. United States patent US 4,238, 757. 1980 Dec 9.
Gotoh, M., Suzuki, M., Kubo, I., Tamiya, E. & Karube, I. Immuno-FET sensor. J Mol Catal 53, 285–292 (1989).
Cui, Y., Wei, Q., Park, H. & Lieber, C. M. Nanowire nanosensors for highly sensitive and selective detection of biological and chemical species. Science 293, 1289–1292 (2001).
Patolsky, F., Zheng, G. & Lieber, C. M. Fabrication of silicon nanowire devices for ultrasensitive, label-free, real-time detection of biological and chemical species. Nat Protoc 1, 1711–1724 (2006).
Stern, E. et al. Label-free immunodetection with CMOS-compatible semiconducting nanowires. Nature 445, 519–522 (2007).
Lee, M. H. et al. Quantitative measurements of C-reactive protein using silicon nanowire arrays. Int J Nanomedicine 3, 117–124 (2008).
Chee, R., Chua, J., Agarwal, A., Wong, S. & Zhang, G. Label-free detection of proteins with surface-functionalized silicon nanowires. The 13th International Conference on Biomedical Engineering (ICBME), IFMBE Proceedings 23, 838–841 Singapore, December 3–6, 2008.
Zheng, G., Gao, X. P. A. & Lieber, C. M. Frequency domain detection of biomolecules using silicon nanowire biosensors. Nano Lett 10, 3179–3183 (2010).
Gao, X. P. A., Zheng, G. & Lieber, C. M. Subthreshold regime has the optimal sensitivity for nanowire FET biosensors. Nano Lett 10, 547–552 (2010).
Bergveld, P. The future of biosensors. Sens Actuators A 56, 65–73 (1996).
Shin, K.-S., Lee, K., Kang, J. Y. & Chui, C. O. Novel T-channel nanowire FET with built-in signal amplification for pH sensing. IEEE Int Electron Dev Mtg (IEDM), Tech Dig 599–602, 10.1109/IEDM.2009.5424288 Baltimore, MD, December 7–9, 2009.
Shin, K.-S., Lee, K., Park, J.-H., Kang, J. Y. & Chui, C. O. Schottky Contacted Nanowire Field-Effect Sensing Device with Intrinsic Amplification. IEEE Electron Dev Lett 31, 1317–1319 (2010).
Shoorideh, K. & Chui, C. O. On the origin of enhanced sensitivity in nanoscale FET-based biosensors. Proc. Natl Acad. Sci. USA 111, 5111–5116 (2010).
Brattsand, M. & Egelrud, T. Purification and characterization of interleukin 1 beta from human plantar stratum corneum. Evidence of interleukin 1 beta processing in vivo not involving interleukin 1 beta convertase. Cytokine 10, 506–513 (1998).
Kolb, M., Margetts, P. J., Anthony, D. C., Pitossi, F. & Gauldie, J. Transient Expression of IL-1β induces acute lung injury and chronic repair leading to pulmonary fibrosis. J Clin Invest 107, 1529–1536 (2001).
Wang, X. et al. Use of a –fibrogenic mechanism-based predictive toxicological approach for tiered testing and decision analysis of carbonaceous nanomaterials. ACS Nano 9, 3034–3043 (2015).
Lin, T. et al. Label-free detection of protein-protein interactions using a calmodulin-modified nanowire transistor. Proc. Natl. Acad. Sci. USA 107, 1042–1052 (2010).
MacDougall, D. et al. Guidelines for data acquisition and data quality evaluation in environmental chemistry. Anal Chem 52, 2242–2249 (1980).
Pui, T.-S., Agarwal, A., Ye, F. & Chen, P. Nanoelectronic detection of triggered secretion of pro-inflammatory cytokines using CMOS compatible silicon nanowires. Biosens Bioelectron 26, 2746–2750 (2011).
Ge, S., Kojio, K., Takahara, A. & Kajiyama, T. Bovine serum albumin adsorption onto immobilized organotrichlorosilane surface: influence of the phase separation on protein adsorption patterns. J Biomater Sci Polym Ed. 9, 131–150 (1998).
Nel, A. et al. Nanomaterial toxicity testing in the 21st century: use of a predictive toxicological approach and high-throughput screening. Acc. Chem. Res. 46, 607–621 (2013).
Wang, X. et al. Quantitative techniques for assessing and controlling the dispersion and biological effects of multiwalled carbon nanotubes in mammalian tissue culture cells. ACS Nano 4, 7241–7252 (2010).
Wang, X. et al. Dispersal state of multiwalled carbon nanotubes elicits profibrogenic cellular responses that correlate with fibrogenesis biomarkers and fibrosis in the murine lung. ACS Nano 5, 9772–9787 (2011).
Zhang, H. et al. Use of metal oxide nanoparticle band gap to develop a predictive paradigm for oxidative stress and acute pulmonary inflammation. ACS Nano 6, 4349–4368 (2012).
Ji, Z. et al. Designed synthesis of CeO2 nanorods and nanowire for studying toxicological effects of high aspect ratio nanomaterials. ACS Nano 6, 5366–5380 (2012).
Li, R. et al. Surface charge and cellular processing of covalently functionalized multiwall carbon nanotubes determine pulmonary toxicity. ACS Nano 7, 2352–2368 (2013).
Wang, X. et al. Pluronic F108 coating decreases the lung fibrosis potential of multiwall carbon nanotubes by reducing lysosomal injury. Nano Lett. 12, 3050–3061 (2012).
Dostert, C. et al. Innate immune activation through Nalp3 inflammasome sensing of asbestos and silica. Science 320, 674–677 (2008).
Jin, C. & Flavell, R. A. Molecular mechanism of NLRP3 inflammasome activation. J. Clin. Immunol. 30, 628–631 (2010).
Wang, X. et al. Use of coated silver nanoparticles to understand the relationship of particle dissolution and bioavailability to cell and lung toxicological potential. Small 10, 385–398 (2014).
Acknowledgements
This work was supported in part by the National Science Foundation (NSF) Grant ECCS 1128673 to C.O.C. and the NSF and Environmental Protection Agency (EPA) under Cooperative Agreement Number DBI 1266377 to UC CEIN. Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of NSF or EPA. This work has not been subjected to EPA review and no official endorsement should be inferred.
Author information
Authors and Affiliations
Contributions
Y.M., H.M. and C.O.C. designed research. C.O.C. guided research. Y.M. and K.-S.S. performed SELFA experiments and analyzed the data. Z.J. synthesized and characterized the ENMs. X.W. and H.M. performed all other experiments and analyzed the data. Y.M., H.M. and C.O.C. wrote the paper.
Ethics declarations
Competing interests
C.O.C. has a commercial interest with Selfa, Inc. The University of California has filed patents on technologies described herein.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mao, Y., Shin, KS., Wang, X. et al. Semiconductor Electronic Label-Free Assay for Predictive Toxicology. Sci Rep 6, 24982 (2016). https://doi.org/10.1038/srep24982
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24982
This article is cited by
-
Gradient-based impedance synthesis for breast and lung cancer cell screening deploying planar and nano-structured electrodes
Medical & Biological Engineering & Computing (2021)
-
Point-of-care-ready nanoscale ISFET arrays for sub-picomolar detection of cytokines in cell cultures
Analytical and Bioanalytical Chemistry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.