Abstract
To determine whether a C. elegans bioassay could predict mammalian developmental activity, we selected diverse compounds known and known not to elicit such activity and measured their effect on C. elegans egg viability. 89% of compounds that reduced C. elegans egg viability also had mammalian developmental activity. Conversely only 25% of compounds found not to reduce egg viability in C. elegans were also inactive in mammals. We conclude that the C. elegans egg viability assay is an accurate positive predictor, but an inaccurate negative predictor, of mammalian developmental activity. We then evaluated C. elegans as a tool to identify mechanisms affecting toxicological outcomes among related compounds. The difference in developmental activity of structurally related fungicides in C. elegans correlated with their rate of metabolism. Knockdown of the cytochrome P450s cyp-35A3 and cyp-35A4 increased the toxicity to C. elegans of the least developmentally active compounds to the level of the most developmentally active. This indicated that these P450s were involved in the greater rate of metabolism of the less toxic of these compounds. We conclude that C. elegans based approaches can predict mammalian developmental activity and can yield plausible hypotheses for factors affecting the biological potency of compounds in mammals.
Similar content being viewed by others
Introduction
Ensuring the safety to humans of the chemicals they may be exposed to is of critical importance to chemical companies, regulatory authorities and the public. It is in the interest of chemical companies researching new active ingredients (AI) to identify adverse toxicological outcomes as soon as possible and avoid wasted investment in unsafe or unregisterable chemical products. One approach is to test new AI earlier in research programmes using the standard, guideline, mammalian toxicological tests required by regulators to determine toxicological outcomes. However, this implies a substantial increase in the number of mammals used which is undesirable for ethical and economic reasons. Therefore much research has investigated alternative experimental systems that have fewer of these concerns including in silico modelling1, cell-based systems2, vertebrate systems of reduced concern e.g. Zebrafish3 as well as invertebrate model systems4,5. Developmental toxicity, where a chemical adversely affects the biological processes of development from egg to adult, is of concern to the agrochemical industry. Developmental biology has been extensively studied in invertebrate model systems making them obvious candidates for the study of developmental toxicity.
The nematode Caenorhabditis elegans is exceptionally well studied and many researchers have used it as a model for different forms of toxicity5,6,7. Its small size, short life cycle and ease of maintenance and culturing make it a viable model for high-throughput screening, while the array of genetic tools available for use with it enable further investigations into the causes of toxicity. Several studies have used C. elegans as a model for: the neurotoxicity of xenobiotics8, neurodegeneration9, genotoxicity10, and germline toxicity5 and all found relevance of the model to man11. For example, the toxicity of a group of organophosphates was shown to correlate between C. elegans and mammals12. C. elegans has also been used to investigate the basis of the toxicity of ethanol13, volatile anaesthetics14 and other drugs15,16,17.
C. elegans development is fully described18,19 and the underlying genetic mechanisms controlling development are well understood and often conserved with those found in mammals20. Embryogenesis, from fertilization to egg hatching takes approximately 13–14 h at 20 °C and produces 671 cells (113 of which die by apoptosis) forming the L1 C. elegans larva19. Numerous developmental processes occur, leading to cell fate specification, tissue formation and morphogenesis. Scoring egg viability by counting the number of eggs that hatch as a proportion of those laid is therefore a convenient and quantitative measure of the success of these developmental processes. Chemicals or other exogenous factors affecting developmental biology are likely to affect egg viability.
Chemical toxicity results from a xenobiotic molecule adversely affecting a process or function upon which a toxicological outcome is contingent. Typically this will be the consequence of a biochemical interaction between the small molecule and an endogenous protein or proteins involved in this process or function. Therefore a major determinant of toxicology is the potency of this biochemical interaction. But this is not the only determinant, also important is the distribution and abundance of the small molecule in the organism and its consequent availability with respect to the target protein(s) driving the toxicological outcomes. The importance of absorption, distribution, metabolism and excretion on the interaction between small molecules and organisms has long been recognised21. Of these, metabolism has been extensively researched, not least because of its importance to the efficacy of pharmaceuticals. An important class of metabolising enzyme is the Cytochrome P450 enzymes which are a superfamily of NADPH-dependent monooxygenases that catalyse the Phase I metabolism of xenobiotics such as pesticides22. The C. elegans genome contains 77 intact cytochrome P450 genes23. Differences in toxicity among compounds could be caused by differences in affinity for a single P450 that metabolises both compounds, metabolism of compounds by more than one P450 or changes in gene expression of P450 gene(s) responsible for metabolism of one or both compounds.
In this study we assessed the utility of C. elegans for toxicological investigations, and in particular for generating hypotheses relevant to human safety. We did this in two ways. First, we screened diverse pesticide chemistry, including compounds with mammalian developmental activity in the ToxRef database24, and measured their developmental activity in C. elegans. This allowed us to estimate the correlation of chemically-induced developmental activity between nematodes and mammals and therefore the predictive power of one system on the other. Definitions of toxicity (including developmental toxicity), which are considered in the registration and labelling of commercial products, vary among jurisdictions and change over time. Even fundamental concepts such as the relative importance of ‘risk’ and ‘hazard’ are debated in this context25 and can cause controversy26. This makes it hard to identify a suitable set of universally accepted, developmentally toxic standards which is needed to evaluate predictive approaches such as the one we describe. To overcome this, we chose to work on compounds that were reported simply to have more potent biological activity in developing, mammalian embryos than in adults, from the ToxRef database24. While such compounds cannot be considered developmentally toxic on the basis of these data alone, they have activity on the developing, early life stages of mammals, which could result in developmental toxicity, We reasoned that a tool that predicted developmental activity and so the possibility of developmental toxicity, could be useful in prioritizing and directing subsequent toxicological investigation. Whether such a compound was ultimately classified as developmentally toxic would depend on these subsequent studies and on the regulatory definitions of toxicity in relevant jurisdictions. In the second part of our study, we focussed on a closely related series of proprietary pyridazine and imidazole fungicides recently dropped from Syngenta’s research portfolio because developmental toxicity was observed with some examples of the series. We asked whether C. elegans could identify factors underlying the toxicology of the series and suggest approaches to, in principle, redesign molecules with improved toxicological profiles.
Results
The egg viability assay in C. elegans as a screen for developmental toxicity in mammals
We assessed the utility of C. elegans as a system used to screen for compounds with developmental activity. We selected 72 pesticide compounds from the ToxRef database as a test set24,27. This database contains summarised results from studies submitted to the US Environmental Protection Agency as part of the registration of pesticides. The initial build contained 1318 records of prenatal developmental studies, conducted on rats or rabbits. Endpoints recorded included maternal effects such as body weight gain, food and water consumption, fertility and pregnancy, as well as foetal effects such as foetal weight reduction, skeletal variations, malformations and other pathologies24. The database provides a lowest effect level (LEL) for both maternal toxicity and developmental effects (a measure of toxicological potency to the embryo) in mammals.
If a compound had a lowest effect level for developmental effects at a lower concentration than for adult toxicity it was considered to be developmentally active because it had effects on embryo development in the absence of effects on the mother. We considered effects occurring at doses when the mother was sick might be indirect maternal effects rather than developmental effects per se. Using these criteria 57 of the selected compounds were developmentally active; the remaining fifteen compounds were negative controls. A full list of compounds is provided as supplementary material (Supplementary Table 1). The compounds represent diverse structures and mechanisms of action including insecticides, fungicides and herbicides.
These compounds were then tested to determine if they affected C. elegans egg viability. L4 C. elegans were exposed to compounds for 48 h and allowed to lay eggs. The adults were then removed and the number of unhatched eggs still remaining after 24 h recorded. If the number of unhatched eggs significantly exceeded control levels the compound was considered to reduce egg viability (and therefore to be developmentally active) in C. elegans. Control levels were a mean number of unhatched eggs of 1.41 per well with a standard deviation of 1.72. Differences from control were measured by t-test (p < 0.001).
Nineteen compounds reduced egg viability in C. elegans, of which seventeen were from the group defined as developmentally active in mammals (Table 1). The positive predictivity of this assay is therefore 89%. In other words 89% of compounds found to be developmentally toxic in our assay in C. elegans are also developmentally active in mammals. Analysis of the complete ToxRef database shows the percentage of compounds found to be developmentally active in mammals ~18% 24. Based on this finding, our assay improves this prediction markedly. If a compound is active in our assay it is ~5 fold (89% versus 18%) more likely to be developmentally active in mammals compared to this “baseline” expectation.
However only 25% of compounds found to not to affect egg viability in C. elegans are also not developmentally active in mammals. The negative predictivity of the assay is therefore low, relative to the positive predictivity. Based on these data, we conclude that a positive result in the assay is likely to accurately predict mammalian developmental activity, while a negative result in this assay only weakly predicts that a compound will not be developmentally active in mammals. Therefore the majority of mammalian developmentally active compounds will be inactive in this assay; however a compound that is active in the assay is likely to be developmentally active in mammals.
The egg viability assay in C. elegans as a tool for investigating differential toxicity across a related series of compounds
Commercial synthetic chemistry research typically produces a series of analogue compounds all structurally related to an initial lead compound. The purpose of this synthetic effort is to understand the impact of structural modifications on the properties of the chemical. This can then enable the rational design of molecules with desirable properties, which may include an improved toxicological profile. Therefore having demonstrated the ability of a C. elegans egg viability assay to identify mammalian developmentally active compounds within a diverse chemical collection, we now looked at the utility of this assay within a series of closely related compounds. The compounds we chose are a series of proprietary pyridazine and imidazole fungicides which disrupt microtubule dynamics28,29.
Seventeen of these compounds were tested in the egg viability assay (Fig. 1). These compounds were selected by Syngenta as having commercial potential and therefore suitable for further research. Compounds 2, 3, 10 and 15 have been shown to cause teratogenicity in rats. Compounds 5 and 6 have not shown clear developmental toxicity in the same preliminary tests (Supplementary Table 2). There is no mammalian data for the remaining compounds. All the compounds were biologically active in C. elegans. We considered that the mechanism of action in C. elegans was likely to be related to that in fungi i.e. the disruption of microtubule function. Fungicidal chemicals acting in this way have previously been shown to also be active on C. elegans30.
We found that, as in mammals, examples from the chemical series induced developmental toxicity (measured as egg viability) in C. elegans though the exact pattern of toxicity for different analogues varied between species (Fig. 2). In C. elegans, most showed a similar (within one order of magnitude) No Effect Level (NOEL) for both maternal and developmental toxicity. However compounds 6 and 7 showed no developmental toxicity but were maternally lethal. Conversely compounds 8, 13, 16 & 17 showed no maternal lethality but were developmentally toxic. Additionally compound 1 had a developmental NOEL two orders of magnitude higher than its maternal NOEL. This variation in induced developmental toxicity presented an opportunity to investigate the mechanism(s) underlying the developmental toxicity of this series of compounds in C. elegans.
The maternal activity and developmental activity in C. elegans of a series of compounds 1-17 (See Fig. 1) tested in the egg viability assay. The blue columns show the No Effect Level (NOEL) for developmental activity i.e. the highest concentration at which no significant (p < 0.001) effect on egg viability was observed. The red columns show the NOEL for maternal activity: the highest concentration at which no adult toxicity was observed. Where no column is present no significant activity was observed at any dose.
Differences in metabolic stability in C. elegans between related compounds
We selected six compounds for further study based on their differing toxicological effects on C. elegans. These fell into three groups. Group A comprised compound 3 and compound 4 which are pyridazine compounds that showed high levels of both maternal toxicity and egg toxicity. Group B comprised compound 6 and compound 7 which are imidazole compounds that showed low egg toxicity and high/medium maternal toxicity. Group C comprised compound 16 and compound 17 which are also pyridazine compounds and these showed low maternal toxicity and medium egg toxicity.
A possible cause of the toxicological differences between compounds could result from variations in bioavailability, e.g. differential metabolism, affecting the exposure of either the egg or adult to the compound. To address this, we measured the rate of metabolism of one compound from each of the three groups (Groups A to C, Fig. 1 compounds 3, 6, 17, respectively) by investigating its metabolic stability in nematodes over a 24 h period (Fig. 3). The data are expressed as a percentage of recovered compound at time 24 h vs. time 0 h. The C. elegans metabolism assay and LC-MS analysis of extracts is described in detail in the Methods section. Overall, compound 6 had the greatest loss in 24 h; compound 3 had the least. These were significantly different (p = 0.013) levels of metabolism. Compound 17 showed an intermediate level of metabolism, closest to compound 6. So the compound (6) showing the highest developmental toxicity NOEL (i.e. the least developmentally toxic) is the least metabolically stable and the compound (3) with the lowest developmental toxicity NOEL (the most developmentally toxic) shows the lowest rate of metabolism after 24 h. Compound 17 is intermediate for both measures. We conclude that reduced developmental toxicity among the test compounds is associated with increased rate of loss of the compound through metabolism. We then investigated the mechanism of increased rate of metabolism.
Data obtained by LC-MS analysis of samples containing compounds 3, 6, and 17. Compound 6 is metabolised to a greater extent in 24 h than Compound 3. The data are expressed as a percentage of recovered compound at time 24 h vs. time 0 h. The columns show the average ± s.e. of n = 3. *indicates p < 0.05 from a t-test.
The cytochrome P450 genes cyp-35A2-5 and cyp-35C1 are upregulated by compound 6 and compound 17 but not compound 3
Xenobiotics are known to induce the expression of metabolic enzymes31,32. Therefore the differential metabolism we observed among analogues might result either from their intrinsic susceptibility to metabolism or from their ability to induce metabolic gene expression. We performed a microarray to establish which genes were altered in expression in response to a 48 h exposure to the six, selected compounds.
We find that the cytochrome P450s cyp-35A2-5 and cyp-35C1 are upregulated by some, but not all of our test compounds (Fig. 4, Supplementary Table 3). They were upregulated to the greatest extent in compound 7 and thereafter in the order compound 6> compound 17> compound 16, except that only cyp-35A3 and cyp-35C1 showed significant upregulation in response to compound 16 and cyp-35A3 showed slightly greater upregulation in compound 17 than compound 6. None of these genes were upregulated at all in compound 3 and compound 4. This mirrors the C. elegans developmental toxicity data for these compounds. Therefore, we asked whether these metabolic genes were involved in the faster metabolism of the less developmentally toxic of these compounds.
(a) Fold upregulation of cytochrome P450 genes in response to three compounds. Only genes showing detectable expression are shown (b) Fold upregulation of cyp-35A3, cyp-35A4, cyp-35A5 and cyp-35C1 in all six compounds. Two oligonucleotides in the microarray targeted the cyp-35C1 gene, both showed a similar pattern of induction. See Supplementary Table 3 for full fold change data of all differently expressed cytochrome P450 genes.
RNAi knockdown of cyp-35A3 and cyp-35A4 together causes C. elegans eggs to fail to hatch after exposure to compound 6 or compound 7
We wanted to determine whether cytochrome P450s were involved in the observed differences in developmental toxicity of these compounds and if so which ones. We targeted 58 cytochrome P450 encoding genes with RNAi and determined the effect of knockdown on the developmental toxicity of compounds 6 and 7 (Table 2). To keep the experiment manageable we knocked down the expression of genes in groups of up to 3 (as previously described33), and scored the subsequent effects on egg hatching following exposure to the test compounds (see Methods). RNAi knockdown of most of the cytochrome P450s tested had no effect on the developmental toxicity caused by either compound. Simultaneous knock down of cyp-35A2, cyp-35A5 and cyp-35C1 resulted in a small, non-significant (6, 0.5 μg/ml, p = 0.114, 7, 50 μg/ml p = 0.203) increase in developmental toxicity which we did not investigate further. Only simultaneous knockdown of cyp-35A3 and cyp-35A4 showed a significant increase in developmental toxicity (6, 0.5 μg/ml, p = 0.004, 7, 50 μg/ml p = 2.53 × 10−6). 79% of eggs exposed to 50 μg/ml of compound 7 and 89% of eggs on 0.5 μg/ml of compound 6 did not hatch. This was compared to 9% and 6% respectively in controls in which the compound was present in the absence of the RNAi treatment. In controls in which the RNAi treatment was present in the absence of the compound the rate was 1%.
Separate knockdown of each gene individually also resulted in reduced developmental toxicity suggesting that either it is influenced by both enzymes or that RNAi targeted to one cytochrome P450 has effects on another (Fig. 5). Interestingly, we also included compound 4 in these experiments and found no evidence that these enzymes affected the toxicological effects of this compound (Supplementary Fig. 1). We conclude that the expression of cyp-35A3 and/or cyp-35A4 is a major determinant of the developmental toxicity of compounds 6 and 7 in C. elegans.
We used 0.5 μg/ml and 50 μg/ml doses of compounds 6 & 7 respectively. These doses induce mild maternal toxicity in C. elegans (N2) See Fig. 2. C. elegans developmental toxicity is greatly increased in (cyp-35A3 RNAi and cyp-35A3 RNAi together) compared to N2. C. elegans developmental toxicity is also increased in response to cyp-35A3 RNAi and cyp-35A4 RNAi separately but not to as great an extent. The columns show mean ± s.e. of n = 4.
Discussion
We evaluated a C. elegans based approach for toxicological research. We first asked whether C. elegans could be used as an alert for the potential of a research compound to cause developmental effects in mammals. Second, we asked whether C. elegans could be used to reveal mechanisms driving the toxicological effects of compounds, mechanisms that, if understood, might enable mitigation of the effects.
For the first component, we find that the positive predictive power of the C. elegans egg viability assay we employed is surprisingly high: 89% of compounds found to be developmentally active in C. elegans by this measure are also developmentally active in mammals (as noted previously whether a compound would be classified as developmentally toxic would depend on subsequent experiments and regulatory oversight). A strength of our assay is that it includes both the mother (a C. elegans hermaphrodite) and the developing embryo. Once laid the eggshell will likely limit chemical ingress to the embryo34, the assay therefore models embryonic exposure via maternal exposure, as in mammalian tests. Furthermore, the assay records the toxicity to the egg relative to the toxicity to the mother, we suspect this relative toxicity measure controls for the effects of scale that could otherwise confound correlations between C. elegans and mammalian effects.
However the negative predictivity of the assay was low: 25% of compounds found not to be toxic to C. elegans were also not developmentally active in mammals. We record only one endpoint, egg viability and it is possible that other C. elegans assays, looking for additional developmental perturbations would identify compounds missed by the assay reported here; such perturbations could include those observed as developmental phenotypes by geneticists35. However accuracy will always be limited by intrinsic differences between mammals and nematodes, while C. elegans shares many developmental processes with mammals, it does not share them all. Processes associated with the formation of structures not present in C. elegans, the skeleton for example, can be only incompletely represented in C. elegans at best and this may underly the weak negative predictivity we observed.
An example of a chemical research project dropped due to adverse toxicological outcomes is the one that produced the pyridazine and imidazole fungicides examined in the second component of our study. Here we investigated the potential for C. elegans to provide mechanistic insights into toxicology that could, in principle, be exploited to design less toxic compounds. We show that the expression of the genes cyp-35A3 and/or cyp-35A4 are required for the reduced toxicity of compounds 6 and 7 while having no impact on the biological activity of other compounds from the chemical series. Biological differences between close chemical analogues are hard to predict and are valuable in revealing subtle effects of structure on biological activity within closely related compounds.
Several studies have investigated the upregulation of P450 enzymes in C. elegans in response to various xenobiotic compounds36,37,38,39,40. The CYP-35 genes in particular have been shown to be strongly inducible38. Fewer studies have tried to identify the enzyme that metabolises a given xenobiotic. In one example, the enzymes cyp-14A and cyp-34A6 were identified as the major contributors to the metabolism of PCB52 in C. elegans by directly measuring the formation of hydroxylated metabolites whose production required the expression of these enzymes33.
Genetic interactions between cytochrome P450 encoding genes and xenobiotic compounds such as those we and others have observed, may arise for different reasons. Firstly compounds may act directly on cytochrome P450s to deliver their toxicological outcome i.e. they may themselves be the target of the compound. Several molecules are known to inhibit cytochrome P450s, including piperonyl butoxide (PBO)41. Second, cytochrome P450s may act to metabolise the compound to a more or less biologically active metabolite and so modulate toxicological outcomes of the original compound. For example the toxicity of the organophosphate fenitrothion and its actions on its target, acetylcholine esterase, was shown to be reduced by knockout of cyp-35A2. This was taken to indicate that this P450 was involved in its biotransformation to the active form42. Thirdly, the effect may be indirect: cytochrome P450s have endogenous functions including the metabolism of fat into which lipophilic compounds may partition and so be sequestered away from their target proteins. Under these circumstances changes in fat metabolism might therefore indirectly affect toxicity by altering the sequestration of toxic compounds. Knockout mutants of genes of the cyp-35A subfamily have been shown to have reduced fat storage43,44, which has been implicated in their role in the toxicity of PCB5233. However they have also been shown to be involved in xenobiotic metabolism42.
We cannot formally distinguish these possibilities on the basis of our experiments. That said our knowledge of the mechanism of action of this compound series does not suggest they are toxic because of direct effects on cytochrome P450 enzymes (they are microtubule disruptors). Furthermore we find no correlation between the logP (a measure of lipophilicity) of the compounds and toxicological potency which does not suggest that partition into fat stores explains the toxicological differences among these compounds in C. elegans (Fig. 6). Rather, we suggest that cyp-35A3 and/or cyp-35A4 are enzymes involved in the metabolism of compounds 6 and 7, and that the reduced toxicity of these compounds compared to the closely related compounds 3 and 4 is due to their reduced bioavailability as a result of their greater rate of metabolism by these enzymes.
The cytochrome P450s cyp-35A2-5 and cyp-35C1 were upregulated in response to compound 6 and compound 7 which showed the lowest levels of developmental toxicity in C. elegans. This upregulation was clearest in cyp-35A3, cyp-35A4 and cyp-35C1. It is likely that the differential bioavailability of these compounds might be due to their upregulation of cyp-35A3 and/ or cyp-35A4 which therefore metabolises them faster. However cyp-35C1 which is massively upregulated in response to these compounds does not appear to play a role in their toxicity. This reflects what was found by Schäfer et al. in their study of the metabolism of PCB52. They found that the enzymes cyp-14A and cyp-34A6 metabolised PCB5233. In earlier studies PCB52 had induced the expression of many different P450s including cyp-14A3, cyp-34A10, cyp-35A and cyp-35C136,37,38 but no induction of expression of cyp-34A6 has been reported. Therefore in this case the induction of cytochrome P450 genes including cyp-35C1 was not indicative of them being involved in the metabolism of the compound. However cyp-14A3 was both induced by, and involved in the metabolism of PCB52. In addition cyp-35A2 has been shown to be both induced by, and involved in the toxicity of, the compound fenitrothion42.
In mammalian systems, where more is known about the responses of cytochrome P450 to xenobiotics, there is no automatic assumption that an inducer or inhibitor of a P450 will be metabolised by it. This is certainly sometimes the case, for instance chronic exposure to ethanol will upregulate CYP2E1 which is the enzyme that metabolises ethanol45. However several P450 inducers or inhibitors such as paroxetine induce/inhibit more P450s than just the one they are metabolised by46. Together with the observations described here, this underlines the importance of combining genetics with analytical chemistry, and with measures of biological activity in the whole organism, to be sure of the functional contributions of metabolic enzymes.
The industry-wide impact of unintended toxicological outcomes during agrochemical research and development has not been calculated, but is certainly significant. In the pharmaceutical industry, non-clinical toxicology (which includes adverse findings in animal tests) is estimated as the most frequent (40%) cause of attrition in the drug development pipeline47. Therefore, a substantial improvement in pipeline efficiency would be achieved if compounds likely to fail through non-clinical toxicology were identified earlier and either dropped or redesigned. One way to achieve this would be to perform toxicity testing earlier in the pipeline, but, using conventional approaches, this would inevitably lead to increased animal testing and is therefore unacceptable. Only the use of predictive tools, such as those we describe here, offer practical means to achieve earlier assessment of toxicology. Many groups are currently investigating this possibility using various approaches such as in silico modelling1 and cell-based systems2, as well as using other model organisms such as the slime mould Dictyostelium discoideum and the zebrafish Danio rerio3,48.
Once the risk of adverse toxicological outcomes in mammals has been identified, hypotheses on the factors determining these outcomes may ultimately help to avert the risk through chemical design. We show that metabolism and the actions of particular cytochrome P450 enzymes are determinants of C. elegans developmental toxicity which is itself predictive for mammalian developmental activity. An appeal of performing such studies in whole organisms is that mechanisms can be directly linked to toxicological endpoins in the study organism: we show that the function of P450s in C. elegans is associated with egg viability when exposed to particular compounds. Whether metabolism by orthologous enzymes affects the developmental toxicity caused by these compounds in mammals, is beyond the scope of this study and is therefore not known. But, more generally, an effect of cytochrome P450 mediated metabolism on the biological potency of compounds in mammals has been frequently observed49 and we suggest that it is at least plausible that our findings in C. elegans would be relevant to mammals. If so, then designing chemical analogues with the metabolic properties of compounds 6 and 7 would reduce the risk of mammalian developmental toxicity.
In summary, we propose that the egg viability assay in C. elegans we describe can be a valuable component of predictive approaches for mammalian developmental toxicity and that C. elegans can be used to develop mechanistic hypotheses about effects, including toxicological endpoints, relevant to mammals.
Methods
Egg viability assay
We have defined developmental toxicity in C. elegans as a reduction in egg viability. The egg viability assay was performed in 24 well plates containing 0.5 ml NGM agar per plate and seeded with 25 μl E. coli OP50. AI was added to the plates in 30 μl of solvent (10%DMSO, 50%IPA and 40% H2O) per well. Initial tests were conducted at final concentrations of 500, 50, 5 and 0 μg/ml (this last was the vehicle control). However if a compound was inactive at all concentrations it was repeated at 1000 μg/ml and if it was lethal at all concentrations it was repeated at lower concentrations (by ten-fold dilution) until no effect was seen.
Five L4 worms per well were added to the plate and left for 48 h at 20 °C. At this point they were scored for adult toxicity (alive/sick/dead) and removed. The eggs that had been laid were left for 24 h at 20 °C to hatch. The number of eggs per well that remained unhatched was recorded.
By coincidence the mean number of unhatched eggs found in control (solvent only) wells was 1.41, with a standard deviation of 1.72, in both the egg viability assay on the ToxRef compounds (n = 124) and the egg viability assay on the fungicide compounds (n = 52) separately. Compounds were considered to be developmentally active if the mean number of unhatched eggs per well found in response to a given dose of the compound was significantly greater than control (measured by two-tailed Student’s t-test, p < 0.001) (n = 2−6).
For the egg viability assay on the fungicide compounds the no effect levels (NOEL) of the compounds were calculated. The no effect level for developmental toxicity was the highest concentration tested at which the mean number of unhatched eggs was not significantly greater than control (measured by t-test, p < 0.001). The no effect level for maternal toxicity was the highest concentration tested at which the adults appeared indistinguishable from controls. Initial tests were performed on a wide range of doses and then repeated at relevant doses close to the NOEL. Therefore while over the whole dose range n = 2−6 for the doses closest to the NOEL n = 4−6.
The egg viability assay was altered for the RNAi screen to allow the extent of toxicity at a single dose under different conditions to be compared. Two concentrations of each compound were used. These were; compound 6 0.5 and 0.05 μg/ml, and compound 7 50 and 5 μg/ml. L4 were left for 24 h to lay eggs. After 24 h they were removed and the number of eggs laid was counted. These eggs were then left for 24 h to hatch and the number of unhatched eggs was counted. The results were expressed as the percentage of the eggs laid that did not hatch.
C. elegans metabolism assay
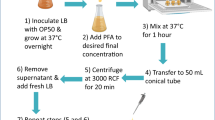
C. elegans were cultivated in liquid bulk culture for one week50. Nematodes were treated with imidacloprid, at a rate of 500 μg/ml, 48 h prior to treatment, to induce cytochrome P450 expression. Healthy nematodes were separated by sucrose floatation, washed with 0.1 M cold NaCl at least three times and resuspended in M9. Following centrifugation at 1500 rpm, the supernatant was removed and the nematodes were used in the metabolism assay immediately.
The C. elegans metabolism assay involved nematodes (100 μl of bulk culture pellet), added to 24 well plates containing the AI (5 μL in DMSO to make a final concentration of 5 μg/ml) and M9 (to a final volume of 500 μL). Separate control plates contained an additional 100 μl M9 instead of nematodes. Further control plates were prepared in the same way but without treatment with the AI (this was a vehicle control). The plates were shaken continuously for 24 h at 20 °C. The metabolism assay was stopped at 0 h or 24 h by the addition of 500 μl acetonitrile to each well, and the plates were frozen.
Nematode lysis was conducted by the following method. The 24 well plates were defrosted at room temperature and the contents of each well were pipetted into an eppendorf, which was frozen in liquid nitrogen and defrosted immediately in the sonicator bath. The samples were homogenised by 2 × 20s cycles with a FastPrep FP120 (Bio101/Savant) then centrifuged at 10,000 rpm for 15 mins to separate solid debris. The supernatant from each sample was transferred into an HPLC vial and all extracts were analysed by LC-MS. If the extracts could not be analysed immediately they were stored at 4 °C overnight and allowed to warm to room temperature prior to LC-MS analysis. The control samples, without AI, were pooled. The contents of the two plates of control samples, containing C. elegans, or saline alone were used to make blank controls and calibration curves.
Liquid Chromatography – Mass Spectrometry (LC-MS) Analysis
Reversed-phase UPLC analysis was carried out using a ACQUITY UPLC system (Waters, Elstree, UK) and a ACQUITY UPLC BEH C18 column (1.7 μm; 50 × 2.1 mm; Waters, Elstree, UK) with a mobile phase mixture of 0.2% formic acid (A) and acetonitrile (B). During the complete 6-min chromatographic cycle time the linear gradient program was as follows: initial 5% B held for 0.5 min, 5% B increasing to 95% by 4.5 min, 95% B held between 4.5 and 4.9 min, then reduced to 5% B in 0.1 min and 5% B between 5.0 and 6.0 min. The injection volume was 5 μL. A constant flow rate and temperature of 0.7 ml/min and 40 °C, respectively, were maintained throughout the run and the mobile phase was split before reaching the electrospray ionisation mass spectrometry interface. Mass spectrometric analysis was performed with a Micromass ZQ (Waters, Elstree, UK) spectrometer. The instrument was operated in positive ion mode employing single-ion recording (SIR) mode at the molecular ion mass [M + H]+ of each compound; inter-scan delay 0.1s and dwell 0.05s. Matrix-matched standard solutions of each AI were analysed alongside the metabolism assay extracts and data processing was performed using MassLynx (Waters). The data are expressed as percent of recovered compound vs. time 0. The compounds tested were 3, 6 and 17.
Microarrays
Mixed stage C. elegans were exposed to AI on plates for 48 h. The compounds were added to the plates in the solvent mixture 10% DMSO 50% IPA 40% H2O to the following final concentrations: compound 3 0.5 μg/ml, compound 4 0.5 μg/ml, compound 6 0.5 μg/ml, compound 7 50 μg/ml, compound 16 5 μg/ml and compound 17 10 μg/ml. These concentrations were chosen as being ones in which the compounds caused effects on egg hatching but not adult lethality. The final concentration of solvent was 1%. Three biological replicates were used per compound. The worms were washed and total RNA was extracted using Trizol.
500 ng total RNA per sample was used to create labeled aRNA target using the Affymetrix 3′ IVT Express Kit. 12.5 μg of each of the resulting aRNAs was fragmented and hybridized to the Affymetrix GeneChip® C. elegans Genome Array, and then washed and stained using the GeneChip® Hybridization, Wash, and Stain Kit. The arrays were scanned using the Affymetrix GeneChip® Scanner 3000 7G, and the signal intensity of probe hybridization was processed using the Affymetrix® GeneChip® Command Console® (AGCC) Software.
Statistical analysis of microarray data were performed using R software (version 3.1)51 and the affy52, affycoretools53, statmod and Bioconductor Limma54 packages. Raw data was initially assessed and normalized using the robust multichip analysis (RMA) algorithm. Differential gene expression between groups was then determined by fitting a linear model to the data using lmFit with subsequent comparisons made using the makeContrasts function. Transcripts with a q-value55 of less than 0.05 were classed as significantly differentially regulated. Further analysis was performed using Expressionist from GeneData.
RNAi knockdown
RNAi knockdown was performed by feeding using the C. elegans RNAi v1.1 Feeding Library from Open Biosystems which is derived from the C. elegans ORFeome Library. This is in the form of glycerol stocks of E. coli with each strain expressing dsRNA against one C. elegans open reading frame. Strains were grown up in LB containing ampicillin and were used to seed 5 cm and 24 well NGM agar plates containing ampicillin and IPTG. These plates were then induced by being placed at 37 °C overnight before use.
Worms were bleached to recover isolated eggs50. These eggs were added to the 5 cm RNAi plates and placed at 15 °C for four days to reach L4. After four days AI was added to the 24 well RNAi plates as described for the egg viability assay. The L4 were then picked onto the 24 well AI containing RNAi plates for the egg viability assay.
Additional Information
How to cite this article: Harlow, P. H. et al. The nematode Caenorhabditis elegans as a tool to predict chemical activity on mammalian development and identify mechanisms influencing toxicological outcome. Sci. Rep. 6, 22965; doi: 10.1038/srep22965 (2016).
References
Hewitt, M., Ellison, C. M., Enoch, S. J., Madden, J. C. & Cronin, M. T. Integrating (Q)SAR models, expert systems and read-across approaches for the prediction of developmental toxicity. Reprod Toxicol 30, 147–160, doi: 10.1016/j.reprotox.2009.12.003 (2010).
Li, H. et al. Use of the ES-D3 cell differentiation assay, combined with the BeWo transport model, to predict relative in vivo developmental toxicity of antifungal compounds. Toxicol In Vitro 29, 320–328, doi: 10.1016/j.tiv.2014.11.012 (2015).
Ball, J. S. et al. Fishing for teratogens: a consortium effort for a harmonized zebrafish developmental toxicology assay. Toxicol Sci 139, 210–219, doi: 10.1093/toxsci/kfu017 (2014).
Boyd, W. A. et al. Developmental Effects of the ToxCast Phase I and II Chemicals in and Corresponding Responses in Zebrafish, Rats, and Rabbits. Environ Health Perspect, doi: 10.1289/ehp.1409645 (2015).
Allard, P., Kleinstreuer, N. C., Knudsen, T. B. & Colaiacovo, M. P. A C. elegans screening platform for the rapid assessment of chemical disruption of germline function. Environ Health Perspect 121, 717–724, doi: 10.1289/ehp.1206301 (2013).
Meyer, D. & Williams, P. L. Toxicity Testing of Neurotoxic Pesticides in Caenorhabditis elegans. J Toxicol Environ Health B Crit Rev 17, 284–306, doi: 10.1080/10937404.2014.933722 (2014).
Cui, Y., McBride, S. J., Boyd, W. A., Alper, S. & Freedman, J. H. Toxicogenomic analysis of Caenorhabditis elegans reveals novel genes and pathways involved in the resistance to cadmium toxicity. Genome Biol 8, R122, doi: 10.1186/gb-2007-8-6-r122 (2007).
Wolozin, B., Saha, S., Guillily, M., Ferree, A. & Riley, M. Investigating convergent actions of genes linked to familial Parkinson's disease. Neurodegener Dis 5, 182–185, doi: 10.1159/000113697 (2008).
Kitagawa, N. et al. The role of the presenilin-1 homologue gene sel-12 of Caenorhabditis elegans in apoptotic activities. J Biol Chem 278, 12130–12134, doi: 10.1074/jbc.M212058200 (2003).
Wang, S. et al. Cadmium-induced germline apoptosis in Caenorhabditis elegans: the roles of HUS1, p53, and MAPK signaling pathways. Toxicol Sci 102, 345–351, doi: 10.1093/toxsci/kfm220 (2008).
Leung, M. C. et al. Caenorhabditis elegans: an emerging model in biomedical and environmental toxicology. Toxicol Sci 106, 5–28, doi: 10.1093/toxsci/kfn121 (2008).
Cole, R. D., Anderson, G. L. & Williams, P. L. The nematode Caenorhabditis elegans as a model of organophosphate-induced mammalian neurotoxicity. Toxicol Appl Pharmacol 194, 248–256, doi: 10.1016/j.taap.2003.09.013 (2004).
Mitchell, P. et al. A differential role for neuropeptides in acute and chronic adaptive responses to alcohol: behavioural and genetic analysis in Caenorhabditis elegans . PLoS One 5, e10422, doi: 10.1371/journal.pone.0010422 (2010).
Kayser, E. B., Morgan, P. G. & Sedensky, M. M. GAS-1: a mitochondrial protein controls sensitivity to volatile anesthetics in the nematode Caenorhabditis elegans . Anesthesiology 90, 545–554 (1999).
de Boer, R. et al. Caenorhabditis elegans as a Model System for Studying Drug Induced Mitochondrial Toxicity. PLoS One 10, e0126220, doi: 10.1371/journal.pone.0126220 (2015).
Kullyev, A. et al. A genetic survey of fluoxetine action on synaptic transmission in Caenorhabditis elegans. Genetics 186, 929–941, doi: 10.1534/genetics.110.118877 (2010).
Ward, A., Walker, V. J., Feng, Z. & Xu, X. Z. Cocaine modulates locomotion behavior in C. elegans. PLoS One 4, e5946, doi: 10.1371/journal.pone.0005946 (2009).
Sulston, J. E. & Horvitz, H. R. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev Biol 56, 110–156 (1977).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans . Dev Biol 100, 64–119 (1983).
Priess, J. Notch signaling in the C. elegans embryo, In WormBook, (ed. The C. elegans Research Community) doi: 10.1895/wormbook.1.4.1 (2005).
Pellegatti, M. Preclinical in vivo ADME studies in drug development: a critical review. Expert Opin Drug Metab Toxicol 8, 161–172, doi: 10.1517/17425255.2012.652084 (2012).
Lindblom, T. H. & Dodd, A. K. Xenobiotic detoxification in the nematode Caenorhabditis elegans . J Exp Zool A Comp Exp Biol 305, 720–730, doi: 10.1002/jez.a.324 (2006).
The-C.elegans-sequencing-consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Knudsen, T. B. et al. Profiling the activity of environmental chemicals in prenatal developmental toxicity studies using the U.S. EPA's ToxRefDB. Reprod Toxicol 28, 209–219, doi: 10.1016/j.reprotox.2009.03.016 (2009).
Lofstedt, R. Communicating food risks in an era of growing public distrust: three case studies. Risk Anal 33, 192–202, doi: 10.1111/j.1539-6924.2011.01722.x (2013).
Dekant, W. & Kehrer, J. P. Scientifically unfounded precaution drives European Commission's recommendations on EDC regulation, while defying common sense, well-established science and risk assessment principles. Toxicol Lett 223, A1–4, doi: 10.1016/j.toxlet.2013.07.010 (2013).
Knudsen, T. B. et al. Activity profiles of 309 ToxCast chemicals evaluated across 292 biochemical targets. Toxicology 282, 1–15, doi: 10.1016/j.tox.2010.12.010 (2011).
Lamberth, C. et al. Synthesis and fungicidal activity of tubulin polymerisation promoters. Part 2: pyridazines. Bioorg Med Chem 20, 2803–2810, doi: 10.1016/j.bmc.2012.03.035 (2012).
Lamberth, C. et al. Synthesis and fungicidal activity of tubulin polymerisation promoters. Part 3: imidazoles. Bioorg Med Chem 21, 127–134, doi: 10.1016/j.bmc.2012.10.052 (2013).
Driscoll, M., Dean, E., Reilly, E., Bergholz, E. & Chalfie, M. Genetic and molecular analysis of a Caenorhabditis elegans beta-tubulin that conveys benzimidazole sensitivity. J Cell Biol 109, 2993–3003 (1989).
Jones, L. M., Flemming, A. J. & Urwin, P. E. NHR-176 regulates cyp-35d1 to control hydroxylation-dependent metabolism of thiabendazole in Caenorhabditis elegans. Biochem J 466, 37–44, doi: 10.1042/bj20141296 (2015).
Jones, L. M., Rayson, S. J., Flemming, A. J. & Urwin, P. E. Adaptive and specialised transcriptional responses to xenobiotic stress in Caenorhabditis elegans are regulated by nuclear hormone receptors. PLoS One 8, e69956, doi: 10.1371/journal.pone.0069956 (2013).
Schäfer, P., Müller, M., Krüger, A., Steinberg, C. E. W. & Menzel, R. Cytochrome P450-dependent metabolism of PCB52 in the nematode Caenorhabditis elegans. Arch. Biochem. Biophys. 488, 60–68 (2009).
Johnston, W. L., Krizus, A. & Dennis, J. W. The eggshell is required for meiotic fidelity, polar-body extrusion and polarization of the C. elegans embryo. BMC Biol 4, 35, doi: 10.1186/1741-7007-4-35 (2006).
Brenner, S. The genetics of Caenorhabditis elegans . Genetics 77, 71–94 (1974).
Menzel, R. et al. Cytochrome P450s and short-chain dehydrogenases mediate the toxicogenomic response of PCB52 in the nematode Caenorhabditis elegans . J Mol Biol 370, 1–13, doi: 10.1016/j.jmb.2007.04.058 (2007).
Menzel, R., Rodel, M., Kulas, J. & Steinberg, C. E. CYP35: xenobiotically induced gene expression in the nematode Caenorhabditis elegans . Arch Biochem Biophys 438, 93–102, doi: 10.1016/j.abb.2005.03.020 (2005).
Menzel, R., Bogaert, T. & Achazi, R. A systematic gene expression screen of Caenorhabditis elegans cytochrome P450 genes reveals CYP35 as strongly xenobiotic inducible. Arch Biochem Biophys 395, 158–168, doi: 10.1006/abbi.2001.2568 (2001).
Lewis, J. A., Szilagyi, M., Gehman, E., Dennis, W. E. & Jackson, D. A. Distinct patterns of gene and protein expression elicited by organophosphorus pesticides in Caenorhabditis elegans . BMC Genomics 10, 202, doi: 10.1186/1471-2164-10-202 (2009).
Chakrapani, B. P., Kumar, S. & Subramaniam, J. R. Development and evaluation of an in vivo assay in Caenorhabditis elegans for screening of compounds for their effect on cytochrome P450 expression. J Biosci 33, 269–277 (2008).
Farnham, A. W. In Piperonyl Butoxide (ed Jones, D. G. ) Ch. 12, 199–214 (Academic Press, 1998).
Roh, J. Y. & Choi, J. Cyp35a2 gene expression is involved in toxicity of fenitrothion in the soil nematode Caenorhabditis elegans . Chemosphere 84, 1356–1361, doi: 10.1016/j.chemosphere.2011.05.010 (2011).
Aarnio, V. et al. Caenorhabditis elegans Mutants Predict Regulation of Fatty Acids and Endocannabinoids by the CYP-35A Gene Family. Front Pharmacol 2, 12, doi: 10.3389/fphar.2011.00012 (2011).
Ashrafi, K. et al. Genome-wide RNAi analysis of Caenorhabditis elegans fat regulatory genes. Nature 421, 268–272 (2003).
Meskar, A., Plee-Gautier, E., Amet, Y., Berthou, F. & Lucas, D. [Alcohol-xenobiotic interactions. Role of cytochrome P450 2E1]. Pathol Biol (Paris) 49, 696–702 (2001).
Preskorn, S. H. Debate resolved: there are differential effects of serotonin selective reuptake inhibitors on cytochrome P450 enzymes. J Psychopharmacol 12, S89–97 (1998).
Waring, M. J. et al. An analysis of the attrition of drug candidates from four major pharmaceutical companies. Nat Rev Drug Discov 14, 475–486, doi: 10.1038/nrd4609 (2015).
Rodriguez-Ruiz, A., Marigomez, I., Boatti, L. & Viarengo, A. Dictyostelium discoideum developmental cycle (DDDC) assay: a tool for Hg toxicity assessment and soil health screening. Sci Total Environ 450-451, 39–50, doi: 10.1016/j.scitotenv.2013.01.060 (2013).
Guengerich, F. P. Cytochrome P450 and Chemical Toxicology. Chemical Research in Toxicology 21, 70–83, doi: 10.1021/tx700079z (2007).
Stiernagle, T. Maintenance of C. elegans, In WormBook, (ed. The C. elegans Research Community) doi: 10.1895/wormbook.1.101.1 (2006).
R. Core Team R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/ (2013).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315, doi: 10.1093/bioinformatics/btg405 (2004).
MacDonald, J. W. Afycoretools: Functions useful for those doing repetitive analyses with Affymetrix GeneChips. Bioconductor, Fred Hutchinson Cancer Research Center, Seattle, U. S. A. URL http://www.bioconductor.org/packages/release/bioc/html/affycoretools.html(2008).
Smyth, G. K. In Bioinformatics and Computational Biology Solutions using R and Bioconductor (ed Carey, V., Gentleman, R., Dudoit, S., Irizarry, R., Huber, W. ) Ch. 5, 397–420 (Springer, 2005).
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat Med 9, 811–818 (1990).
Acknowledgements
Some strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). We thank Dr Phil Botham and Dr Dick Lewis for useful discussions on the manuscript.
Author information
Authors and Affiliations
Contributions
P.H.H., R.A.C. and A.J.F. conceived the experiments, P.H.H. conducted the experiments, S.J.P. developed the LC-MS analysis methods, S.D. and E.B. conducted the microarrays, S.W. assisted in analysis of the microarrays, C.L. provided the compounds, P.H.H. analysed the results. P.H.H. and A.J.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.H.H., S.J.P., S.D., E.B., C.L., R.A.C. and A.J.F., are employees of Syngenta. S.W. is an employee of General Bioinformatics. The work was funded by Syngenta.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Harlow, P., Perry, S., Widdison, S. et al. The nematode Caenorhabditis elegans as a tool to predict chemical activity on mammalian development and identify mechanisms influencing toxicological outcome. Sci Rep 6, 22965 (2016). https://doi.org/10.1038/srep22965
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep22965
This article is cited by
-
CYP35 family in Caenorhabditis elegans biological processes: fatty acid synthesis, xenobiotic metabolism, and stress responses
Archives of Toxicology (2022)
-
Acute, reproductive, and developmental toxicity of essential oils assessed with alternative in vitro and in vivo systems
Archives of Toxicology (2021)
-
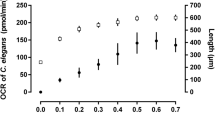
Oxygen consumption rate of Caenorhabditis elegans as a high-throughput endpoint of toxicity testing using the Seahorse XFe96 Extracellular Flux Analyzer
Scientific Reports (2020)
-
Indigenous Preparations of Bryonia laciniosa, Quercus infectoria, Putranjiva roxburghii and Mesua ferrea Induce Developmental Toxicity in C. elegans
Proceedings of the National Academy of Sciences, India Section B: Biological Sciences (2020)
-
Comparative metabolism of xenobiotic chemicals by cytochrome P450s in the nematode Caenorhabditis elegans
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.