Abstract
The rock-hosted, oceanic crustal aquifer is one of the largest ecosystems on Earth, yet little is known about its indigenous microorganisms. Here we provide the first phylogenetic and functional description of an active microbial community residing in the cold oxic crustal aquifer. Using subseafloor observatories, we recovered crustal fluids and found that the geochemical composition is similar to bottom seawater, as are cell abundances. However, based on relative abundances and functional potential of key bacterial groups, the crustal fluid microbial community is heterogeneous and markedly distinct from seawater. Potential rates of autotrophy and heterotrophy in the crust exceeded those of seawater, especially at elevated temperatures (25 °C) and deeper in the crust. Together, these results reveal an active, distinct, and diverse bacterial community engaged in both heterotrophy and autotrophy in the oxygenated crustal aquifer, providing key insight into the role of microbial communities in the ubiquitous cold dark subseafloor biosphere.
Similar content being viewed by others
Introduction
Oceanic crust encompasses the largest aquifer on Earth, with a liquid volume equal to approximately 2% of the ocean’s volume1. It also harbors a substantial reservoir of microbial life that may influence global-scale biogeochemical cycles2. To date, knowledge of microbial life in basaltic crust is derived primarily from the study of crustal fluids from the warm ridge flank in the NE Pacific where anaerobic and thermophilic microbial lifestyles dominate3,4,5 and from the examination of basaltic rocks collected from mid-ocean spreading ridges6,7,8. However, very little is known about the oceanic crustal aquifer in low-temperature ridge flank hydrothermal systems, where circulating fluids are predicted to be cold (<20 °C) and oxygenated9. North Pond is a sedimented basin along the Mid-Atlantic ridge that is relatively well studied in terms of its geology and hydrology, and may serve as a model for thousands of similar sediment basins flanking slow-spreading mid-ocean ridges10. The sediment layer (<200 m thick) at North Pond provides an effective barrier to seawater-basalt exchange, while the surrounding ridges of exposed rocky outcrops facilitate the exchange of fluids between the deep ocean and the crustal aquifer. Previous work has shown that fluid flows rapidly in a horizontal direction through the porous oceanic crust beneath North Pond11,12, and oxygen profiles of the sediment column revealed that the deepest sediment contains higher concentrations of oxygen than shallower sediment, providing evidence that the basaltic aquifer is oxygenated2,13. Less well constrained are what microbial communities reside in these fluids and how this microbial life impacts biogeochemical cycling in the crustal aquifer and overlying ocean.
CORK (Circulation Obviation Retrofit Kit) subseafloor observatories, installed through scientific ocean drilling programs (e.g., Integrated Ocean Drilling Program, IODP), allow access to subseafloor fluids and enable the characterization of microbial life in low-temperature ridge-flank hydrothermal systems14. These CORK observatories are installed in several locations, including in the warm (64 °C), 3.5 million-year-old ridge flank of the Juan de Fuca Ridge and in the cold (<20 °C), 8 million-year-old ridge flank at North Pond. New CORK observatories in two drill holes were installed at North Pond in November 2011 during IODP Expedition 33615,16 and the first crustal fluids for microbiological analysis were collected from these boreholes six months later using a remotely operated vehicle (ROV). These CORK observatories are designed and constructed to minimize contamination and serve studies that utilize basaltic crustal fluids, allowing for samplers to be deployed within the seafloor at multiple isolated depth intervals as well as at the CORK wellhead15,16. Specifically, the CORK observatory in U1382A has a packer seal in the bottom of the casing and monitors/samples a single zone in uppermost oceanic crust extending from 90 to 210 mbsf (meters below seafloor). Hole U1383C was equipped with a three-level CORK observatory that spans a zone of thin basalt flows with intercalated limestone (~70–146 mbsf), a zone of glassy, thin basaltic flows and hyaloclastites (146–200 mbsf), and a lowermost zone (~200–331.5 mbsf) of more massive pillow flows with occasional hyaloclastites in the upper part15. Both CORK observatories consist of fiberglass casing within the oceanic crust15. Here we present the initial microbiological and geochemical characterization of the crustal fluids from cold, oxygenated igneous crust at North Pond on the western flank of the Mid Atlantic Ridge. We examined the geochemical and microbial signatures of crustal fluids to resolve the extent of geochemical transformations during passage through the crust and the presence and activities of microbes living in the circulating subseafloor fluids.
Results
General chemical characteristics of North Pond crustal fluids
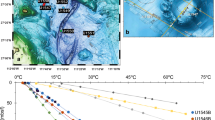
Crustal fluids were retrieved from the single horizon at IODP Hole U1382A and from the shallow, middle, and deep horizons at IODP Hole U1383C (Fig. 1). Slightly lower levels of dissolved oxygen were detected in crustal fluids from both boreholes than in bottom seawater (Table 1). We attempted to measure hydrogen sulfide and iron species at the well head using in situ voltammetry, as well as on deck using a shipboard voltammetic analyzer, but they were not detected (detection limit 0.1 μM for HS- and 1 μM for Fe(II)). Hydrogen sulfide, hydrogen, and methane were also below the limits of detection by the shipboard mass spectrometer on deck (limits of detection ~500 nM for H2, ~500 nM for CH4, ~1 nM H2S). Concentrations of the dissolved elements Si, V, Cs, Ba, and U differed between crustal fluids and bottom seawater, with crustal fluids having more V, Si, Cs, Ba but less U than bottom seawater (Table 2). In contrast, concentrations of dissolved Sr, B, Mo, Mn, Fe, Li, Rb, S (Sulfate), Na, Ca, Mg, K, N2, CO2, and pH were indistinguishable within analytical uncertainties between crustal fluids and seawater (Tables 1 and 2). Dissolved organic carbon (DOC) concentrations in basement fluids were lower than those in background seawater, with the highest values in borehole U1382A and the lowest values in borehole 1383C. Total dissolved nitrogen (TDN) was similar between seawater and borehole 1383C, with elevated values in borehole U1382A (Table 1). A high heterogeneity of particles was detected on the GFF filters and 3 to 9 L of crustal fluids could be filtered onto a single membrane before clogging. Particulate organic carbon (POC) ranged from 0.4 to 1.8 μmol L−1 in crustal fluids, relatively more abundant than POC in bottom seawater (0.1 to 0.3 μmol L−1, Table 3). Particulate nitrogen (PN) content was 0.01 to 0.03 μmol L−1 in crustal fluids, much lower than PN in bottom seawater (0.2 to 0.5 μmol L−1, Table 3). As a result, the POC/TN molar ratios of the particulate organic matter from basement fluids ranged from 27 to 62, while those of bottom seawater were 7 to 8.
Map (A) and diagram (B) of CORK observatories at North Pond. Crustal fluids were extracted from IODP Holes U1382A and U1383C for microbial community analysis in this study. Adapted with permission from15.
Cell densities
All fluid samples, including crustal fluids and bottom seawater, ranged from 1.4 to 2.2 × 104 cells per ml (Table 4). Quantitative PCR detected 260 to 7500 copies of bacterial 16S rRNA genes per ng extracted DNA (Table 4). Archaeal 16S rRNA genes could not be consistently amplified and are not included in the results. Approximately 97% of cells in U1383C hybridized with the bacterial-specific FISH probe EUB338, while slightly lower proportions of cells hybridized with EUB338 in fluids from U1382A and bottom seawater (93% and 91%, respectively, Table 4). Cells hybridizing with the archaea-specific FISH probe were only observed very rarely on the filters (i.e. once every other counting grid).
Incubations with 13C-labeled carbon sources
Incubation of crustal fluids and bottom seawater with labeled carbon sources demonstrated the presence of cells capable of utilizing the isotopically labeled carbon sources through both autotrophic and heterotrophic activity at 5 or 25 °C (Fig. 2; Supplementary Table 1A,B). Potential autotrophic activity, as determined by the incorporation of labeled bicarbonate at atmospheric pressure, ranged from 320 to 4100 pmol C ml−1 day−1 (Fig. 2A). Potential heterotrophic activity, determined by the incorporation of labeled acetate at atmospheric pressure, ranged from 6 to 104 pmol C per ml per day (Fig. 2B). Overall, both autotrophic and heterotrophic incorporation of 13C was higher in the incubated crustal fluids than in bottom seawater. In most cases, the rates of autotrophy and heterotrophy were higher at 25 °C than at 5 °C. This trend was most pronounced in samples from the deepest horizon of U1383C, where very little activity was detected at 5 °C. The two shallower crustal fluids had higher levels of both autotrophic and heterotrophic growth when incubated at 5 °C than the deeper crustal fluids from U1383C at 5 °C.
Rates of carbon transformation, including autotrophic (A) and heterotrophic (B) growth, calculated from stable isotope incubation experiments. Incubations with stable isotopes were conducted at both 5 °C (black bars) and 25 °C (grey bars). Incubated fluids include: deep Atlantic bottom seawater (SW) and crustal fluids retrieved from IODP Holes U1382A and U1383C.
Microbial community composition via Illumina 16S rRNA and 16S rRNA gene sequencing
Genomic DNA and total RNA were successfully extracted and bacterial 16S rRNA and 16S rRNA genes were amplified from all crustal fluid samples and from bottom seawater. Approximately 72 ng of DNA was recovered for each fluid sample. Only genomic DNA was extracted from the drilling mud that was used during drilling operations to help wash cuttings from the borehole17, as RNA was not preserved at the time of collection. Archaeal 16S rRNA genes were not amplifiable from genomic DNA extractions. A total of 14,656,528 quality-filtered sequences of the hypervariable v6 region were obtained for bacterial amplicon libraries. At the 3% clustering level, a total of 33,287 OTUs were detected. All samples, including crustal fluids, bottom seawater, and drilling mud, were dominated by Proteobacteria, which constituted as much as 90% of reads (Fig. 3). Gammaproteobacteria made up 30 to 57% of the total reads and Epsilonproteobacteria were common only in crustal fluids (Fig. 3). The Orders Alteromonadales and Chromatiales were the most commonly detected Gammaproteobacteria, while most of the Epsilonproteobacteria belonged to the Campylobacterales (Supplementary Fig. 1). Alphaproteobacteria and Deltaproteobacteria were detected in higher relative abundances in seawater than in crustal fluids. Overall, more than 80% of taxa in either the DNA or RNA fraction of crustal fluid samples were also detected in bottom seawater. While there was considerable overlap in the identity of taxa in the crustal fluids and in the bottom seawater, the relative abundances of different groups reveal a distinct crustal fluid bacterial community structure (Supplementary Fig. 2).
The three most abundant genera in all crustal fluid samples were Colwellia, Sulfurimonas, and Acidiferrobacter (Supplementary Fig. 3). These three genera were also detected in bottom seawater or in the drilling mud at very low levels. A total of 1,502 OTUs were assigned to the genus Colwellia, and the dominant Colwellia OTUs were different between boreholes U1382A and U1383C (Fig. 4). No OTUs in bottom seawater or drilling mud DNA fractions were assigned to Colwellia. A total of 1,143 OTUs were assigned to the genus Sulfurimonas, again with the most abundant OTU varying between holes and depth horizons (Fig. 4). Less than 1% of the reads in bottom seawater and drilling mud were assigned to Sulfurimonas. A total of 477 OTUs were assigned to the genus Acidiferrobacter. All borehole samples were dominated by a single Acidiferrobacter OTU (Fig. 4). No OTUs in bottom seawater or drilling mud DNA fractions were assigned to Acidiferrobacter. In all samples, reads assigned to Acidiferrobacter were in higher abundance in the RNA fraction than in the DNA fraction. In contrast, the most abundant group in the DNA fraction of bottom seawater was the Deltaproteobacteria clade SAR324 (Fig. 3). Chloroflexi clade SAR202 was the most abundant group in the RNA fraction of bottom seawater. The most abundant group in the drilling mud, which is prepared with surface seawater, was Alteromonas.
Proportion of sequencing reads assigned to the five most abundant OTUs (Operational Taxonomic Units) of Colwellia, Sulfurimonas, and Acidiferrobacter in crustal fluids from IODP Holes U1382A and U1383C, in comparison to bottom seawater and drilling mud. Each column in the heatmap represents a unique OTU.
While many taxa occurred at relatively low abundances in the DNA and RNA fractions (Supplementary Table 2), the dominant genera, Colwellia, Sulfurimonas, and Acidiferrobacter, were usually detected in higher abundances in the RNA fraction. Overall, the community composition of the DNA fraction was significantly different from the RNA fraction in crustal fluids (ANOSIM statistic R = 0.68, p = 0.003) and OTUs assigned to Colwellia, Sulfurimonas, and Acidiferrobacter contributed at least 12% of this difference. Other groups that contributed most to the difference between RNA and DNA fractions included Mariprofundus, JTB255 Marine Benthic Group, Alteromondaceae C1-B045, and Marinobacter, all of which were generally detected at higher abundances in the RNA fraction.
Metagenomic results
Details of assembly statistics and annotation results can be found in the Supplementary Online Material (SOM). 671 putative full-length 16S rRNA genes were assembled from the identified 16S rRNA fragments, with 167 ± 75 putative genes per sample. Based on the length-normalized abundance estimates determined by EMIRGE, many of the putative 16S rRNA genes represent organisms with <1% abundance (8.2–64.9% of putative genes) (Supplementary Fig. 4). For the bottom water sample, most of the groups >1% abundance belonged to the Alphaproteobacteria, while U1382A possessed a nearly equivalent abundance of Epsilonproteobacteria (25%) and Gammaproteobacteria (22%). The U1383C deep sample was almost entirely assigned to the Epsilonproteobacteria (84%), while the U1383C shallow sample had more sequences assigned to the Gammaproteobacteria compared to the Epsilonproteobacteria (Supplementary Fig. 4).
Using the essential marker genes described in Albertsen et al.18, 12,774 putative marker genes were identified from the North Pond samples. These putative markers genes were assigned taxonomies at the Class level, if available, and used to recruit sequences from size-normalized metagenomic libraries. In general, the number of putative markers without an assignment at the Class level was small (12–36%), as was the percentage of markers assigned to groups with <1% abundance (2–5%). The metagenome sequences of the bottom water predominantly represented groups within the Alphaproteobacteria, while in U1382A, Epsilonproteobacteria and Gammaproteobacteria were represented in nearly equivalent proportions (Fig. 5). The deep U1383C sample recruited almost exclusively sequences to Epsilonproteobacteria markers (80%), with a few other minor bacterial groups. The shallow U1383C sample had a larger proportion of Gammaproteobacteria sequences (41%) than any other taxonomic group (Fig. 5). The taxonomic assignment based on essential marker genes was similar to the taxonomic assignments from amplicon sequencing of the 16S rRNA gene v6 region, with the exception of U1383C Deep, where the metagenomic profile indicates ~75% Epsilonproteobacteria, while amplicon sequencing suggests they are present at ~15% (Fig. 5).
Putative coding DNA sequences (CDS) from the North Pond IMG annotations were matched to various essential genes for the dominant known carbon fixation pathways. Sequences were recruited from size-normalized metagenomes to each putative carbon fixation gene to determine which forms of carbon fixation potentially dominate in each sample from the aquifer (Fig. 6). There was no indication of the reductive acetyl-CoA pathway in any of the North Pond samples. Genes related to the 3-hydroxypropionate bicycle and the 3-hydroxypropionate/4-hydrocybutyrate cycle were detected in some of the samples, but with very low representation in the libraries. However, in the crustal fluid samples, both reductive Citric acid (rTCA) and Calvin-Benson-Bassham (CBB) cycles were relatively abundant, with the rTCA cycle being most highly represented in the U1383C deep horizon (Fig. 6). Conversely, in the shallow horizon at U1383C, the genes associated with the CBB cycle were the more abundant, with two forms of RuBisCo and only minor contributions from ATP-citrate lyase (aclAB) (Fig. 6). Assignment of putative taxonomies to the putative aclAB genes demonstrates that for the crustal fluids, the predominant organisms associated with the rTCA cycle belong to the Epsilonproteobacteria, which were not identified in the bottom seawater sample (Supplementary Fig. 5). In some cases, taxonomic assignments of the putative CDS could be assigned to the Order Campylobacterales and Family Helicobacteraceae, where the Sulfurimonas lineage resides (Supplementary Fig. 5).
Percent of reads annotated to genes for 4 carbon fixation pathways for the 4 metagenomes from IODP Holes U1382A, U1383C, and bottom seawater. kor, 2-oxoglutarate ferredoxin oxidoreductase; acl, ATP-citrate lyase; mcl, malyl-CoA lyase; abfD, 4-hydroxybutyryl-CoA dehydratase; rbc, ribulose-bisphosphate carboxylase; prk, phosphoribulokinase; rbcL and cbbM, ribulose-1,5-bisphosphate carboxylase/oxygenase.
Using the fraction of putative CDS assigned to TIGRFAMs as a signature for potential large-scale functional differences for each North Pond and several publicly available metagenomes, a principal component analysis was performed to visualize how the various metagenomes relate to each other (Supplementary Fig. 6). The crustal fluid North Pond samples cluster together away from the corresponding bottom water sample. Further, metagenomes representing oxic, surface-influenced sediment (south Pacific abyssal sediment, 5 cmbsf), anoxic, deep sediment (Arctic ridge, 75 cmbsf) and metal-rich hydrothermal plume waters (Guaymas and Abe Lau), were all distinct from the North Pond crustal fluid samples (Supplementary Fig. 6).
Discussion
The geochemical attributes of these fluids suggests a high degree of connectivity between the cold crustal aquifer at North Pond and deep Atlantic seawater. Many of the dissolved major and minor elements and other geochemical tracers in crustal fluids were indistinguishable in concentration, and in some cases isotopic composition, from seawater. It should also be noted that both of these boreholes were drilled ~6 months prior to sampling, a procedure that involves injecting surface seawater into the hole along with periodic flushes of drilling mud. Additionally, both holes were located within 50 m of open boreholes that penetrated into the basement and continued to accept an influx of bottom seawater until they were sealed in 1997 and 201211,19,20. Each of these factors may impact the local hydrology and geochemistry in the crustal aquifer, but the extent of that impact is difficult to constrain. Nevertheless, the data herein do reveal that the fluids sampled are, at the very least, partly if not mostly derived from the formation. For example, there are indicators that crustal fluids are reacting with rocks and/or sediments within the aquifer, resulting in dissolved concentrations of elements that are distinct from background seawater, such as the lower uranium concentrations in U1383C crustal fluids than in seawater, that is consistent with the removal of U into minerals during water-rock reactions21. The higher concentrations of dissolved silica in the crustal fluids may result from either diffusive exchange with sediment pore waters or water-rock reactions22. Sediment contains opal, which has the highest solubility of all Si-bearing minerals in the crust, and has been shown to impact borehole conditions22,23. Alternatively, water-rock reactions at the low temperatures within these boreholes could also produce dissolved silica without a measurable calcium or magnesium anomaly24. Similarly, the distinct increases in the concentrations of vanadium, cesium, and barium are likely due to diffusive exchange with sediment pore waters22. Finally, oxygen concentrations were lower than background seawater in all horizons at U1383C, but higher than in the basal sediments2,13. This suggests that abiotic and/or biotic consumption of oxygen occurs within the aquifer, either from the oxidation of reduced minerals and microbial oxygen consumption in basalt or from diffusive losses to the overlying sediment pore waters where it is likely consumed by microbes as described in Ziebis et al.13. The DOC concentrations in the North Pond crustal fluids are consistent with those previously reported for the crustal fluids from eastern flank of the Juan de Fuca Ridge, demonstrating that ridge flank aquifer is a sink for DOC in the deep ocean25,26. The magnitude of decrease in DOC concentrations in U1382A and U1383C (all three horizons) crustal fluids relative to bottom seawater DOC concentrations is very similar to the amount of loss of oxygen from bottom seawater (~7 and ~25 μM), suggesting that oxidation of DOC may be a possible explanation to the loss of oxygen. POC and POC/TN ratio were higher in crustal fluids relative to bottom seawater, suggesting differences in particulate organic carbon and nitrogen cycling in the subseafloor compared to the deep ocean, perhaps reflecting the consumption of readily available carbon and nitrogen sources and the accumulation of recalcitrant particulate organic carbon in the subseafloor27,28,29.
Stable isotopic experiments demonstrate that some microbes resident in the North Pond fluids are poised to perform both autotrophic and heterotrophic growth when provided with inorganic carbon and a simple organic acid as carbon sources. This is in contrast to recent stable isotopic experiments with subsurface basalts from the same North Pond boreholes (U1382A, U1383C) that found carbon fixation rates were below detection30. The contrasting results may reflect the different incubation conditions used, as our work used more bicarbonate, with a lower fraction of labeled carbon, or it may reflect the loss of biomass from the basalts during the decontamination process. Previous studies of acetate respiration in sediment cores from North Pond showed generally low rates in most samples (<200 pmol respired acetate cm−3 d−1), but spikes of higher respiration (up to 10,600 pmol cm−3 d−1) at different depths that corresponded to a change in the lithology31. In addition, the authors detected no lag time in the utilization of acetate, indicating that some North Pond sediment microbes, while existing in an oligotrophic habitat, are poised to take advantage of labile carbon sources as they become available. Similarly, we measured generally low potential rates (up to 104 pmol respired acetate cm−3 d−1) of acetate respiration in crustal fluids. While rates of potential heterotrophy in shallow crustal fluids exceeded that of deep crustal fluids and bottom seawater, potential autotrophy in the deepest horizon at 25 °C exceeded all others. These results suggest differences in the functional capacity of microbial communities in the different depth horizons and that autotrophy may be occurring in this community. This may be due to limitations in organic carbon abundance and quality, or other factors that have yet to be determined. Data from previously deployed thermistor arrays at North Pond suggests basement temperatures up to 20 °C at approximately 600 mbsf11, and we saw that the potential rate of carbon utilization increased with elevated temperatures, which likely reflects increased reaction kinetics or the increased activity of microbes that are adapted to warmer temperatures, such as those found with increasing depth in the crust.
The 16S rRNA and 16S rRNA gene amplicon sequencing from crustal fluids, seawater, and drilling mud, together with the metagenomic analysis of select samples, revealed a diverse microbial community in the North Pond crustal fluids that is distinct from deep seawater communities. The crustal fluids are dominated by Alpha- Epsilon-, and Gammaproteobacteria unlike the population structure seen in sediments, deep seawater, or hydrothermal habitats (as reviewed in32). Amplicon sequencing identified some key subseafloor organisms in the aquifer, including one potential group of iron-oxidizing bacteria, Acidiferrobacter. While there is only one named isolate of Acidiferrobacter (an iron- and sulfur-oxidizing diazotroph formerly known as Acidothiobacillus)33, this genus has been detected in many different habitats in the deep sea including sediment, basalt, hydrothermal habitats, and deep seawater32. Although iron levels in crustal fluids were below the detection limit, microbes in the subseafloor likely access iron through attachment to basalt rocks. Other putative iron oxidizers, including Mariprofundus and Marinobacter, were also detected in the 16S rRNA amplicon libraries at higher abundance than in the DNA fractions, suggesting iron-based energy metabolism may be important in the crustal aquifer. Colwellia was the most abundant Gammaproteobacteria detected. This group is among the common denizens of the pelagic ocean32 and cultivated isolates of Colwellia include obligate piezophiles and psychrophiles34,35,36,37,38. Colwellia has previously been isolated through aerobic enrichment cultures from basalt, but were not found exclusively in basalt-hosted environments35. Despite the ubiquity of Colwellia in deep seawater, the high relative abundance of this group in crustal fluids rather than in bottom seawater at North Pond suggests that it may be favored by conditions found in the aquifer. Finally, the presence and high relative abundance of Sulfurimonas, particularly in the deepest North Pond crustal fluids, is especially striking. Sulfurimonas is a widespread resident of deep-sea habitats, including marine sediment, oxygen minimum zones, hydrothermal vents and plumes, ridge flanks, and inactive sulfide chimneys39,40,41. Sulfurimonas was also detected in sediment from North Pond collected during IODP Exp 336 in 2009 (B. Orcutt, personal communication). This interaction between sediment and the crustal aquifer is also evident in the presence of oxygen in the sediment at the sediment-basement interface at North Pond2, which would support microbes like Sulfurimonas that preferentially straddle gradients between oxic and anoxic habitats. However, in the deepest horizon of U1383C, there are different Sulfurimonas OTUs than those seen in the shallow horizon of U1383C or at U1382A, and the metagenomic analysis showed that the deepest horizon is dominated by Sulfurimonas. This suggests that at depth in the crust, far away from the overlying sediment, fluids and their microbial communities may be isolated, perhaps due to reduced permeability of oceanic crust as cracks are filled with clays and alteration minerals, thus influencing the biogeochemical state and microbial communities within crustal fluids42.
Furthermore, clear differences in the relative abundance of dominant genera in the DNA and RNA fractions of North Pond crustal fluids were detected using amplicon sequencing of the bacterial 16S rRNA and 16S rRNA gene. The most dominant groups in the RNA fraction may reflect the groups that are most active in these crustal fluids, or may reflect groups with more ribosomes per cell. In either case, the most abundant genera in the RNA fraction likely have the highest potential for protein synthesis. While any functional assignments based on taxonomy are purely speculative, it is possible that North Pond Colwellia are heterotrophic and contribute to the observed transformation of acetate. Likewise, both the Sulfurimonas and Acidiferrobacter that dominate the RNA fraction of North Pond crustal fluids have the potential to be autotrophic and contribute to the carbon fixation documented by the stable isotopic experiments. The metagenomic analysis of carbon fixation genes provides further support for a community capable of autotrophy in the aquifer, particularly among the Sulfurimonas deep in the aquifer.
The differences in the microbial community structure between North Pond crustal fluids and bottom seawater provide several significant insights into the ecology of the basalt-hosted biosphere. First, although there are similar cell densities in warm, reducing, and highly altered (64 °C) crustal fluids collected from the Juan de Fuca ridge flank5 and cold, oxic, and unaltered crustal fluids at North Pond, the two sites have vastly different microbial communities. This is not surprising given the substantial differences in the chemical characteristics of these fluids. The data herein also reveal that the microbial communities in the North Pond aquifer are notably different in both taxonomic and functional composition than those in the deep Atlantic bottom water. However, this is surprising in light of the thermal and chemical similarities of the aquifer fluid and the bottom water. This suggests that the crustal structure (e.g., permeability, hydrogeology, effective porosity) and/or composition (e.g., mineralogy, bulk composition) may be important factors in shaping this subsurface microbial community. In addition, our results suggest that autotrophy occurs in the North Pond crustal fluids, likely supported by the availability of electron donors and acceptors such as iron and sulfur in the basaltic crust. This could also alleviate competition for labile organic carbon, which is likely to be sparse in these deep oligotrophic waters. Future metagenomics analysis will focus on determining the potential energy acquisition pathways for the dominant organisms and what metabolisms drive these communities. Together, the observed differences in potential rates of both heterotrophic and autotrophic carbon utilization between the horizons in U1383C, as well as the differences in the distribution of strains and carbon fixation genes of the dominant taxa (especially Sulfurimonas), provide compelling support for the hypothesis that crustal fluids may be more isolated with depth at North Pond. Genomic reconstruction of key subseafloor lineages will help assess the evolutionary relationship, functional capacity, and potentially the physical connection, if any, of microbes from different depths beneath the seafloor.
These first results from the oxygenated crustal aquifer contribute to our broader understanding of the deep subseafloor, as well as carbon processing in the cold, dark biosphere. We know that the vast flow of fluid exchanging between ocean basins and crustal reservoirs plays an important role in mediating transport of heat and solutes in the oceans, and this study provides the first insight into microbial life and its potential impact on biogeochemical cycles in the global cold, oxygenated crustal aquifer.
Materials and Methods
Sample collection
Crustal fluids were collected from the single horizon at U1382A and from the shallow, middle, and deep horizons at U1383C15 using an ROV-based pumping and filtration system tailored for microbial sampling. The mobile pumping system (or MPS)43 consists of a plumbing flow path that is primed with sterile DI water prior to deployment and includes sensors for measuring real time flow rate (Seametrics), temperature (Seabird), dissolved oxygen and temperature (Aanderaa), and redox analytes (O2, HS-/H2S, Fe(II), using in situ voltammetry). The MPS was mounted on the ROV Jason II and attached to one of the connectors on the wellhead that emerges via an umbilical to the hydrologic horizon of interest within the crustal aquifer. The MPS was used to pump fluids up a 1.25 cm inner diameter Tefzel tubing that is part of the CORK’s umbilical system. Individual CORK fluid delivery lines were flushed using the MPS at a rate of ~4 liters per minute for at least 3 times volume of the fluid delivery line (~20–30 minutes) prior to diverting fluid flow to six 15 L foil-lined tedlar™ sample bags (Jensen Inert Products) that were acid cleaned and sterilized using gamma irradiation prior to deployment. Oxygen concentrations and temperatures were measured every minute during flushing, and fluids were not sampled until at least 15–30 minutes of stable, reproducible measurements were observed, indicating a fully flushed fluid delivery system and access to pristine crustal fluids. In addition, ~5 L of fluid from each of the three horizons in U1383C was filtered in situ onto 47 mm SUPOR filters in pancake-style filter holders (McLane Inc). These filtered samples were preserved in situ using a reservoir of RNA Later (Qiagen) that is part of the MPS pumping system43. Organic particles for carbon and nitrogen analysis were filtered in situ onto 25 mm glass microfiber filter (GFF). Once recovered, ten liters of each sample was filtered onto a 0.22 μm Sterivex-GP filter at 5 °C for microbial analysis. Similarly, bottom seawater was collected by CTD at 100 m above the sea floor and filtered in the same manner. In situ and ship-based filters were fixed at 4 °C for 18 hours with RNA Later immediately after filtering or upon recovery, then frozen at −80 °C until nucleic acid extractions. A frozen sample of drilling mud was obtained from U1382A during IODP Exp. 33616.
Aliquots of recovered fluids were placed in acid-washed 1000 mL high-density polyethylene (HDPE) bottles, 500 and 1000 mL glass bottles, and 10 to 50 mL glass ampoules for shore-based geochemical analyses. Some of these aliquots were acidified with trace metal clean 6N HCl to a final pH of 1.8. Other aliquots were frozen for nutrient analysis and preserved with a dilution of 1:200 with saturated HgCl2.
Chemistry methods
Concentrations of the major ions in seawater (Na, Mg, Ca, S [sulfate], K, and Sr) were determined using an Inductively Coupled Plasma-Optical Emission Spectrometer (ICP-OES) and by diluting samples 1:200 with ultra-pure water (yielding a precision of 1% at the bottom seawater value). Concentrations of B, Mn, Fe, Si, and Li were also measured with an ICP-OES diluted to 1:25 to yield a precision of ~1 to 2% at the bottom seawater value, with the exception of Mn and Fe which have bottom seawater concentrations that are below detection (<0.1 μmol kg−1). At a concentration of 4.5 μmol kg−1 the precision is 2%. An identical precision is determined for Fe measurements at concentrations of 4.4 μmol kg−1. Concentrations of the trace ions V, Ba, Mo, Rb, Cs, and U were determined using an inductively coupled plasma - mass spectrometer (ICP-MS) with a dilution of 1:75 and an analytical precision of 3% at the bottom seawater value. Concentrations of Si, nitrate, ammonium, and phosphate also were determined colorimetrically44 with a precision of about 2% . Chlorinity was determined by potentiometric titration with AgNO3 with a precision of 0.2% of the seawater concentration.
Dissolved volatiles (H2, H2S, CH4 and CO2) were determined shipboard by analyzing 4 L of fluid directly from unopened, unaltered tedlar™ gastight bags (or deep water samples collected via CTD casts) using a Stanford Research Systems Inc. quadropole mass spectrometer equipped with a custom-membrane inlet and a peristaltic pump and Norprene™ tubing45. Values were estimated by analyzing representative discrete spectra of deep bottom water and borehole water, and using existing calibration curves to determine the relative differences among samples. Bottom seawater concentrations are derived from existing literature. Volatile concentrations shown are ±10%. On deck, discrete samples were also analyzed using voltammetry with an inline flow cell fluid path for comparison in situ voltammetry. pH was determined in triplicate using a double junction electrode and a pH meter (Radiometer Inc).
Dissolved organic carbon (DOC) and total dissolved nitrogen (TDN) were measured by high-temperature (680 °C) combustion using a Shimadzu TOC-L analyzer at the SOEST Laboratory for Analytical Biogeochemistry, University of Hawaii. Samples analyzed include those filtered in situ for crustal fluids as well as the CTD samples filtered on deck. Samples were acidified to pH < 2 within the autosampler syringe and were purged with nitrogen to remove inorganic carbon. Five to six replicate analyses were performed using an injected sample volume of 150 μL. The detection limits of DOC and TDN were about 2 and 1.5 μM, respectively. Two consensus reference materials CRM, from University of Miami46, deep seawater and low carbon water, were used extensively before, between, and after sample analysis to monitor analytical accuracy. The analytical reproducibility for DOC and TDN is better than 1.1 μM and 0.2 μM, respectively, by repeated analysis of four deep seawater CRM. Carbon and nitrogen concentrations of particulate organic matter were measured by the Biogeochemical Stable Isotope Facility at School of Ocean and Earth Science and Technology, University of Hawaii. A few drops of sulfurous acid (6–9% of H2SO3) were added to wet the filter and to remove inorganic carbon from the filter. The acidified filters were transferred to a 60 °C oven for 24 hours. The dried filters were then transferred into a tin capsule before placing within a high-temperature combustion CN elemental analyzer (Costech, ECS 4010) connected in-line with a Mass Spectrometer (ThermoFinnigan Delta XP interfaced with a ConFloIV) for analysis of their nitrogen isotopic compositions. The detection limit for carbon and nitrogen content is 10 μg-C and 0.3 μg-N, respectively. The limit for reliable nitrogen isotopic determination is 10 μg-C and 10 μg-N, respectively.
Microbial cell counts with DAPI and CARD-FISH
Whole crustal fluids and bottom seawater were fixed with 3.7% formaldehyde for cell counts. Up to 19.8 ml of fixed fluids were filtered onto a 0.2 μm GTBP polycarbonate filter (Millipore Inc), stained with DAPI (4′,6′-diamidino-2-phenylindole; Sigma), and counted via epiflourescent microscopy47. For catalysed reported deposition fluorescence in situ hybridization (CARD-FISH), cells were filtered onto 0.2 μm GTTP polycarbonate filters (Millipore Inc) and fixed with 2% paraformaldehyde, rinsed with milliQ H2O, air dried and stored at −20 °C until further use. Cells on filters were hybridized with HRP-labeled 16S rRNA targeted oligonucleotide probes EUB33848, ARCH91549 and NON33848 (Biomers GmbH, Ulm, Germany), and the signal was amplified as described elsewhere50 using Alexa 488® tyramides (Invitrogen). The permeabilization step of the protocol before probe hybridization was modified, such that the cells on the filters were first permeabilized with Proteinase K (0.005 U μl −1 in 0.05 M EDTA, 0.1 M Tris-HCl, at pH 8) for 30 minutes at 37 °C. Filters were then washed in 50 ml 1X PBS at room temperature, followed by a second permeabilization treatment with Lysozyme (106 U ml−1, in 0.05 M EDTA, 0.1 M Tris-HCl, at pH 8) for 30 minutes at 37 °C. After signal amplification, all cells were counterstained with DAPI and counted via epiflourescent microscopy.
Quantitative PCR (qPCR)
The relative abundance of bacterial and archaeal 16S rRNA genes was determined by qPCR assays as previously described51,52. To generate standards, plasmid DNA was extracted from Axial Seamount low-temperature diffuse vent clone libraries, purified, and linearized using the WizardPlus SV Minipreps DNA Purification System (Promega Inc). Standards were constructed by mixing equal amounts of four bacterial plasmids for the quantification of bacterial 16S rRNA gene and two archaeal plasmids for the quantification of archaeal 16S rRNA gene. A 1:10 dilution series of the plasmid mixtures beginning with an initial concentration of 0.06 ng μl−1 (bacteria) and 0.10 ng μl−1 (archaea) was used to produce standard curves with R2 values >0.991 and with efficiency ranging from 93 to 96%. Each 20 μl reaction contained KAPA PROBE FAST ABI Prism® 2X qPCR Master Mix (Kapa Biosystems Inc), forward and reverse primers at optimized concentrations of either 3 nM (bacteria) or 4 nM (archaea), optimized probe concentrations of either 2.5 nM (bacteria) or 5.0 nM (archaea), DEPC-treated water, and 2 μl of DNA template. Triplicate reactions were performed on a StepOne Plus Real Time PCR System (Applied Biosystems Inc) for each sample and for no template controls. Cycles began with initial denaturation for 3 min at 96 °C, followed by 40 cycles of 15 s at 96 °C and 3 min at 59 °C. STEPONE software version 2.2.2 (Applied Biosystems Inc) was used to analyze the results.
Incubations with 13C-labeled carbon sources
To determine potential rates of autotrophic and heterotrophic metabolism within crustal aquifer fluids and deep Atlantic bottom water, fluids were incubated with either 13C-labeled bicarbonate (autotrophy) or 13C-labeled acetate (heterotrophy) at atmospheric pressure. Sterile, butyl stoppered 25-ml Balch tubes were prepared under N2 gas in an anaerobic chamber with resazurin (20 μM final concentration) with a mix of either unlabeled bicarbonate and 10% 13C-labeled bicarbonate (to a final concentration of 1.8 mM NaHCO3 and 0.2 mM NaH13CO3) or unlabeled acetate and 10% 13C-labeled acetate (to a final concentration of 13.5 μM C2H3NaO2 and 1.5 μM 13C2H3NaO2). At sea, the pre-amended Balch tubes were filled with 20 mL of freshly sampled fluids using a sterile syringe, with overpressure released by insertion of a second hypodermic needle. Incubated tubes were monitored for color changes and incubations were stopped when an observable color change in the resazurin indicated a change in the redox potential inside the tube (ie. consumption of oxygen). Sterile controls were set up as described above but with an additional filter (0.2 μm pore size) inserted between the syringe outlet and the hypodermic needle. Tubes were incubated in the dark at either 5 or 25 °C. Incubations were stopped at distinct time intervals by addition of either 0.5 mL of a 1 M NaOH solution (for incubations with bicarbonate) or 4 mL of a 20% zinc acetate solution (for incubations with acetate). Tubes were stored at −20 °C until further processing.
For analysis of 13C-labeled biomass, tubes were thawed and the residual pool of bicarbonate or acetate was removed by acidification to a pH of 2 by adding 25% HCl (molecular grade) while stirring and sparging with N2 for at least 30 minutes. The entire volume was then filtered through a pre-combusted glass fiber filter (25 mm diameter, 0.7 μm particle retention, Whatman, UK). The filters were dried in a desiccator overnight and stored at 5 °C until further processing. Filters were weighed into tin capsules and analyzed for 13C/12C ratios with an automated Isotope Cube elemental analyzer (Elementar, Germany) interfaced to a Delta Advantage isotope ratio mass spectrometer (Thermo, Germany). Rates of potential autotrophic metabolism (carbon fixation from 13C-labeled bicarbonate) and heterotrophic metabolism (degradation of 13C-labeled acetate) were calculated from δ13C of the theoretical carbon pool at the start and the end of the incubations (time intervals of not more than 13 days). For this, the transfer of 13C between pools was calculated, according to the following equation:

where δ is the isotopic ratio ((Rsample/Rstandard − 1) · 1000), V is the volume of the incubation (20 mL) and C is the theoretical concentration of the carbon pool based on added labeled carbon sources and assumed endogenous dissolved inorganic carbon (DIC) and dissolved organic carbon (DOC). For incubations with 13C-labeled bicarbonate we assumed a concentration of 2.3 mM endogenous DIC in addition to the added mix of 13C-labeled bicarbonate, thus decreasing the amount of label in the substrate pool. For incubations with 13C-labeled acetate we assumed a concentration of 45 μM endogenous DOC in addition to the added 13C-labeled acetate.
Nucleic acid extraction, Illumina tag sequencing, and analysis
Sterivex filters and 47 mm flat filters were cut into two equal pieces using sterile technique. Total genomic DNA was extracted from one half using a phenol chloroform method as previously described53 and RNA was extracted from the other half with a mirVana miRNA isolation kit (Ambion Inc) preceded by a bead beating step using RNA Powersoil beads (MoBio). Extracted RNA was treated with Turbo DNase (Ambion Turbo DNA-free kit) and converted to cDNA with an Applied Biosystems (ABI) High Capacity RNA to cDNA kit prior to amplicon library preparation. Total genomic DNA was extracted from approximately 1 g of drilling mud using a MoBio UltraClean® Soil DNA Isolation Kit. The V6 region of 16S rRNA and 16S rRNA genes was amplified in triplicate for each sample with previously reported primers designed for archaea and bacteria54 that were modified to include indices and barcodes compatible with the Illumina HiSeq1000 platform rather than 454 Life Sciences Adapters55. Triplicate PCR amplifications were pooled for each sample, cleaned with a Qiagen MinElute kit, and quanitified by PicoGreen assay on a Turner Biosystems spectrophotometer. Fifty nanograms of each cleaned amplicon library was then size selected with a 2% agarose PippinPrep cassette to produce a narrow range of fragment sizes from 200 to 240 bp for sequencing and cleaned again to remove agarose. Equimolar amounts of pooled amplicon libraries and a metagenomic library were run in the same lane to avoid known difficulties of sequencing low-complexity amplicon libraries with Illumina56.
Paired Illumina sequencing reads were quality filtered to remove any reads containing ambiguous nucleotides and only pairs with perfectly overlapping reads were used for further analysis55. Raw sequencing reads are publicly available through the NCBI under BioProject PRJNA280201. Sequences were clustered at 97% similarity with a minimum word length of 30, using usearch57. Taxonomy was assigned by global alignment for sequence taxonomy (GAST)58 with the SILVA 111 database59. Analysis of similarities (ANOSIM) and similarity percentage (SIMPER) were performed in R using VEGAN v2.0-860. Community structure was analyzed in R with phyloseq61 and plotted with ggplot262 and phinch63.
Metagenomic sequencing
DNA from the Sterivex samples collected from bottom seawater, U1382A, U1383C Shallow, and U1383C Deep was sheared to 175 bp using a Covaris S-series sonicator. Libraries were constructed using the Ovation Ultralow Library DR multiplex system (Nugen) following manufacturer’s instructions. Paired-end sequencing was performed on an Illumina HiSeq 1000 at the W.M. Keck sequencing facility at the Marine Biological Laboratory. Details of sequence trimming and assembly are described in the SOM. Raw metagenomic sequencing reads are publicly available through the NCBI under BioProject PRJNA280201.
Initial putative CDS were determined for contigs generated from each sample using Prodigal v2.6.1 (parameters: -m -p meta -q)64. Prodigal was used to generate putative CDS for the sediment metagenomes used for functional comparison (see below). Prodigal-predicted CDS for the NP samples were compared to publicly available deep-sea metagenomes. Additional metagenomes from deep-sea environments that also used Illumina HiSeq were accessed from IMG for comparison to the NP samples as described in the SOM. Additional comparative analyses of these metagenomes are described in the SOM. Each of the NP samples was processed through the IMG/M annotation pipeline (GOLD Analysis Project ID: Bottom water - Ga0071103; 1382A - Ga0071100; 1383C Deep - Ga0071101; 1383C Shallow - Ga0071102)65. IMG/M annotations were used for all searches involving metabolisms of interest and assignments of phylogenetic markers.
Full-length 16S rRNA genes were assembled and identified from each North Pond metagenome using Meta-RNA66 and EMIRGE67 as described in the SOM.
Carbon fixation genes and their taxonomic affiliation were determined in each North Pond metagenome as described in the SOM. Utilizing a method detailed previously18, the putative CDS from the NP sample IMG annotations were searched against a database of 100 essential phylogenetic markers using Hidden Markov models (HMMs) as described in the SOM.
Additional Information
How to cite this article: Meyer, J. L. et al. A distinct and active bacterial community in cold oxygenated fluids circulating beneath the western flank of the Mid-Atlantic ridge. Sci. Rep. 6, 22541; doi: 10.1038/srep22541 (2016).
Change history
16 April 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Johnson, H. P. & Pruis, M. Fluxes of fluid and heat from the oceanic crustal reservoir. Earth and Planet. Sci. Lett. 216, 565–574 (2003).
Orcutt, B. N. et al. Oxygen consumption rates in subseafloor basaltic crust derived from a reaction transport model. Nature Commun. 4, 2539 (2013).
Cowen, J. P. et al. Fluids from aging ocean crust that support microbial life. Science 299, 120–123 (2003).
Huber J. A., Johnson, H. P., Butterfield, D. A. & Baross, J. A. Microbial life in ridge flank crustal fluids. Environ. Microbiol. 8, 88–99 (2006).
Jungbluth, S. P., Grote, J., Lin, H.-T., Cowen, J. P. & Rappé, M. S. Microbial diversity within basement fluids of the sediment-buried Juan de Fuca Ridge flank. ISME J. 7, 161–172 (2013).
Lysnes, K. et al. Microbial community diversity in seafloor basalt from the Arctic spreading ridges. FEMS Microbiol. Ecol. 50, 213–230 (2004).
Mason, O. U. et al. Prokaryotic diversity, distribution, and insights into their role in biogeochemical cycling in marine basalts. ISME J. 3, 231–242 (2009).
Santelli, C. M., Edgcomb, V. P., Bach, W. & Edwards, K. J. The diversity and abundance of bacteria inhabiting seafloor lavas positively correlate with rock alteration. Environ. Microbiol. 11, 86–98 (2009).
Edwards, K. J., Fisher, A. T. & Wheat, C. G. The deep subsurface biosphere in igneous ocean crust: Frontier habitats for microbiological exploration. Front. Microbiol. 3, 1–11 (2012).
Schmidt-Schierhorn, F., Kaul, N., Stephan, S. & Villinger, H. Geophysical site survey results from North Pond (Mid-Atlantic Ridge). In Edwards, K.J., Bach, W., Klaus, A. and the Expedition 336 Scientists, Proc. IODP, 336: Tokyo (Integrated Ocean Drilling Program Management International, Inc.). doi: 10.2204/iodp.proc.336.107.2012 (2012).
Becker, K., Bartetzko, A. & Davis, E. E. Leg 174B Synopsis: Revisiting Hole 395A for logging and long-term monitoring of off-axis hydrothermal processes in young oceanic crust. Proc. ODP, Sci. Results 174B, 1–13 (2001).
Langseth, G. & Yon, P. Heat and fluid flux through sediment on the Western flank of the Mid-Atlantic Ridge: A hydrogeological study of North Pond. Geophys. Res. Lett. 19, 517–520 (1992).
Ziebis, W. et al. Interstitial fluid chemistry of sediments underlying the North Atlantic gyre and the influence of subsurface fluid flow. Earth Planet. Sci. Lett. 323–324, 79–91 (2012).
Wheat, C. G. et al. Fluid sampling from oceanic borehole observatories: design and methods for CORK activities (1990–2010). In Fisher, A.T., Tsuji, T., Petronotis, K., and the Expedition 327 Scientists, Proc. IODP, 327: Tokyo (Integrated Ocean Drilling Program Management International, Inc.). doi:10.2204/iodp.proc.327.109.2011 (2011).
Edwards, K. J. et al. Design and deployment of borehole observatories and experiments during IODP Expedition 336, Mid-Atlantic Ridge flank at North Pond. In Edwards, K.J., Bach, W., Klaus, A., and the Expedition 336 Scientists, Proc. IODP, 336: Tokyo (Integrated Ocean Drilling Program Management International, Inc.). doi:10.2204/iodp.proc.336.109.2012 (2012).
Edwards, K. J., Bach, W. & Klaus, A. Mid-Atlantic Ridge microbiology: initiation of long-term coupled microbiological, geochemical, and hydrological experimentation within the seafloor at North Pond, western flank of the Mid-Atlantic Ridge. IODP Sci. Prosp. 336, doi: 10.2204/iodp.sp.336.2010 (2012).
Masui, N., Morono, Y. & Inagaki, F. Microbiological assessment of circulation mud fluids during the first operation of riser drilling by the deep-earth research vessel Chikyu. Geomicrobiol. J. 25, 274–282 (2008).
Albertsen, M. et al. Genome sequences of rare uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat. Biotechnol. 31, 533–538 (2013).
Morin, R., Hess, A. & Becker, K. In situ measurement of fluid flow in Holes 395A and 534A. Geophys. Res. Lett. 19, 509–512 (1992).
Wheat, C. et al. CORK-Lite: Bringing legacy boreholes back to life. Sci. Drill. 14, 39–43 (2012).
Pirajano F. Hydrothermal Processes and Mineral Systems (Springer Netherlands, 2009).
Wheat, C. G. et al. Subseafloor seawater-basalt-microbe reactions: Continuous sampling of borehole fluids in a ridge flank environment. Geochem. Geophys. Geosyst. 11 doi: 10.1029/2010GC003057 (2010).
Wheat, C. G. et al. Venting formation fluids from deep sea boreholes in a ridge flank setting: ODP Sites 1025 and 1026. Geochem. Geophys. Geosyst. 5, doi: 10.1029/2004GC000710 (2004).
Seyfried, W. Seawater-basalt interaction from 25°–300 °C and 1-500 bars: implications for the origin of submarine metal-bearing hydrothermal solutions and regulation of ocean chemistry. Ph.D. Thesis, Univ. Southern California, 216 pp (1977).
Lang, S. Q., Butterfield, D. A., Lilley, M., Johnson, H. P. & Hedges, J. Dissolved organic carbon in ridge-axis and ridge-flank hydrothermal systems. Geochem. Cosmochim. Acta 70, 3830–3842 (2006).
Lin, H.-T., Hsieh, C. C., Cowen, J. P. & Rappé, M. S. Data report: dissolved and particulate organic carbon in the deep sediments of IODP Site U1363 near Grizzly Bear seamount. In Fisher, A.T., Tsuji, T., Petronotis, K., and the Expedition 327 Scientists, Proc. IODP, 327: Tokyo (Integrated Ocean Drilling Program Management International, Inc.). doi:10.2204/iodp.proc.327.202.2015 (2015).
Blair, N. et al. Carbon isotopic fractionation in heterotrophic microbial metabolism. Appl. Environ. Microbiol. 50, 996–1001 (1985).
Jahnke, L. L., Summons, R. E., Hope, J. M. & Des Marais, D. J. Carbon isotopic fractionation in lipids from methanotrophic bacteria II: the effects of physiology and environmental parameters on the biosynthesis and isotopic signatures of biomarkers. Geochem. Cosmochim. Acta 63, 79–93 (1999).
Meyers, P. A. Preservation of elemental and isotopic source identification of sedimentary organic matter. Chem. Geol. 114, 289–302 (1994).
Orcutt, B. N. et al. Carbon fixation by basalt-hosted microbial communities. Front. Microbiol. 6, 904 (2015).
Picard, A. & Ferdelman, T. Linking microbial heterotrophic activity and sediment lithology in oxic, oligotrophic sub-seafloot sediments of the North Atlantic Ocean. Front. Microbiol. 2, 263 (2011).
Orcutt, B. N., Sylvan, J. B., Knab, N. J. & Edwards, K. J. Microbial ecology of the dark ocean above, at, and below the seafloor. Microbiol. Molecul. Biol. Rev. 75, 361–422 (2011).
Hallberg, K., Hedrich, S. & Johnson, D. Acidiferrobacter thiooxydans, gen. nov. sp. nov.; an acidophilic, thermo-tolerant, facultatively anaerobic iron- and sulfur-oxidizer of the family Ectothiorhodospiraceae. Extremophiles 15, 271–279 (2011).
Deming, J., Somers, L., Straube, W., Swartz, D. & Macdonell, M. Isolation of an obligately barophilic bacterium and description of a new genus, Colwellia gen-nov. System. Appl. Microbiol. 10, 152–160 (1988).
Mason, O. U. et al. The phylogeny of endolithic microbes associated with marine basalts. Environ. Microbiol. 9, 2539–2550 (2007).
Methé, B. A. et al. The psychrophilic lifestyle as revealed by the genome sequence of Colwellia psychrerythraea 34H through genomic and proteomic analyses. Proc. Natl. Acad. Sci. USA 102 10913–10918 (2005).
Nakagawa, S. et al. Distribution, phylogenetic diversity and physiological characteristics of epsilon-Proteobacteria in a deep-sea hydrothermal field. Environ. Microbiol. 7, 1619–1632 (2005).
Nogi, Y., Hosoya, S., Kato, C. & Horikoshi, K. Colwellia piezophila sp. nov., a novel piezophilic species from deep-sea sediments of the Japan Trench. Int. J. Syst. Evol. Microbiol. 54, 1627–1631 (2004).
Sylvan, J. B., Toner, B. M. & Edwards, K. J. Life and death of deep-sea vents: Bacterial diversity and ecosystem succession on inactive hydrothermal sulfides. mBio 3, 1–10 (2012).
Sylvan, J., Pyenson, B., Rouxel, O., German, C. G. & Edwards, K. J. Time-series analysis of two hydrothermal plumes at 9°50’N East Pacific Rise reveals distinct, heterogeneous bacterial populations. Geobiol. 10, 178–192 (2012).
Han, Y. & Perner, M. The globally widespread genus Sulfurimonas: versatile energy metabolisms and adaptations to redox clines. Front. Microbiol. 6, 989 doi: 10.3389/fmicb.2015.00989 (2015).
Fisher, A. T. & Becker, K. Channelized fluid flow in oceanic crust reconciles heat-flow and permeability data. Nature 403, 71–74 (2000).
Cowen, J. P. et al. Advanced instrument system for real-time and time-series microbial geochemical sampling of the deep (basaltic) crustal biosphere. Deep-Sea Res I 61, 43–56 (2012).
Caspers, H., Parsons, T. R., Maita, Y. & Lalli, C. M. A Manual of Chemical and Biological Methods for Seawater Analysis. (Pergamon Press, 1984).
Girguis, P. R. & Childress, J. J. Metabolite uptake, stoichiometry, and chemoautotrophic function of the hydrothermal vent tubeworm Riftia pachyptila: Responses to environmental variations in substrate concentrations and temperature. J. Exp. Biol. 209, 3516–3528 (2006).
Dickson, A., Sabine, C. & Christian, J. Guide to best practices for ocean CO2 measurements. (PICES Science Report, 2007).
Porter, K. & Feig, Y. The use of DAPI for identification and enumeration of bacteria and blue-green algae. Limnol. Oceanogr. 25, 943–948 (1980).
Amann, R. I., Krumholz, L. & Stahl, D. A. Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J. Bacteriol. 172, 762–770 (1990).
Stahl, D. & Amann, R. Development and application of nucleic acid probes in bacteria systematics In Nucleic Acid Techniques in Bacterial Systematics (eds Stackebrandt, E. & Goodfellow, M. ). (John Wiley and Sons, 1991).
Pernthaler, A., Pernthaler, J. & Amann, R. Fluorescence In Situ Hybridization and Catalyzed Reporter Deposition for the identification of marine bacteria. Appl Environ Microbiol 68, 3094–3101 (2002).
Nadkarni, M. A., Martin, F. E., Jacques, N. A. & Hunter, N. Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiol. 148, 257–266 (2002).
Takai, K. & Horikoshi, K. Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl Environ. Microbiol. 66, 5066–5072 (2000).
Sogin, M. L. et al. Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc. Nat. Acad. Sci. USA 103, 12115–12120 (2006).
Huber, J. A. et al. Microbial population structures in the deep marine biosphere. Science 318, 97–100 (2007).
Eren, A. M., Vineis, J. H., Morrison, H. G. & Sogin, M. L. A filtering method to generate high quality short reads using Illumina paired-end technology. Plos One 8, 6–11 (2013).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6, 1621–1624 (2012).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Huse, S. M. et al. Exploring microbial diversity and taxonomy using SSU rRNA hypervariable tag sequencing. Plos Genetics 4, (11) e1000255. doi: 10.1371/journal.pgen.1000255 (2008).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, 590–596 (2013).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
McMurdie, P. J. & Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8 (4), e61217 doi: 10.1371/journal.pone.0061217 (2013).
Wickham, H. ggplot2: elegant graphics for data analysis. (2009).
Bik, H., Bu, S. & Grubbs, W. Phinch: An interactive, exploratory data visualization framework for environmental sequence data. http://biorxiv.org/content/early/2014/10/03/009944.
Hyatt, D., Chen, G.-L., Larimer, F. W. & Hauser, L. J. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinfor. 11, 119 (2010).
Markowitz, V. M. et al. The integrated microbial genomes (IMG) system. Nucleic Acids Res 34, D344–D348 (2006).
Huang, Y., Gilna, P. & Li, W. Identification of ribosomal RNA genes in metagenomic fragments. Bioinformatics 25, 1338–1340 (2009).
Miller, C. S., Baker, B. J., Thomas, B. C., Singer, S. W. & Banfield, J. F. EMIRGE: reconstruction of full-length ribosomal genes from microbial community short read sequencing data. Genome Biol. 12, R44 (2011).
Bates, N. R., Michaels, A. F. & Knap, A. H. Alkalinity changes in the Sargasso Sea: geochemical evidence of calcification? Marine Chemistry 51, 347–358 (1996).
Acknowledgements
We thank the crews of the R/V Merian and ROV Jason, Wolfgang Bach, Beth Orcutt, and Heiner Villinger for outstanding support in accomplishing our field programs. Ship time was generously provided by the German Science Foundation (DFG). Katrina Edwards led the multi-disciplinary efforts at North Pond and her contributions were critical to the success of this project. James Cowen’s pioneering work in deep-sea instrumentation development was key to sample collection for this project as well. Both are missed immensely. This work was supported by NSF OCE1062006 to JAH, NSF OCE1061934 to PRG, and NSF OCE1061827 to BTG. The Center for Dark Energy Biosphere Investigations (C-DEBI) (OCE-0939564), a National Science Foundation-funded Science and Technology Centers of Excellence also supported the participation of CGW, THL, JPC, JAH, BT, and CHH, as well as JLM and UJ through C-DEBI Postdoctoral Fellowships. This is C-DEBI contribution number 204.
Author information
Authors and Affiliations
Contributions
J.A.H., B.T.G. and P.R.G. designed the experimental plan. J.L.M. and U.J. wrote the manuscript, with major contributions from J.A.H., B.T.G. and P.R.G. and approval of all authors. All authors contributed to sample collection and/or data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Meyer, J., Jaekel, U., Tully, B. et al. A distinct and active bacterial community in cold oxygenated fluids circulating beneath the western flank of the Mid-Atlantic ridge. Sci Rep 6, 22541 (2016). https://doi.org/10.1038/srep22541
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep22541
This article is cited by
-
Time-series transcriptomics from cold, oxic subseafloor crustal fluids reveals a motile, mixotrophic microbial community
The ISME Journal (2021)
-
Deep microbial proliferation at the basalt interface in 33.5–104 million-year-old oceanic crust
Communications Biology (2020)
-
Nitrifier abundance and diversity peak at deep redox transition zones
Scientific Reports (2019)
-
Iron-rich Smectite Formation in Subseafloor Basaltic Lava in Aged Oceanic Crust
Scientific Reports (2019)
-
Subseafloor life and its biogeochemical impacts
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.