Abstract
Transposable elements (TEs), or transposons, play an important role in adaptation. TE insertion can affect host gene function and provides a mechanism for rapid increases in genetic diversity, particularly because many TEs respond to environmental stress. In the current study, we show that the transposition of a heat-activated retrotransposon, ONSEN, generated a mutation in an abscisic acid (ABA) responsive gene, resulting in an ABA-insensitive phenotype in Arabidopsis, suggesting stress tolerance. Our results provide direct evidence that a transposon activated by environmental stress could alter the genome in a potentially positive manner. Furthermore, the ABA-insensitive phenotype was inherited when the transcription was disrupted by an ONSEN insertion, whereas ABA sensitivity was recovered when the effects of ONSEN were masked by IBM2. These results suggest that epigenetic mechanisms in host plants typically buffered the effect of a new insertion, but could selectively “turn on” TEs when stressed.
Similar content being viewed by others
Introduction
Plants cannot relocate to avoid stressful environmental conditions; therefore, climatic variation is a major environmental stressor and exerts strong selective pressure on plant populations1. In response, plants must either possess phenotypic plasticity or generate enough genetic diversity to adapt2.
Transposable elements (TEs) are a major source of genetic variation because their insertions create mutations that can affect coding regions or cause genomic rearrangements3,4,5,6. Furthermore, environmental stress activates TEs in plants5,7,8,9,10,11,12,13,14,15,16, potentially triggering the genetic diversity required to evolve adaptations. However, we currently know little about the exact mechanisms behind stress-activated TEs, including whether they are successfully inherited by future generations.
Although TE-generated mutations are important for adaptive evolution, excessive genomic changes can be harmful. Therefore, plants have also evolved processes that can suppress TE activity. A well-studied epigenetic regulation of TEs is RNA-directed DNA methylation (RdDM)17,18. In Arabidopsis, small interfering RNAs (siRNAs) are synthesized from RNA polymerase IV (PolIV)19,20,21 and eventually form an RNA-induced silencing complex that is involved in DNA methyltransferase recruitment, leading to the de novo methylation of target TEs22,23,24,25.
Previously, we reported that heat stress activates the retrotransposon ONSEN in Arabidopsis. Moreover, a transgenerational transposition was observed in the progeny of heat-stressed nrpd1, a line of mutants that lack functional PolIV (NRPD1 is a subunit of PolIV)14. Other experiments in Arabidopsis created ONSEN-integrated progeny in heat-stressed nrpd1 mutants14,26. Because ONSEN insertions generally alter gene expression, we hypothesized that ONSEN’s gene-targeting transposition could generate novel stress-responsive regulatory genes. In the present study, we induced ONSEN transposition in nrpd1 mutants via heat stress to determine stress tolerance in ONSEN-integrated progenies. We used a plant hormone, abscisic acid (ABA), that plays an important role in plant responses to environmental stress27. The application of exogenous ABA has been used to mimic osmotic stress. Increased ABA levels induce the expression of many genes that possibly play multifaceted roles in response to osmotic stress28. Nearly 10% of the protein coding-genes in Arabidopsis are regulated by ABA29.
We investigated the mechanism of ABA-insensitive phenotypes in F2 progenies by focusing on the transcriptional regulation of ABA-responsive genes that possessed an ONSEN insertion. In Arabidopsis, some intronic TEs within transcribed genes are marked by repressive epigenetic modifications and splicing of the TE-containing intron is promoted by the nuclear protein INCREASE IN BONSAI METHYLATION2 (IBM2)30,31. IBM2 controls one of the ONSEN copies AT1G11265, embedded in the intron of F-box protein AT1G11270 30. Here, we show that ONSEN-integrated progenies in Arabidopsis are stress tolerant and the effects of an ONSEN insertion are regulated by the host plant.
Results
Transgenerational transposition of ONSEN resulted in ABA-insensitive plants
The ONSEN-integrated progeny in heat-stressed nrpd1 mutants were seeded on an MS medium containing ABA. ABA is closely associated with cellular dehydration processes in seed maturation and vegetative growth. We also tested heat, salt and cold stress. The phenotype we analyzed with ABA was the clearest result of environmental stress responses. The ONSEN-integrated mutants that exhibited ABA-insensitive phenotypes were screened 10 d after germination. Out of 8,000 seeds, we found two ABA-insensitive mutants. Mutant phenotypes do not differ from the wild type under normal conditions (Fig. 1a). However, in the presence of ABA the mutants exhibited elevated ABA-resistance (Fig. 1b). A Southern blot analysis of the ABA-insensitive progenies revealed that ONSEN copies were inserted into multiple loci (Fig. 1c), suggesting that a new ONSEN insertion could affect the expression of an ABA-responsive gene.
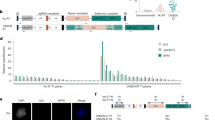
ABA-insensitive progenies with ONSEN copies.
(a,b) Phenotypes of young seedlings growing under normal conditions (a) and under 2 μM ABA (b). 13–7; ONSEN-integrated mutant; wt, wild type; nrpd1, original nrpd1 mutant line. (c) Southern blot analysis revealed that several new copies of ONSEN were detected in ABA-insensitive progenies (13–7). Each lane represents an individual progeny produced from a single heat-stressed nrpd1. The leftmost column represents a wild-type progeny growing under normal conditions. (d) Mapping of ONSEN-integrated genes in a mutant line. Twenty new copies were mapped within genes on the Arabidopsis genome in line 13–7. The loci names with red letters indicate the new insertions, while the black letters indicate the original eight ONSEN copies.
ONSEN insertions are associated with euchromatic genes
To identify the ONSEN insertion sites, we analyzed the whole genome sequences of the two stress-insensitive mutants using a next generation sequencer. In one mutant (13–7), new ONSEN insertions were identified in 24 loci of the sequenced genome (Fig. 1d, see Supplementary Table S1): 20 were located within genes and 14 were inserted within exons. In the other mutant (19–4), the new copies were inserted into 21 loci: 13 were located within genes and eight were inserted within exons (see Supplementary Fig. S1, Supplementary Table S2). We then assessed the ONSEN-targeted genes based on all annotated genes in the TAIR10 database (see Supplementary Table S3) to understand ONSEN target site preferences. The average size, exon number and exon size of the targeted genes did not differ from the reference genes. However, the average intron size of the targeted genes was significantly smaller than that of the reference genes. These results indicate that a new ONSEN copy can target genes independent of DNA sequence. Instead, the transposon may be directed by gene structure on euchromatic regions.
ONSEN target sites
A heat-activated gene is a possible candidate for an ONSEN target site because transgenerational transpositions occurred in heat-stressed plants. To test this hypothesis, we compared heat-stressed versus normal expression levels of ONSEN-targeted genes in both wild type and nrpd2 mutants (see Supplementary Table S4). We chose nrpd2 mutants because plants deficient in NRPD2 (a subunit of PolIV) are hypersensitive to heat exposure32. Published microarray data of genome-wide gene expression in heat-stressed wild type and nrpd2 plants were used for the analysis. In nrpd2, ONSEN-targeted gene expression levels were not upregulated by heat stress; rather, 6 out of 33 target genes were down-regulated (see Supplementary Table S4). Although heat activation of ONSEN-targeted genes in nrpd1 was not analyzed, these results indicate that ONSEN transposition does not require heat activation of the target site.
We previously reported that ONSEN insertion must take place in flowers33. To determine whether the insertions identified in the current study correspond to genes that are activated during flower development, we analyzed genes that are targeted by ONSEN using microarray data (see Supplementary Table S5). The results showed that the genes were not transcribed at a specific stage during flower development and did not correspond to the time when transposition events previously resulted in stable inheritance.
Transposition of heat-activated ONSEN into ABA-responsive genes
To identify the gene(s) responsible for the ABA-insensitive phenotype, we focused on ONSEN insertions in ABA-responsive genes. Whole genome sequencing data of the 13–7 line revealed an ONSEN insertion in the first intron of ABSCISIC ACID-INSENSITIVE5 (ABI5) in the 13–7 line: located on chromosome 2 in position 15,207,390 on an AGI (Arabidopsis Genome Initiative) map, ABI5 encodes a basic leucine zipper transcription factor involved in ABA signaling during seed maturation and germination34,35,36,37 (see Supplementary Fig. S2a). In the 19–4 line, an ONSEN insertion was detected on an exon of ABSCISIC ACID-INSENSITIVE4 (ABI4). Located on chromosome 2 in position 16,797,945 of an AGI map, ABI4 is a member of the APETALA2 (AP2) domain family of transcriptional regulators. Mutant abi4 alleles have been identified in a variety of screens including ABA-resistant germination and salt-resistant germination38,39,40 (see Supplementary Fig. S2b).
To confirm an ONSEN insertion in the ABI5- and ABI4-coding regions, PCR was conducted to amplify the target region. The results indicate that a full-length ONSEN was integrated into both ABI5 and ABI4 (see Supplementary Fig. S2c,d).
The ONSEN family in the A. thaliana ecotype Columbia consists of eight full-length copies distributed over chromosomes 1, 3 and 5. To determine which member of the family was transposed into ABI5 in 13–7 and ABI4 in 19–4, a 700-bp sequence of the long terminal repeat (LTR) and the coding region of the inserted ONSEN were scored for element-specific single nucleotide polymorphisms that distinguish the eight genomic templates. We found that the ONSEN insertion sequence in 13–7 and 19–4 was assigned to AT1G11265 and AT3G61330, respectively (see Supplementary Fig. S3). These two copies contained intact LTRs and are capable of forming extrachromosomal copies that can potentially integrate into new genomic sites41.
An ONSEN insertion disrupted ABA-responsive genes in the mutant progenies
To understand how ONSEN insertions affect genes, genome-wide gene expression analyses were performed on the ONSEN-integrated lines using a microarray. The microarray analyses identified 6,520 and 7,262 genes that were expressed differentially under ABA stress in 13–7 and 19–4, respectively (expression change was at least 1.5-fold for upregulated genes and 0.67-fold or lower for down-regulated genes, FDR <0.05, Fig. 2a). We also found that 4,702 genes (72%) in 13–7 and 4,891 genes (67%) in 19–4 overlapped with the ABA-responsive genes in nrpd1 (Fig. 2a). Most of the differentially expressed genes in the two mutant lines exhibited low ABA-sensitivity (Fig. 2b), suggesting that both 13–7 and 19–4 were critically deficient in processes involving ABA response. ABI4 and ABI5 are both transcription factors that regulate seed-specific ABA-inducible genes during germination37,39,42. 13–7 and 19–4 exhibited ABA-insensitive phenotypes at germination that were consistent with the phenotype in abi5 and abi4 mutants (Fig. 2c). In addition, 72 out of 95 genes and 28 out of 59 genes that were directly regulated by ABI4 and ABI5 were suppressed in 19–4 and 13–7, respectively (see Supplementary Fig. S4). The probability frequency of ABI5/ABI4 targets or differential ABA-response genes in 13–7 and 19–4 was examined by a two-tailed binomial test. The statistical test indicated that the representation of the overlap between ABI5/ABI4 targets and the differential ABA-response genes in the 13–7 and 19–4 lines was significantly high (p-value: 2.2 × 10−1616 and 1.4 × 10−1747 respectively). Previous research investigating the suppression of ABI4 and ABI5 expression has shown similar results in downstream genes43. A mutation in ABI4 was shown to increase salt tolerance44,45,46. To determine the salt tolerance in an ONSEN insertion mutant, we applied salt stress to the seedlings of 19–4. The tested mutant plants led to elongate root compared to wild type when the seedlings were growing on top of agar-solidified medium containing 100 mM NaCl (Fig. 3). In summary, ONSEN insertion in these lines created mutant alleles of the affected genes.
Genome-wide gene expression analysis and ABA insensitive phenotype of the ONSEN-integrated lines.
(a) Venn diagram of differentially expressed genes in 13–7, 19–4 and nrpd1 under ABA stress. Microarray analysis identified significant differentially expressed (expression change was 1.5-fold or 0.67-fold, FDR <0.05) genes of 13–7 and 19–4 compared with parental nrpd1 under ABA treatment (green and red). ABA-responsive genes were identified by comparing ABA treated-nrpd1 with non-treated-nrpd1 (yellow, expression change was 1.5-fold or 2/3-fold, FDR <0.05). (b) Expression heat map of significant differentially expressed genes in 13–7 and 19–4. Genes that exhibited significant differential expression in (a) were used. The heat map shows the expression levels in wild type, nrpd1, 13–7 and 19–4, with or without ABA treatment. (c) ABA-insensitive phenotypes at germination. The phenotype of abi5 and abi4 were similar to 13–7 and 19–4 respectively.
Salt-tolerance phenotype of 19–4.
(a) Two-week old seedlings of 19–4 and abi4 mutant showed root elongation under 100mM NaCl. (b) Comparison of root length of two-week old seedlings growing under salt stress (n = 10). Statistic analysis showed the root length of 19–4 and abi4 mutant was significantly longer than that of wild type. Asterisks mark significant difference (P < 0.05).
The intragenic insertion of ONSEN was cloaked by IBM2
The ABA insensitive phenotype was inherited in the self-fertilizing progeny of 13–7 (see Supplementary Fig. S5), but not when 13–7 was crossed with the wild type (F2 population; see Supplementary Fig. S6b). To test potential trans-effects, 13–7 was backcrossed with nrpd1. Of 112 seedlings, 8 showed an ABA-insensitive phenotype and the ABA-insensitive segregant contained the ONSEN insertion within ABI5; however 20 out of 100 of the ONSEN insertion was detected in ABI5 in the F2 (see Supplementary Fig. S7). In contrast, about 25% (24 out of 100) of the F2 population from a cross between 19–4 and the wild type was ABA-insensitive (see Supplementary Fig. S6d).
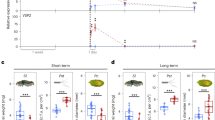
ONSEN was inserted into the first intron of ABI5 in 13–7 and into the single exon of ABI4 in 19–4 (see Supplementary Fig. S2a,b). To determine epigenetic regulation of intragenic ONSEN, we crossed 13–7 and 19–4 with an IBM2 mutant (ibm2). The resultant F2 progeny was genotyped at IBM2, NRPD1 and the ONSEN insertion in ABI5 (13–7) and in ABI4 (19–4). The F3 progenies derived from the genotyped parents were then exposed to ABA for analysis of ABA-insensitivity. For the F3 progeny of 13–7, an ABA-insensitive phenotype was observed 87 out of 100 seedlings in 13–7 ibm2, although 13–7 and 13–7 nrpd1 were ABA sensitive (Fig. 4a). In contrast, the F3 progeny of 19–4 exhibited an ABA-insensitive phenotype in 19–4 (96 out of 99 seedlings), 19–4 ibm2 (97 out of 100 seedlings) and 19–4 nrpd1 (90 out of 100 seedlings) (Fig. 4a). To confirm a relationship between the phenotype and the transcription of ABA-responsive genes, we used quantitative real-time PCR to analyze transcript levels in the F3 progenies. We found that ONSEN insertions disrupted ABI4 transcripts in 19–4, 19–4 ibm2 and 19–4 nrpd1 (Fig. 4b), while ABI5 transcript levels were significantly recovered in 13–7 and 13–7 nrpd1 (Fig. 4b).
ABA-insensitive phenotype and a transcription level of ABI4 and ABI5 in the F3 progeny.
(a) Phenotype of F3 progenies derived from progenies of a cross between an ABA-insensitive mutant (13–7 or 19–4) and an IBM2 mutant (ibm2) growing under 2 μM ABA. 13–7, 19–4: F3 progenies that contain an ONSEN insertion in ABI5 and ABI4, respectively. 13–7 ibm2, 19–4 ibm2: F3 progenies that contain a mutation in ibm2 and an ONSEN insertion in ABI5 or ABI4, respectively. 13–7 nrpd1, 19–4 nrpd1: F3 progenies that contain a mutation in nrpd1 and an ONSEN insertion in ABI5 or ABI4, respectively. (b) Transcript levels of ABI5 and ABI4 in the F3 progenies. Transcript levels of ABI5 and ABI4 in F3 progenies (derived from the progeny of a cross between 13–7 or 19–4 and an ibm2) growing under 2 μM ABA.
Next, we used reverse-transcription PCR to amplify ONSEN-inserted regions in the F3 progeny and detect RNA expression. We found that the first intron was spliced out in 13–7 and 13–7 nrpd1, similar to the wild type, but the expected product was not amplified in 13–7 ibm2 (Fig. 5a). To reveal whether transcription or splicing was affected in the insertion line, we conducted 3′ Rapid Amplification of cDNA Ends (RACE) of ABI5 in the F3 progeny. The results showed that proportions of shorter forms of transcript variants were increased in 13–7 ibm2 (Fig. 5b). These results suggest that IBM2 successfully promoted RNA expression on the ONSEN-inserted regions in 13–7 nrpd1 and 13–7, but in 13–7 ibm2, ONSEN insertions may cause transcription elongation and/or splicing defects.
RNA expression at the ONSEN insertion and DNA methylation of the 5′ and 3′ flanking regions of the ONSEN insertion.
(a) RT-PCR analyses of transcripts for ABI5. A PCR product of the ONSEN insertion site was not amplified in the 13–7 ibm2 and 13–7 parent line because of transcription elongation defects. In contrast, the first intron was spliced out in 13–7 and 13–7 nrpd1 of F3, similarly to the wild type. ABI5 1st intron; the first intron that ONSEN was inserted in 13–7. 18S; 18S rRNA gene. Genomic DNA (gDNA) was used as a template. The red arrowhead indicates an ONSEN insertion. (b) A 3′ Rapid Amplification of cDNA Ends (RACE) of ABI5 in the F3 progeny. The red arrowhead indicates an ONSEN insertion and black arrows indicate primers used in 3′ RACE. (c) Bisulfite sequence analysis of 5′ and 3′ flanking regions of the ONSEN insertion in F3 progenies (derived from the progeny of a cross between 13–7 and an ibm2). 13–7: F3 progeny that contains an ONSEN insertion in ABI5. 13–7 ibm2: F3 progeny that contains an ONSEN insertion in ABI5 and a mutation in IBM2. 13–7 nrpd1: F3 progeny that contains an ONSEN insertion in ABI5 and a mutation in NRPD1.
Finally, we examined the participation of DNA methylation in the epigenetic regulation of ONSEN insertion into ABI5. We observed an increase in methylation of the 500-bp 5′ and 3′ regions of the ONSEN insertion when 13–7 was crossed with ibm2 mutants. Interestingly, the methylation level of the F1 between 13–7 and nrpd1 showed a re-establishment of cytosine methylation at CHH (where H = A, T, or C) sites (see Supplementary Fig. S8). The re-establishment of cytosine methylation might responsible for the unexpected segregation of ABA insensitive progeny by the cross of 13–7 with nrpd1. The presence of CHH methylation in an nrpd1 background suggests other DNA methylation pathways to establish the de novo DNA methylation besides the canonical RdDM. In the F3 progeny of 13–7, cytosine methylation at CHH sites in both 13–7 ibm2 and 13–7 were similar, whereas methylation was significantly reduced in 13–7 nrpd1 (Fig. 5c). These results suggest that the IBM2-mediated transcriptional recovery of ABI5 and related ABA-sensitive phenotypes in the F3 progeny were independent of epigenetic regulation through the RdDM pathway.
Heat-activation of the intragenic insertion of ONSEN
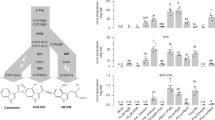
It has previously been shown that new ONSEN insertions are able to activate genes in close proximity under heat stress that renders genes sensitive to environmental stimuli14. To understand the effect of the ONSEN transcript, we analyzed the transcript level of ABI5 on the ONSEN inserted region and downstream of ONSEN insertion in the 13–7 line. The transcript of ABI5 on the inserted region was not detected under heat-stress conditions (Fig. 6a). However, even without ABA stress, the ABI5 transcript level on the downstream of the ONSEN insertion was upregulated under heat-stress conditions (Fig. 6b). We also analyzed the transcript level of ABI5 on the re-activated backcross ONSEN insertion line of ABI5 in the F3 progeny (13–7 IBM2 NRPD1). The transcript level of ABI5 on the downstream of ONSEN insertion was also upregulated under heat-stress conditions, although the transcript was not detected on the inserted region (Fig. 6a,b). To determine whether the ONSEN transcript confers adaptive advantages to overcome ABA stress, we analyzed the ABA sensitivity of the ONSEN insertion line of ABI5 in the F3 progeny subject to heat stress under ABA stress. In the F3 progeny derived from 13–7, an ABA-insensitive phenotype was observed in 13–7 ibm2, although 13–7 and 13–7 nrpd1 were ABA sensitive (Fig. 6c). The results indicated that heat stress lead to a reactivation of ONSEN in the inserted region; however, the upregulation upon heat-treatment might be the result of ectopic aberrant transcripts that did not affect the functional transcript of ABI5.
Transcriptional state of ABI5 and ABA sensitivity with heat stress.
Relative transcript level of ABI5 without ABA stress was analyzed on the ONSEN inserted region (a) and on the downstream of the insertion (b). Values are relative to ABA-stressed wild-type seedlings. wt; wild-type seedlings, nrpd1; nrpd1 mutant, 13–7; parental 13–7 line, 13–7 BC; F3 progeny that contains an ONSEN insertion in ABI5. (c) ABA-insensitive phenotype of ABI5 in the F3 progeny subjected to heat stress. 13–7: F3 progeny that contains an ONSEN insertion in ABI5. 13–7 ibm2: F3 progeny that contains an ONSEN insertion in ABI5 and a mutation in IBM2. 13–7 nrpd1: F3 progeny that contains an ONSEN insertion in ABI5 and a mutation in NRPD1.
Discussion
Our data have directly demonstrated that a stress-activated transposon can produce ABA-insensitive phenotypes in Arabidopsis. This outcome is supported by previous studies in other species46. In some rice strains, the DNA transposon, mping, rapidly increased in every generation47 and acted as an enhancer that rendered adjacent genes stress-inducible48. In soybean, the insertion of retrotransposon SORE-1 into a paralog of phytochrome A resulted in photoperiod insensitivity, allowing for soybean cultivation at high latitudes, where the growing season is limited49. Although the ABA-insensitive phenotypes generated here suggest stress tolerance, more data is required to verify whether these plants actually experience fitness gains. However, it is possible that ONSEN transpositions could facilitate the occurrence of beneficial mutations and may contribute to adaptive evolution.
We also found evidence that ONSEN activation was controlled by an siRNA-mediated epigenetic regulation: nrpd1 mutants with a deficient RdDM pathway exhibited increased ONSEN heat activation and a transgenerational transposition was observed in these mutants. This finding suggests the involvement of DNA demethylation in ONSEN activation, similar to findings in maize demonstrating that temperature stress results in the selective demethylation of transposons50. However, heat-activated ONSEN transcription is independent of demethylation14,41. We thus propose that siRNA-mediated regulation of ONSEN may occur at levels other than transcription. For example, the epigenetic activation of Athila retrotransposons in Arabidopsis produces siRNA854, which regulates the formation of a stress granule-related protein on post-transcriptional and translational levels51. More research is required before we can fully understand the precise mechanism of siRNA-mediated ONSEN regulation in Arabidopsis.
New ONSEN transpositions occurred in euchromatic regions in nrpd1 mutants. The target site preference of transposons has been reported in other plants48,52,53,54, including Arabidopsis55. The ONSEN target genes have on average a smaller intron compared with the genome-wide distribution. Genes with short introns minimize the cost of transcription and other molecular processes such as splicing56. ONSEN might select a target gene by splicing efficiency. Since the mutations in the RdDM components affect DNA methylation in euchromatin57,58, epigenetic modifications on the ONSEN target site may also be important for integration-site preferences. Unfortunately, the technical difficulties of isolating and analyzing single ONSEN-activated cells prevented us from obtaining more data on site preference mechanisms.
The insertion of ONSEN into ABA-responsive genes suggests a role for the transposon in influencing adaptive stress responses, as ABA plays a pivotal role in the latter59. We demonstrated salt tolerance in the ONSEN inserted lines that indicated the adaptation to environmental stress. Although more research is needed to investigate the actual stress responses of an ONSEN-integrated population, data available in maize indicates that TEs are involved in the regulation of gene responses to abiotic stress60.
Finally, self-fertilization of ABA-insensitive mutants resulted in progenies that inherited ABA-insensitivity (see Supplementary Fig. S5), while crossing mutants with the wild type recovered ABA sensitivity (see Supplementary Fig. S6). We observed an increase in methylation when ABA-insensitive mutants were crossed with ibm2 mutants (see Supplementary Fig. S8). Together, these results suggest that an NRPD1-heterozygous F1 hybrid could establish an epigenetic state that recruits IBM2 on the ONSEN insertion, which could be maintained in the F2 progenies independent of NRPD1, to promote proper splicing of the intron harboring ONSEN (Fig. 7). Consistent with this hypothesis, the loss of heterochromatic epigenetic modifications in intronic TEs affects the transcription of associated genes31, as we observed in ibm2.
A model of ABA response in an ONSEN-integrated mutant.
ONSEN is inserted into an intron of an ABA-responsive gene. (a) An ABA-insensitive mutant is crossed with nrpd1. The ABA-insensitive phenotype appears in an F2 segregant that is homozygous on the ONSEN insertion locus. (b) An ABA-insensitive mutant is crossed with a wild-type plant. IBM2 is recruited on the ONSEN insertion locus by RdDM and ONSEN insertion is masked in the F1 and F2 segregants, resulting in ABA-sensitive phenotypes.
The ONSEN insertion in ABI5 was masked by IBM2 and the reactivation of ONSEN by heat-stress did not affect the functional transcription of ABI5. It is necessary to analyze how many stress-induced TEs acquired an adaptive advantage in the new insertion site and how many were affected by IBM2-mediated regulation of the host genome.
To conclude, the present study investigated whether the environment can directly induce genetic and epigenetic diversity for selection to act upon or whether it is simply a selective force to which existing diversity responds. We found evidence that an environmental stress-induced transposition changes the structure of the host genome and influences gene expression. The generation of mutations by ONSEN could play a role in the adaptive evolution of Arabidopsis. Future studies could expand upon these findings by investigating the biological significance (i.e., adaptive function) of epigenetic variations associated with TEs.
Methods
Plant material and growing conditions
The A. thaliana plants used in the experiments included the wild type, nrpd1 mutants61 and ibm2 mutants30. The seedlings were grown on Murashige and Skoog (MS) plates under continuous light at 21 °C. All wild-type and mutant plants were A. thaliana ecotype Columbia.
SOLiD Sequencing
The sequencing library preparations and high-throughput sequencing were performed according to the manufacturer’s instructions. Total purified genomic DNA samples (1 μg) were processed into pair-endo sequencing libraries using the SOLiD fragment library construction kit (Life Technologies). After quantification by qPCR, libraries were amplified onto beads using emulsion PCR, deposited on slides and sequenced using the SOLiD 4 sequencing system (Life Technologies). Each sample was distinguished by adding a unique “barcode” sequencing adaptor #1–16 using the SOLiD fragment library Barcode Kit (Life Technologies). Fragments were sequenced 50 and 35 bp from each end. The sequencing data from the experiment are available from the NIH short reads archive (accession GSE73097).
ABA treatments
For ABA screening, 7-d-old seedlings were grown on MS plates containing 1 μM ABA. ABA-insensitive phenotypes of the progenies were analyzed on MS plates containing 2 μM ABA. For quantitative real-time polymerase chain reaction (qRT-PCR) and microarray analysis, RNA was extracted from 7-d-old seedlings grown on MS plates containing 2 μM ABA.
Salt treatments
For salt-stress experiments, seedlings were grown on Murashige and Skoog (MS) plates that contained 100 mM NaCl incubated in a near vertical position under continuous light at 21 °C.
Southern blot analysis
Arabidopsis genomic DNA was isolated using a Nucleon PhytoPure DNA extraction kit (GE Healthcare Life Science). Southern blots were performed following previously described protocol62. We detected hybridization signals in a high-SDS hybridization buffer, using a radio-labeled ONSEN-specific probe that was generated with the Megaprime DNA Labeling System (GE Healthcare Life Science)63.
Microarray analysis
Microarrays experiments were performed using an Agilent DNA Microarray Scanner G2539A ver. C and all procedures and data analyses followed manufacturer protocol, except where noted. We used three biological replicates in the experiments. cRNA was labeled using a Low Input Quick Amp Labeling Kit (Agilent) and hybridized to an Agilent custom array platform (Design ID = 034592). cRNA probes were designed based on expression regions of the TAIR10 genome, using previous tiling-array and RNA-Seq analyses64,65,66.
We normalized the signals of microarray probes to the 75th percentile. We then set the following criteria to determine significance in differential gene expression: 1) changes in expression level must either be greater than 1.5-fold or less than 0.67-fold and 2) the results of a Student’s t-test, adjusted with a Benjamini Hochberg FDR67, must yield P < 0.05.
The gene expression heat map was built using the heatmap.2 package in R ver. 2.1.12 (R Core Team). Hierarchical gene clusters were built using complete linkage clustering under Euclidean distance, with Z-scores calculated from log2 normalized values. Heat map coloration was dependent on the Z-scores. Arabidopsis microarray expression profiling data is available in the GEO (GSE71951).
Quantitative real-time PCR (qRT-PCR)
To analyze the gene expression of ABI4 and ABI5, total RNA was extracted from 50 seedlings using TRI Reagent (Sigma T9424), following supplier protocol. Around 3–5 μg of total RNA was treated with RQ1 RNase-free DNase (Promega) and reverse-transcribed using the ReverTraAce qPCR RT Kit (TOYOBO FSQ-101). qRT-PCR was performed using the Applied Biosystems 7300 Real Time PCR System with the THUNDERBIRD SYBR qPCR Mix (TOYOBO QPS-201). Primers used for qRT-PCR are listed in Supplementary Table S6. We performed three biological replicates and determined the standard deviation of the replicates.
Reverse transcription-PCR (RT-PCR)
RNA was isolated from 50 7-d-old seedlings with TRI Reagent (Sigma). We used the OneStep RT-PCR Kit (QIAGEN) with gene-specific primers (see Supplementary Table S6). The thermocycling profile was as follows: 30 min at 50 °C; 15 min at 95 °C; 30 cycles of 94 °C (30 s), 55 °C (30 s) and 72 °C (1 min); and 7 min at 72 °C for ABI5 and 30 min at 50 °C; 15 min at 95 °C; 25 cycles of 94 °C (30 s), 55 °C (30 s) and 72 °C (30 s); and 7 min at 72 °C for 18srRNA. 3′ Rapid Amplification of cDNA Ends was performed using oligo dT primers followed by the first round of PCR using gene-specific primers and the first oligo-dT-specific primer (see Supplementary Table S6). Amplified fragments were diluted and further used for the second round of PCR using another gene-specific primer and a second oligo-dT-specific primer (see Supplementary Table S6).
DNA methylation analysis
For bisulfite sequencing analysis68, 0.25–1 μg of heat-denatured genomic DNA in 20 μl H2O was incubated with 1/9 the volume of 3 M NaOH, for 20 min at 37 °C. Next, 275 μl 10 M bisulfite solution was added to the denatured DNA sample and incubated at 70 °C for 1 h. Bisulfite-treated DNA was purified and desulfonated using an EZ DNA Methylation Kit (Zymo Research), following manufacturer protocol. We used 2 μl DNA as a template in PCR. Primers for the analysis are listed in Supplementary Table S6. Twenty clones were sequenced for each region.
Additional Information
How to cite this article: Ito, H. et al. A Stress-Activated Transposon in Arabidopsis Induces Transgenerational Abscisic Acid Insensitivity. Sci. Rep. 6, 23181; doi: 10.1038/srep23181 (2016).
References
Hoffmann, A. A. & Sgro, C. M. Climate change and evolutionary adaptation. Nature 470, 479–485, 10.1038/nature09670 (2011).
Chevin, L. M., Lande, R. & Mace, G. M. Adaptation, Plasticity and Extinction in a Changing Environment: Towards a Predictive Theory. Plos Biol. 8, 10.1371/journal.pbio.1000357 (2010).
Biemont, C. & Vieira, C. Genetics - Junk DNA as an evolutionary force. Nature 443, 521–524, 10.1038/443521a (2006).
Casacuberta, E. & Gonzalez, J. The impact of transposable elements in environmental adaptation. Mol. Ecol. 22, 1503–1517, 10.1111/mec.12170 (2013).
Pecinka, A. et al. Epigenetic Regulation of Repetitive Elements Is Attenuated by Prolonged Heat Stress in Arabidopsis. Plant Cell 22, 3118–3129, 10.1105/Tpc.110.078493 (2010).
Kazazian, H. H. Mobile elements: Drivers of genome evolution. Science 303, 1626–1632, 10.1126/Science.1089670 (2004).
Kumar, A. & Bennetzen, J. L. Plant retrotransposons. Annu. Rev. Genet. 33, 479–532, 10.1146/annurev.genet.33.1.479 (1999).
Beguiristain, T., Grandbastien, M. A., Puigdomenech, P. & Casacuberta, J. M. Three Tnt1 subfamilies show different stress-associated patterns of expression in tobacco. Consequences for retrotransposon control and evolution in plants. Plant Physiol. 127, 212–221, 10.1104/Pp.127.1.212 (2001).
Takeda, S., Sugimoto, K., Otsuki, H. & Hirochika, H. A 13-bp cis-regulatory element in the LTR promoter of the tobacco retrotransposon Tto1 is involved in responsiveness to tissue culture, wounding, methyl jasmonate and fungal elicitors. Plant J. 18, 383–393, 10.1046/J.1365-313x.1999.00460.X (1999).
Kimura, Y. et al. OARE-1, a Ty1-copia retrotransposon in oat activated by abiotic and biotic stresses. Plant Cell Physiol. 42, 1345–1354, 10.1093/Pcp/Pce171 (2001).
Suoniemi, A., AnamthawatJonsson, K., Arna, T. & Schulman, A. H. Retrotransposon BARE-1 is a major, dispersed component of the barley (Hordeum vulgare L) genome. Plant Mol. Biol. 30, 1321–1329, 10.1007/Bf00019563 (1996).
Woodrow, P., Pontecorvo, G., Ciarmiello, L. F., Fuggi, A. & Carillo, P. Ttd1a promoter is involved in DNA-protein binding by salt and light stresses. Mol. Biol. Rep. 38, 3787–3794, 10.1007/S11033-010-0494-3 (2011).
Salazar, M., Gonzalez, E., Casaretto, J. A., Casacuberta, J. M. & Ruiz-Lara, S. The promoter of the TLC1.1 retrotransposon from Solanum chilense is activated by multiple stress-related signaling molecules. Plant Cell Rep. 26, 1861–1868, 10.1007/S00299-007-0375-Y (2007).
Ito, H. et al. An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress. Nature 472, 115–U151, 10.1038/Nature09861 (2011).
Ito, H., Yoshida, T., Tsukahara, S. & Kawabe, A. Evolution of the ONSEN retrotransposon family activated upon heat stress in Brassicaceae. Gene 518, 256–261, 10.1016/j.gene.2013.01.034 (2013).
Wessler, S. R. Turned on by stress. Plant retrotransposons. Curr. Biol. 6, 959–961, 10.1016/S0960-9822(02)00638-3 (1996).
Gao, Z. et al. An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation. Nature 465, 106–109, 10.1038/nature09025 (2010).
Wierzbicki, A. T., Haag, J. R. & Pikaard, C. S. Noncoding transcription by RNA polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 135, 635–648, 10.1016/j.cell.2008.09.035 (2008).
Onodera, Y. et al. Plant nuclear RNA polymerase IV mediates siRNA and DNA methylation-dependent heterochromatin formation. Cell 120, 613–622, 10.1016/j.cell.2005.02.007 (2005).
Kanno, T. et al. Atypical RNA polymerase subunits required for RNA-directed DNA methylation. Nat. Genet. 37, 761–765, 10.1038/ng1580 (2005).
Pontier, D. et al. Reinforcement of silencing at transposons and highly repeated sequences requires the concerted action of two distinct RNA polymerases IV in Arabidopsis. Genes Dev. 19, 2030–2040, 10.1101/gad.348405 (2005).
Mosher, R. A., Schwach, F., Studholme, D. & Baulcombe, D. C. PolIVb influences RNA-directed DNA methylation independently of its role in siRNA biogenesis. Proc. Natl. Acad. Sci. USA 105, 3145–3150, 10.1073/pnas.0709632105 (2008).
Zhang, X., Henderson, I. R., Lu, C., Green, P. J. & Jacobsen, S. E. Role of RNA polymerase IV in plant small RNA metabolism. Proc. Natl. Acad. Sci. USA 104, 4536–4541, 10.1073/pnas.0611456104 (2007).
Cao, X. & Jacobsen, S. E. Role of the arabidopsis DRM methyltransferases in de novo DNA methylation and gene silencing. Curr. Biol. 12, 1138–1144, 10.1016/S0960-9822(02)00925-9 (2002).
Matzke, M. A. & Birchler, J. A. RNAi-mediated pathways in the nucleus. Nat. Rev. Genet. 6, 24–35, 10.1038/nrg1500 (2005).
Matsunaga, W., Kobayashi, A., Kato, A. & Ito, H. The effects of heat induction and the siRNA biogenesis pathway on the transgenerational transposition of ONSEN, a copia-like retrotransposon in Arabidopsis thaliana. Plant Cell Physiol. 53, 824–833, 10.1093/pcp/pcr179 (2012).
Cutler, S. R., Rodriguez, P. L., Finkelstein, R. R. & Abrams, S. R. Abscisic Acid: Emergence of a Core Signaling Network. Annu. Rev. Plant Biol. 61, 651–679, 10.1146/annurev-arplant-042809-112122 (2010).
Yamaguchi-Shinozaki, K. & Shinozaki, K. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 57, 781–803, 10.1146/annurev.arplant.57.032905.105444 (2006).
Nemhauser, J. L., Hong, F. X. & Chory, J. Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126, 467–475, 10.1016/j.cell.2006.05.050 (2006).
Saze, H. et al. Mechanism for full-length RNA processing of Arabidopsis genes containing intragenic heterochromatin. Nat. Commun. 4, 10.1038/Ncomms3301 (2013).
Le, T. N., Miyazaki, Y., Takuno, S. & Saze, H. Epigenetic regulation of intragenic transposable elements impacts gene transcription in Arabidopsis thaliana. Nucl. Acids Res. 43, 3911–3921, 10.1093/nar/gkv258 (2015).
Popova, O. V., Dinh, H. Q., Aufsatz, W. & Jonak, C. The RdDM Pathway Is Required for Basal Heat Tolerance in Arabidopsis. Mol. Plant 6, 396–410, 10.1093/Mp/Sst023 (2013).
Matsunaga, W. et al. A small RNA mediated regulation of a stress-activated retrotransposon and the tissue specific transposition during the reproductive period in Arabidopsis. Front. Plant Sci. 6, 10.3389/Fpls.2015.00048 (2015).
Brocard, I. M., Lynch, T. J. & Finkelstein, R. R. Regulation and role of the Arabidopsis Abscisic Acid-Insensitive 5 gene in abscisic acid, sugar and stress response. Plant Physiol. 129, 1533–1543, 10.1104/Pp.005793 (2002).
Finkelstein, R. R., Gampala, S. S. L. & Rock, C. D. Abscisic acid signaling in seeds and seedlings. Plant Cell 14, S15–S45, 10.1105/Tpc.010441 (2002).
Bensmihen, S., Giraudat, J. & Parcy, F. Characterization of three homologous basic leucine zipper transcription factors (bZIP) of the ABI5 family during Arabidopsis thaliana embryo maturation. J. Exp. Bot. 56, 597–603, 10.1093/Jxb/Eri050 (2005).
Finkelstein, R. R. & Lynch, T. J. The arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12, 599–609 (2000).
Finkelstein, R. R. Mutations at 2 New Arabidopsis Aba Response Loci Are Similar to the Abi3 Mutations. Plant J. 5, 765–771, 10.1046/J.1365-313x.1994.5060765.X (1994).
Finkelstein, R. R., Wang, M. L., Lynch, T. J., Rao, S. & Goodman, H. M. The Arabidopsis abscisic acid response locus ABI4 encodes an APETALA2 domain protein. Plant Cell 10, 1043–1054, 10.1105/Tpc.12.4.599 (1998).
Soderman, E. M., Brocard, I. M., Lynch, T. J. & Finkelstein, R. R. Regulation and function of the arabidopsis ABA-insensitive4 gene in seed and abscisic acid response signaling networks. Plant Physiol. 124, 1752–1765, 10.1104/Pp.124.4.1752 (2000).
Cavrak, V. V. et al. How a Retrotransposon Exploits the Plant’s Heat Stress Response for Its Activation. Plos Genet. 10, 10.1371/Journal.Pgen.1004115 (2014).
Lopez-Molina, L., Mongrand, B., McLachlin, D. T., Chait, B. T. & Chua, N. H. ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J. 32, 317–328, 10.1046/J.1365-313x.2002.01430.X (2002).
Reeves, W. M., Lynch, T. J., Mobin, R. & Finkelstein, R. R. Direct targets of the transcription factors ABA-Insensitive(ABI)4 and ABI5 reveal synergistic action by ABI4 and several bZIP ABA response factors. Plant Mol. Biol. 75, 347–363, 10.1007/S11103-011-9733-9 (2011).
Tezuka, K., Taji, T., Hayashi, T. & Sakata, Y. A novel abi5 allele reveals the importance of the conserved Ala in the C3 domain for regulation of downstream genes and salt tolerance during germination in Arabidopsis. Plant Signal Behav. 8, e23455, 10.4161/psb.23455 (2013).
Shkolnik-Inbar, D., Adler, G. & Bar-Zvi, D. ABI4 downregulates expression of the sodium transporter HKT1;1 in Arabidopsis roots and affects salt tolerance. Plant J. 73, 993–1005, 10.1111/tpj.12091 (2013).
Quesada, V., Ponce, M. R. & Micol, J. L. Genetic analysis of salt-tolerant mutants in Arabidopsis thaliana. Genetics 154, 421–436 (2000).
Naito, K. et al. Dramatic amplification of a rice transposable element during recent domestication. Proc. Natl. Acad. Sci. USA 103, 17620–17625, 10.1073/Pnas.0605421103 (2006).
Naito, K. et al. Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461, 1130–U1232, 10.1038/Nature08479 (2009).
Kanazawa, A., Liu, B. H., Kong, F. J., Arase, S. & Abe, J. Adaptive Evolution Involving Gene Duplication and Insertion of a Novel Ty1/copia-Like Retrotransposon in Soybean. J. Mol. Evol. 69, 164–175, 10.1007/S00239-009-9262-1 (2009).
Steward, N., Kusano, T. & Sano, H. Expression of ZmMET1, a gene encoding a DNA methyltransferase from maize, is associated not only with DNA replication in actively proliferating cells, but also with altered DNA methylation status in cold-stressed quiescent cells. Nucleic Acids Res. 28, 3250–3259 (2000).
McCue, A. D., Nuthikattu, S., Reeder, S. H. & Slotkin, R. K. Gene Expression and Stress Response Mediated by the Epigenetic Regulation of a Transposable Element Small RNA. Plos Genet. 8, 10.1371/Journal.Pgen.1002474 (2012).
SanMiguel, P. et al. Nested retrotransposons in the intergenic regions of the maize genome. Science 274, 765–768, 10.1126/Science.274.5288.765 (1996).
Tsukahara, S. et al. Centromere-targeted de novo integrations of an LTR retrotransposon of Arabidopsis lyrata. Genes Dev. 26, 705–713, 10.1101/gad.183871.111 (2012).
Cresse, A. D., Hulbert, S. H., Brown, W. E., Lucas, J. R. & Bennetzen, J. L. Mu1-Related Transposable Elements of Maize Preferentially Insert into Low Copy Number DNA. Genetics 140, 315–324 (1995).
Peterson-Burch, B. D., Nettleton, D. & Voytas, D. F. Genomic neighborhoods for Arabidopsis retrotransposons: a role for targeted integration in the distribution of the Metaviridae. Genome Biol. 5, 10.1186/Gb-2004-5-10-R78 (2004).
Castillo-Davis, C. I., Mekhedov, S. L., Hartl, D. L., Koonin, E. V. & Kondrashov, F. A. Selection for short introns in highly expressed genes. Nat. Genet. 31, 415–418, 10.1038/ng940 (2002).
Huettel, B. et al. Endogenous targets of RNA-directed DNA methylation and Pol IV in Arabidopsis. EMBO J. 25, 2828–2836, 10.1038/Sj.Emboj.7601150 (2006).
Zemach, A. et al. The Arabidopsis Nucleosome Remodeler DDM1 Allows DNA Methyltransferases to Access H1-Containing Heterochromatin. Cell 153, 193–205, 10.1016/J.Cell.2013.02.033 (2013).
Fujita, Y., Fujita, M., Shinozaki, K. & Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 124, 509–525, 10.1007/S10265-011-0412-3 (2011).
Makarevitch, I. et al. Transposable elements contribute to activation of maize genes in response to abiotic stress. PLoS Genet. 11, e1004915, 10.1371/journal.pgen.1004915 (2015).
Herr, A. J., Jensen, M. B., Dalmay, T. & Baulcombe, D. C. RNA polymerase IV directs silencing of endogenous DNA. Science 308, 118–120, 10.1126/science.1106910 (2005).
Miura, A. et al. Genomic localization of endogenous mobile CACTA family transposons in natural variants of Arabidopsis thaliana. Mol. Genet. Genomics 270, 524–532, 10.1007/s00438-003-0943-y (2004).
Church, G. M. & Gilbert, W. Genomic sequencing. Proc. Natl. Acad. Sci. USA 81, 1991–1995 (1984).
Kawaguchi, S. et al. Positional correlation analysis improves reconstruction of full-length transcripts and alternative isoforms from noisy array signals or short reads. Bioinformatics 28, 929–937, 10.1093/Bioinformatics/Bts065 (2012).
Matsui, A. et al. Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array. Plant Cell Physiol. 49, 1135–1149, 10.1093/Pcp/Pcn101 (2008).
Okamoto, M. et al. Genome-wide analysis of endogenous abscisic acid-mediated transcription in dry and imbibed seeds of Arabidopsis using tiling arrays. Plant J. 62, 39–51, 10.1111/J.1365-313x.2010.04135.X (2010).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate - a Practical and Powerful Approach to Multiple Testing. J. Roy. Stat. Soc. B. Met. 57, 289–300 (1995).
Shiraishi, M. & Hayatsu, H. High-speed conversion of cytosine to uracil in bisulfite genomic Sequencing analysis of DNA methylation. DNA Res. 11, 409–415, 10.1093/Dnares/11.6.409 (2004).
Acknowledgements
This work was supported by a grant from Cooperative Research Grant of the Genome Research for BioResource, NODAI Genome Research Center, Tokyo University of Agriculture, the MEXT as part of Joint Research Program implemented at the Institute of Plant Science and Resources, Okayama University, the National Institute of Genetics Cooperative Research Program (2014-A), JST-PRESTO, Grants-in-Aid for JSPS Fellows (14J02452) and Grants-in-Aid for Scientific Research in Innovative Areas (2511970103). This project was also supported partly by grants from RIKEN and the Japan Science and Technology Agency (JST), Core Research for Evolutionary Science and Technology (CREST) and by the Japan Advanced Plant Science Research Network to M.S.
Author information
Authors and Affiliations
Contributions
H.I., J.M.K. and M.S. planned and designed the research. Y.H., M.T., J.I. and T.M. performed next generation sequencing. T.A.E., H.T., Y.H., S.T. and T.T. performed bioinformatics analyses. A.M. performed expression profiling. H.S. and Y.H. performed whole genome bisulfite sequence. W.M. and S.M. perform quantitative RT-PCR. H.S. performed 3′ Rapid Amplification of cDNA Ends. Y.M. performed Southern blot analysis. T.K. and A.K. initiated and guided the project. H.I., J.M.K., H.S. and A.M. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ito, H., Kim, JM., Matsunaga, W. et al. A Stress-Activated Transposon in Arabidopsis Induces Transgenerational Abscisic Acid Insensitivity. Sci Rep 6, 23181 (2016). https://doi.org/10.1038/srep23181
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23181
This article is cited by
-
Investigation of Tos17 LTR retrotransposon movements in rice (Oryza sativa L.) under nickel and boron stress
Cereal Research Communications (2024)
-
Ex vitro Morpho-Physiological Screening of Drought Tolerant Sugarcane Epimutants Generated Via 5-Azacytidine and Imidacloprid Treatments
Tropical Plant Biology (2022)
-
Identification and characterization of genes for drought tolerance in upland rice cultivar ‘Banglami’ of North East India
Molecular Biology Reports (2022)
-
Long terminal repeats (LTR) and transcription factors regulate PHRE1 and PHRE2 activity in Moso bamboo under heat stress
BMC Plant Biology (2021)
-
Robustification of GWAS to explore effective SNPs addressing the challenges of hidden population stratification and polygenic effects
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.