Abstract
Methane is the second most abundant greenhouse gas (GHG), with nearly 60% of emissions derived from anthropogenic sources. Microbial conversion of methane to fuels and value-added chemicals offers a means to reduce GHG emissions, while also valorizing this otherwise squandered high-volume, high-energy gas. However, to date, advances in methane biocatalysis have been constrained by the low-productivity and limited genetic tractability of natural methane-consuming microbes. Here, leveraging recent identification of a novel, tractable methanotrophic bacterium, Methylomicrobium buryatense, we demonstrate microbial biocatalysis of methane to lactate, an industrial platform chemical. Heterologous overexpression of a Lactobacillus helveticus L-lactate dehydrogenase in M. buryatense resulted in an initial titer of 0.06 g lactate/L from methane. Cultivation in a 5 L continuously stirred tank bioreactor enabled production of 0.8 g lactate/L, representing a 13-fold improvement compared to the initial titer. The yields (0.05 g lactate/g methane) and productivity (0.008 g lactate/L/h) indicate the need and opportunity for future strain improvement. Additionally, real-time analysis of methane utilization implicated gas-to-liquid transfer and/or microbial methane consumption as process limitations. This work opens the door to develop an array of methanotrophic bacterial strain-engineering strategies currently employed for biocatalytic sugar upgrading to “green” chemicals and fuels.
Similar content being viewed by others
Introduction
Methane (CH4), the primary component of natural gas and anaerobic digestion-derived biogas, offers a promising, high-volume petroleum replacement for fuel and chemical bioprocesses. Recent advances in gas-recovery technologies have facilitated access to previously inaccessible natural gas reserves, while biogas generated from anaerobic digestion of waste streams offers a versatile, renewable CH4 source. However, the gaseous state of CH4 makes for a lack of compatibility with current transportation and industrial manufacturing infrastructure, limiting its utilization as a transportation fuel and intermediate in biochemical processes. Importantly, CH4 is also the second most abundant greenhouse gas (GHG), with nearly 60% of emissions derived from anthropogenic sources1. Microbial conversion of CH4 to value-added chemicals using natural CH4-consuming bacteria offers valorization potential2,3,4, while reducing GHG emissions.
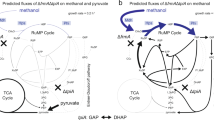
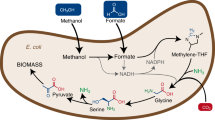
Obligate methanotrophic bacteria (methanotrophs) are a unique group of microorganisms capable of utilizing CH4 or methanol (CH3OH) as their sole carbon and energy source. These bacteria use the enzyme methane monooxygenase (MMO) to convert CH4 to CH3OH, which is further oxidized to formaldehyde (CH2O), formate (CHOOH) and CO2. Depending on the metabolic arrangement, CH4-derived carbon is assimilated at the level of CH2O (via the Ribulose-monophosphate cycle), methylene tetrahydrofolate and CO2 (Serine cycle), or CO2 (Calvin cycle)5,6. In the past, methanotrophs have been exploited for the conversion of CH4 to an array of products7, including bioprotein8,9, polyhydroxybutyrate10, carotenoids11,12,13, vitamins14 and CH3OH15,16. However, advances in CH4 biocatalysis and methanotroph strain engineering have largely been limited by the low-productivity of methanotroph cultures and lack of genetic tools for use in these organisms3,7,17.
Recently, an active Embden–Meyerhof–Parnas (EMP) pathway was identified in novel gammaproteobacterial methanotrophs that are resistant to the toxic components of natural gas and biogas18,19,20,21 and a set of genetic tools, including expression vectors, have been developed for the halotolerant, alkaliphilic methanotrophic bacteria Methylomicrobium buryatense21,22. Given the conserved nature of their downstream metabolic machinery, conventional industrial strain-engineering routes from sugars to biochemical intermediates and products can potentially be paralleled in these methanotrophs. Here, we report microbial biocatalysis of methane to an industrial platform chemical, lactate, a precursor to the biodegradable polylactide (PLA) polymer used in bioplastics. We demonstrate effective genetic engineering strategies in a methanotrophic bacterium, enabling production of lactate from both CH4 and CH3OH as sole carbon sources. The presented route circumvents competition with food substrates, such as corn, utilized in conventional sugar-based lactate production and offers a potentially transformational path to concurrent mitigation of GHG emissions and biological CH4 upgrading.
Results
M. buryatense tolerance to lactate
In order to assess potential end product inhibition on bacterial growth, we first examined M. buryatense tolerance to increasing concentrations of sodium lactate in NMS2 medium22 containing CH3OH as the sole carbon source. Growth inhibition was observed at concentrations above 0.5 g lactate/L (Fig. 1), though this was not due to changes in the pH of the alkaline medium. These data suggest production of lactate from CH4 or CH3OH is feasible in M. buryatense, but also indicate that it may be difficult to achieve high lactate titers without addressing lactate toxicity.
Lactic acid minimum inhibitory concentration against M. buryatense.
Growth of M. buryatense in NMS2 medium supplemented with 1% CH3OH (v/v) and increasing concentrations of sodium lactate. Cultures were inoculated at OD600 = 0.1. The data represent the mean OD600 ± SEM of biological triplicates. ***p < 0.001.
Construction of an inducible broad host-range vector for fine-tuned gene expression in Methylomicrobium
Genetic tools for methanotrophic bacteria are currently limited, making heterologous gene expression and knockout difficult in these organisms. Although constitutive promoters functional in M. buryatense have recently been characterized22, an inducible promoter has yet to be identified for use in this genus. In order to facilitate regulated, heterologous gene expression in the M. buryatense, we constructed an inducible, broad-host range vector, pCAH01, (Fig. 2a) by fusing the tetracycline promoter/operator (tetp/o) from pASK75 with the IncP-based pAWP78 vector that can be replicated by Methylomicrobium spp.22,23. Anhydrotetracycline (aTc) is a tetracycline derivative commonly used as an inducer of the tetp/o in bacteria24, which can exhibit antimicrobial activity at higher concentrations. We observed no effect on M. buryatense growth in the presence of 0.1–1.0 μg aTc/mL, whereas 2.5–10 μg aTc/mL inhibited bacterial growth (Fig. 2b). Experiments using GFP fluorescence as a readout of promoter activity indicated tightly controlled tetp/o-mediated gene expression in M. buryatense after induction with sub-lethal concentrations of the aTc inducer (Fig. 2c). Importantly, the tetp/o did not show any “leaky” gene expression in the absence of inducer, making it a promising tool for conditional gene expression/knock-out studies in methanotrophic bacteria that replicate vectors containing the oriV origin of replication.
Characterization of the inducible, broad host range vector pCAH01.
(a) pCAH01 was constructed by fusing the IncP-based origin of pAWP78 with the tetracycline promoter/operator (Ptet) of pASK75 using Gibson assembly. ahp (kanamycin resistance), bla (ampicillin resistance), tetR (transcriptionally-fused tetracycline repressor), oriV/oriT (IncP-based origin of replication/transfer), trfA (oriV replication initiation protein). (b) Antimicrobial activity of the tetp/o inducer, anhydrotetracycline, against wild-type M. buryatense. (c) tetp/o-dependent induction of GFP in M. buryatense grown in 1% CH3OH (v/v) NMS2 medium supplemented with increasing concentrations of anhydrotetracycline. Relative fluorescence was calculated by comparing fluorescence in pCAH01::emGFP samples to fluorescence in uninduced controls. The data in (b,c) represent the mean ± SEM from two independent experiments (n = 4).
Engineering of M. buryatense for lactate production
We next employed pCAH01 to demonstrate methanotrophic biocatalysis targeting production of lactate. The biosynthetic conversion of the glycolytic intermediate pyruvate to lactate is catalyzed by an NADH-dependent lactate dehydrogenase (LDH) enzyme. Given that gammaproteobacterial methanotrophs have high flux of C1 substrates through pyruvate19, we hypothesized that heterologous LDH expression would facilitate CH4 biocatalysis to lactate in M. buryatense. Heterologous, codon-optimized LDH genes from Escherichia coli, Bifidobacterium longum and Lactobacillus helveticus, whose corresponding LDHs have been used for the production of optically pure lactate25,26,27, as well as the native M. buryatense ldh gene, were placed under control of the tetp/o with a 3′ Flag-tag to track protein expression (Table S1). Western blots using an anti-Flag antibody showed detectable LDH expression in all engineered strains post-induction (Fig. 3a).
LDH expression and lactate accumulation in engineered strains of M. buryatense.
Engineered M. buryatense harboring the pCAH01 empty vector (c) or ectopically expressing the native LDH (Mb), or heterologous, codon-optimized L. helveticus LDH (Lh), B. longum LDH (Bl), or E. coli LDHA (Ec) with C-terminal Flag tags were grown in shake flasks with 1% CH3OH (v/v). (a) Anti-Flag immunoblot confirmation of LDH expression 24 h post-induction. (b) Lactate titers in culture supernatants of induced strains 72 h post-inoculation/induction from aerobic shake flask cultures as measured by HPLC. (c) Growth (black), CH3OH consumption (red) and lactate accumulation (green) in shake flasks cultures of the Lactobacillus helveticus LDH-overexpressing methanotroph (pLhldh). The data in (b,c) represent the mean ± SEM from at least two independent experiments (n = 2–4). ***p < 0.001.
We evaluated lactate production by the LDH-overexpressing strains under aerobic growth conditions since CH4 biocatalysis is oxygen dependent. Engineered strains expressing the B. longum or L. helveticus LDH accumulated lactate in the medium of CH3OH-grown aerobic shake flask cultures, whereas lactate production was not observed in strains overexpressing the native M. buryatense LDH, E.coli LDH, an empty vector control strain (Fig. 3b), or uninduced controls (data not shown). The L. helveticus LDH-expressing methanotroph (pLhldh) produced significant levels of lactate (65 ± 1 mg/L after 72 h, p < 0.001 compared to wild-type) during the initial screen; therefore, it was selected for further analysis. The pLhldh strain produced negligible lactate (~3 mg/L) during the first 24 h of growth, although it consumed 2.5 g/L CH3OH (Fig. 3c) and possessed significant LDH activity compared to wild-type M. buryatense during this period of growth (Table 1). In contrast, this strain accumulated 71 ± 15 mg lactate/L with a 0.018 ± 0.005 g lactate/g CH3OH yield between 24–72 h. Similar lactate titers were observed in CH4-grown shake-flask cultures (62 ± 35 mg/L, Figure S1).
Bioconversion of methane to lactate by an engineered methanotrophic biocatalyst
We next evaluated growth and lactate production by pLhldh in both 0.5 L (Fig. 4a,b) and 5 L (Fig. 4c,d) continuously stirred tank bioreactors with constant CH4 feed (20% v/v CH4 in air) and increased nitrate, phosphate and trace elements in the medium to support enhanced cell growth. As shown in Fig. 4a, a control strain carrying an empty pCAH01 vector and the pLhldh strain grew to high cell densities (OD600 ~17 and 15 after 96 h, respectively) in modified NMS2 medium at 0.5 L scale. The pLhldh strain produced 0.12 g lactate/L after 96 h of growth while no detectable lactate was produced by the induced control strain (Fig. 4b).
Growth and lactate production in continuously stirred tank methane bioreactors.
Growth and lactate accumulation were monitored by engineered M. buryatense harboring an empty vector (pCAH01, open symbols/bars), or ectopically expressing heterologous Lactobacillus helveticus LDH (pLhLDH, closed symbols/bars) grown in a 0.5 L bioreactor (0.3 L culture volume, (a,b) or a 5.0 L bioreactor (3.0 L culture volume, (c,d) with continuous methane feed. The data represent the mean ± SEM from at least two independent experiments (n = 2–4).
High cell-density cultures were also obtained in the 5 L bioreactor (OD600 ~25 for both control and pLhldh strains, Fig. 4c). Here, the induced control strain accumulated low lactate titers (0.038 ± 0.0007 g lactate/L) in the larger bioreactor (Fig. 4d), presumably from native LDH activity. Lactate produced by the pLhldh strain reached titers of 0.808 ± 0.343 g lactate/L after 96 h, representing a 21-fold increase in lactate production over the control strain (Fig. 4d). Notably, these lactate titers coincide with the maximum lactate concentration tolerated by this organism (Fig. 1). To determine the lactate yield from CH4, we measured real-time CH4 consumption in the 0.5 L and 5 L bioreactor (Fig. 5a). The pLhldh strain consumed 4.06 g CH4 and 49.03 g CH4 after 96 h of growth in the 0.5 L and 5 L bioreactor, respectively (Fig. 5b), which was similar to the control strain (data not shown). Under our experimental conditions, a maximum 0.05 g lactate/g CH4 yield and 0.008 g/L/h volumetric productivity was observed from the pLhldh strain in a 5 L bioreactor (Table 2). Of note, 50–60% of consumed CH4 was utilized for biomass synthesis while additional carbon was lost to formate and acetate, which were detected in the medium of pCAH01 and pLhldh strains under these growth conditions (Figure S2).
Methane consumption by engineered M. buryatense.
(a) Real-time or (b) total methane consumption after 96 h by an engineered M. buryatense strain expressing heterologous Lactobacillus helveticus LDH (pLhLDH) in a 0.5 L or 5.0 L bioreactor. Data in (a,b) are representative data from at least two experiments.
Effect of lactate production on biosynthesis of potential methanotroph lipid-fuel precursors
Methanotrophs are being considered for production of biomass-derived fuels from CH4 due to their ability to accumulate intracellular lipids2,3,4. Based on this interest, we evaluated the effect of lactate production on cellular lipid concentration. Fatty acid methyl ester (FAME) analysis indicated that carbon was not diverted from potential methanotroph-derived lipid fuel precursors while concurrently producing lactate, as both the pLhldh and control strains displayed similar cellular lipid content and profiles, composed primarily of hexadecanoic and hexadecenoic acids (Fig. 6a,b). Collectively, these data support the potential for concurrent methane biocatalysis to a platform chemical and fuel precursors from residual biomass.
Fatty acid methyl ester (FAME) compositional analysis (a) % FAME based on dry cell weight and (b) representative fatty acid composition (%, w/w) of M. buryatense harboring an empty vector (pCAH01), or ectopically expressing heterologous, Lactobacillus helveticus LDH (pLhLDH) after 96 h of growth in a 5.0 L bioreactor.
Discussion
CH4 from natural gas is currently flared or vented globally, resulting in large greenhouse gas emissions2 and revenue and energy losses2,7. Additionally, CH4-rich biogas derived from energy crops, crop residues, biofuel residues (such as stillage and glycerol), manure and other organic waste streams, through anaerobic digestion in agricultural/food waste digesters, waste water treatment plants (WWTP) and landfills, offers a versatile, high-volume, renewable source of CH428,29.
Biological production of value-added chemicals from CH4 represents a path to concurrently mitigate greenhouse gas emissions and utilize an abundant-yet-underutilized feedstock. Halotolerant alkaliphilic methanotrophs are promising biocatalysts for this purpose, displaying relatively rapid growth rates, high culture densities, recalcitrance to toxic components in biogas and natural gas and possessing well-characterized central metabolic pathways18,19,30. Based on these characteristics, we employed M. buryatense for the bioconversion of CH4 to the platform chemical, lactate. We constructed an inducible broad host-range gene expression vector containing the tetracycline promoter/operator, which can be used in an array of methanotrophs that replicate the oriV origin of replication. Using these tools, we demonstrated microbial conversion of CH4 to lactate, a high-volume biochemical precursor predominantly utilized for the production of bioplastics, by an engineered methanotroph expressing a heterologous Lactobacillus LDH. The data presented herein provide proof-of-concept for bioconversion of CH4 to an industrial platform chemical using an engineered methanotrophic bacterium, but also highlight some of the hurdles associated with CH4 biocatalysis, such as CH4 assimilation rate (discussed further below).
The maximum lactate titer produced by the pLhldh M. buryatense reached 1.3 g lactate/L. Currently, lactate produced from pure sugars or lignocellulosic hydrolysates utilizing metabolically-engineered industrial microbes reach titers approximately 100-fold greater than those achieved here31,32, leaving significant room for further M. buryatense metabolic engineering. Notably, however, this titer is over one order of magnitude higher than those achieved for any previously reported engineered methanotrophic bioproduct13 and comparable to titers achieved for an array of fuels and chemicals from biomass-derived substrates in proof-of-principle investigations33,34,35.
Our data show that CH4 uptake by the organism limits productivity in a continuous stirred tank bioreactor. Indeed, the engineered pLhldh strain only assimilated ~2% (g/g) of the supplied CH4 in a 5 L bioreactor (Fig. 5), potentially due to limited gas-to-liquid transfer and/or methanotrophic methane oxidation. Moreover, M. buryatense converts 50–60% (g/g) of the assimilated CH4 to biomass (Figure S2). Considering that >10% (g/g) carbon is converted to other excreted products, including acetate, formate and CO2 (Figure S2), 60% (g/g) of carbon is converted to biomass (Figure S2) and it is estimated that up to 25% of carbon is utilized to synthesize exopolysaccharide36, ~5% (g/g) of carbon (~0.6 g CH4/24 h based on Fig. 5B) is available for conversion to lactate in a process coupled to cell growth. Based on these metrics (Figure S3), >75% of the available carbon was converted to the target product under our experimental conditions. Given the large proportion of carbon flux diverted to non-target products, bioprocess designs that decouple biomass and bioproduct accumulation (such as 2-stage fermentation or cell immobilization), novel bioreactors and hypothesis-driven strain-engineering strategies will be required to optimize CH4 assimilation and flux to product. Ongoing studies are focused on increasing the tolerance of M. buryatense to lactate and rational strain engineering to increase CH4 biocatalysis by enhancing rates of CH4 consumption, carbon conversion efficiency and carbon flux to pyruvate. Technoeconomic analysis is also currently underway to define the minimum productivity metrics required for a gas-to-liquid bioprocess to be competitive with conventional sugar-based lactate production.
Additional potential and hurdles associated with bioconversion of methane to liquid fuels has recently been extensively reviewed2,3,4,7. Several methanotrophic bacteria possess relatively high lipid content due to the accumulation of intracytoplasmic membrane to accommodate the particulate methane monooxygenase37. This intrinsically high lipid content makes these organisms attractive platforms for production of fatty-acid derived fuels2. Indeed, the relatively high biomass yield and lipid content of M. buryatense presented here support that these organisms have potential for lipid-derived fuels. Importantly, lactate production did not alter methanotroph lipid content or speciation. This suggests that the pLhldh strain maintains flux to lipids while also producing lactate, either by increasing flux to pyruvate or diverting additional acetyl-CoA to lipid biosynthesis. However, it is worth noting that we did observe a modest decrease in pLhldh growth compared to the control strain (Fig. 4), presumably due to elevated LDH activity. Thus, an increase in lactate titer may have a more pronounced effect on cell growth since LDH could deplete the pyruvate pool. Nonetheless, data presented here indicate future studies targeting co-production of fuels and chemicals using methanotrophic bacteria are warranted.
CH4 biocatalysis offers a means to concurrently liquefy and upgrade CH4, enabling its utilization in conventional transportation and industrial manufacturing infrastructure. This work also raises the possibility of syngas-derived CH3OH valorization38, expanding the renewable substrates available for methanotrophic biocatalysis. Producing chemicals and fuels from CH4 expands the suite of products generated from biorefineries, municipalities and agricultural operations, with the potential to increase revenue and reduce greenhouse gas emissions. By integrating this process into a conventional biorefinery, new opportunities for recycling and other cost reductions will become apparent.
Materials and Methods
Plasmid construction and transformation
Strains and plasmids used in this study are presented in Table S2. Plasmids for heterologous gene expression were constructed using 2X Gibson Assembly Mix from New England Biolabs (Ipswich, MA) following the manufacturers protocol. Polymerase chain reactions were performed using Q5 High-Fidelity Polymerase from New England Biolabs and primers (Table S3) purchased from Integrated DNA Technologies (Coralville, IO). The inducible, broad-host range vector pCAH01 was constructed by fusing the tetracycline promoter/operator (tetp/o) from pASK75 with the IncP-containing pAWP78 backbone22,23. Codon-optimized LDH genes from B. longum, L. helveticus and E.coli (Table S1) were synthesized by Integrated DNA Technologies. Synthetic ldh genes and the native M. buryatense ldh (METBUDRAFT_3726) were amplified with a 3′ Flag-tag via PCR and cloned directly downstream of the tetp/o, generating plasmids pCAH01::MbldhFlag, pCAH01::LhldhFlag, pCAH01::BlldhFlag, pCAH01::EcldhFlag. Final constructs were confirmed by sequence analysis (Genewiz, South Plainfield, NJ). Escherichia coli Zymo 5a (Zymo Research, Irvine, CA) was used for cloning and plasmid propagation and E. coli S17-1 λpir was used as the conjugation donor strain. E.coli strains were grown at 37 °C in Luria-Bertani (LB) broth supplemented with 50ug/mL of kanamycin. Plasmid constructs were transformed into M. buryatense via conjugation as previously described22. Positive transformants selected on NMS2 agar containing 50 μg/mL of kanamycin were confirmed using plasmid-specific primers in polymerase chain reactions.
Cultivation and growth parameters
M. buryatense 5GB1S were routinely cultured in NMS2 medium at 30 °C with orbital shaking at 175 rpm as previously described22,39. Strains were grown in sealed 1 L glass serum bottles (Kimble Chase, Vineland, NJ) with 20% (v/v) CH4 in air, or 500 mL baffled flasks supplemented with 1% CH3OH (v/v). Serum bottle and shake flask cultures were inoculated at OD600 = 0.01 with plate-harvested biomass. LDH expression was induced by adding 0.5–2.0 μg/mL anhydrotetracycline (aTc, Sigma-Aldrich, St. Louis, MO) at the time of inoculation. To determine the minimum inhibitory concentration, 1% CH3OH (v/v) NMS2 was supplemented with increasing concentrations of aTc or sodium lactate (Sigma-Aldrich) and growth was monitored by measuring the OD600 using a spectrophotometer.
GFP fluorescence
A synthetic emerald GFP (Life Technologies, Carlsbad, CA) open reading frame was amplified by PCR and cloned into pCAH01 via Gibson assembly to generate pCAH01::emGFP. M. buryatense harboring the pCAH01::emGFP vector was subcultured (OD600 = 0.01) in 1% CH3OH (v/v) NMS2 and induced with increasing concentrations of aTc. After 24 h of induction, 100 μL of culture samples were aliquoted into a Nunc™ F96 MicroWell™ Plate (Thermo Scientific, Waltham, MA) and fluorescence was measured (485 nmex/520 nmem) using a FLUOstar Omega fluorometer (BMG LABTECH, Cary, NC).
Western blotting
5GB1S harboring pCAH01::MbldhFlag, pCAH01::LhldhFlag, pCAH01::BlldhFlag, pCAH01::EcldhFlag, or the empty pCAH01 vector were grown in NMS2 supplemented with 1% CH3OH (v/v) and induced with 0.5 μg/mL aTc. Samples were taken at 24 h post-induction, pelleted, resuspended in lysis buffer and disrupted by sonication. Samples normalized to 500 ng total protein were resolved using 12% (v/v) SDS-PAGE, transferred electrophoretically to a PVDF membrane and immunoblotted with the anti-FLAG M2 monoclonal antibody (Sigma-Aldrich).
Enzyme Assays
Cell free extracts for LDH activity measurements were prepared by sonication of logarithmically growing M. buryatense or L. helveticus cultures with or without LDH induction. Protein concentrations were determined with the Pierce 660nm Protein Assay Reagent (Life Technologies) using bovine serum albumin as a protein standard. Reactions were initiated by the addition of 50–500 μg lysate to reaction buffer consisting of 0.2 M Tris HCl, pH 7.5, 10 mM NADH and 30 mM sodium pyruvate at 30 °C. LDH activities are expressed in units (U) per milligram of protein where one U was defined as the amount of enzyme required to oxidize 1 μmol of NADH per min measured at A340nm.
Bioreactor fermentations
CH4 fermentations were performed in a 0.5 L Biostat-Q Plus (Sartorius Stedim Biotech) or 5.0 L BioFlo batch bioreactor (New Brunswick Scientific, Edison, NJ) containing NMS2 medium supplemented with 8X KNO3, 2X phosphate buffer and 4X trace element solution to support high cell growth. The bioreactor was inoculated at OD600 = 2 with a CH4-grown seed culture grown in NMS2 containing 2X KNO3. 2 μg/mL aTc was immediately added to induce LDH expression. The temperature was maintained at 30 °C and mixing was achieved by using a bottom marine impeller and mid-height Rushton impeller at 500 rpm. A continuous flow rate of 300 ccm (0.5 L) or 300αcm (5.0 L) 20% (v/v) methane in air was maintained. Antifoam was added as needed. CH4 off-gas was measured every 20 min for the duration of bioreactor fermentations by using an infrared-based BlueSens methane detector (Herten, Germany). CH4 consumption was determined by calculating the difference between off-gas detected before and after inoculation. % CH4 consumption was converted to weight based on CH4 density and the flow rate (56.64 g CH4/24 h, 0.5 L reactor; 566.4 g CH4/24 h, 5.0 L reactor).
Compositional analysis of culture supernatants
High pressure liquid chromatography (HPLC) was used to detect lactate, formate, acetate and CH3OH in culture supernatants. At the indicated time, the OD600 was measured and a 1 mL sample was taken for HPLC analysis. Culture supernatant was filtered using a 0.2 μm syringe filter or 0.5 mL 10 K MWCO centrifuge tube (Life Technologies) and then separated using a model 1260 HPLC (Agilent, Santa Clara, CA) and a cation H HPx-87H column (Bio-Rad). A 0.1 mL injection volume was used in 0.01 N sulfuric acid with a 0.6 mL/min flow rate at 55 °C. DAD detection was measured at 220 nm and referenced at 360 nm and organic acid concentrations were calculated by regression analysis compared to known standards.
FAME analysis
Whole biomass lipid content was measured as fatty acid methyl esters (FAMEs), as described previously40. Briefly, 7 to 10 mg of lyophyilized biomass (dried overnight at 40 °C under vacuum) was homogenized with 0.2 mL of chloroform:CH3OH (2:1, v/v) and the resulting solubilized lipids were transesterified in situ with 0.3 mL of HCl:CH3OH (5%, v/v) for 1 h at 85 °C in the presence of a known amount of tridecanoic acid (C13) methyl ester as an internal standard. FAMEs were extracted with hexane (1 mL) at room temperature for 1 h and analyzed by gas chromatography:flame ionization detection (GC:FID) on a DB-WAX column (30 m × 0.25 mm i.d. and 0.25 μm film thickness).
Statistical analysis
Data between two groups were analyzed using an unpaired, two-tailed t-test. Determination of statistical significance between multiple comparisons was achieved using one-way analysis of variance (ANOVA) followed by a Dunnett’s post-test. Data were considered statistically significant when p < 0.05.
Additional Information
How to cite this article: Henard, C. A. et al. Bioconversion of methane to lactate by an obligate methanotrophic bacterium. Sci. Rep. 6, 21585; doi: 10.1038/srep21585 (2016).
References
Anderson, B. et al. Methane and Nitrous Oxide Emissions From Natural Sources, U.S. Environ. Prot. Agency, Washington, D. C. (2010).
Fei, Q. et al. Bioconversion of natural gas to liquid fuel: opportunities and challenges. Biotechnol. Adv. 32, 596–614 (2014).
Conrado, R. J. & Gonzalez, R. Chemistry. Envisioning the bioconversion of methane to liquid fuels. Science 343, 621–623 (2014).
Haynes, C. A. & Gonzalez, R. Rethinking biological activation of methane and conversion to liquid fuels. Nat. Chem. Biol. 10, 331–339 (2014).
Anthony, C. The biochemistry of methylotrophs. (Academic Pr, 1982).
Chistoserdova, L., Kalyuzhnaya, M. G. & Lidstrom, M. E. The expanding world of methylotrophic metabolism. Annu. Rev. Microbiol. 63, 477–499 (2009).
Strong, P. J., Xie, S. & Clarke, W. P. Methane as a resource: can the methanotrophs add value? Environ. Sci. Technol. 49, 4001–4018 (2015).
Bothe, H. et al. Heterotrophic bacteria growing in association with Methylococcus capsulatus (Bath) in a single cell protein production process. Appl. Microbiol. Biotechnol. 59, 33–39 (2002).
Harrison, D. E. & Hamer, G. C1 compounds as substrates for the production of single-cell protein. Biochem. J. 124, 78P (1971).
Khosravi-Darani, K., Mokhtari, Z.-B., Amai, T. & Tanaka, K. Microbial production of poly(hydroxybutyrate) from C1 carbon sources. Appl. Microbiol. Biotechnol. 97, 1407–1424 (2013).
Ye, R. W. et al. Construction of the astaxanthin biosynthetic pathway in a methanotrophic bacterium Methylomonas sp. strain 16a. J Ind Microbiol Biotechnol 34, 289–299 (2007).
Tao, L. et al. Expression of bacterial hemoglobin genes to improve astaxanthin production in a methanotrophic bacterium Methylomonas sp. Appl. Microbiol. Biotechnol. 74, 625–633 (2007).
Sharpe, P. L. et al. Use of transposon promoter-probe vectors in the metabolic engineering of the obligate methanotroph Methylomonas sp. strain 16a for enhanced C40 carotenoid synthesis. Appl. Environ. Microbiol. 73, 1721–1728 (2007).
Ivanova, E. G., Fedorov, D. N., Doronina, N. V. & Trotsenko, I. A. [Production of vitamin B12 in aerobic methylotrophic bacteria]. Mikrobiologiia 75, 570–572 (2006).
Duan, C., Luo, M. & Xing, X. High-rate conversion of methane to methanol by Methylosinus trichosporium OB3b. Bioresour. Technol. 102, 7349–7353 (2011).
Hwang, I. Y. et al. Biocatalytic conversion of methane to methanol as a key step for development of methane-based biorefineries. J. Microbiol. Biotechnol. 24, 1597–1605 (2014).
Kalyuzhnaya, M. G., Puri, A. W. & Lidstrom, M. E. Metabolic engineering in methanotrophic bacteria. Metabolic Engineering 29, 142–152 (2015).
Kaluzhnaya, M., Khmelenina, V. & Eshinimaev, B. Taxonomic Characterization of New Alkaliphilic and Alkalitolerant Methanotrophs from Soda Lakes of the Southeastern Transbaikal Region and description of Methylomicrobium buryatense sp.nov. Syst. Appl. Microbiol. 24, 166–176 (2001).
Kalyuzhnaya, M. G. et al. Highly efficient methane biocatalysis revealed in a methanotrophic bacterium. Nat Commun 4 (2013).
Vuilleumier, S. et al. Genome sequence of the haloalkaliphilic methanotrophic bacterium Methylomicrobium alcaliphilum 20Z. J. Bacteriol. 194, 551–552 (2012).
Khmelenina, V. N. et al. Draft Genome Sequence of Methylomicrobium buryatense Strain 5G, a Haloalkaline-Tolerant Methanotrophic Bacterium. Genome Announc 1, e00053–13 (2013).
Puri, A. W. et al. Genetic tools for the industrially promising methanotroph Methylomicrobium buryatense. Appl. Environ. Microbiol. 5, 1775–81. (2014). doi: 10.1128/AEM.03795-14.
Skerra, A. Use of the tetracycline promoter for the tightly regulated production of a murine antibody fragment in Escherichia coli. Gene 151, 131–135 (1994).
Henard, C. A., Bourret, T. J., Song, M. & Vázquez-Torres, A. Control of redox balance by the stringent response regulatory protein promotes antioxidant defenses of Salmonella. J. Biol. Chem. 285, 36785–36793 (2010).
Ilmén, M., Wiebe, M. G. & Ruohonen, L. L-lactic acid production from D-xylose with Candida sonorensis expressing a heterologous lactate dehydrogenase encoding gene. Microb. Cell 13, 107 (2014).
Utrilla, J. et al. Engineering and adaptive evolution of Escherichia coli for D-lactate fermentation reveals GatC as a xylose transporter. Metabolic Engineering 14, 469–476 (2012).
Ishida, N. et al. Efficient production of L-Lactic acid by metabolically engineered Saccharomyces cerevisiae with a genome-integrated L-lactate dehydrogenase gene. Appl. Environ. Microbiol. 71, 1964–1970 (2005).
Cuéllar, A. D. & Webber, M. E. Cow power: the energy and emissions benefits of converting manure to biogas. Environ. Res. Lett. 3, 034002 (2008).
Cuéllar, A. D. & Webber, M. E. Wasted Food, Wasted Energy: The Embedded Energy in Food Waste in the United States. Environ. Sci. Technol. 44, 6464–6469 (2010).
Khmelenina, V. N., Kalyuzhnaya, M. G., Starostina, N. G., Suzina, N. E. & Trotsenko, Y. A. Isolation and Characterization of Halotolerant Alkaliphilic Methanotrophic Bacteria from Tuva Soda Lakes. Curr Microbiol 35, 257–261 (1997).
Eiteman, M. A. & Ramalingam, S. Microbial production of lactic acid. Biotechnol. Lett. 37, 955–972, doi: 10.1007/s10529-015-1769-5 (2015).
Abdel-Rahman, M. A., Tashiro, Y. & Sonomoto, K. Recent advances in lactic acid production by microbial fermentation processes. Biotechnol. Adv. 31, 877–902 (2013).
Linger, J. G. et al. Lignin valorization through integrated biological funneling and chemical catalysis. Proceedings of the National Academy of Sciences 111, 12013–12018 (2014).
Blazeck, J. et al. Metabolic engineering of Yarrowia lipolytica for itaconic acid production. Metabolic Engineering 32, 66–73 (2015), doi: 10.1016/j.ymben.2015.09.005.
Peralta-Yahya, P. P. et al. Identification and microbial production of a terpene-based advanced biofuel. Nat Commun 2, 483 (2011).
Malashenko, Y. R., Pirog, T. P., Romanovskaya, V. A., Sokolov, I. G. & Grinberg, T. A. Search for Methanotrophic Producers of Exopolysaccharides. Applied Biochemistry and Microbiology 37, 599–602 (2001).
Brantner, C. A., Remsen, C. C. & Owen, H. A. Intracellular localization of the particulate methane monooxygenase and methanol dehydrogenase in Methylomicrobium album BG8. Arch. Microbiol. 178, 59–64 (2002).
Bogorad, I. W. et al. Building carbon–carbon bonds using a biocatalytic methanol condensation cycle. Proceedings of the National Academy of Sciences 111, 15928–15933 (2014).
Ojala, D. S., Beck, D. & Kalyuzhnaya, M. G. Genetic Systems for Moderately Halo (alkali) philic Bacteria of the Genus Methylomicrobium. Methods Enzymol. 495, 99–118 (2011).
Laurens, L. M. L., Quinn, M., Van Wychen, S., Templeton, D. W. & Wolfrum, E. J. Accurate and reliable quantification of total microalgal fuel potential as fatty acid methyl esters by in situ transesterification. Anal Bioanal Chem 403, 167–178 (2012).
Acknowledgements
We would like to thank Michelle Reed, Kelsey Ramirez, Deb Hyman, Lisa Warner, Nick Sweeney and Qiang (John) Fei, all of the National Renewable Energy Laboratory, for technical assistance. Our thanks to Mary Lidstrom and Aaron Puri of the University of Washington for providing the 5GB1S strain and pAWP78 vector. This project was funded by the Office of Energy Efficiency and Renewable Energy, Bioenergy Technologies Office, WBS # 2.3.2.102, United States Department of Energy.
Author information
Authors and Affiliations
Contributions
C.A.H. and M.T.G. conceived and designed the experiments. C.A.H. and H.S. performed the experiments. C.A.H. and M.T.G. analyzed the data. N.D., P.T.P. and M.G.K. contributed intellectually and provided materials. C.A.H. and M.T.G. wrote the manuscript. All authors edited the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Henard, C., Smith, H., Dowe, N. et al. Bioconversion of methane to lactate by an obligate methanotrophic bacterium. Sci Rep 6, 21585 (2016). https://doi.org/10.1038/srep21585
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep21585
This article is cited by
-
Regulation of acetate tolerance by small ORF-encoded polypeptides modulating efflux pump specificity in Methylomonas sp. DH-1
Biotechnology for Biofuels and Bioproducts (2023)
-
Phylogenetic and metabolic diversity of microbial communities performing anaerobic ammonium and methane oxidations under different nitrogen loadings
ISME Communications (2023)
-
Microbial engineering strategies to utilize waste feedstock for sustainable bioproduction
Nature Reviews Bioengineering (2023)
-
Methane Based Continuous Culture of Methylosinus trichosporium for Production of Poly-3-hydroxybutyrate Using Membrane Recycle System
Biotechnology and Bioprocess Engineering (2023)
-
Methanol bioconversion in Methylotuvimicrobium buryatense 5GB1C through self-cycling fermentation
Bioprocess and Biosystems Engineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.