Abstract
Lipid rafts are membrane nanodomains that facilitate important cell functions. Despite recent advances in identifying the biological significance of rafts, nature and regulation mechanism of rafts are largely unknown due to the difficulty of resolving dynamic molecular interaction of rafts at the nanoscale. Here, we investigate organization and single-molecule dynamics of rafts by monitoring lateral diffusion of single molecules in raft-containing reconstituted membranes supported on mica substrates. Using high-speed interferometric scattering (iSCAT) optical microscopy and small gold nanoparticles as labels, motion of single lipids is recorded via single-particle tracking (SPT) with nanometer spatial precision and microsecond temporal resolution. Processes of single molecules partitioning into and escaping from the raft-mimetic liquid-ordered (Lo) domains are directly visualized in a continuous manner with unprecedented clarity. Importantly, we observe subdiffusion of saturated lipids in the Lo domain in microsecond timescale, indicating the nanoscopic heterogeneous molecular arrangement of the Lo domain. Further analysis of the diffusion trajectory shows the presence of nano-subdomains of the Lo phase, as small as 10 nm, which transiently trap the lipids. Our results provide the first experimental evidence of non-uniform molecular organization of the Lo phase, giving a new view of how rafts recruit and confine molecules in cell membranes.
Similar content being viewed by others
Introduction
Phase transition and domain formation in biological membranes are fascinating phenomena that play critical roles in many membrane functions1,2,3,4. Among the various phase separations, coexistence of liquid-ordered (Lo) and liquid-disordered (Ld) phases is of particular interest as it underlies the concepts of lipid rafts in biological membranes5,6. Lipid rafts are nanoscopic membrane domains enriched in saturated lipids of high melting temperature (e.g., sphingolipids) and cholesterol, which actively participate in various biological processes, including signaling transduction5, membrane trafficking6 and viral entry7. Previous studies have suggested that the relatively ordered molecular structure of lipid rafts can selectively recruit and trap lipids and membrane proteins of similar molecular properties8,9 and thus, rafts are postulated as local reaction centers for protein clustering and activations5,6. Characterization of lipid raft domains is therefore of great importance in order to understand fundamental regulation mechanism of membrane mobility and its relationship to membrane functions.
Since the molecular composition of cell membranes is highly diverse, inherent properties of lipid rafts have been extensively studied in simplified model membranes of ternary lipid compositions where the mimetic rafts of Lo phase coexists with non-raft Ld phase10,11,12,13. In this simple system, it has been found that the Lo phase consists largely of saturated lipids of high melting temperature and cholesterol, sharing many properties of lipid rafts in the cell membranes10. The term “liquid-ordered” is coined to describe a special phase of lipids with ordered and extended acyl chains, but it still exhibits the liquid-like lateral and rotational mobility11. Cholesterol plays a unique role in the formation of Lo phase by disturbing the translational order in the crystalline (gel) state of the lipids14. Despite many properties of the Lo phase have been learned15,16, their implications to rafts in cell membranes remain unclear. Previous measurements indicated that the putative rafts in cell plasma membranes are nano-sized5,6. Therefore, ideally, characterization of the Lo phase should be performed with nanometer spatial resolution so that it provides nanoscopic details that are more relevant to rafts in the cell membranes. Unfortunately, such nanoscopic measurement in aqueous environment has been challenging. Furthermore, biological membrane is a dynamic fluidic system in which molecules undergo fast lateral diffusion due to thermal fluctuations: single molecules diffuse by a few nanometers in one microsecond in the membrane. Therefore, interaction between individual membrane molecules with the Lo domains at the nanoscale is expected to be very transient and its detection requires high temporal resolution of microseconds. Finally, single-molecule sensitivity is another desired capability in order to observe short-lived and rare interactions that can be obscured via ensemble average. Putting together, investigation of molecular dynamics of rafts calls for a technique with single-molecule sensitivity, nanometer spatial resolution and microsecond temporal resolution, which has been technically challenging for existing techniques.
Scattering-based single-particle tracking (SPT) is a promising technique for high-speed localization of single membrane molecules. Using 40 nm gold nanoparticles (GNPs) as labels with bright-field detection, single lipids and membrane proteins were tracked in the plasma membrane with 17 nm spatial precision at 40 kHz from which membrane compartmentalization of hundreds of nanometers was discovered17. Recently, a novel concept of interferometric detection of scattering signal (iSCAT) has been introduced that shows promise of further improving the sensitivity and spatiotemporal resolution of scattering-based SPT18,19,20,21,22,23,24,25. The interferometric detection makes it possible to circumvent the limitation of signal-to-noise ratio (SNR) set by the electronic noise of high-speed detectors, reaching the optimal sensitivity required for super localization of very small particle via intrinsic scattering24. By iSCAT microscopy, detection and localization of single nanoparticles, ranging from metallic particles to virus particles have been demonstrated20,21,22,23,24,25. The stable and unsaturated scattering signal allows for long-term detection with unlimited observation time. In this work, we use ultrahigh-speed iSCAT SPT at 50 kHz to investigate single-molecule diffusion in raft-containing reconstituted bilayer membranes. The capability of tracking single lipid molecules at the nanoscale via 20 nm GNP tags without introducing labeling artifacts is carefully examined. The iSCAT microscopy allows for continuous observation of single lipid molecules partitioning into and escaping from the raft-mimetic Lo domains with nanometer spatial precision and microsecond temporal resolution. From the detailed trajectories of single lipids exploring the raft-containing membranes, we observe deviation from Brownian motion of lipid molecules in the Lo domain in the microsecond timescale, showing the first experimental evidence of the nanoscopic heterogeneous molecular arrangement of the Lo phase. Our finding also reveals the existence of nano-subdomains in the Lo phase that strongly influence motion of molecules. These nano-subdomains are expected to be critical for recruiting and confining raft-associated molecules in lipid rafts of cell membranes.
Results
High-speed nanoscopic tracking of single lipid molecules by iSCAT microscopy
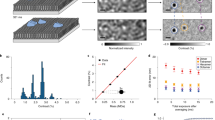
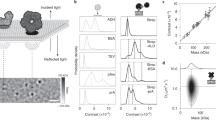
Small GNP of 20 nm in diameter is used to label lipid molecule in the membrane via streptavidin and biotin for optical imaging and tracking (Fig. 1a). The iSCAT microscope (see Methods and Fig. S1) captures scattering image of single GNP via interferometry at 50 kHz. Benchmark measurements were performed by tracking single probe lipid (biotin-cap-DPPE) diffusing laterally in a homogeneous DiPhyPC membrane of Ld phase prepared on an atomically flat mica substrate (see Methods). We chose DiPhyPC instead of other commonly used unsaturated phospholipids in order to avoid photo-oxidation effect of the lipid molecules that may modify the mobility of the lipids26. Multiple particles can be simultaneously visualized on the membrane (see Movie S1) and their diffusion trajectories were obtained by SPT (see Methods). Figure 1b shows a representative diffusion trajectory consisting of more than 155,000 steps. The iSCAT image of 20 nm GNP has a SNR ~20 (Fig. 1c), providing a spatial localization precision of 3 nm (Fig. S2). The long and accurate trajectory together with 20 μs temporal resolution allows us to analyze its diffusion characteristics with high accuracy. We observe perfect Gaussian step size distributions in two lateral directions, implying the measurement provides sufficient data for statistical analysis (Fig. 1d). To assure the labeling scheme and the measurement are reliable, we collected a large amount of diffusion data of 62 trajectories (each of which is longer than 6,000 steps) and calculated their individual time-averaged mean squared displacements (MSD) as a function of time interval Δt and their diffusion rates  (see Fig. 1ef and Methods). We measured similar diffusion rates from individual particles (1.48 ± 0.39 μm2 s−1), suggesting high uniformity of the membrane and the particles. To analyze the diffusion characteristics, the MSD was fitted by a model of anomalous diffusion27:
(see Fig. 1ef and Methods). We measured similar diffusion rates from individual particles (1.48 ± 0.39 μm2 s−1), suggesting high uniformity of the membrane and the particles. To analyze the diffusion characteristics, the MSD was fitted by a model of anomalous diffusion27:
Visualization of single lipid diffusion in homogeneous bilayer membrane via iSCAT SPT with 3 nm spatial precision at 50 kHz.
(a) Schematics of 20 nm GNP attaching to single lipid molecule at the headgroup in a membrane for high-speed iSCAT tracking. The particle and the bilayer membrane are plotted at the same scale. (b) Diffusion trajectory of single lipid molecule in a homogeneous membrane recorded at 50 kHz for 3.1 s (see also Movie S1). (c) An iSCAT snapshot of a 20 nm GNP diffusing on the membrane captured at 50 kHz. The spot size is ~300 nm (full width at half maximum) and the SNR is 20. The center of the spot is determined by SPT and the precision of determining the center position is 3 nm. (d) Step size distributions in two lateral directions at 20 μs timescale both show perfect Gaussian distributions. (e) Time-average MSD of 62 diffusion trajectories (grey lines) and their ensemble average (thick black line), showing uniform Brownian diffusion. The red line indicates the linear dependence of the MSD on the time interval Δt as for perfect Brownian diffusion. (f) Histogram of the measured diffusion rate ( = 1.48 ± 0.39 μm2 s−1). (g) Histogram of measured anomalous exponent α. Narrow peak around α = 1 indicates the diffusion in homogeneous membrane is nearly Brownian.
= 1.48 ± 0.39 μm2 s−1). (g) Histogram of measured anomalous exponent α. Narrow peak around α = 1 indicates the diffusion in homogeneous membrane is nearly Brownian.

where  is the anomalous transport coefficient,
is the anomalous transport coefficient,  is the anomalous exponent and
is the anomalous exponent and  is an offset due to the dynamic localization error28. The histogram of the fitted values of
is an offset due to the dynamic localization error28. The histogram of the fitted values of  shows a narrow peak at 0.99 (Fig. 1g), implying the diffusion was nearly Brownian (
shows a narrow peak at 0.99 (Fig. 1g), implying the diffusion was nearly Brownian ( = 1 when diffusion is perfectly Brownian). The observed Brownian diffusion indicates that the membrane is homogeneous over the length scales from nanometers to micrometers and the supporting substrate does not introduce spatially heterogeneous effects to the membrane. Brownian diffusion of biotin-cap-DPPE was also observed in homogeneous membranes of unsaturated phospholipids DOPC via GNP labeling and iSCAT SPT (data not shown). We further verified that the mass loading of 20 nm GNP was negligible, as the GNP showed the same diffusion rate as single dye-labeled streptavidin when labeled to biotin-cap-phospholipid (Fig. S3). More about the validity of tracking single lipid molecules by localizing GNP labels are discussed in Supplementary Information.
= 1 when diffusion is perfectly Brownian). The observed Brownian diffusion indicates that the membrane is homogeneous over the length scales from nanometers to micrometers and the supporting substrate does not introduce spatially heterogeneous effects to the membrane. Brownian diffusion of biotin-cap-DPPE was also observed in homogeneous membranes of unsaturated phospholipids DOPC via GNP labeling and iSCAT SPT (data not shown). We further verified that the mass loading of 20 nm GNP was negligible, as the GNP showed the same diffusion rate as single dye-labeled streptavidin when labeled to biotin-cap-phospholipid (Fig. S3). More about the validity of tracking single lipid molecules by localizing GNP labels are discussed in Supplementary Information.
Diffusion of single lipid molecules in raft-containing membranes
Using high-speed iSCAT SPT, we investigated diffusion of single lipid molecules in raft-containing membranes at high spatiotemporal resolution. The raft-containing bilayer membranes were prepared from a lipid mixture of DiPhyPC/DPPC/cholesterol (40:40:20 mol%) and a trace amount of lipophilic dye DiI-C18 (see Methods). Micro-sized Lo domains were observed via fluorescence (Fig. 2a) from which the domain boundaries were determined (Fig. 2b). We noticed that the Lo domains can slowly drift, rearrange and merge over time as reported previously29. Within our short observation time of one minute, no changes of the shape and location of the Lo domains were observed in the fluorescence image. Interestingly, the Lo domains also appear in iSCAT image with low contrast (~1%) due to the slight mismatch of refractive index of the Ld and Lo phases, as reported recently30. We then added GNPs to label the probe lipid (biotin-cap-DPPE) and followed their motions via iSCAT SPT at 50 kHz. Each high-speed iSCAT recording took a few seconds and we were able to record many videos within one minute before the shape of the Lo domain changed noticeably. Note that the precise localization of the GNP was performed on background-free iSCAT images where the non-uniform stationary background due to the illumination, the membrane domains and the heterogeneous pixel responses was removed (Methods). The recorded diffusion trajectories of the single probe lipids were superimposed on the fluorescence image of the same sample area indicating the location of the raft-mimetic Lo domains (Fig. 2c). The stable scattering signal from the GNP provides the opportunity to capture precise and continuous diffusion trajectories of single probe lipids exploring both the Ld and Lo domains and therefore enables investigation of how the Lo domain influences diffusion of lipid molecules. In our recorded data, more than 90% of the trajectories explore both the Ld and Lo domains. We emphasize that the capability of observing single particle exploring both domains is valuable as it allows for direct comparison of diffusion characteristics of the same particle in both domains and therefore possible systematic artifacts introduced by the inevitable differences between individual probe nanoparticles (e.g., size, shape, orientation of the crosslinker molecule, etc.) can be avoided. Figure 2d shows a segment of the diffusion trajectory of a lipid molecule repeatedly escaping from and entering the Lo domain (see also Movie S2). Close-up views of the trajectories in Fig. 2e,f reveal distinct diffusion behaviors in the Lo and Ld domains. It is apparent that the transient diffusion rate of the probe lipid changes abruptly when it crosses the domain boundary (Fig. 2g, see Methods for the calculation of transient diffusion rate). From this trajectory, we observe fast diffusion of 1.19 μm2 s−1 in the Ld domain and slow diffusion of 0.13 μm2 s−1 in the Lo domain. Our experiment also clearly illustrates the fact that while the morphology of the Lo domain does not change within the observation time, individual molecules in the membrane undergo rapid diffusive motion and they even constantly cross the domain boundary.
Continuous observation of single lipid molecule diffusion in raft-containing membrane at 50 kHz.
(a) Fluorescence raw image of micron-sized raft-mimetic Lo domains (appear as dark areas) in the membrane. (b) Areas of the Lo and Ld domains defined from the fluorescence image. (c) Ten representative diffusion trajectories of single lipid molecules in the raft-containing membrane recorded at 50 kHz where each of them explores both the Ld and Lo domains. The parts of trajectories in the Ld and Lo domains are colored in red and blue respectively, while the domain boundaries are marked by the dashed lines. (d) A segment of diffusion trajectory of a lipid molecule repeatedly leaving from and entering the raft domain (see Movie S2). Domain boundary is plotted as the black line. The lipid was initially in the Lo domain (marked as I); it left the Lo domain and re-entered the Lo domain shortly (marked as II) and it left the Lo domain again at the end. (e,f) Close-up views of the trajectory shown in (d), in the Lo domain (e) and in the Ld domain (f) respectively, showing distinct diffusion behaviors. (g) Transient diffusion rate of the lipid calculated from the trajectory in (d). The diffusion rates in the Lo and Ld domains are very different: slow diffusion of 0.13 μm2 s−1 and fast diffusion of 1.19 μm2 s−1 were measured in the Lo and Ld domains respectively.
Anomalous molecular diffusion in the Lo phase
We recorded 166 and 88 diffusion trajectories in the Ld and Lo domains respectively and calculated their individual time-averaged MSD (Fig. 3). Each trajectory is longer than 1,000 steps. In total, more than 2.3 × 106 and 1.7 × 106 steps were measured in the Ld and Lo domains respectively, providing a large amount of statistical data for accurate characterization of diffusion. The MSD measured in the Ld domain scales linearly with the time interval Δt, indicating the diffusion is Brownian. On the other hand, the MSD measured in the Lo domain shows subdiffusion in the microsecond timescale. A crossover from subdiffusion to apparent Brownian motion in the Lo domain was observed at ~1 ms with the MSD value of ~ μm2, i.e., 20 × 20 nm2. It indicates that the molecular diffusion in the Lo domain experiences heterogeneity over a length scale <20 nm in the microsecond timescale. At longer timescale of milliseconds, the effect of heterogeneity is averaged out and thus apparent Brownian motion is observed. We stress that experiment without sufficient temporal resolution would not be able to detect subdiffusion in the Lo domain but measure apparent Brownian diffusion of slower diffusion rate. We performed the same iSCAT SPT measurement at a lower speed of 1 kHz under otherwise identical conditions and the diffusion in the Ld and Lo domains was indeed both Brownian (Fig. S4 and Movie S3). The observed anomalous subdiffusion in microsecond timescale at a length scale <20 nm is truly due to the nanoscopic substructures in the Lo domain; it is not systematic artifact (due to GNP labeling or the transient interaction with the supporting substrate) because from the MSD data (Fig. 1e), Brownian diffusion is measured in the benchmark experiment of homogeneous membranes at a length scale as small as 10 × 10 nm2 where no confinement is detected. The histograms of the anomalous exponent
μm2, i.e., 20 × 20 nm2. It indicates that the molecular diffusion in the Lo domain experiences heterogeneity over a length scale <20 nm in the microsecond timescale. At longer timescale of milliseconds, the effect of heterogeneity is averaged out and thus apparent Brownian motion is observed. We stress that experiment without sufficient temporal resolution would not be able to detect subdiffusion in the Lo domain but measure apparent Brownian diffusion of slower diffusion rate. We performed the same iSCAT SPT measurement at a lower speed of 1 kHz under otherwise identical conditions and the diffusion in the Ld and Lo domains was indeed both Brownian (Fig. S4 and Movie S3). The observed anomalous subdiffusion in microsecond timescale at a length scale <20 nm is truly due to the nanoscopic substructures in the Lo domain; it is not systematic artifact (due to GNP labeling or the transient interaction with the supporting substrate) because from the MSD data (Fig. 1e), Brownian diffusion is measured in the benchmark experiment of homogeneous membranes at a length scale as small as 10 × 10 nm2 where no confinement is detected. The histograms of the anomalous exponent  and the diffusion rate
and the diffusion rate  of individual trajectories measured in the Ld and Lo domains are plotted in Fig. S5, where the diffusion in the Lo domain is slow (
of individual trajectories measured in the Ld and Lo domains are plotted in Fig. S5, where the diffusion in the Lo domain is slow ( = 0.24 ± 0.12 μm2 s−1) and subdiffusive (
= 0.24 ± 0.12 μm2 s−1) and subdiffusive ( = 0.75 ± 0.16), while diffusion in the Ld domains is fast (
= 0.75 ± 0.16), while diffusion in the Ld domains is fast ( = 1.43 ± 0.50 μm2 s−1) and much more Brownian (
= 1.43 ± 0.50 μm2 s−1) and much more Brownian ( = 0.92 ± 0.08). The measured diffusion rates in the two phases are similar to what were measured previously with dye molecules DiD by fluorescence correlation spectroscopy (FCS)29. Nevertheless, we note that the diffusion rate varies between different types of probe molecules, possibly because of their different charges resulting in different strength of electrostatic interaction with the substrate29. Anomalous subdiffusion was also observed in the membranes of ternary lipid mixture of DOPC/brain sphingomyelin/cholesterol (40:40:20 mol%) (Fig. S6), suggesting the heterogeneous molecular organization in the Lo phase is general, not lipid-specific.
= 0.92 ± 0.08). The measured diffusion rates in the two phases are similar to what were measured previously with dye molecules DiD by fluorescence correlation spectroscopy (FCS)29. Nevertheless, we note that the diffusion rate varies between different types of probe molecules, possibly because of their different charges resulting in different strength of electrostatic interaction with the substrate29. Anomalous subdiffusion was also observed in the membranes of ternary lipid mixture of DOPC/brain sphingomyelin/cholesterol (40:40:20 mol%) (Fig. S6), suggesting the heterogeneous molecular organization in the Lo phase is general, not lipid-specific.
Time-averaged MSD measured in the Ld and Lo domains from 20 μs to 20 ms.
166 and 88 trajectories were measured from the Ld and Lo domains respectively and their MSDs were individually calculated and plotted. The red and blue thick lines are their corresponding ensemble averages. In this double logarithmic plot of MSD, the slope represents the value of anomalous exponent  . Two black solid lines show the slopes of 0.75 and one. Single lipid molecules diffuse freely in the Ld domain, as the slopes are nearly one. On the other hand, the lipid molecules undergo anomalous subdiffusion in the Lo domains at the microsecond timescale, indicating the molecular diffusion in the Lo domain experiences nanoscopic heterogeneity within microseconds. The effect of heterogeneity is averaged out at longer timescale and thus apparent Brownian motion is observed at longer time interval (>1 ms).
. Two black solid lines show the slopes of 0.75 and one. Single lipid molecules diffuse freely in the Ld domain, as the slopes are nearly one. On the other hand, the lipid molecules undergo anomalous subdiffusion in the Lo domains at the microsecond timescale, indicating the molecular diffusion in the Lo domain experiences nanoscopic heterogeneity within microseconds. The effect of heterogeneity is averaged out at longer timescale and thus apparent Brownian motion is observed at longer time interval (>1 ms).
Transient confinements of saturated lipids in nano-subdomains of the Lo phase
To gain more insights of the observed subdiffusion, we looked carefully into the diffusion trajectories in the Lo domains. As shown in Fig. 2e, there are numerous clusters of localizations in the trajectories measured in the Lo domain. Further analysis shows that many of these clusters are transient confinements of the lipid in nano-sized zones (see Methods)31. We analyzed the size of confinements and the residence time in each confinement from all trajectories measured in the Lo domains (see Methods, Fig. 4ab). We set relatively high criteria (99% confidence) for detecting transient confinement so that false detection of stochastic confinement-like Brownian motion is mostly avoided (Fig. 4ab). Our analysis shows that there are considerable amount of nano-confinements in the Lo phase. The average residence time is 0.62 ms and the average size of the confinement is 32 nm in diameter. We note that there could be much more confinements that are very transient and of very small sizes and their reliable detection requires measurements of even higher spatiotemporal resolution. In fact, our MSD data of the Lo domains (Fig. 3) show anomalous diffusion spanning the length scale from (6 nm)2 to (20 nm)2, implying the membrane organization of the Lo domain is heterogeneous over the nanoscale. As the probe lipid being tracked in our experiment is the saturated lipid biotin-cap-DPPE, these nano-subdomains which transiently trap the probe lipid are likely to be clusters of closely packed saturated lipids DPPC of similar molecular structures and properties. Based on our experimental observation, we propose a potential model of nanoscopic molecular arrangement in the Lo domain (Fig. 4c): inside the Lo domain, there exist numerous subdomains smaller than tens of nanometers that strongly influence the molecular diffusion. This model has been suggested by the nuclear magnetic resonance (NMR) spectroscopy studies showing that the formation of the Lo phase is mainly governed by the interaction between saturated lipids of high melting temperature (e.g., sphingolipids) and therefore nano-sized clusters may exist in the Lo domain32. Molecular dynamic simulation has also predicted anomalous diffusion and substructures of saturated lipids in the Lo phase at nanosecond timescale33,34.
Detection of transient confinements in the Lo domain due to nanoscopic substructures.
(a) Size distributions of the nano-confinements (in diameter) detected from the trajectories measured in the Lo domains (in blue) and from the simulated trajectories of Brownian diffusion (in gray). See Methods for the details of detection of nano-confinements. Considerable amount of nano-confinements of 32 ± 10 nm are detected in the Lo domains. (b) Distributions of the residence time in each confinement. The blue histogram shows the experimental results measured from the Lo domains, while the gray histogram shows the control of simulated Brownian diffusion. Single lipid molecules are transiently confined in nano-sized zones inside the raft Lo domain for less than 1 ms. (c) Schematics of the proposed nanoscopic molecular dynamics and organization in the Lo domain. The molecular arrangement in the Lo domain is not homogeneous at the nanoscale. There exist numerous subdomains of tens of nanometers, potentially clusters of closely packed saturated lipids of high melting temperature. These subdomains strongly influence diffusion of molecules. For illustration purpose, an artificial diffusion trajectory is plotted showing fast and free diffusion in the Ld phase (red) and transient confinements in the nano-subdomains of the Lo phase (blue).
Discussion
The fact that the molecular organization in the Lo phase is heterogeneous at the nanoscale is striking. It has been known that the cholesterol in the Lo phase disturbs the packing of lipids that are otherwise in gel phase14. The phase diagram of lipid mixture is changed after adding cholesterol, revealed by NMR spectroscopy and differential scanning calorimetry (DSC) measurements35,36. The intermediate phase state induced by the cholesterol, e.g., the Lo phase, is speculated to play critical role in biological membranes via reducing the tendency of the lipids to form separate Ld and gel phases37 and at the same time stabilizing putative functional nanodomains in cell membranes (rafts)38. However, there has been no experimental data showing how cholesterol disturbs the packing of lipids at the nanoscale and at the same time sustains the integrity of the Lo domain. In this work, we show explicitly that there are numerous nano-subdomains in the Lo phase. Although we do not have further evidence to confirm exactly what those nano-subdomains are consisted of at the present stage, based on our observation of nano-confinements in the trajectories, they are likely to be densely packed saturated lipids DPPC (the lipid that has higher melting temperature in the ternary mixture) which have similar molecular structures as our probe lipids DPPE. Cholesterol and a low density of DiPhyPC (the lipid of lower melting temperature) fill the gaps between the nano-subdomains. Molecular dynamic simulation has also indicated the existence of nano-clusters of saturated lipids in the Lo domains34.
We notice that there have been several measurements of the topography of the Ld and Lo coexisting supported bilayer membranes by using atomic force microscopy (AFM)29,39,40. However, in those AFM studies, no nano-substructures were observed in the Lo phase. We think it may be because of the following reasons. First, detection of the nano-substructures of the Lo phase requires high-resolution AFM measurement. We estimate the height difference between the nano-substructures and the surrounding area to be <0.8 nm, considering the height difference between Ld and Lo phases is ~0.8 nm29,39,40. Regarding the lateral size, according to our MSD data and transient confinement analysis, the size of the substructures is smaller than tens of nanometers. As mentioned in the text, we believe that there are smaller subdomains that are not detected by the technique presented in this work. In order to explicitly map the dense and fine substructures of the Lo phase, resolution of subnanometer may be required. Second, the nano-substructures may be dynamic which requires high-speed AFM to resolve them. Third, the nano-substructures may be fragile and therefore perturbation introduced by the AFM tip during measurement should be minimized. As the sensitivity and spatiotemporal resolution of AFM technique has been improved rapidly in recent years, high-resolution and high-speed AFM with subnanometer resolution41 and other near field techniques show promise of providing complementary evidence of the molecular organization of the Lo phase.
It is worth noting that a very similar study was conducted aiming to resolve the nanoscopic molecular organization of the Lo phase by using a superresolution optical technique of stimulated emission depletion – fluorescence correlation spectroscopy (STED-FCS)42. That measurement did not detect anomalous diffusion in the Lo phase and concluded that the slow diffusion of the Lo phase is mainly due to the tight molecular packing rather than causing anomalous subdiffusion over a length scale >40 nm. The high-speed iSCAT SPT data presented in this work further extend the detection resolution down to a few nanometers and show that at the smaller length scale (<30 nm) there are nano-substructures of the Lo phase that strongly influence lipid diffusion, causing subdiffusion via transient confinements. High spatiotemporal resolution is ultimately necessary in order to resolve nanoscopic structures in dynamic membrane systems.
While our measurements were performed in the Lo domains of model membranes, the measured inherent molecular interaction and the resulting molecular organization at the nanoscale are expected to be closely related to that of lipid rafts in cell membranes. It has been shown that biological membranes are heterogeneous at the nanoscale43,44 and the size of lipid rafts could be smaller than tens of nanometers5,6. Considering the nano-sized subdomains of the Lo phase observed in our experiments, molecular organization of rafts in cell membranes could be highly heterogeneous. Also, we reveal that molecular dynamics in the Lo phase is strongly influenced by the nano-subdomains and therefore these nano-substructures may readily be responsible for determining the partition of molecules in lipid rafts. The raft-associated proteins could be trapped between subdomains of densely packed saturated lipids and surrounded by high density of cholesterol. The nanoscopic membrane environment created by the nano-subdomain of rafts is thus unique, which illustrates the essentials of lipid rafts of Lo phase whose roles cannot be fulfilled solely by the gel-phase or Ld-phase domains.
In summary, by using ultrahigh-speed iSCAT optical technique together with small nanoparticles as labels, we have investigated single molecule diffusion in bilayer model membranes with nanometer spatial precision and microsecond temporal resolution. Diffusion of single nanoparticles on supported bilayer membranes has been studied by high-speed iSCAT microscopy before22,23. In the previous work, anomalous diffusion, multimobility and also nano-confinements of the particles were observed due to the multiple lipid binding sites on the particle, clustering of the lipids and the substrate effects, which complicate the investigation of single-molecule dynamics in the membrane. Addressing these issues, in this work, we promote the probability of monovalent binding by saturating the multiple binding sites on the particle. In addition, we prepared bilayer membranes on atomically flat mica substrate, which significantly reduces the heterogeneous substrate effect. We also used a well-behaved probe lipid, biotin-cap-DPPE, that does not form clusters and diffuses freely in homogeneous membrane without transient confinements. Motion of the single particles faithfully represents the diffusion of single molecules with nanometer precision as supported by the data of Brownian diffusion observed in the homogeneous membrane down to microsecond timescale. We have further employed iSCAT SPT to investigate molecular dynamics and nanoscopic organization of raft-mimetic Lo domains. We have found transient confinements of lipid molecules due to the inherent nano-substructures of the Lo domains that have not been resolved before, providing a new view of molecular organization in the Lo phase at the nanoscale. These nano-subdomains of the Lo phase are likely to be responsible for governing the selection, partition and confinement of raft-associated molecules in raft domains of cell membranes.
Methods
iSCAT microscope
In iSCAT microscope, illumination in wide field was made by focusing a 532 nm laser beam (Finesse Pure, Laser Quantum) at the back focal plane of a high NA microscope objective (UPLSAPO 100X, NA 1.4 Olympus). Part of the excitation (~0.6%) was reflected at the mica-water interface due to the refractive index mismatch, serving as a coherent reference beam for iSCAT imaging. The transmitted light illuminated the GNP. The backscattered signal from the GNP together with the reflective reference beam was collected by the same objective and imaged onto a high-speed CMOS camera (Phantom v711, Vision Research). The overall optical magnification of iSCAT imaging was 416.7, corresponding to pixel resolution of 48 × 48 nm2 per pixel. Destructive interference between the scattering signal of GNP and the reference beam results in a dark spot in the iSCAT image. We recorded iSCAT videos at 50 kHz for a few seconds during which GNPs diffuse on the membrane. Excitation intensity was ~10 kW cm−2, causing a minor temperature increase on the surface of GNP by ~0.3 K. The iSCAT microscope was equipped with epi-fluorescence imaging in parallel. Fluorescence images were taken by a sensitive camera (EMCCD; iXon Ultra 897, Andor) at 1 Hz simultaneously with iSCAT imaging. The overall optical magnification of fluorescence imaging was 166.7, corresponding to pixel resolution of 96 × 96 nm2 per pixel. The iSCAT and fluorescence images were aligned by localizing immobile 100 nm fluorescent particles on coverglass simultaneously in both channels before the membrane experiments. The relative sub-pixel displacement between the iSCAT and fluorescence images is corrected in the final overlay of the fluorescence image of the membrane domains and the iSCAT trajectories. The precision of the overlay is ~4 nm, limited by the localization precision of the fluorescent particles.
Preparation of reconstituted supported bilayer membranes on mica and GNP labeling
We used mainly two types of lipid bilayer membranes in this work. The raft-containing membrane consists of 40:40:20 mol% of 1,2-diphytanoyl-sn-glycero-3-phosphocholine (DiPhyPC), 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and cholesterol (Chol). The homogeneous membrane was made by DiPhyPC. Both types of membranes composed of 1 mol% biotinylated probe lipid analogues (1,2-dipalmitoyl-sn-glycero-3-phosphoethanolamine-N-(cap biotinyl); biotin-cap-DPPE) and 0.1 mol% fluorescent carbocyanine dyes. Since both phospholipids (DiPhyPC and DPPC) are saturated, the membranes have less chance to be photo-oxidized during measurement. These lipid molecules and cholesterol were purchased from Avanti (Avanti Polar Lipids Inc., Alabaster, Alabama, USA.) The orange—red fluorescent dye, 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate (DiI-C18), was purchased from Life Science. Note that DiI-C18 is a lipophilic membrane stain that prefers to partition in the Ld phase of membranes. Therefore, the dark regions of the fluorescence images represented Lo domains.
The supported lipid bilayers (SLBs) were formed by vesicles fusion method45. All lipids were dissolved in chloroform and they were dried to a film by nitrogen and kept under vacuum for at least 1 hour right before use. The dried lipid film was hydrated to a lipid concentration of 1 mg mL−1 in bilayer buffer solution (10 mM HEPES, 200 mM NaCl2 and 2 mM CaCl2, PH 7.0) for 1 hour at 60 °C. Multilamellar vesicles (MLVs) were then formed by vortexing the lipid solution. This cloudy solution was diluted by buffer solution to 0.25 mg mL−1 and further sonicated for 10 minutes at 30% power in a tip sonicator at 60 °C (Q700, Qsonica LLC., USA). Through this process, the MLVs were broken into small unilamellar vesicles (SUVs). Next, the lipid solution was centrifuged for 20 minutes at 16,000 g and the SUVs in the supernatant were carefully extracted without the debris and the remaining MLVs at the bottom. The clear solution of SUVs were stored at 4 °C and used within 48 hours.
To form SLBs, a mica disc (Ted Pella Inc, USA) was first glued on a cleaned coverslip with the optical adhesive (Norland Products Inc, USA). Then, the SUV solution was introduced to a freshly cleaved thin mica disc (a few microns thick) in a dish-type chamber (Live Cell Instrument, Seoul, Korea) for at least 90 minutes at 60 °C. The SLBs were gently washed with buffer solution to remove excess SUVs. Note that all the procedures were done at room temperature while preparing homogeneous membranes. Finally, 20 nm GNPs functionalized with streptavidin (BBI Solutions, Cardiff, U.K.) were added to the SLB for labeling the biotinylated probe lipid analogue at the hydrophilic headgroup. The streptavidin as the crosslinker is a tetramer that is able to bind to more than one biotin-cap-lipid in the membrane. In order to make monovalent labeling more probable, we estimated the number of available streptavidin binding sites on the GNP and then saturated them by mixing an excess amount of biotin molecules into the GNP colloidal solution before introducing them to membrane labeling. Stochastically, there should be one or no available binding site for most GNPs.
Single-particle tracking (SPT)
In order to localize the GNP with nanometer spatial precision, it is necessary to start with a clean iSCAT image of the GNP with a highly uniform background. Therefore, the inhomogeneous pixel response of the CMOS camera, the inevitably non-uniform light illumination and the weak iSCAT signal of the membrane domains need to be removed. These effects are relatively stationary within our observation timescale, especially compared with the fast diffusing GNP on a membrane. We extracted a stationary background from a stack of images by calculating the temporal median value of each pixel of the image22. Background-free iSCAT images were obtained after correcting the extracted stationary background from the raw images. After background removal, a diffraction-limited point spread function as a dark spot of a GNP can be clearly visualized. The center of the dark spot in every frame was determined by fitting the measured point spread function with a two-dimensional Gaussian function with a home-written code in MATLAB. Diffusion trajectories were obtained by connecting the corresponding localizations in the consecutive frames. Typical localization precision of single 20 nm GNP diffusing on the membrane recorded at 50 kHz is 3 nm (see Fig. S2).
Calculation of time averaged MSD for characterizing diffusion behaviors
For every trajectory, the two-dimensional MSD as a function of time interval Δt ranging from 20 μs to 20 ms was calculated according to

where  is the position of the particle at time
is the position of the particle at time  and
and  is the length of the trajectory. The diffusion rate,
is the length of the trajectory. The diffusion rate,  , is calculated from the first two data points of the MSD46, which refers to the microscopic diffusion rate measured at 20 μs time interval.
, is calculated from the first two data points of the MSD46, which refers to the microscopic diffusion rate measured at 20 μs time interval.
To analyze whether diffusion deviates from Brownian motion, the MSD data were fitted by a model of anomalous diffusion as shown in Eq. (1) where  , the anomalous transport coefficient,
, the anomalous transport coefficient,  , the anomalous exponent and
, the anomalous exponent and  , an offset are the three fitting parameters. Notice that the offset
, an offset are the three fitting parameters. Notice that the offset  in the MSD is the result of dynamic localization uncertainty, determined by the localization precision of the particle, the diffusion rate and the exposure time28. Error in estimating the value of
in the MSD is the result of dynamic localization uncertainty, determined by the localization precision of the particle, the diffusion rate and the exposure time28. Error in estimating the value of  can cause misinterpretation of the data for anomalous diffusion, which is especially critical in high-speed measurements where the true displacements can be comparable to the dynamic localization uncertainty. We found that the most reliable way of finding the value of
can cause misinterpretation of the data for anomalous diffusion, which is especially critical in high-speed measurements where the true displacements can be comparable to the dynamic localization uncertainty. We found that the most reliable way of finding the value of  is to fit it as a free parameter for individual MSD. After subtracting the offset from the MSD, the double logarithmic plot of the corrected MSD versus time interval has a slope of
is to fit it as a free parameter for individual MSD. After subtracting the offset from the MSD, the double logarithmic plot of the corrected MSD versus time interval has a slope of  . Based on the value of
. Based on the value of  , the diffusion characteristics of the particle can be classified:
, the diffusion characteristics of the particle can be classified:  for Brownian diffusion,
for Brownian diffusion,  and
and  indicating sub- and superdiffusion, respectively.
indicating sub- and superdiffusion, respectively.
Furthermore, to statistically elaborate the dynamic motion of multiple particles we ensemble averaged the MSD data by weighting the data according to the length of each trajectory,

Here, MSDi(Δt) is the MSD for the  th trajectory with
th trajectory with  steps and
steps and  (Δt) is the ensemble averaged MSD of N trajectories with
(Δt) is the ensemble averaged MSD of N trajectories with  steps in total.
steps in total.
To estimate a transient diffusion constant, we chopped the original trajectory into segments of 146 steps and computed the diffusion rate from the MSD of each segment.
Detection of transient confinements
Transient confinements occurring in the Lo domains were detected by the algorithm introduced by Simson et al.31. Briefly, we calculate the probability level L of a lipid of diffusion rate  staying within a region of radius
staying within a region of radius  within a time window
within a time window  as
as

where  is defined by the point in the time window with the largest displacement from the starting point. In our analysis, we chose the time window to be 25 steps. Transient confinements were defined as the probability level
is defined by the point in the time window with the largest displacement from the starting point. In our analysis, we chose the time window to be 25 steps. Transient confinements were defined as the probability level  surpassed a critical probability level
surpassed a critical probability level  . In our work, the value of
. In our work, the value of  was chosen to be one, corresponding to 99% confidence that the detected transient confinement is indeed restricted non-Brownian motion. The residence time in a confinement is defined as the duration when
was chosen to be one, corresponding to 99% confidence that the detected transient confinement is indeed restricted non-Brownian motion. The residence time in a confinement is defined as the duration when  is successively larger than
is successively larger than  . In order to minimize the disturbance of inevitable localization errors in the detection, we calculate the moving average of the raw probability level as a function of time with an averaging window of 10 steps. Following the procedures described above, we reliably detected a large number of events of transient confinement in the Lo domain. To further affirm the detected confinements in the Lo domains are not rare stochastic behavior of Brownian motion, we performed the identical detection with the computer-generated Brownian diffusion of the same trajectory length, diffusion rate
. In order to minimize the disturbance of inevitable localization errors in the detection, we calculate the moving average of the raw probability level as a function of time with an averaging window of 10 steps. Following the procedures described above, we reliably detected a large number of events of transient confinement in the Lo domain. To further affirm the detected confinements in the Lo domains are not rare stochastic behavior of Brownian motion, we performed the identical detection with the computer-generated Brownian diffusion of the same trajectory length, diffusion rate  and localization errors. We confirmed that the detected transient confinements from simulated Brownian motion is negligible, indicating that the confinements measured in the Lo domains were real.
and localization errors. We confirmed that the detected transient confinements from simulated Brownian motion is negligible, indicating that the confinements measured in the Lo domains were real.
Additional Information
How to cite this article: Wu, H.-M. et al. Nanoscopic substructures of raft-mimetic liquid-ordered membrane domains revealed by high-speed single-particle tracking. Sci. Rep. 6, 20542; doi: 10.1038/srep20542 (2016).
References
Hurley, J. H., Boura, E., Carlson, L.-A. & Różycki, B. Membrane budding. Cell 143, 875–887 (2010).
Surma, M. A., Klose, C. & Simons, K. Lipid-dependent protein sorting at the trans-Golgi network. BBA – Mol. Cell Biol. L. 1821, 1059–1067 (2012).
Simons, K. & Toomre, D. Lipid rafts and signal transduction. Nat. Rev. Mol. Cell Biol. 1, 31–39 (2000).
Fessler, M. B. & Parks, J. S. Intracellular lipid flux and membrane microdomains as organizing principles in inflammatory cell signaling. J. Immunol. 187, 1529–1535 (2011).
Lingwood, D. & Simons, K. Lipid rafts as a membrane-organizing principle. Science 327, 46–50 (2010).
Simons, K. & Gerl, M. J. Revitalizing membrane rafts: new tools and insights. Nat. Rev. Mol. Cell Biol. 11, 688–699 (2010).
Chazal, N. & Gerlier, D. Virus entry, assembly, budding and membrane rafts. Microbiol. Mol. Biol. Rev. 67, 226–237 (2003).
Zacharias, D. A., Violin, J. D., Newton, A. C. & Tsien, R. Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Eggeling, C. et al. Direct observation of the nanoscale dynamics of membrane lipids in a living cell. Nature 457, 1159–1162 (2009).
Dietrich, C. et al. Lipid rafts reconstituted in model membranes. Biophys. J. 80, 1417–1428 (2001).
Veatch, S. L. & Keller, S. L. Seeing spots: Complex phase behavior in simple membranes. BBA – Mol. Cell Res. 1746, 172–185 (2005).
McConnell, H. M. & Vrljic, M. Liquid-liquid immiscibility in membranes. Annu. Rev. Biophys. Biomolec. Struct. 32, 469–492 (2003).
Elson, E. L., Fried, E., Dolbow, J. E. & Genin, G. M. Phase separation in biological membranes: integration of theory and experiment. Annu. Rev. Biophys. 39, 207–226 (2010).
Hjort Ipsen, J., Karlström, G., Mourtisen, O. G., Wennerström, H. & Zuckermann, M. J. Phase equilibria in the phosphatidylcholine-cholesterol system. BBA – Biomembranes 905, 162–172 (1987).
Filippov, A., Orädd, G. & Lindblom, G. Lipid lateral diffusion in ordered and disordered phases in raft mixtures. Biophys. J. 86, 891–896 (2004).
Kahya, N., Scherfeld, D., Bacia, K., Poolman, B. & Schwille, P. Probing lipid mobility of raft-exhibiting model membranes by fluorescence correlation spectroscopy. J. Biol. Chem. 278, 28109–28115 (2003).
Fujiwara, T., Ritchie, K., Murakoshi, H., Jacobson, K. & Kusumi, A. Phospholipids undergo hop diffusion in compartmentalized cell membrane. J. Cell Biol. 157, 1071–1081 (2002).
Lindfors, K., Kalkbrenner, T., Stoller, P. & Sandoghdar, V. Detection and spectroscopy of gold nanoparticles using supercontinuum white light confocal microscopy. Phys. Rev. Lett. 93, 037401 (2004).
Jacobsen, V., Stoller, P., Brunner, C., Vogel, V. & Sandoghdar, V. Interferometric optical detection and tracking of very small gold nanoparticles at a water-glass interface. Opt. Express 14, 405–414 (2006).
Kukura, P. et al. High-speed nanoscopic tracking of the position and orientation of a single virus. Nat. Methods 6, 923–927 (2009).
Ortega-Arroyo, J. & Kukura, P. Interferometric scattering microscopy (iSCAT): new frontiers in ultrafast and ultrasensitive optical microscopy. Phys. Chem. Chem. Phys. 14, 15625–15636 (2012).
Hsieh, C. L., Spindler, S., Ehrig, J. & Sandoghdar, V. Tracking single particles on supported lipid membranes: multimobility diffusion and nanoscopic confinement. J. Phys. Chem. B 118, 1545–1554 (2014).
Spillane, K. M. et al. High-speed single-particle tracking of GM1 in model membranes reveals anomalous diffusion due to interleaflet coupling and molecular pinning. Nano Lett. 14, 5390–5397 (2014).
Lin, Y.-H., Chang, W.-L. & Hsieh, C.-L. Shot-noise limited localization of single 20 nm gold particles with nanometer spatial precision within microseconds. Opt. Express 22, 9159–9170 (2014).
Zhang, L. et al. Interferometric detection of single gold nanoparticles calibrated against TEM size distributions. Small 11, 3550–3555 (2015).
Ayuyan, A. G. & Cohen, F. S. Lipid peroxides promote large rafts: effects of excitation of probes in fluorescence microscopy and electrochemical reactions during vesicle formation. Biophys. J. 91, 2172–2183 (2006).
Saxton, M. J. & Jacobson, K. Single-particle tracking: applications to membrane dynamics. Annu. Rev. Biophys. Biomolec. Struct. 26, 373–399 (1997).
Michalet, X. Mean square displacement analysis of single-particle trajectories with localization error: Brownian motion in an isotropic medium. Phys. Rev. E 82, 041914 (2010).
Chiantia, S., Ries, J., Kahya, N. & Schwille, P. Combined AFM and two-focus SFCS study of raft-exhibiting model membranes. Chem Phys Chem 7, 2409–2418 (2006).
de Wit, G., Danial, J. S. H., Kukura, P. & Wallace, M. I. Dynamic label-free imaging of lipid nanodomains. Proc. Natl. Acad. Sci. USA 112, 12299–12303 (2015).
Simson, R., Sheets, E. D. & Jacobson, K. Detection of temporary lateral confinement of membrane proteins using single-particle tracking analysis. Biophys. J. 69, 989–993 (1995).
Yasuda, T., Tsuchikawa, H., Murata, M. & Matsumori, N. Deuterium NMR of raft model membranes reveals domain-specific order profiles and compositional distribution. Biophys. J. 108, 2502–2506 (2015).
Jeon, J.-H., Monne, H. M.-S., Javanainen, M. & Metzler, R. Anomalous diffusion of phospholipids and cholesterols in a lipid bilayer and its origins. Phys. Rev. Lett. 109, 188103 (2012).
Sodt, A. J., Sandar, M. L., Gawrisch, K., Pastor, R. W. & Lyman, E. The molecular structure of the liquid-ordered phase of lipid bilayers. J. Am. Chem. Soc. 136, 725–732 (2014).
Vist, M. R. & Davis, J. H. Phase equilibria of cholesterol/dipalmitoylphosphatidylcholine mixtures: deuterium nuclear magnetic resonance and differential scanning calorimetry. Biochemistry 29, 451–464 (1990).
McMullen, T. P. W., Lewis, R. N. A. H. & McElhaney, R. N. Cholesterol–phospholipid interactions, the liquid-ordered phase and lipid rafts in model and biological membranes. Curr. Opin. Colloid Interface Sci. 8, 459–468 (2004).
McElhaney, R. N. The use of differential scanning calorimetry and differential thermal analysis in studies of model and biological membranes. Chem. Phys. Lipids 30, 229–259 (1982).
Suzuki, K. G. N. et al. Transient GPI-anchored protein homodimers are units for raft organization and function. Nat. Chem. Biol. 8, 774–783 (2012).
Chiantia, S., Kahya, N., Ries, J. & Schwille, P. Effects of ceramide on liquid-ordered domains investigated by simultaneous AFM and FCS. Biophys. J. 90, 4500–4508 (2006).
Lin, W.-C., Blanchette, C. D. & Longo, M. L. Fluid-phase chain unsaturation controlling domain microstructure and phase in ternary lipid bilayers containing GalCer and cholesterol. Biophys. J. 92, 2831–2841 (2007).
Eghiaian, F., Rico, F., Colom, A., Casuso, I. & Scheuring, S. High-speed atomic force microscopy: Imaging and force spectroscopy. FEBS Lett. 588, 3631–3638 (2014).
Honigmann, A., Mueller, V., Hell, S. W. & Eggeling, C. STED microscopy detects and quantifies liquid phase separation in lipid membranes using a new far-red emitting fluorescent phosphoglycerolipid analogue. Faraday Discuss. 161, 77–89 (2013).
Kusumi, A. et al. Dynamic organizing principles of the plasma membrane that regulate signal transduction: commemorating the fortieth anniversary of Singer and Nicolson’s fluid-mosaic model. Annu. Rev. Cell Dev. Biol. 28, 215–250 (2012).
Vicidomini, G. et al. STED-FLCS: an advanced tool to reveal spatiotemporal heterogeneity of molecular membrane dynamics. Nano Lett. 15, 5912–5918 (2015).
Brian, A. A. & McConnell, H. M. Allogeneic stimulation of cytotoxic T cells by supported planar membranes. Proc. Natl. Acad. Sci. USA 81, 6159–6163 (1984).
Michalet, X. & Berglund, A. J. Optimal diffusion coefficient estimation in single-particle tracking. Phys. Rev. E 85, 061916 (2012).
Acknowledgements
This work is supported by the Nano Program of Academia Sinica and the Ministry of Science and Technology, Taiwan (grants no. 102-2112-M-001-002-MY3).
Author information
Authors and Affiliations
Contributions
H.M.W., Y.H.L. and C.L.H. conceived and designed experiments. H.M.W., Y.H.L., T.C.Y. and C.L.H. performed experiments and analyzed the results. H.M.W. and C.L.H. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wu, HM., Lin, YH., Yen, TC. et al. Nanoscopic substructures of raft-mimetic liquid-ordered membrane domains revealed by high-speed single-particle tracking. Sci Rep 6, 20542 (2016). https://doi.org/10.1038/srep20542
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20542
This article is cited by
-
Molecular substructure of the liquid-ordered phase formed by sphingomyelin and cholesterol: sphingomyelin clusters forming nano-subdomains are a characteristic feature
Biophysical Reviews (2022)
-
Low-flux scanning electron diffraction reveals substructures inside the ordered membrane domain
Scientific Reports (2020)
-
Permeability of membranes in the liquid ordered and liquid disordered phases
Nature Communications (2019)
-
Interferometric scattering microscopy reveals microsecond nanoscopic protein motion on a live cell membrane
Nature Photonics (2019)
-
Complex dynamics at the nanoscale in simple biomembranes
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.