Abstract
Although BRAFV600E mutation is associated with adverse clinical outcomes in patients with colorectal cancer (CRC), response and resistance mechanisms for therapeutic BRAFV600E inhibitors remains poorly understood. In the present study, we demonstrate that selective BRAFV600E inhibition activates AMP-activated protein kinase (AMPK), which induces autophagy as a mechanism of therapeutic resistance in human cancers. The present data show AMPK-dependent cytoprotective roles of autophagy under conditions of therapeutic BRAFV600E inhibition and AMPK was negatively correlated with BRAFV600E-dependent activation of MEK-ERK-RSK signaling and positively correlated with unc-51-like kinase 1 (ULK1), a key initiator of autophagy. Furthermore, selective BRAFV600E inhibition and concomitant suppression of autophagy led to the induction of apoptosis. Taken together, present experiments indicate that AMPK plays a role in the survival of BRAFV600E CRC cells by selective inhibition and suggest that the control of autophagy contributes to overcome the chemoresistance of BRAFV600E CRC cells.
Similar content being viewed by others
Introduction
Although outcomes in patients with colorectal cancers (CRC) have improved over the last decade, poor prognoses remain for some subtypes of CRC1. In particular, mutations in valine 600 (V600) of the BRAF oncogene occur in approximately 7% of all human cancers, including approximately 10% of CRC1,2. Moreover, BRAF mutations are associated with adverse clinical outcomes in patients with CRC, with a 70% increase in mortality in patients with metastatic CRC harboring BRAFV600E mutations compared with those carrying wild-type BRAF3,4. Therefore, novel therapeutic strategies for patients with BRAF mutant CRC are critically needed.
Although a selective RAF inhibitor was recently approved by the Food and Drug Administration for the treatment of metastatic melanomas harboring BRAFV600E mutations, response rates to selective BRAF inhibitors vary between tumor types. While selective BRAF inhibitors have produced response rates of approximately 50%–80% in patients with BRAFV600E mutant melanomas5, a selective BRAF inhibitor alone has proven disappointingly ineffective in CRCs harboring BRAFV600E mutations. Multiple studies have investigated the underlying mechanisms of resistance of BRAFV600E CRC to selective BRAF inhibitors, including KRAS and BRAF amplifications and MEK1 mutations6. Other studies have shown that EGFR-mediated reactivation of the mitogen-activated protein kinase (MAPK) pathway, PIK3CA mutations and PTEN loss may also contribute to selective resistance to BRAF inhibitors7. However, the relative correlations with these resistance mechanisms and clinical outcomes remain poorly understood. Therefore, elucidating the underlying mechanisms of resistance to selective BRAF inhibitors may lead to new therapeutic strategies for CRCs harboring the BRAFV600E mutation.
Autophagy has been described as a mechanism of resistance for cancer cells under conditions of therapeutic stress in numerous human cancers, including CRC. Autophagy is an intracellular bulk degradation system in which cytoplasmic components, including organelles, are directed to the lysosome/vacuole by a membrane-mediated process8. Autophagy is thought to be initiated under nutrient-limited conditions by a conserved kinase complex containing the unc-51-like kinase 1 (ULK1) and ULK2 and the subunits autophagy-related gene 13 (Atg13) and FAK family kinase-interacting protein of 200 (FIP200)9. Although autophagy is activated under chemotherapy or radiation stresses10,11, subsequent influences on cancer cell death or survival remain controversial. However, numerous reports indicate that the activation of autophagy promotes cancer cell survival after exposure to chemotherapy or radiation therapy and inhibition of autophagy can be a valuable strategy for cancer therapy.
Autophagy is a complicated regulatory process that involves numerous upstream regulating signaling pathways, including the PI3K-Akt-mammalian target of rapamycin (mTOR) pathway; liver kinase B1 (LKB1)-AMP-activated protein kinase (AMPK)-mTOR pathway; and p53, Beclin1 and Bcl-2 pathways12 and, to a limited extent, MAPK signaling pathway. Whether autophagy is required for BRAFV600E CRC remains unclear, evidence suggests that it is important for BRAFV600E melanomas13,14. Interestingly, previous studies report a molecular relationship between LKB1-AMPK and RAF-MEK-ERK pathways in melanomas harboring the BRAFV600E mutation15,16. However, to the best of our knowledge, no previous studies have examined the molecular linkage between the BRAFV600E mutation and selective BRAF inhibitor-induced autophagy in BRAFV600E CRC. Considering the potential roles of AMPK-related cellular signaling pathways, such as the MEK-ERK pathway, we hypothesized that AMPK interacts with the MEK-ERK pathway to induce autophagy in BRAFV600E CRC.
In the present study, we report elevated levels of autophagy after exposure to selective BRAF inhibitors in BRAFV600E CRC cells. Subsequently, the roles of selective BRAF inhibitor-induced autophagy, the effects of autophagy inhibition by small-interfering RNAs (siRNAs) or a pharmacological inhibitor and the mechanistic link between BRAFV600E mutation and autophagy in BRAFV600E CRC cell lines were studied. Our findings indicate that selective BRAF inhibitor-induced AMPK phosphorylation coordinates control of autophagy and tumor chemoresistance in BRAFV600E CRC cells.
Experimental Procedures
Reagents and antibodies
Selective BRAF inhibitors PLX4032 (also known as Vemurafenib, AXON Medchem, catalogue #1624; AdooQ BioScience Catagog Num A10739) and PLX4720 (AXON Medchem, #1474) and Chloroquine (CQ) (Focus Biomolecules, #10-2473; SIGMA-ALDRICH, C6628) were used. The antibodies for Western blotting are as follows: the microtubule-associated protein 1 light chain 3 (LC3) (Cell Signaling Technology, CST, #2775); anti-Atg13 (CST, #13468); anti-Atg7 (CST, #2631); anti-phospho-mTOR (Ser2448) (CST, #2971); anti-mTOR (CST, #2972); anti-phospho-AMPKα (Thr172) (CST, #2535); anti-AMPKα (CST, #5832); anti-phospho-MEK1/2 (Ser221) (CST, #2338); anti-phospho-Erk1/2 (Thr202/Tyr204) (CST, #4370); anti-phospho-p90RSK (T359/S363) (Abcam, ab32413); anti-phospho-LKB1 (Ser428) (Abcam, ab63473); anti-phospho-Raptor (Ser792) (CST, #2083); anti-phospho-ULK1 (Ser555) (CST, #5869); anti-phospho-ULK1 (Ser757) (CST, #6888); anti-ULK1 (CST, #8054).
Cell lines and cell culture
Human CRC cell lines HT29, RKO and Caco2 were obtained from ATCC (Manassas, VA) and melanoma cell line A375 was given from Dr. Kikuchi (Division of Gene Therapy Science, Graduate School of Medicine, Osaka University). All cell lines were cultured with Dulbecco’s modified Eagle medium (DMEM, Sigma-Aldrich) supplemented 10% fetal bovine serum (FBS, Thermo Fisher Scientific), 100 U/mL penicillin and grown at 37 °C in a humidified atmosphere of 95% air, 5% CO2. BRAF inhibitors were dissolved in dimethyl-sulphoxide (DMSO) and were diluted with medium before use. Final concentration of DMSO was 0.1%.
Small interfering RNA and transfection
The oligonucleotide small interfering RNA (siRNA) specifically targeting Atg13 (Custom Select, #:4390827) and negative control siRNA (Silence Select, #:4390843) were purchased from Life Technologies. Atg7 siRNA (SI02655373) was purchased from QIAGEN. Target sequences were as follows; sense Atg13, GAGUUUGGAUAUACCCUUUtt, antisense Atg13: AAAGGGUAUAUCCAAACUCgt; sense Atg7, CAGUGGAUCUAAAUCUCAATT, antisense Atg7, UUGAGAUUUAGAUCCACUGAT. Cells were transfected with control siRNA, Atg13 siRNA, or Atg7 siRNA using Lipofectamine RNAiMax (Life Technologies) according to the manufacturer’s instructions. After incubation of 48 hours, the medium was changed to DMEM containing selective BRAF inhibitors or DMSO. We determined the efficiency of siRNA-mediated protein knockdown by Western blot. For the BRAF inhibitors sensitivity, cells were seeded in 96-well plates at a density of 2,000 cells 100 μL per well and transfected with 10 μM of either control siRNA, Atg13 siRNA, or Atg7 siRNA. The cytotoxicity of selective BRAF inhibitors was determined by MTT and apoptosis assays.
Cell viability and combination index (CI)
Cell viability was determined by MTT assay. Cells were seeded into 96-well plates at a density of 5,000 cells 100 μL per well and treated with selective BRAF inhibitors, CQ or their combination. After treatment, 10 μL of the cell-counting solution (Cell counting kit-8, Dojindo Molecular Technologies) was added to each well and incubated in a humidified 5% CO2 atmosphere at 37 °C for 2 h. The absorbance at 450 nm was determined using iMarkTM microplate absorbance reader (Bio-Rad Laboratories) to study the proliferation.
Drug interactions and dose-effect relationships were analyzed according to the reported method17,18, with minor modifications. For drug combinations, the combination index (CI) was calculated by the classic isobologram equation. The previous studies recommend the use of cytotoxic agents at IC50, but this strategy would not have been feasible in the case of RKO cells. Thus we determined CI at 50% cell death in HT29 cells and at 30% cell death in RKO cells. The CI was defined as follows: CI = [(D)A/A+B/(D)A/(D)A]+[(D)B/A+B/(D)B/(D)B] where (D)A and (D)B is the concentrations of drug A or B alone giving a 30% or 50% reduction in cell viability compared to a control. (D)A+B is the concentration of drug A and B in combination producing a 30% or 50% reduction in cell viability compared to a control. The IC30 or 50 values were calculated for monotherapy. The ratio for combination of reagents was determined on the basis of IC30 or 50 values. We used a cut off for the CI of 0.8. We identified a synergistic effect when CIs were smaller than 0.8; an antagonistic effect when they were greater than 1.2; and an additive effect when they were between 0.8 and 1.2.
Western blot analysis
Cell lysates was harvested from the cell lines using radio immunoprecipitation assay (RIPA) buffer (Thermo Fisher Scientific) with protease inhibitors, phosphatase inhibitors and EDTA (Thermo Fisher Scientific). Aliquots of protein were electrophoresed on SDS/PAGE Tris·HCl gels (Bio-Rad Laboratories). The proteins were separated by electrophoresis prior to transfer to PVDF membranes (Bio-Rad Laboratories) and incubated with primary antibodies overnight at 4 °C, followed by incubation with HRP-linked anti-rabbit or anti-mouse IgG (GE Healthcare Biosciences) at a dilution of 1:100,000 for 1 h at room temperature. The antigen–antibody complex was detected with the ECL Prime Western Blotting Detection Kit (GE Healthcare Biosciences). The intensity of the blots was quantified by densitometry analysis using Image lab software version 5.0 (Bio-Rad Laboratories).
Apoptosis assay
Apoptotic rate was determined by flow cytometry with the Annexin V-fluorescein isothiocyanate (FITC) apoptosis detection kit (Biovision). Briefly, cells were collected by accutase and suspended with 500 mL of 1× binding buffer and then treated with 5 μL of Annexin V–FITC and 2.5 μL PI. After incubation of 10 minutes on ice, each sample was analyzed immediately using the BD FACSAria™ IIu instrument (BD Bioscience). We defined apoptotic cells as follow: unstained cells were classified as “live”; cells stained by Annexin V only were “early apoptotic”; cells stained by both Annexin V and PI were “late apoptotic”; and cells stained by PI only were “dead” cells.
Immunocytochemistry for LC3 localization
Cells were seeded in 35-mm diameter ibidi dishes. After treatment of 24 h, cells were washed with 1 × PBS and fixed with 4% paraformaldehyde for 15 minutes at room temperature. Subsequently, cells were blocked by blocking-one for 1 h at room temperature and then incubated overnight with primary antibodies, anti-LC3 antibody (1:400 diluted in PBS with 0.1% Tween). Cells were washed with 1 × BSA buffer, followed by incubation with secondary antibodies, Alexa Fluor 488 conjugated anti-rabbit antibody (1:1000 diluted in PBS with 0.1% Tween), for 1 h at room temperature. Cells were washed with 1 × PBS and then mounted with DAPI (Life technologies). Cells were visualized using a confocal laser scanning microscope (Olympus FluoView FV1000) at an objective of ×200.
Xenograft models
Animal studies were conducted in strict accordance with the principles and procedures approved by the Committee on the Ethics of Animal Experiments of Osaka University. For xenograft models, 7-week-old female mice (BALB/c–nu/nu) were purchased from CLEA Japan, Inc. HT29 (5.0 × 106) cells in a total volume of 200 μL DMEM/Matrigel (4:1 (v/v) suspension) were injected subcutaneous into the flanks. When the diameter of the subcutaneous tumor reached about 100 mm3, tumorbearing animals were randomly assigned to four groups that were administered Control (Vehicle plus PBS), CQ alone, PLX4032 alone, or Combination (PLX4032 plus CQ). Vehicle solution contained 5% DMSO and 1% methyl cellulose. PLX4032 was formulated in 5% DMSO, 1% methyl cellulose and dosed at 50 mg/kg twice daily by oral gavage. CQ was dissolved in PBS and intraperitoneally administered daily at the dose of 60 mg/kg/day. Tumor volume was measured with calipers and calculated using the formula V = (ab2)/2, where a is the largest diameter and b is the smallest diameter. Mice were sacrificed 24 hours after the last treatment. The tumors were weighed and tumor lysates were subjected to Western blot analysis.
Statistical analysis
Data were collected from at least three independent experiments. Data are presented as mean ± SD. Comparisons between two groups were performed by unpaired t test using JMP pro version 11 (SAS Institute, Cary, N.C., USA). A P value < 0.05 was considered significant.
Results
Autophagy is induced in BRAFV600E CRC cells after treatment with selective BRAF inhibitors
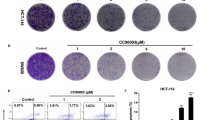
To examine the effects of selective BRAF inhibitors on autophagy, we monitored the induction of autophagy in BRAFV600E HT29 and RKO CRC cells after treatment with selective BRAF inhibitors. Initially, we assessed the expression of light chain 3 (LC3), a mammalian homolog of yeast Atg8, using western blotting. During autophagy, the cytoplasmic form of LC3-I is converted to the membrane-bound lipidated form LC3-II, as detected by electrophoretic mobility shifts8. On the basis of a previous study13, A375 melanoma cells were used as a positive control (Supplementary Fig. 1A,B left panel) and dose dependency of this response was examined in HT29 cells in the presence of drugs at 0.1, 1 and 10 μM. Although LC3-II expression was not clearly increased in the presence of 0.1 μM PLX4032, the expression of LC3-I was decreased, indicating the modest activation of autophagosome formation. Accordingly, LC3-II and LC3-II/LC3-I ratios were clearly increased at 1 and 10 μM PLX4032 or PLX4720 (Fig. 1A and Supplementary Fig. 1C left panel) and in complimentary experiments, changes in autophagic activity were visually examined using confocal microscopy. In particular, PLX4032-induced punctate staining of LC3-II was increased in HT29 cells (Fig. 1B left panel) and LC3 conversion was observed as early as 2–6 h after treatment with 10 μM PLX4032 and gradually increased until 24 h (Fig. 1C,D left panel). In further experiments, 1 and 10 μM PLX4032 or PLX4720 increased LC3-II expression and LC3-II/LC3-I ratios in RKO cells (Fig. 1A,B right panel and Supplementary Fig. 1C right panel). Moreover, at 10 μM PLX4032, increased LC3-II expression was significant at 2 h after the initiation of treatment and was stable until 24 h in RKO cells (Fig. 1C,D right panel). These data indicate that LC3-II accumulates dose and time dependently in CRC cell lines. In contrast, no significant changes in LC3-II expression were observed after identical treatments in BRAFwild-type Caco2 cells (Supplementary Fig. 1A, B right panel). Subsequently, to clarify whether accumulation of autophagosomes following treatments with selective BRAF inhibitors reflects induction of autophagosome formation or inhibition of autophagosome degradation, BRAF inhibitor-treated cells were exposed to chloroquine (CQ). In these experiments, accumulation of LC3-II was further enhanced, indicating that the accumulation of LC3-II in the presence of selective BRAF inhibitors follows the activation of autophagosome formation rather than the inhibition of autophagosome degradation (Fig. 1E).
Selective BRAF inhibitors induce autophagy in BRAFV600E CRC cells.
(A) HT29 and RKO cells were treated with DMSO and 0.1, 1, or 10 μM PLX4032 for 24 h. LC3-II/LC3-I ratios were determined using Image lab software version 5.0 (Bio-Rad Laboratories). (B) HT29 and RKO cells were treated with DMSO and 0.1, 1, or 10 μM PLX4032 for 24 h. Cells were visualized using a confocal laser microscope (Olympus FluoView FV1000; objective, ×200). Green dots correspond with accumulated LC3 in autophagosomes and nuclei are stained with DAPI (blue). Scale bars indicate 10μm. (C) HT29 and RKO cells were treated with DMSO for 24 h and 10 μM PLX4032 for 2, 6, 12 and 24 h and LC3-II expression and LC3-II/LC3-I ratios were determined using Western blot analyses. (D) HT29 and RKO cells were treated with DMSO for 24 h and 10 μM PLX4032 for 2, 6, 12 and 24 h. Cells were visualized using a confocal laser microscope (Olympus FluoView FV1000; objective,×200). Green dots correspond with accumulated LC3 in autophagosomes and nuclei are stained with DAPI (blue). Scale bars indicate 10μm. (E) HT29 cells were treated with or without 10 μM PLX4032 in the presence or absence of 25 μM CQ for 24 h and Western blot analyses of LC3 turnover were performed.
Inhibition of selective BRAF inhibitor-induced autophagy by pharmacological or siRNA inhibitors sensitizes BRAFV600E CRC cells to selective BRAF inhibitors
To elucidate the role of selective BRAF inhibitor-induced autophagy, autophagy was inhibited using the pharmacological inhibitor CQ or using RNA interference of Atg13 or Atg7. After exposure of cells to PLX4032 and CQ, which blocks downstream autophagy pathways, drug interactions and dose-effect relationships were assessed using a CI according to the methods of Chou and Talalay17,18. Initially, the effects of monotherapy were examined in HT29 cells at CQ concentrations of 0, 0.01, 0.1, 1, 10, 100 and 1000 μM and whereas 1 μM CQ had almost no inhibitory effect after 24 h, CQ concentrations of more than 1 μM inhibited proliferation in a dose-dependent manner (Supplementary Fig. 2A). Subsequently, combined treatments of HT29 cells resulted in improved cytotoxic effects at 24 h for every regimen compared with respective monotherapies (Fig. 2A). Moreover, analysis of combined effects revealed synergy with a CI value of 0.523 ± 0.15 (Fig. 2B). In subsequent experiments using RKO and A375 cells, combination treatments improved the cytotoxic effects of all monotherapeutic regimens at 24 h (Fig. 2C and Supplementary Fig. 2B,C) and analysis of combined effects revealed synergy with CI values of 0.577 ± 0.19 and 0.505 ± 0.15, respectively (Fig. 2D and Supplementary Fig. 2D). In further experiments, the extent of apoptosis was investigated at 24 h after single and combination treatments with PLX4032 or PLX4720 and CQ using apoptosis assays. These experiments revealed significantly higher percentages of early and late apoptotic cells after combination treatments in all cell lines (Fig. 2E and Supplementary Fig. 2E).
Pharmacological inhibition of autophagy by CQ increases sensitivity of BRAFV600E CRC cells to PLX4032.
(A) HT29 cells were treated with or without 0.1, 1, or 10 μM PLX4032 in the presence or absence of 0, 5, 25, or 50 μM CQ for 24 h. Cell viability was determined using MTT assays. (B) Isobologram analysis of interactions between PLX4032 and CQ in HT29 cells. (C) RKO cells were treated with or without 0.1, 1, or 10 μM PLX4032 in the presence or absence of 0, 5, 25, or 50 μM CQ for 24 h. Cell viability was determined using MTT assays. (D) Isobologram analysis of interactions between PLX4032 and CQ in RKO cells. (E) HT29 and RKO cells were treated with PLX4032 or PLX4720 at 10 μM in the presence or absence of CQ at 25 μM for 24 h. Apoptosis rates were determined using flow cytometry and cells were defined as early or late apoptotic cells. Data are presented as means and standard deviations of at least three independent experiments; *p < 0.05.
To elucidate associated mechanisms, we downregulated autophagy by targeting Atg13 or Atg7 which was gene to be associated with autophagy. Total protein expressions of Atg13 or Atg7 were stable in the presence of PLX4032 at all doses and times (Supplementary Fig. 3A–D) but was significantly decreased after treatment with targeted siRNA. Moreover, knockdown of Atg13 or Atg7 suppressed LC3-II expression in selective BRAF inhibitor-treated cells (Fig. 3A–D and Supplementary Fig. 3E,F), indicating that autophagy is suppressed under physiological conditions. Atg13 or Atg7 knockdown also enhanced selective BRAF inhibitor-induced apoptosis and decreased cell viability in HT29 and RKO cells (Fig. 3E–H).
Inhibition of autophagy by Atg13 or Atg7 siRNA increases sensitivity of BRAFV600E CRC cells to PLX4032.
(A,B) HT29 cells were transfected with Atg13 siRNA or control siRNA and were then treated with DMSO and PLX4032 or PLX4720 at 10 μM for 24 h. LC3 and Atg13 expression was determined using western blot analyses. (C,D) HT29 and RKO cells were transfected with Atg7 siRNA or control siRNA and were then treated with DMSO or PLX4032 at 10 μM for 24 h. LC3 and Atg7 expression was determined using western blot analyses. (E,F) HT29 and RKO cells were transfected with control siRNA, Atg13 siRNA, or Atg7 siRNA and were treated with DMSO, PLX4032, or PLX4720 at 10 μM for 24 h. Cell viability was then determined using MTT assays. (G,H) Apoptosis assay in HT29 and RKO cells transfected with control siRNA, Atg13 siRNA, or Atg7 siRNA. Apoptotic rates were determined using flow cytometry and early and late apoptotic cells were distinguished. Data are presented as means and standard deviations of at least three independent experiments; *p < 0.05.
AMPK phosphorylation is required for selective BRAF inhibitor-induced autophagy
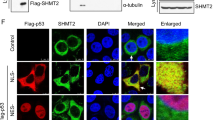
Autophagy can be induced by alterations in various growth factor signaling pathways within the tumor microenvironment and the major direct signaling regulators of autophagy include mTORC1, AMPK and specific components of the ER stress response19. To investigate mechanisms by which selective BRAF inhibitors induce autophagy, protein expression of these molecules was determined using western blotting. Although no significant changes in phospho-mTOR (ser2448) expression were observed after treatment with PLX4032, compared with DMSO-treated cells, PLX4032 increased the phosphorylation of AMPK (Thr172) after 24 h in HT29 and RKO cells (Fig. 4A). Phosphorylation of AMPK (Thr172) was also increased in the presence of increased LC3II/LC3I ratios following treatments with inhibitors of BRAFV600E. In addition, AMPK activity was elevated at early time points after treatment with PLX4032, indicating that AMPK activation is associated with PLX4032-induced autophagy in BRAFV600E CRC cells. Additionally, combination treatments with PLX4032 or PLX4720 and CQ increased AMPK phosphorylation (Thr172) at 24 h in HT29 cells (Supplementary Fig. 4A,B).
AMPK phosphorylation is required for ULK1 function during PLX4032-induced autophagy.
(A) Western blot assays of AMPKα (Thr172) and mTOR (Ser2448) phosphorylation and LC3 expression in HT29 and RKO cells after treatment with DMSO for 24 h and 10 μM PLX4032 for 2, 6, 12, or 24 h. (B) Western blot analyses of phospho-AMPKα (Thr172), phospho-ULK1 (Ser555), phospho-Raptor (Ser792), phospho-ULK1 (Ser757) and LC3 in HT29 cells after treatment with DMSO and 10 μM PLX4032 for 24 h. (C) Western blot analyses of phospho-AMPKα (Thr172), phospho-ULK1 (Ser555) and LC3 in HT29 cells after treatment with DMSO for 24 h and 10 μM PLX4032 for 2, 6, 12 and 24 h. (D) Western blot analyses of phospho-AMPKα (Thr172), phospho-ULK1 (Ser555), phospho-Raptor (Ser792), phospho-ULK1 (Ser757) and LC3 in RKO cells after treatment with DMSO and 10 μM PLX4032 for 24 h. (E) Western blot analyses of phospho-AMPKα (Thr172), phospho-ULK1 (Ser555), phospho-Raptor (Ser792), phospho-ULK1 (Ser757) and LC3 in RKO cells after treatment with DMSO for 24h and 10 μM PLX4032 for 2, 6, 12 and 24 h.
In further experiments, associations between AMPK and selective BRAF inhibitor-induced autophagy were investigated in HT29 and RKO cells. Several previous studies report roles of AMPK in the control of autophagy20 and show involvement of the mTORC1 pathway following phosphorylation of the regulatory associated protein mTOR (Raptor) and the tuberous sclerosis complex 2 (TSC2). However, whereas inhibition of mTORC1 correlates with increased autophagy, AMPK directly activates ULK1, which is a key initiator of autophagy. In present experiments, AMPK was markedly phosphorylated at approximately 6 h after treatments with inhibitors of BRAFV600E and ULK1 phosphorylation and (Ser555) autophagic activity were increased at 12 h compared with DMSO-treated cells. Moreover, changes in degrees of AMPK and ULK1 phosphorylation were time dependent (Fig. 4B,C) and similar changes were observed after exposure to PLX4720 (Supplementary Fig. 5A,B). In contrast, the effects of mTOR-dependent phosphorylation on the AMPK–ULK1 interaction did not produce significant changes in Raptor (Ser792) or ULK1 (Ser757) phosphorylation after 24 h treatments of HT29 cells (Supplementary Fig. 5C). These results show that selective BRAF inhibitors induce autophagy in HT29 cells, likely through direct AMPK-ULK1 interactions. In contrast with experiments in HT29 cells, PLX4032 treatments led to increased phosphorylation of Raptor (Ser792) and ULK1 (Ser757) in RKO cells (Fig. 4D,E). These data indicate that AMPK stimulates autophagy by inhibiting mTORC1 at the level of Raptor (Ser792) in addition to stimulating direct AMPK-ULK1 interactions in RKO cells.
Downregulation of oncogenic BRAF signaling stimulates AMPK via the downstream MEK-Erk-p90RSK kinase signaling cascade
Interactions between AMPK and RAF-MEK-ERK pathways have been documented in various human cancer cells, including colon cancer15,16. Accordingly, negative regulation of AMPK in BRAFV600E CRC cells suggests that AMPK or its upstream kinases may be targets of the MEK-ERK-RSK signaling cascade. Thus, to evaluate the relevance of AMPK inhibition by MEK-ERK-RSK signaling, phosphorylation of MEK-Erk-p90RSK and AMPK was determined in HT29 cells. Although untreated HT29 cells showed strong MEK-Erk-p90RSK phosphorylation but no AMPK phosphorylation, PLX4032-treated HT29 cells showed strong AMPK phosphorylation and no MEK-Erk-p90RSK phosphorylation. These data suggest an inverse correlation between AMPK and MEK-Erk-p90RSK signaling pathways (Fig. 5A). Moreover, MEK-Erk-p90RSK was inactivated after only 2 h treatments and remained ineffective for 24 h and AMPK phosphorylation was markedly increased, further suggesting inactivation of MEK-Erk-p90RSK signaling (Fig. 5B). Similar changes were also observed after treatments with PLX4720 (Supplementary Fig. 6A,B), with a corresponding negative correlation between AMPK and MEK-Erk-p90RSK phosphorylation levels in RKO cells (Fig. 5C). The present data suggest that the activity of AMPK is negatively regulated by the BRAFV600E mutation, presumably via downstream MEK-Erk-p90RSK kinase signaling.
Downregulation of BRAF signaling by PLX4032 activates AMPK by inactivating LKB1 (Ser428).
(A) Western blot analyses of phospho-MEK1/2 (Ser221), phospho-Erk1/2, phospho-p90RSK (Ser363), phospho-AMPKα (Thr172) and phospho-LKB1 (Ser428) in HT29 cells after treatment with DMSO and 10 μM PLX4032 for 24 h. (B) Western blot analyses of phospho-MEK1/2 (Ser221), phospho-Erk1/2, phospho-p90RSK (Ser363), phosphor-AMPKα (Thr172) and phospho-LKB1 (Ser428) in HT29 cells after treatment with DMSO for 24 h and 10 μM PLX4032 for 2, 6, 12, or 24 h. (C) Western blot analyses of phospho-MEK1/2 (Ser221), phospho-Erk1/2, phospho-p90RSK (Ser363) and phospho-AMPKα (Thr172) in RKO cells after treatment with DMSO and 10 μM PLX4032 for 24 h. (D) Western blot analyses of phospho-AMPKα (Thr172) and phospho-LKB1 (Ser428) in RKO cells after treatment with DMSO and 10 μM PLX4032 for 24 h.
Because phosphorylation of LKB1 at Ser428 (Ser431 in mouse) by p90RSK was previously demonstrated in vitro and in vivo21, we determined whether the effects of selective BRAF inhibitors on AMPK activity are dependent on LKB1. In these experiments, degrees of LKB1 (Ser428) and MEK-Erk-p90RSK phosphorylation were determined in HT29 cells within 24 h exposure to PLX4032. Accordingly, downregulation of BRAFV600E expression by PLX4032 led to decreased LKB1 phosphorylation at Ser428 and increased phosphorylation of AMPK at Thr172 (Fig. 5A,B). These data were corroborated by experiments in RKO cells (Fig. 5D), indicating that selective BRAF inhibitors phosphorylate AMPK by disinhibiting LKB1.
Autophagy inhibition chloroquine enhances the anti-tumor effect of PLX4032 in vivo
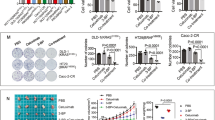
To further determine the therapeutic benefit of inhibiting autophagy in combination with PLX4032, mice with HT29 xenograft were randomly assigned to four groups when the volume of the subcutaneous tumor reached approximately 100 mm3: Control (CON), CQ alone, PLX4032 alone, or PLX4032 + CQ (Fig. 6A). As shown in Fig. 6B–D, CQ alone had no effect on the growth of tumors and PLX4032 alone displayed moderate anti-tumor activity. In contrast, combination of CQ with PLX4032 significantly reduced tumor growth compared to the PLX4032 alone. Furthermore, no significant weight loss was observed by combination treatments (Fig. 6E).
Autophagy inhibition chloroquine enhances the anti-tumor effect of PLX4032 in HT29 xenograft models.
(A) A typical case in each group. (B) Tumors from nude mice in each group. (C–E) Tumor volume, tumor weight and body weight in each group. (F) Tumor lysate was subjected to Western blot for LC3 and βActin. (G) Tumor lysate was subjected to immunoblotting for phospho-MEK1/2 (Ser221), phospho-Erk1/2, phospho-p90RSK (Ser363), phospho-AMPKα (Thr172), LC3 and βActin. BRAF mutant CRC xenografts derived from HT29 cells were treated with Control (vehicle plus PBS), PLX4032 alone (50 mg/kg, twice daily), CQ alone (60 mg/kg, daily), or combination therapy (PLX4032 plus CQ) for 13 day (Day0 – Day12). Average percent change in tumor volume relative to initial tumor volume is shown. Error bars represent SD. Asterisks represent p < 0.05 for combined PLX4032 plus CQ vs. all other treatment groups.
We then investigated whether PLX4032 induces autophagy in HT29 xenografted tumors. Consistent with the in vitro results, tumors treated with PLX4032 or PLX4032 plus CQ displayed accumulation of LC3-II on western blot compared with those from vehicle-treated mice (Fig. 6F). In addition, the LC3-II/LC3-I ratio and expression of LC3-II are much higher in PLX4032 plus CQ group than that in PLX4032 or CQ alone. We next investigated the mechanism of PLX4032 induced autophagy in vivo. Consistence with the in vitro findings, PLX4032 induced autophagy through inhibition of MEK-Erk-p90RSK signaling pathways and phosphorylation of AMPK. (Fig. 6G). These findings suggest that PLX4032 induced autophagy through inhibition of MEK-Erk-p90RSK signaling pathways and phosphorylation of AMPK and inhibition of autophagy by CQ can potently enhance the anti-tumor effect of PLX4032 in vivo.
Discussion
In this study, autophagy was induced by phosphorylated AMPK in BRAFV600E CRC cells after exposure to selective BRAF inhibitors and provided protection against the cytotoxic effects of selective BRAF inhibitors. Here we show three complementary lines of evidence that autophagy is induced in BRAFV600E CRC cells after exposure to selective BRAF inhibitors. Subsequently, functional roles of selective BRAF inhibitor-induced autophagy were demonstrated and molecular associations between the BRAFV600E mutation and treatment-induced autophagy were elucidated. To the best of our knowledge, the cytotoxic roles of selective BRAF inhibitor-induced autophagy have not been examined in BRAFV600E CRC cells.
The present experiments showed convincing evidence that autophagy is induced after exposure of BRAFV600E CRC cells to selective BRAF inhibitors. In particular, LC3 is widely used to monitor autophagy and LC3-II expression usually correlates closely with numbers of autophagosomes. Hence, assessments of autophagosome numbers were performed using light microscopic and biochemical methods as previously described8 and showed dose-dependent increases in punctate LC3 staining. Moreover, LC3-I conversion to LC3-II was increased, resulting in the accumulation of LC3-II after exposure of cells to selective BRAF inhibitors. However, accumulations of autophagosomes are not always indicative of increased autophagy and may alternatively reflect blockade of autophagosomal maturation and completion of the autophagy pathway8. Hence, LC3 turnover assays were performed in the presence of CQ and showed evidence that selective BRAF inhibitors increase numbers of autophagosomes and activate autophagic flux.
In subsequent experiments, we investigated functional roles of selective BRAF inhibitor-induced autophagy using CQ as a specific pharmacological inhibitor of autophagy. CQ has been widely used for many years without significant toxicity in malaria prophylaxis and has been implicated as a potential adjunct to anticancer treatments. In the present study, CQ inhibited autophagy, increased numbers of apoptotic cells and decreased cell viability. In accordance with the requirement of Atg13 in complex with ULK1 and FIP200 for initiation of autophagy22, specific siRNA against Atg13 increased cell death markedly as treatments with CQ. Similar changes were also observed after treatment with targeted Atg7, which is an autophagy-related gene. Autophagy is known to serve as a cytoprotective mechanism that allows escape from anticancer treatment-induced cell death and can compromise the efficacy of therapeutic interventions in cancer cells23. The present data suggest that both initial inhibition of autophagy by Atg13 or Atg7 knockdown and subsequent inhibition by CQ sensitizes BRAFV600E CRC cells by converting the autophagic process into an apoptotic process. To further determine the therapeutic benefit of inhibiting autophagy in combination with PLX4032, we examined whether selective BRAF inhibitor-induced autophagy can be therapeutically targeted using appropriate xenograft models. In HT29 xenograft models, although PLX4032 monotherapy was shown to be superior to that of CON or CQ monotherapy, greater efficacy was achieved with combination therapy. Furthermore, there were no adverse effects detected in mice treated with PLX4032 plus CQ. Thus, whereas autophagy can protect cancer cells from cytotoxic treatments, its inhibition can improve the effects of selective BRAF inhibitors in BRAFV600E CRC cells.
Finally, AMPK activation was central to BRAF inhibitor-induced autophagy, indicating a mechanistic linkage that may be exploited in the treatment of inhibitor resistant BRAFV600E CRC cells. AMPK acts as a cellular sensor for energy balance status and reports showing that AMPK activation suppresses cell growth and proliferation have warranted speculation that AMPK contributes to tumor suppressor pathways under stress conditions24,25,26. However, several reports associate AMPK with autophagy27,28,29,30 and corresponding protection against cytotoxic treatments, creating controversy about the role of AMPK in BRAFV600E CRC. In the present experiments, exposure to selective BRAF inhibitors activated AMPK in conjunction with clearly increased autophagic activity and subsequent inhibition of autophagy sensitized BRAFV600E CRC cells to selective BRAF inhibitors. Hence, AMPK likely facilitates escape from cell death following treatments with selective BRAF inhibitors as an inducer of autophagy during the early phase. However, whereas direct interactions of the AMPK-ULK1 signaling pathway were observed in HT29 and RKO cells, selective BRAF inhibitors also induced autophagy by inhibiting mTORC1 at the level of Raptor (Ser792) in RKO cells. Moreover, subsequent experiments showed that AMPK expression is relatively low in BRAFV600E cells, potentially reflecting relative MEK-ERK-RSK kinase signaling activities in these cells. In recent studies, reduced AMPK levels were shown in some cases of hepatocellular carcinomas31 and BRAFV600E was shown to regulate AMPK signaling by inhibiting Lkb1/Stk11 in melanoma cells15,16. Moreover, LKB1 was found to be the key upstream kinase for activation of AMPK32. Accordingly, the present data show an inverse correlation between the activities of MEK-ERK-RSK signaling and AMPK in BRAFV600E CRC cells, suggesting that BRAF mutations activate phospho-MEK, Erk and p90RSK without activating AMPK (Fig. 7 upper panel) and that inhibition of BRAF using selective BRAF inhibitors leads to inactivation of phospho-MEK, Erk and p90RSK and activation of AMPK (Fig. 7 lower panel). Intriguingly, the present data also suggested that activation of AMPK by the BRAFV600E mutant protein is impaired due to phosphorylation of LKB1 and loss of LKB1 activity following inhibition of BRAF led to activation of AMPK (Fig. 7). Although these observations require confirmation in further studies of the roles of LKB1 and AMPK, consequent improvements in understanding of selective BRAF inhibitor-induced autophagy may lead to improved therapeutic strategies for patients with BRAFV600E CRC.
In summary, selective BRAF inhibitors induced autophagy in BRAFV600E CRC cells. Inhibition of autophagy protected cancer cells from the cytotoxic effects of selective BRAF inhibitors, in part, by promoting apoptosis in the presence of selective BRAF inhibitors. Critically, the present observations demonstrate that AMPK activity is sensitive to MEK-Erk-p90RSK signaling in BRAFV600E CRC cells. Further clarification of the mechanisms associated with this molecular linkage in BRAFV600E CRC cells may facilitate the development of novel combination therapies with improved efficacy for patients with BRAFV600E CRC.
Additional Information
How to cite this article: Sueda, T. et al. BRAFV600E inhibition stimulates AMP-activated protein kinase-mediated autophagy in colorectal cancer cells. Sci. Rep. 6, 18949; doi: 10.1038/srep18949 (2016).
References
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
Kopetz, S. et al. Improved survival in metastatic colorectal cancer is associated with adoption of hepatic resection and improved chemotherapy. J. Clin. Oncol. 27, 3677–3683 (2009).
Montagut, C. & Settleman, J. Targeting the RAF-MEK-ERK pathway in cancer therapy. Cancer Lett. 283, 125–134 (2009).
Hutchins, G. et al. Value of mismatch repair, KRAS and BRAF mutations in predicting recurrence and benefits from chemotherapy in colorectal cancer. J. Clin. Oncol. 29, 1261–1270 (2011).
Flaherty, K. T. et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N. Engl. J. Med. 363, 809–819 (2010).
Lito, P., Rosen, N. & Solit, D. B. Tumor adaptation and resistance to RAF inhibitors. Nat. Med. 19, 1401–1409 (2013).
Boussemart, L. et al. eIF4F is a nexus of resistance to anti-BRAF and anti-MEK cancer therapies. Nature 513, 105–109 (2014).
Mizushima, N., Yoshimori, T. & Levine, B. Methods in mammalian autophagy research. Cell 140, 313–326 (2010).
Mizushima, N. The role of the Atg1/ULK1 complex in autophagy regulation. Curr. Opin. Cell Biol. 22, 132–139 (2010).
Mathew, R., Karantza-Wadsworth, V. & White, E. Role of autophagy in cancer. Nat. Rev. 7, 961–967 (2007).
Levine, B. Unraveling the role of autophagy in cancer. Autophagy 2, 65–66 (2006).
Chen, S. et al. Autophagy is a therapeutic target in anticancer drug resistance. Biochim. Biophys. Acta 1806, 220–229 (2010).
Ma, X. H. et al. Targeting ER stress-induced autophagy overcomes BRAF inhibitor resistance in melanoma. J. Clin. Invest. 124, 1406–1417 (2014).
Xie, X., Koh, J. Y., Price, S., White, E. & Mehnert, J. M. Atg7 Overcomes senescence and promotes growth of BrafV600E-driven melanoma. Cancer Discov. 5, 410–423 (2015).
Zheng, B. et al. Oncogenic B-RAF negatively regulates the tumor suppressor LKB1 to promote melanoma cell proliferation. Molecular Cell 33, 237–247 (2009).
Esteve-Puig, R., Canals, F., Colome, N., Merlino, G. & Recio, J. A. Uncoupling of the LKB1-AMPKalpha energy sensor pathway by growth factors and oncogenic BRAF. PLOS ONE 4, e4771 (2009).
Chou, T. C. & Talalay, P. Generalized equations for the analysis of inhibitions of Michaelis-Menten and higher-order kinetic systems with two or more mutually exclusive and nonexclusive inhibitors. Eur. J. Biochem. 115, 207–216 (1981).
Chou, T. C. & Talalay, P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv. Enzym. Regul. 22, 27–55 (1984).
Amaravadi, R. K. et al. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. 17, 654–666 (2011).
Hardie, D. G. AMPK and autophagy get connected. EMBO J. 30, 634–635 (2011).
Sapkota, G. P. et al. Phosphorylation of the protein kinase mutated in Peutz-Jeghers cancer syndrome, LKB1/STK11, at Ser431 by p90(RSK) and cAMP-dependent protein kinase, but not its farnesylation at Cys(433), is essential for LKB1 to suppress cell vrowth. J. Bio. Chem. 276, 19469–19482 (2001).
Ganley, I. G. et al. ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J. Bio. Chem. 284, 12297–12305 (2009).
Hoyer-Hansen, M. & Jaattela, M. Autophagy: an emerging target for cancer therapy. Autophagy 4, 574–580 (2008).
Hardie, D. G. AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat. Rev. Mol. Cell. Biol. 8, 774–785 (2007).
Hardie, D. G. AMP-activated protein kinase: an energy sensor that regulates all aspects of cell function. Gene. Dev. 25, 1895–1908 (2011).
Hardie, D. G., Ross, F. A. & Hawley, S. A. AMPK: a nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell. Biol. 13, 251–262 (2012).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell. Biol. 13, 132–141 (2011).
Lee, J. W., Park, S., Takahashi, Y. & Wang, H. G. The association of AMPK with ULK1 regulates autophagy. PLOS ONE 5, e15394 (2010).
Shang, L. et al. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc. Natl. Acad. Sci. USA. 108, 4788–4793 (2011).
Zheng, L. et al. Prognostic significance of AMPK activation and therapeutic effects of metformin in hepatocellular carcinoma. Clin. Cancer Res. 19, 5372–5380 (2013).
Hardie, D. G. The AMP-activated protein kinase pathway—new players upstream and downstream. J. Cell. Sci. 117, 5479–5487 (2004).
Acknowledgements
We thank the members of our laboratories for their helpful discussions. In particular, we thank H. Yoshimori, M. Hamasaki and the members of The Yoshimori Lab of the Department of Genetics, Osaka University Graduate school of Medicine. This work was partly supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology; P-DIRECT; Ministry of Health, Labour and Welfare. Partial support was received as institutional endowments from Taiho Pharmaceutical Co., Ltd., the Evidence Based Medical (EBM) Research Center, Chugai Co., Ltd. and Yakult Honsha Co., Ltd. No funding bodies were involved in data collection or analysis, the decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
D.S., K.K., H.Y., T.S., Y.D., M.M. and H.I. conceived the concept; T.S., K.K., M.K., N.N., J.K., H.C., H.T., N.H., J.N., T.H., I.T. and T.M. performed the experiments, analyzed the data and prepared the figures; T.S., K.K. and H.I. wrote the paper; All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sueda, T., Sakai, D., Kawamoto, K. et al. BRAFV600E inhibition stimulates AMP-activated protein kinase-mediated autophagy in colorectal cancer cells. Sci Rep 6, 18949 (2016). https://doi.org/10.1038/srep18949
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18949
This article is cited by
-
Pharmacological inhibition of Ref-1 enhances the therapeutic sensitivity of papillary thyroid carcinoma to vemurafenib
Cell Death & Disease (2022)
-
Targeted Therapy for Melanomas Without BRAF V600 Mutation
Current Oncology Reports (2022)
-
In silico high throughput screening and in vitro validation of a novel Raf/Mek dual inhibitor against colorectal carcinoma
Biologia (2022)
-
The MAPK and AMPK signalings: interplay and implication in targeted cancer therapy
Journal of Hematology & Oncology (2020)
-
Colorectal cancer cells respond differentially to autophagy inhibition in vivo
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.