Abstract
Recent research interest in phytochemicals has consistently driven the efforts in the metabolic engineering field toward microbial production of various carotenoids. In spite of systematic studies, the possibility of using C30 carotenoids as biologically functional compounds has not been explored thus far. Here, we generated 13 novel structures of C30 carotenoids and one C35 carotenoid, including acyclic, monocyclic and bicyclic structures, through directed evolution and combinatorial biosynthesis, in Escherichia coli. Measurement of radical scavenging activity of various C30 carotenoid structures revealed that acyclic C30 carotenoids showed higher radical scavenging activity than did DL-α-tocopherol. We could assume high potential biological activity of the novel structures of C30 carotenoids as well, based on the neuronal differentiation activity observed for the monocyclic C30 carotenoid 4,4′-diapotorulene on rat bone marrow mesenchymal stem cells. Our results demonstrate that a series of structurally novel carotenoids possessing biologically beneficial properties can be synthesized in E. coli.
Similar content being viewed by others
Introduction
Isoprenoids and their derivatives such as sterols, terpenes, dolichols, quinines, etc. are the most abundant class of secondary metabolites with diverse functions in microbial, plant and animal metabolism1. The building block of isoprenoid derivative compounds is isopentenyl diphosphate (IPP), which is synthesized via two non-homologous pathways: the mevalonate pathway and the 1-deoxy-d-xylulose 5-phosphate/2-C-methyl-d-erythritol 4-phosphate (DOXP/MEP) pathway2,3. Terpenoids and carotenoids are the most industrially important isoprenoid derivatives that are used as cosmeceuticals, flavors, colorants and pharmaceutical compounds.
Many efforts for microbial production of valuable compounds including isoprenoids and polyketides have been made using metabolic engineering4,5,6,7. Most of these studies have focused on providing an improved pool of precursors such as IPP in heterologous hosts including E. coli and yeasts. More recent studies have focused on the optimization of metabolic flux for enhanced target production to avoid accumulation of harmful intermediates and control rate-limiting steps in the heterologous host8. To this end, synthetic and systems biology approaches have been adapted to control pathway enzymes at both the transcriptional and translational levels with synthetic biological parts including engineered promoters, ribosome binding sites and genetic circuits9,10.
Carotenoids are derivatives of isoprenoids that play diverse roles in nature11. Thus far, more than 700 carotenoids have been isolated and identified12. Carotenoids gained considerable research attention in the past owing to their usability as natural pigments; however, their high antioxidative and anticarcinogenic activities and potential use as cosmeceutical or pharmaceutical compounds have only been revealed more recently13,14,15,16. Animals obtain carotenoids through their diet, via consumption of fruits and vegetables. After oxidative cleavage of the dietary carotenoids (including xanthins), the bioactive cleavage products, such as apocarotenoids, are used in retinal formation and transcription system activation17. In addition, apocarotenoids are known to act as anticancer agents and cellular modulators of the retinoic acid receptors, retinoid X receptors, peroxisome proliferator-activated receptors and estrogen receptors17. In a recent study, we proposed an anticancer mechanism of two major carotenoids, crocin and crocetin, found in the dried, dark red stigmas of Crocus sativus, which has been traditionally used since ancient times in Southwest Asia for the treatment of some diseases and as a flavoring and coloring agent18. These two carotenoids are C20 structures generated by the cleavage of C40 carotenoids, the most abundant carotenoid structures in nature including lycopene, β-carotene and astaxanthin. Some bacteria produce C50 or C30 carotenoids; however, these are structurally less diverse than the C40 carotenoids. Until date, only few C30 carotenoid-producing natural sources are known, including Staphylococcus aureus, Methylomonas sp. 16a and some strains of the genus Bacillus19,20,21,22.
Several studies successfully discovered novel carotenoids by applying a combinatorial biosynthesis approach with in vitro evolved carotenogenic enzymes23,24,25,26,27. In vitro evolution of key enzymes, which determine the structure of carotenoid in the early step, has led generation of novel series of carotenoids in E. coli23,24,25,26,27. Recently, Furubayashi and coworkers created non-natural C50 carotenoids through the assembly of moderately selective enzymes engineered by directed evolution in E. coli28. Enzymes commonly show strict substrate specificity; however, carotenogenic enzymes show relatively promiscuous substrate specificity giving rise to novel carotenoid structures in a heterologous host29. Phytoene synthase (CrtB), phytoene desaturase (CrtI), lycopene cyclase (CrtY), 4,4′-diapophytoene synthase (CrtM) and 4,4′-diapophytoene desaturase (CrtN) were evolved in vitro, allowing the biosynthesis of diverse carotenoids in E. coli30. Although extensive work has been performed on the diversification and identification of carotenoid structures, the existence of cyclic C30 carotenoids in nature has not been reported; 4,4′-diapotorulene was the first reported monocyclic C30 carotenoid structure biosynthesized by metabolically engineered E. coli23.
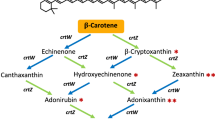
In this study, we created 13 structurally novel C30 carotenoids, including acyclic, monocyclic and bicyclic structures and one novel C35 carotenoid, through combinatorial biosynthesis with natural carotenoid biosynthetic enzymes from various microorganisms and the evolved enzymes, CrtN and CrtY, in E. coli (Fig. 1). Furthermore, we assayed the 2,2-diphenyl-1-picrylhydrazyl (DPPH)-scavenging and neuronal differentiative activities of the C30 carotenoids on rat bone marrow mesenchymal stem cells to explore their potential as bioactive compounds.
Results
Biosynthesis of structurally novel acyclic C30 carotenoids and a C35 carotenoid
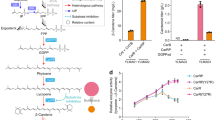
Previously, CrtN mutant clones obtained by directed evolution showed altered desaturation activity in E. coli30. Similarly, we constructed a CrtN mutant library by error-prone PCR upon expression of the substrate, 4,4′-diapophytoene, in E. coli. We screened clones based on the color change to dark reddish, yellowish and pale-yellowish colonies, depending on the number of conjugated double bonds (CDBs) in the chromophore of the carotene backbone, as compared to the reddish wild-type colonies. Their carotenoid profiles were monitored by HPLC to select CrtNr, CrtNy and CrtNz mutant clones showing increased pathway selectivity for 4,4′-diapolycopene, 4,4′-diaponeurosporene and 4,4′-diapo-ζ-carotene, respectively (peaks 1, 2 and 3 in Fig. 2a). Unfortunately, we failed to determine the structure of the CrtN mutants due to the lack of CrtN structural data, even though we performed site-directed mutagenesis (Supplementary Fig. S1) and docking prediction analysis.
Analysis of C30and C35 acyclic carotenoids produced by engineered E. coli cells.
(a) HPLC profiles of the C30 acyclic carotenoids pathway reconstructed in E. coli expressing background pACM-MSA and wild-type and mutant CrtN. (b) HPLC profiles of the C30 carotenoids pathway reconstructed in E. coli expressing pACM-MSA-NSA-ARS and CrtDRC. (c) HPLC profiles of the C35 acyclic carotenoid pathway reconstructed in E. coli expressing pACM-MSA-NSA and CrtEbGC. UV/Vis absorption spectra for compounds corresponding to individual peaks are shown in the upper-right panels. Cell pellets and identified carotenoid structures are shown in the lower-right panel.
In an attempt to create novel carotenoid structures, we first extended the wild-type 4,4′-diapolycopene pathway (pACM-MSA-NSA) using two wild-type enzymes, C40 spheroidene monooxygenase (CrtARC) and C40 1-hydroxycarotenoid 3,4-desaturase (CrtDRC) from Rhodobacter capsulatus, resulting in the generation of two new peaks in the HPLC profile (peaks 4 and 5 in Fig. 2b). Based on combined analysis of retention time, UV/Vis absorption and mass spectra, peaks 4 and 5 were assigned to the acyclic C30 carotenoids 4,4′-diaponeurosporen-6,6′-dione and 4,4′-diapoplycopene-6,6′-dione, respectively (Fig. 2b). Especially, the broadened UV/Vis absorption spectrum and longer λmax (467 nm) observed for the compound of peak 5 supported the presence of 11 fully CDBs and two ketone groups in its structure31.
Next, the 4,4′-diapolycopene pathway was extended with C50 lycopene elongase (CrtEb), which was cloned from Corynebacterium glutamicum. CrtEb catalyzes the sequential addition of two IPPs into the natural substrate lycopene to produce C45 nonaflavuxanthin and C50 flavuxanthin. Extension of the 4,4′-diapolycopene pathway with CrtEb resulted in the production of an elongated acyclic C35 carotenoid, 4,4′-diapononaflavuxanthin (peak 6 in Fig. 2c), assigned on the basis of combined structural analysis as described above.
Biosynthesis of structurally novel cyclic C30 carotenoids
Next, we aimed to create cyclic C30 carotenoids by introducing various bacterial C40 lycopene cyclases (CrtY) into the redesigned C30 carotenoid biosynthesis pathways. As β-ionone ring formation in C30 carotenoids requires the saturated C4-C5 and C4′-C5′ bonds of a carotene backbone, we chose the redesigned 4,4′-diapo-ζ-carotene (pACM-MSA-NzSA) and 4,4′-diaponeurosporene (pACM-MSA-NySA) pathways (Fig. 1). First, expanding upon the redesigned 4,4′-diaponeurosporene pathway, we expressed each C40 CrtY from Salinibacter ruber (CrtYSR), C. glutamicum (CrtYCG), Brevibacterium linens (CrtYBL), Pantoea ananatis (CrtYPN) and Pantoea agglomerans (CrtYPA) to investigate their non-natural substrate promiscuity to form one β-ionone ring in acyclic C30 4,4′-diaponeurosporene. Among them, CrtYBL and CrtYPA produced new HPLC peaks, while the others did not show noticeable differences when compared to control cells expressing empty vector (Fig. 3a). Based on combined structural analysis, we speculated that CrtYBL catalyzed the cyclization of one saturated end of non-natural substrate 4,4′-diaponeurosporene, generating structurally novel monocyclic C30 4,4′-diapotorulene (peak 3 in Fig. 3a). To enhance the selectivity toward the monocyclic 4,4′-diapotorulene pathway, CrtYBL was evolved by error-prone PCR and some mutant CrtYtBL enzymes showing higher selectivity toward 4,4′-diapotorulene (Supplementary Fig. 2) were selected for pathway extension (see next section). Interestingly, unlike the high selectivity of CrtYBL for 4,4′-diapotorulene, CrtYPA showed low specificity toward non-natural short substrates 4,4′-diaponeurosporene and 4,4′-diapo-ζ-carotene and low catalytic efficiency. We hypothesized that CrtYPA cyclized each saturated end of the non-natural substrate 4,4′-diapo-ζ-carotene and one saturated end of the non-natural substrate 4,4′-diaponeurosporene, creating bicyclic C30 4,4′-diapo-β-carotene (peak 4 in Fig. 3a) and monocyclic C30 4,4′-diapotorulene, respectively, albeit at very small amounts. Notably, the formation of bicyclic 4,4′-diapo-β-carotene indicated that CrtYPA had an affinity for 4,4′-diapo-ζ-carotene, one of pathway intermediates in the redesigned 4,4′-diaponeurosporene pathway (Fig. 1).
Functional comparison of CrtYs from various sources in E. coli expressing the 4,4′-diaponeurosporene or 4,4′-diapo-ζ-carotene pathway.
(a) Each C40 CrtY from a different source was coexpressed with the 4,4′-diaponeurosporene pathway, pACM-MSA-NySA, to compare its activity on the C30 backbone. (b) In vivo activity of mutant CrtYtBL, wild-type CrtYBL and CrtYPA on 4,4′-diapo-ζ-carotene was compared in E. coli expressing pACM-MSA-NzSA. UV/Vis absorption spectra for compounds corresponding to individual peaks are shown in the upper-right panel. Identified carotenoid structures are shown in the lower-right panel.
Next, to selectively increase bicyclic 4,4′-diapo-β-carotene production, we coexpressed CrtYBL, CrtYPA and mutant CrtYtBL with the redesigned 4,4′-diapo-ζ-carotene pathway. As expected from the observed affinity of CrtYPA for 4,4′-diapo-ζ-carotene, CrtYPA produced 4,4′-diapo-β-carotene with high selectivity (Fig. 3b). Unexpectedly, CrtYBL also produced a small amount of bicyclic 4,4′-diapo-β-carotene, which was not detected upon coexpression with 4,4′-diaponeurosporene, suggesting that the relative substrate concentration significantly influences the product profile of CrtYBL. Mutant CrtYtBL produced 4,4′-diapo-β-carotene and 4,4′-diapotorulene at a similar ratio, indicating that mutant CrtYtBL had a virtually equal substrate affinity for 4,4′-diapo-ζ-carotene and 4,4′-diaponeurosporene.
Biosynthesis of structurally novel monocyclic C30 carotenoids harboring a modified β-ionone ring
Two β-ionone ring moiety-modifying enzymes, C40 carotene ketolase (CrtOSY) from Synechocystis sp. PCC 6830 and C40 β-carotene hydroxylase (CrtZPA) from P. agglomerans, were individually coexpressed with the 4,4′-diapotorulene pathway (pACM-MSA-NySA-YtBL) to create structurally novel monocyclic C30 carotenoids. Combined structural analysis revealed that CrtZPA catalyzed the hydroxylation of the β-ionone ring in the non-natural substrate 4,4′-diapotorulene, creating 7-hydroxy-4,4′-diapotorulene (peak 4 in Fig. 4a), while CrtOSY catalyzed the addition of a ketone group into the β-ionone ring, creating 8-keto-4,4′-diapotorulene (peak 3 in Fig. 4a).
Analysis of C30 monocyclic carotenoids produced by engineered E. coli cells.
(a) HPLC profiles of the C30 monocyclic carotenoids pathway reconstructed in E. coli expressing background pACM-MSA-NySA-YtBL or pACM-MSA-NySA-YtBL-ZPA and C40 carotenoid modifying enzymes (CrtXPA, CrtWNO, CrtZPA and CrtOSY). (b) HPLC profiles of the C30 monocyclic carotenoids pathways reconstructed in E. coli expressing pACM-MSA-NSA-PSA with CrtYtBL and AldHSA. UV/Vis absorption spectra for compounds corresponding to individual peaks are shown in the upper-right panels. Identified carotenoid structures are shown in the lower-right panels.
The redesigned 7-hydroxy-4,4′-diapotorulene pathway (pACM-MSA-NySA-YtBL-ZPA) was further extended by individually coexpressing C40 carotene ketolase (CrtWNO) from Nostoc sp. PCC 7120 and C40 zeaxanthin glucosyltransferase (CrtXPA) from P. agglomerans. The results indicated that CrtXPA catalyzed the glycosylation of the hydroxyl-β-ionone ring in the non-natural substrate 7-hydroxy-4,4′-diapotorulene, creating 7-hydroxy-4,4′-diapotorulene glucoside (peak 6 in Fig. 4a), while CrtWNO catalyzed the addition of a ketone group into the hydroxyl-β-ionone ring, creating 7-hydroxy-8-keto-4,4′-diapotorulene (peak 5 in Fig. 4a).
Biosynthesis of structurally novel monocyclic C30 carotenoids with a modified acyclic end
Next, we aimed to create novel monocyclic C30 carotenoid structures by modifying the acyclic end of the carotene backbone. We first coexpressed C30 4,4′-diaponeurosporene oxidase (CrtPSA) from S. aureus32 with the redesigned 4,4′-diapotorulene pathway. As expected from the reported low specificity of CrtPSA32, novel C30 4,4′-diapotorulen-4′-al (peak 8 in Fig. 4b) was produced and 4,4′-diaponeurosporen-4′-al accumulated (peak 7 in Fig. 4b). Notably, CrtPSA could compete with YtBL for 4,4′-diaponeurosporene substrate for the production of 4,4′-diaponeurosporen-4′-al and 4,4′-diapotorulene, respectively, which were further modified by the other competing enzymes. The reaction sequence for 4,4′-diapotorulen-4′-al production is unclear, but based on the fact that 4,4′-diapotorulene was not detectable on HPLC, we postulate that YtBL would first produce 4,4′-diapotorulene, to which an aldehyde group was added by CrtPSA, resulting in the formation of 4,4′-diapotorulen-4′-al.
For further extension of the redesigned 4,4′-diapotorulen-4′-al pathway (pACM-MSA-NySA-PSA+pUCM-YtBL), C30 4,4′-diaponeurosporene aldehyde dehydrogenase (AldHSA) from S. aureus32 was expressed. The results suggested that AldHSA catalyzed the oxidation of non-natural substrate 4,4′-diapotorulen-4′-al, creating novel C30 4,4′-diapotorulen-4′-oic acid (peak 10 in Fig. 4b). These novel monocyclic C30 structures could be used as substrates for carotenoid cleavage dioxygenases for the production of novel apocarotenoids33. 1H NMR analysis of 4,4′-diapotorulene was performed to provide structural evidence of 4,4′-diapotorulene derivatives produced in E. coli (Supplementary Fig. 3).
Biosynthesis of structurally novel bicyclic C30 carotenoids
Next, we attempted to extend the bicyclic C30 4,4′-diapo-β-carotene pathway. Similar to the extension of the monocyclic 4,4′-diapotorulene pathway, we individually coexpressed C40 CrtOSY and CrtZPA with the redesigned 4,4′-diapo-β-carotene pathway (pACM-MSA-NzSA-YPA). Combined structural analysis indicated that CrtZPA catalyzed the hydroxylation of the β-ionone ring at each end of the non-natural substrate 4,4′-diapo-β-carotene, creating C30 4,4′-diapo-β-cryptoxanthin (peak 5 in Fig. 5a) through one-step hydroxylation and a very tiny amount of C30 4,4′-diapozeaxanthin (peak 6 in Fig. 5a) through two-step hydroxylation. Unlike the two-step reactions of CrtZPA on the two β-ionone rings, CrtOSY only catalyzed one addition of a ketone group into one β-ionone ring in the non-natural substrate 4,4′-diapo-β-carotene, creating C30 4,4′-diapoechinenone (peak 7 in Fig 5a). The detection of a tiny amount of 4,4′-diapozeaxanthin and the absence of the expected structure 4,4′-diapocanthaxanthin suggested that the catalytic activity of C40 carotenoid-modifying enzymes towards the non-natural short substrate 4,4′-diapo-β-carotene was lower than that towards the other non-natural short substrate 4,4′-diapotorulene.
Analysis of C30 bicyclic carotenoids produced by engineered E. coli cells.
(a) HPLC profiles of the C30 bicyclic carotenoids pathway reconstructed in E. coli expressing background pACM-MSA-NzSA-YPA and C40 carotenoid modifying enzymes (CrtOSY and CrtZPA). Insets: absorption spectra for individual peaks. (b) HPLC profiles of the C30 bicyclic carotenoids pathway reconstructed in E. coli expressing pACM-MSA-NzSA-YPA-ZPA and CrtXPA. UV/Vis absorption spectra for compounds corresponding to individual peaks are shown in the middle panels. Identified carotenoid structures are shown in the right panels.
Finally, we coexpressed C40 CrtXPA with the redesigned 4,4′-diapo-β-cryptoxanthin pathway (pACM-MSA-NzSA-YtBL-ZPA). On the basis of combined structural analysis, we hypothesized that CrtXPA catalyzed the glycosylation of a hydroxyl-β-ionone ring in the non-natural substrate 4,4′-diapo-β-cryptoxanthin, creating C30 4,4′-diapo-β-cryptoxanthin glucoside (peak 8 in Fig. 5b).
Taken together, we created 14 structurally novel carotenoids in recombinant E. coli; their mass spectra are shown in Supplementary Fig. 4.
Radical scavenging activity of structurally novel C30 carotenoids
Most carotenoids typically have antioxidative activity13. Recently, carotenoids are gaining considerable research attention, along with the increasing interest in the role of dietary phytochemicals in human health34. While antioxidative activities of natural C40 carotenoids have been well analyzed, C30 carotenoids have been hardly studied because they are rare in nature and are absent in plants. Therefore, we investigated the radical scavenging activity of five purified, novel C30 carotenoids, including acyclic, monocyclic and bicyclic structures, as well as two previously reported C30 carotenoids (4,4′-diapolycopene and 4,4′-diaponeurosporene) using the DPPH free radical method35,36. DL-α-tocopherol was included as a control compound37. Scavenging reactions were found to be optimal when molar ratios of carotenoid:DPPH between 1:10 and 1:20 were used (data not shown). As expected from the reported antioxidant activities of various C40 carotenoid structures, C30 carotenoids containing higher numbers of CDBs tended to exhibit higher radical scavenging activity than those with fewer CDBs (Fig. 6a), e.g., 4,4′-diapolycopene (13 CDBs) vs. 4,4′-diaponeurosporene (11 CDBs). Notably, 4,4′-diapolycopene-4,4′-dial (13 CDBs+2 aldehyde groups) was the most efficient scavenger, followed by 4,4′-diapolycopene (13 CDBs), 4,4′-diaponeurosporenoic acid (11 CDBs+1 carboxylic group), 4,4′-diaponeurosporenoic acid (11 CDBs+1 aldehyde group) and 4,4′-diaponeurosporene (11 CDBs) (Table 1). Cyclic C30 carotenoids, 4,4′-diapotorulene and 4,4′-diapo-β-carotene, showed lower antioxidative activity than acyclic C30 carotenoids and even lower than the antioxidant control DL-α-tocopherol (Fig. 6a, Table 1). Collectively, the number of CDBs seemed to be one of the most important factors influencing the antioxidative activity of C30 carotenoids and C30 carotenoids having a carboxyl group at the end of the backbone had slightly higher antioxidative activities than those with the same number of CDBs but with an aldehyde end group.
Comparison of DPPH scavenging activity of various C30 carotenoids and morphological changes of rBMSCs treated with varying concentrations of 4,4′-diapotorulene for 7 days.
(a) DL-α-tocopherol was used as a control antioxidant. Values are represented as the mean ± SD from three independent experiments. (b) Cells were treated with varying concentrations of 4,4′-diapotorulene and observed under the microscope for 7 days. Scale bar represents 50 μm.
Neuronal differentiation activity of structurally novel C30 carotenoid 4,4′-diapotorulene
Retinoic acid, an important signaling molecule, has been extensively investigated for its role in inducing neuronal differentiation from various progenitor cells38,39. Because 4,4′-diapotorulen-4′-oic acid is structurally similar to retinoic acid, we assumed that it would also have similar biological activity. However, since the amount of purified 4,4′-diapotorulen-4′-oic acid produced by recombinant E. coli was not sufficient for neuronal differentiation and cytotoxicity assays, 4,4′-diapotorulene was selected for the neuronal differentiation assay in rat bone marrow mesenchymal stem cells (rBMSCs)40.
Cytotoxicity of 4,4′-diapotorulene in the rBMSCs was dose-dependent and approximately 20% of the cells were died after 7 days of treatment with the highest concentration (50 μM) of 4,4′-diapotorulene as compared to control cells (Supplementary Fig. 5). When rBMSCs were treated with varying concentrations of 4,4′-diapotorulene (2, 10, 20 and 50 μM), rBMSCs exhibited bipolar/multipolar or web-shaped morphology characteristic of neuronal cells after a 24-h treatment (Fig. 6b). With longer incubation time (4 and 7 days), the morphological changes enhanced in a dose-dependent manner. These findings indicated the potential neuronal differentiation activity of monocyclic C30 4,4′-diapotorulene on rBMSCs.
Discussion
Metabolic engineering enables the creation of non-natural pathways for novel structures of biotechnological importance, especially by exploiting the promiscuity of biosynthetic pathway enzymes and directed evolution41,42,43,44. Recent research interest in phytochemicals has consistently driven the efforts in the metabolic engineering field toward microbial production of various isoprenoids. In spite of systematic studies on the microbial production of carotenoids, similar to other isoprenoids such as terpenoids, the possibility of using C30 carotenoids, which have not been reported in plant sources, as biologically functional compounds has not been explored thus far. Our study sheds light on the generation of a new series of short C30 carotenoid structures as well as on their potential biological properties such as radical scavenging activity and neuronal differentiation activity. Carotenoids including apo-carotenoids are well known as antioxidative compounds, also interested by their other beneficial biological activities including anticancer activity17. Even though some studies have been made to understand relationship between structure of carotenoids and their anticancer effect, still there is a lack of knowledge on how carotenoids are transferred and modified into apo-carotenoids in the cell45,46. Nevertheless, it is known that complex structures of carotenoids with longer CDBs and/or more functional groups show higher antioxidant activity, which was consistent with our radical scavenging assay (Fig. 6a). Microbial production of apo-carotenoids of biotechnological interest using carotenoid cleavage enzymes is not promising due to low yield and productivity33. Thus, our strategy to produce short C30 carotene-aldehyde/acid derivatives can be served as an alternative method without employing carotenoid cleavage enzymes. Furthermore, as 4,4′-diapotorulene showed neuronal differentiation activity (Fig. 6b) like retinoic acid, an important apo-carotenoid in cell development38, 4,4′-diapotorulene derivatives could open new opportunities for developing bioactive compounds.
Directed evolution of enzymes in early steps in the biosynthesis pathway and combinatorial biosynthesis is a powerful tool for generating diverse novel structures in microbes as described in Introduction section. In this study, CrtN was evolved to optimize 4,4′-diaponeurosporene or 4,4′-diapo-ζ-carotene biosynthesis pathway (Fig. 2a), which could be cyclized into non-natural cyclic C30 carotenoids. Directed evolution of CrtY from B. linens successfully drove cyclization reaction of 4,4′-diaponeurosporene into 4,4′-diapotorulene (Fig. 3a). Through combinatorial biosynthesis by using different CrtY enzymes47, we achieved selective production of 4,4′-diapo-β-carotene from 4,4′-diapo-ζ-carotene (Fig. 3b). After we selectively optimized C30 carotenoid biosynthetic pathways using mutant CrtN and CrtY enzymes, combinatorial biosynthesis with diverse C40 and C50 carotenoid-modifying enzymes found in nature (Supplementary Fig. 6) yielded one acyclic C35 carotenoid and two acyclic, six monocyclic and five bicyclic C30 carotenoids in recombinant E. coli (Fig. 1). Because new types of carotenoid enzymes including carotenoid cleavage enzymes, which can produce biologically more functional apo-carotenoids are continually being discovered and characterized, more structurally novel C30 carotenoids could be generated by using our strategies in the near future, as we had shown in the previous report33. The expanding pool of structurally novel short carotenoids will open up the possibilities for discovery of bioactive compounds, which are potential candidates for drug development and cosmetic applications. However, since this is the first report on the generation of novel C30 and C35 carotenoid structures, more detailed and comprehensive investigation of the biological and physiological activity of C30 carotenoids is warranted. To address this, more quantitative and systems-level studies are needed to selectively direct the metabolic flux towards the formation of novel structures and increase titer of these novel structures in E. coli. Finally, we believe that this work can serve as a basis for investigators handling various carotenoids as candidates of biological active compounds in cosmetics, pharmaceutical, feeds and fine chemical industries.
Materials and Methods
Bacterial strains, plasmids and growth conditions
The bacterial strains and plasmids used in this study are listed in Supplementary Table 1. For gene cloning, the E. coli XL1-Blue strain was grown at 37 °C in Luria-Bertani (LB) medium on a rotary shaker at 250 rpm. For carotenoid production, the E. coli SURE strain29 was grown at 30 °C in Terrific Broth (TB) medium on a rotary shaker at 250 rpm, except for pBBR-aldHSA, which was expressed in XL1-Blue because of kanamycin resistance of the SURE strain. Chloramphenicol (50 μg/mL), ampicillin (100 μg/mL), and/or kanamycin (30 μg/mL) (Sigma) were added as required. P. agglomerans KCTC 2479, R. capsulatus KCTC 2583 and C. glutamicum KCTC 1445 were cultivated in LB medium at 30 °C on a rotary shaker at 250 rpm. S. ruber DSMZ 13855 was grown in medium consisting of 195 g of NaCl, 34.6 g of MgCl2, 49.5 g of MgSO4, 1.25 g of CaCl2, 5 g of KCl, 0.25 g of NaHCO3, 0.625 g of NaBr and 0.1 g of yeast extract per 1 L of double-distilled water at 37 °C for 7 days.
Gene cloning and construction of C30 carotenogenic gene modules
The genes crtY (encoding lycopene cyclase), crtX (glucosyltransferase) and crtD (carotene 3,4-desaturase) were PCR-amplified from genomic DNA of S. ruber, P. agglomerans KCTC 2479 and R. capsulatus KCTC 2583, respectively, using gene-specific primers (Supplementary Table 2). The crtEb (lycopene elongase) and crtY (carotene cyclase) genes were amplified from C. glutamicum KCTC 1445 genomic DNA using gene-specific primers. The PCR products were cloned into pUCM vector29, resulting in pUCM-XAB (where pUCM indicates the vector used, X represents the gene name and the subscript AB in X represents the gene source microorganism; see Supplementary Table 1). C30 carotenogenic gene modules were constructed as described previously32. Briefly, a gene was subcloned from pUCM-XAB into pACYC184 by amplifying the gene together with a modified constitutive lac-promoter, resulting in pACM-XAB (note: sequential assembly of a gene YAB into a module results in pACM-XAB-YAB, for example). PCR amplification was carried out using a DNA Engine Thermal Cycler (Bio-Rad; Hercules, CA, USA) with Vent polymerase (New England Biolabs; Beverly, MA, USA) for cloning, with Taq polymerase (Intron Biotechnology; Seoul, Korea) for random mutagenesis, and, with pfuUltra II fusion polymerase (Agilent Technologies; Palo Alto, CA, USA) for site-directed mutagenesis. Restriction enzymes and T4 DNA ligase were all purchased from New England Biolabs.
Directed evolution of CrtN and CrtY
A mutant library was constructed by error-prone PCR with Taq polymerase by increasing the MgCl2 concentration (up to 2.5 mM) and altering the dNTP ratio (ATP:TTP:CTP:GTP, 1:1:1:4). Error-prone PCR of crtN or crtY in pUCM was carried out under error-prone conditions with plasmid-specific PCR primers (5′-CCGACTGGAAAGCG-3′ and 5′-CGGTGTGAAATACCG-3′) flanking the gene and promoter. The PCR products of crtN and crtY were cloned into the pUCM vector and electrotransformed into recombinant E. coli XL1-Blue harboring pACM-MSA and pACM-MSA-NSA, respectively. Colonies were screened visually on agar plates for color variants after 24 h of incubation (or until color developed) at room temperature. Selected colonies were aerobically grown in TB (4 mL) overnight, pelleted and repeatedly extracted with 2–4 mL acetone. Thin-layer chromatography (TLC) was carried out using a 100% hexane solvent system for initial screening of positive mutant clones. For discrimination of false positive clones, plasmids having mutant a gene were isolated from positive clones and then retransformed into fresh competent E. coli cells harboring pACM-MSA or pACM-MSA-NSA. Sequences of selected crtN and crtY clones were confirmed by Sanger sequencing (Macrogen; Seoul, Korea).
Isolation of carotenoids
Cells were pelleted by centrifugation (4 °C, 4000 rpm) and carotenoids were repeatedly extracted from the pellets with 15 mL acetone until all visible pigments were removed from the pellets. The colored supernatants were pooled after centrifugation (4 °C, 4000 rpm) and concentrated to 5–10 mL using an EZ-2 Plus centrifugal evaporator (Genevac Inc.; Vally Center, NY, USA). An equal volume of ethyl acetate (EtOAc) was added to the concentrated solution that was re-extracted after adding an equal volume of 5 N NaCl solution for salting out. The upper phase, containing the carotenoids, was collected, washed with distilled water, passed through a sodium sulfate column (anhydrous, Bio Basic Inc; ON, Canada) and completely dried using the EZ-2 Plus evaporator. The dried samples were stored at −80 °C until further use.
For purification, the isolated carotenoids were dissolved in 0.2 mL of EtOAc, filtered (0.45-μm GHP membrane, Pall Corporation; NY, USA) and loaded onto a silica gel column, which was pre-equilibrated with a 100% hexane or a 9:1 hexane/EtOAc solvent system. Carotenoids were eluted using the same solvent system with an increasing gradient of EtOAc. Each of the eluted carotenoids was concentrated into a small volume, loaded onto a TLC silica gel plate (Merck Millipore, Billerica, MA, USA) and developed with solvent systems comprising hexane/EtOAc (9:1, v/v) or hexane/EtOAc/methanol (9:1:3, v/v/v). Each carotenoid band was scraped from the TLC plate, followed by elution with methanol. Glassware was used for each step and stray light was blocked with aluminum foil. Purified C30 carotenoids were quantified using a spectrophotometer (SpectraMax Plus384, Molecular Devices; CA, USA) by comparison with the extinction coefficient of known, similar structures of C40 carotenoids48.
Structural analysis of carotenoids
An aliquot (10–20 μL) of the collected fraction or crude extracts was applied to a ZORBAX Eclipse XDB-C18 column (4.6 mm × 150 mm or 4.6 mm × 250 mm, 5.0 μm; Agilent Technologies; Santa Clara, USA) and eluted under isocratic conditions with a solvent system (acetonitrile:methanol:isopropanol, 80:15:5) at a flow rate of 1 mL/min using an Agilent 1200 HPLC system equipped with a photodiode array detector (Agilent Technologies). Chromatograms were recorded at wavelengths of 470, 440 and 400 nm to trace the elution of linear, monocyclic and bicyclic C30 carotenoids, respectively. The mass fragmentation spectra were monitored using both negative and positive ion modes in the mass range of m/z 200 to 600 on a liquid chromatography–mass spectrometer (LC/MS; Agilent 6150) equipped with an atmospheric pressure chemical ionization ion source (Agilent Technologies). The HPLC retention times, UV/Vis absorption spectra and mass fragmentation spectra were all combined for the structural identification of the C30 carotenoids. For NMR analysis of 4,4′-diapotorulene, purified 4,4′-diapotorulene was dissolved in CD3OD and the 1H NMR (600 MHz) spectra were recorded on a Bruker Avance 600 system.
DPPH radical scavenging assay
The 1,1-diphenyl-2-picrylhydrazyl (DPPH) radical scavenging assay for purified C30 carotenoids was performed as described previously37 with a minor modification. Buffered methanol was prepared by mixing 40 mL of 0.1 M acetate solution (pH 5.5) with 60 mL of methanol. Assays were initiated by adding 250 μL of 0.2 M DPPH solution into 200 μL of buffered methanol containing 50 μL of 100 μM carotenoid and incubated in the dark for 1 h. The absorbance at 517 nm was measured using a UV/Vis spectrophotometer (SpectraMax Plus384) against a blank consisting of the reaction mixture without DPPH. dl-α-tocopherol was included as a control. Experiments were performed in triplicate and the data are presented as mean ± standard deviation (SD). IC50 values were calculated after curves were fitted using SigmaPlot 11.0.
Cytotoxicity tests and neurogenesis of rBMSCs
4,4′-Diapotorulene was dissolved in DMSO at a concentration of 100 mM and diluted in culture medium to obtain the desired concentrations (2, 5, 10 and 50 μM). The final concentration of DMSO was kept below 0.02% to maximally eliminate the effect of DMSO. Control cells were incubated in the presence of 0.02% DMSO to compare the effect of 4,4′-diapotorulene. rBMSCs49 were seeded in a 48-well plate at a density of 3 × 103 cells/well in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 100 units/mL penicillin and 100 μg/mL streptomycin. After incubation for 24 h, the culture media were replaced with fresh DMEM supplemented with 10 ng/mL basic fibroblast growth factor and 20% FBS and further incubated overnight. Then, the media were replaced with induction media containing 4,4′-diapotorulene. 4,4′-Diapotorulene solution was prepared at concentrations of 2, 5 and 10 μM in culture medium. For controls, three wells were treated with culture media. At 1, 4 and 7 days, the viability of rBMSCs in the induction media was determined by using MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide], which was converted to formazan that accumulated in the cytoplasm of viable rBMSCs. The viability of rBMSCs was tested in three plates individually and the optical density of each well was determined at 590 nm using a plate reader (E-max, Molecular Device, USA). All experiments were performed at least three times and the results are presented as mean ± SD. To detect neuronal differentiation, digital images of the rBMSCs in the induction media containing 4,4′-diapotorulene were captured using an inverted phase-contrast microscope at 1, 4 and 7 days of incubation.
Additional Information
How to cite this article: Kim, S. H. et al. Generation of structurally novel short carotenoids and study of their biological activity. Sci. Rep. 6, 21987; doi: 10.1038/srep21987 (2016).
References
Roberts, S. C. Production and engineering of terpenoids in plant cell culture. Nat. Chem. Biol 3, 387–395 (2007).
Oldfield, E. & Lin, F. Y. Terpene biosynthesis: Modularity rules. Angew. Chem., Int. Ed. 51, 1124–1137 (2012).
Chandran, S. S., Kealey, J. T. & Reeves, C. D. Microbial production of isoprenoids. Process Biochem. 46, 1703–1710 (2011).
Leonard, E. et al. Combining metabolic and protein engineering of a terpenoid biosynthetic pathway for overproduction and selectivity control. Proc. Natl. Acad. Sci. USA 107, 13654–13659 (2010).
Rodriguez, E., Menzella, H. G. & Gramajo, H. Heterologous production of polyketides in bacteria. Methods Enzymol. 459, 339–365 (2009).
Joseph, A. C., Zachary, L. F. & Mattheos, A. G. K. Trends in microbial synthesis of natural products and biofuels. Advances in Enzymology and Related Areas of Molecular Biology 76, 151–217 (2009).
Horinouchi, S. Combinatorial biosynthesis of plant medicinal polyketides by microorganisms. Curr. Opin. Chem. Biol. 13, 197–204 (2009).
Ajikumar, P. K. et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
Xu, P., Bhan, N. & Koffas, M. A. G. Engineering plant metabolism into microbes: From systems biology to synthetic biology. Curr. Opin. Biotechnol. 24, 291–299 (2013).
Yadav, V. G., De Mey, M., Giaw, L. C., Ajikumar, P. K. & Stephanopoulos, G. The future of metabolic engineering and synthetic biology: Towards a systematic practice. Metab. Eng. 14, 233–241 (2012).
Olson, J. A. Molecular actions of carotenoids. Ann. N. Y. Acad. Sci. 691, 156–166 (1993).
Walter, M. H. & Strack, D. Carotenoids and their cleavage products: Biosynthesis and functions. Nat. Prod. Rep. 28, 663–692 (2011).
Zaini, R. G., Brandt, K., Clench, M. R. & Le Maitre, C. L. Effects of bioactive compounds from carrots (Daucus carota L.), polyacetylenes, beta-carotene and lutein on human lymphoid leukaemia cells. Anti-Cancer Agents Med. Chem. 12, 640–652 (2012).
Tanaka, T., Shnimizu, M. & Moriwaki, H. Cancer chemoprevention by carotenoids. Molecules 17, 3202–3242 (2012).
Korkina, L. & Kostyuk, V. Biotechnologically produced secondary plant metabolites for cancer treatment and prevention. Curr. Pharm. Biotechnol. 13, 265–275 (2012).
Nishino, H., Murakoshi, M., Tokuda, H. & Satomi, Y. Cancer prevention by carotenoids. Arch. Biochem. Biophys. 483, 165–168 (2009).
Sharoni, Y. et al. Carotenoids and apocarotenoids in cellular signaling related to cancer: A review. Mol. Nutr. Food Res. 56, 259–269 (2012).
Kim, S. H., Lee, J. M., Kim, S. C., Park, C. B. & Lee, P. C. Proposed cytotoxic mechanisms of the saffron carotenoids crocin and crocetin on cancer cell lines. Biochem. Cell Biol. 92, 105–111 (2014).
Pelz, A. et al. Structure and biosynthesis of staphyloxanthin from Staphylococcus aureus. J. Biol. Chem, 280, 32493–32498 (2005).
Tao, L., Schenzle, A., Odom, J. M. & Cheng, Q. Novel carotenoid oxidase involved in biosynthesis of 4,4′-diapolycopene dialdehyde. Appl. Environ. Microbiol. 71, 3294–3301 (2005).
Steiger, S., Perez-Fons, L., Fraser, P. D. & Sandmann, G. Biosynthesis of a novel C30 carotenoid in Bacillus firmus isolates. J. Appl. Microbiol. 113, 888–895 (2012).
Maeda, I. Genetic modification in Bacillus subtilis for production of C30 carotenoids. Microbial Carotenoids from Bacteria and Microalgae 892, 197–205 (2012).
Lee, P. C., Momen, A. Z. R., Mijtz, N. B. & Schmidt-Dannert, C. Biosynthesis of structurally novel carotenoids in Escherichia coli. Chem. Biol. 10, 453–462 (2003).
Mijts, B. N., Lee, P. C. & Schmidt-Dannert, C. Identification of a carotenoid oxygenase synthesizing acyclic xanthophylls: Combinatorial biosynthesis and directed evolution. Chem. Biol. 12, 453–460 (2005).
Umeno, D. & Arnold, F. H. Evolution of a pathway to novel long-chain carotenoids. J. Bacteriol. 186, 1531–1536 (2004).
Schmidt-Dannert, C., Umeno, D. & Arnold, F. H. Molecular breeding of carotenoid biosynthetic pathways. Nat. Biotechnol. 18, 750–753 (2000).
Sandmann, G., Albrecht, M., Schnurr, G., Knörzer, O. & Böger, P. The biotechnological potential and design of novel carotenoids by gene combination in Escherichia coli. Trends Biotechnol. 17, 233–237 (1999).
Furubayashi, M. et al. A highly selective biosynthetic pathway to non-natural C50 carotenoids assembled from moderately selective enzymes. Nat. Commun. 6:7534 doi: 10.1038/ncomms8534 (2015).
Kim, S. H., Park, Y. H., Schmidt-Dannert, C. & Lee, P. C. Redesign, reconstruction and directed extension of the Brevibacterium linens C40 carotenoid pathway in Escherichia coli. Appl. Environ. Microbiol. 76, 5199–5206 (2010).
Umeno, D., Tobias, A. V. & Arnold, F. H. Diversifying carotenoid biosynthetic pathways by directed evolution. Microbiol. Mol. Biol. Rev. 69, 51–78 (2005).
Balashov, S. P. et al. Reconstitution of gloeobacter rhodopsin with echinenone: Role of the 4-keto group. Biochemistry 49, 9792–9799 (2010).
Kim, S. H. & Lee, P. C. Functional expression and extension of staphylococcal staphyloxanthin biosynthetic pathway in Escherichia coli. J. Biol. Chem. 287, 21575–21583 (2012).
Heo, J., Kim, S. H. & Lee, P. C. New insight into the cleavage reaction of Nostoc sp. strain PCC 7120 carotenoid cleavage dioxygenase in natural and nonnatural carotenoids. Appl. Environ.Microbiol. 79, 3336–3345 (2013).
Traka, M. H. & Mithen, R. F. Plant science and human nutrition: Challenges in assessing health-promoting properties of phytochemicals. Plant Cell 23, 2483–2497 (2011).
Ohta, T. et al. Loss of Keap1 function activates Nrf2 and provides advantages for lung cancer cell growth. Cancer Res. 68, 1303–1309 (2008).
Müller, L., Fröhlich, K. & Böhm, V. Comparative antioxidant activities of carotenoids measured by ferric reducing antioxidant power (FRAP), ABTS bleaching assay (αTEAC), DPPH assay and peroxyl radical scavenging assay. Food Chem. 129, 139–148 (2011).
Sharma, O. P. & Bhat, T. K. DPPH antioxidant assay revisited. Food Chem. 113, 1202–1205 (2009).
Niederreither, K. & Dollé, P. Retinoic acid in development: Towards an integrated view. Nat. Rev. Genet. 9, 541–553 (2008).
Jones-Villeneuve, E. M., Rudnicki, M. A., Harris, J. F. & McBurney, M. W. Retinoic acid-induced neural differentiation of embryonal carcinoma cells. Mol. Cell. Biol. 3, 2271–2279 (1983).
Kang, K. N. et al. Regeneration of completely transected spinal cord using scaffold of poly(D,L-Lactide-co-Glycolide)/small intestinal submucosa seeded with rat bone marrow stem cells. Tissue Eng., Part A 17, 2143–2152 (2011).
Kim, S. H., Kim, J. H., Lee, B. Y. & Lee P. C. The astaxanthin dideoxyglycoside biosynthesis pathway in Sphingomonas sp. PB304. Appl. Microbiol. Biotechnol. 98, 9993–10003 (2014).
Lee, P. C. & Schmidt-Dannert, C. Metabolic engineering towards biotechnological production of carotenoids in microorganisms. Appl. Microbiol. Biotechnol. 60, 1–11 (2002).
Lee, P. C., Salomon, C., Mijts, B. & Schmidt-Dannert, C. Biosynthesis of Ubiquinone Compounds with Conjugated Prenyl Side Chains. Appl. Environ.Microbiol. 74, 6908–6917 (2008).
Yim, H. et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat Chem Biol. 7, 445–452 (2011).
Albrecht, M., Takaich, S., Steiger, S., Wang, Z. Y. & Sandmann, G. Novel hydroxycarotenoids with improved antioxidative properties produced by gene combination in Escherichia coli. Nat. Biotechnol. 18, 843–846 (2000).
Linnewiel, K. et al. Structure activity relationship of carotenoid derivatives in acrivation of the electrophile/antioxidant response element transcription system. Free Radical Biol. Med. 47, 659–667 (2009).
Song, G. H., Kim, S. H., Choi, B. H., Han, S. J. & Lee, P. C. Heterologous carotenoid-biosynthetic enzymes: Functional complementation and effects on carotenoid profiles in Escherichia coli. Appl. Environ. Microbiol. 79, 610–618 (2013).
Britton, G., Liaaen-Jensen, S. & Pfander, H. Carotenoids Handbook, Volume 1B, G. Britton, L.-J., Pfander eds. Springer Basel AG. (2004).
Cho, M. et al. Induction of Neurogenesis in Rat Bone Marrow Mesenchymal Stem Cells Using Purine Structure-Based Compounds. Mol. Biosys. 5, 609–611 (2009).
Acknowledgements
This work was supported by the Intelligent Synthetic Biology Center of Global Frontier Project funded by the Ministry of Education, Science and Technology (2011-0031968) and by grants from the National Research Foundation of Korea, funded by the Korean Government (2014029244).
Author information
Authors and Affiliations
Contributions
S.H.K., M.S.K., B.Y.L. and P.C.L. conceived and designed the research; S.H.K., M.S.K. and B.Y.L. performed the experiments. S.H.K. constructed the strains development and chemical analysis; MSK performed the cell assay experiments; B.Y.L. performed the N.M.R. analysis; P.C.L. supervised the project. The manuscript was written with the contributions of all authors. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, S., Kim, M., Lee, B. et al. Generation of structurally novel short carotenoids and study of their biological activity. Sci Rep 6, 21987 (2016). https://doi.org/10.1038/srep21987
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep21987
This article is cited by
-
Identification of a novel monocyclic carotenoid and prediction of its biosynthetic genes in Algoriphagus sp. oki45
Applied Microbiology and Biotechnology (2024)
-
4,4′-Diaponeurosporene Production as C30 Carotenoid with Antioxidant Activity in Recombinant Escherichia coli
Applied Biochemistry and Biotechnology (2023)
-
Optimization and multiple in vitro activity potentials of carotenoids from marine Kocuria sp. RAM1
Scientific Reports (2022)
-
Metabolites from halophilic bacterial isolates Bacillus VITPS16 are cytotoxic against HeLa cells
3 Biotech (2021)
-
4–4′ Diaponeurosporenic Acid, the C30 Carotenoid Pigment in Endophytic Pseudomonas Mendocina with Squalene Cyclase Activity
Current Microbiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.