Abstract
MicroRNAs (miRNAs) are important regulators of gene expression. The recent advances in high-throughput sequencing (HTS) technique have greatly facilitated large-scale detection of the miRNAs. However, thoroughly discovery of novel miRNAs from the available HTS data sets remains a major challenge. In this study, we observed that Dicer-mediated cleavage sites for the processing of the miRNA precursors could be mapped by using degradome sequencing data in both animals and plants. In this regard, a novel tool, miRNA Digger, was developed for systematical discovery of miRNA candidates through genome-wide screening of cleavage signals based on degradome sequencing data. To test its sensitivity and reliability, miRNA Digger was applied to discover miRNAs from four organs of Arabidopsis. The results revealed that a majority of already known mature miRNAs along with their miRNA*s expressed in these four organs were successfully recovered. Notably, a total of 30 novel miRNA-miRNA* pairs that have not been registered in miRBase were discovered by miRNA Digger. After target prediction and degradome sequencing data-based validation, eleven miRNA–target interactions involving six of the novel miRNAs were identified. Taken together, miRNA Digger could be applied for sensitive detection of novel miRNAs and it could be freely downloaded from http://www.bioinfolab.cn/miRNA_Digger/index.html.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are a class of small (18–25 nt in length) non-coding RNAs (ncRNAs) acting as negative regulators in gene expression. Previous studies showed that the regulatory activities of miRNAs could be involved in many biological processes, such as organ development1,2, metabolism3, immune response4 and tumorigenesis5,6. Certain miRNAs also showed great potential of being biomarkers for specific diseases7,8. In addition to mechanistic studies on already known miRNAs, discovery of novel miRNAs is becoming a major task and a hot research topic for thoroughly understanding of the miRNA world.
With the advances of high-throughput sequencing (HTS) techniques, a number of bioinformatics tools have been developed for systematical discovery of novel miRNAs based on different principles9, including the miRNA biogenesis mechanism10,11,12, the homology from known miRNAs13,14,15, the energy distribution pattern and Random Forest algorithm16. Although each of these approaches has its own merit and serviceability, several tools were included for performance comparison, in order to provide valuable information for selection of the ideal tool to meet different research needs9. However, accurate detection and quantification of these available tools are challenged by complex properties of the miRNAs and their precursors17, including the existence of miRNA variants (called isomRs), the secondary structures of the miRNA precursors9 and the Dicer-mediated processing of the miRNA precursors18,19. Therefore, thoroughly prediction and identification of the miRNAs, especially for the novel miRNAs, from the huge pool of HTS data remains a challenge for the following functional studies on miRNAs.
Degradome sequencing is a widely used approach for direct and global identification of miRNA-mediated target cleavages20,21,22. The miRNA-mediated cleavage sites could be mapped by using degradome signatures20 and a reversed framework for novel plant miRNA detection has been developed in our previous work, based on the degradome-supported miRNA cleavage loci and the highly complementarity of miRNA and the targets in plants22. Interestingly, several recent reports have showed that a portion of the degradome signatures could be mapped to the ends of the miRNA- and/or the miRNA*-coding loci on the miRNA precursors. Thus, it was speculated that these degradome signatures might be evidences for Dicer-mediated processing of the miRNA precursors20,23. However, no further investigation was conducted. In our previous work, mapping of degradome signatures onto the plant genomes has uncovered massive cleavage signals22. Intriguingly, a portion of the degradome signatures were found to locate on the known miRNA precursors. Although several miRNAs were indicated to regulate their own precursors by cleavages24, it could not explain the existence of the degradome signatures mapped to the ends of the miRNA- and/or the miRNA*-coding loci on the precursors. Further investigations revealed that a great number of degradome signatures on the newly identified miRNA precursors in Arabidopsis were resulted from DCL1 (Dicer-like 1)-mediated cleavages, suggesting that the processing of miRNA precursors could be supported by degradome signatures25. In this regard, degradome sequencing data could be utilized for the systematical identification of DCL1-dependent miRNA and miRNA* loci on their precursors in plants. Notably, as the miRNA-miRNA* duplexes are released by Dicer-mediated processing of miRNA precursors in animals26, the miRNA and miRNA* loci could also be supported by degradome signatures theoretically. In this scenario, the degradome-based evidences of Dicer-mediated cleavages could increase the reliability of systematical detection of novel miRNA loci in both animals and plants.
In this study, the miRNA precursors of Mus musculus and Arabidopsis thaliana registered in miRBase (release 21) were included for degradome-based scanning of Dicer-mediated cleavage signals. We demonstrated that Dicer-mediated processing of miRNA-miRNA* duplexes could be well supported by degradome signatures. Based on the results, a publicly available software, miRNA Digger, was developed for genomic-wide discovery of novel miRNA/miRNA* loci on specific precursors in plants and animals. To demonstrate its sensitivity and reliability, miRNA Digger was applied for the identification of already known and novel miRNAs from different tissues of Arabidopsis.
Results
Known miRNA/miRNA* loci were well supported by degradome signatures produced during Dicer-mediated processing of miRNA precursors
MiRNA biogenesis begins with the transcription of a primary miRNA (pri-miRNA). In animals, the pri-miRNA is recognized and cropped by RNase III enzyme Drosha to produce the pre-miRNA (precursor miRNA) and further processed by another RNase III enzyme, Dicer, to generate a miRNA-miRNA* duplex with 2-nt 3′ overhangs. In plants, this process is carried out by a single RNase III enzyme, Dicer-like 1 (DCL1)26. After processing, one strand (miRNA*) of the short duplex is typically degraded whereas the mature miRNA strand is incorporated into the RISC (RNA-induced silencing complex) to exert regulatory functions27.
According to the previous work, thousands of DCL1-dependent small RNA (sRNA) loci on their precursors were supported by degradome signatures in Arabidopsis25. We hypothesized that during the processing of the miRNA precursors, Dicer cut pri-miRNA at the 5′ and 3′ ends of miRNA and miRNA* and release the processing intermediates which could be detected by degradome sequencing. In another word, the miRNA-miRNA* duplex loci within the corresponding precursors could be supported by degradome signatures.
To test this hypothesis, the known mature miRNAs and their corresponding pre-miRNAs of Mus musculus and Arabidopsis thaliana (miRBase release 21) were recruited. We took advantage of the degradome sequencing data sets from previous studies, including GSM561025, GSM561026, GSM561027, GSM561028, GSM561029 and GSM561030 of Mus musculus [retrieved from GEO (Gene Expression Omnibus); http://www.ncbi.nlm.nih.gov/geo/] and AxIDT_raw, AxIRP_raw, AxSRP_raw, Col_raw, ein5l_raw, TWF_summary and Tx4F_summary of Arabidopsis (retrieved from PARE (Arabidopsis Parallel Analysis of RNA Ends); http://mpss.udel.edu/at_pare/)28. The degradome data sets were pre-treated as described in “Materials and Methods” and the degradome tags were mapped onto the pre-miRNAs of Mus musculus and Arabidopsis thaliana respectively for extraction of the degradome-based cleavage evidences (see details in “Materials and Methods”). As the Dicer-mediated cleavages locate at 5′ or 3′ ends of miRNAs and miRNA*s on the corresponding precursors, the degradome-supported Dicer cleavage loci could be detected by mapping the degradome tags to pre-miRNAs. Therefore, we searched for the miRNA/miRNA* loci with degradome signatures mapped at either 5′ or 3′ ends which were considered as the Dicer-mediated cropping sites.
The results revealed that, 66 miRNA-miRNA* duplex loci including 123 known mature miRNAs in Mus musculus, corresponding to 96.1% of known miRNAs from the 69 corresponding precursors which contain detectable degradome signatures, were well supported by degradome signatures mostly located at the 5′ end of the miRNAs/miRNA*s (Fig. 1A, Supplementary Table S1). As shown in Fig. 1C and Supplementary Table S1, 140 miRNA-miRNA* duplex loci including 150 known mature miRNAs, corresponding to 74.3% of known miRNAs from 182 miRNA precursors which contain detectable degradome signatures, were found to be supported by the degradome signatures mostly located at the 5′ end of the miRNAs in Arabidopsis as well. However, a fraction of degradome signatures were found to offset from the 5′ or 3′ ends of miRNA-miRNA* duplexes by 1 to 3 nt (Fig. 1B,D). This phenomenon might be caused by Dicer-mediated alternative cropping of pre-miRNAs17. Taken together, the above results demonstrated that a great portion of the miRNA/miRNA* loci could be well supported by the degradome signatures.
Statistic analysis of the miRNA-miRNA* duplex loci supported by degradome signatures.
The frequency of degradome signatures occurred at different ends ① (5′ end of miRNA), ② (3′ end of miRNA), ③ (5′ end of miRNA*) and ④ (3′ end of miRNA*)(marked by red arrows) of recovered known miRNAs within the corresponding precursors in Mus musculus (A) and Arabidopsis (C) were estimated and present in the figures. If two or more degradome signatures were detected locating at 5′/3′ ends of a known miRNA-miRNA* duplex, then these cleavage sites were considered have equal distribution on the discovery of this miRNA-miRNA* duplex. The frequency of degradome signatures occurred at the migration sites of 5′/3′ ends of known miRNA-miRNA* duplexes on their pre-miRNAs in Mus musculus (B) and Arabidopsis (D) were evaluated and profiled. Each ends of duplexes were set as cleavage site “0” and the migration sites located at its upstream are set as −1, −2 and −3 and downstream are set as +1,+2 and +3.
Overview of miRNA Digger pipeline
Based on the above observation, we developed miRNA Digger, a software for genome-wide mining of novel miRNA-miRNA* loci in animals and plants, by using the degradome-supported Dicer cropping sites as the “guidepost”. An overview of the miRNA Digger algorithm is shown in Fig. 2. Briefly, the miRNA Digger starts from the scanning of the degradome signatures on both sense and antisense strands of the genomic sequences. Then, it extracts potential miRNA precursors based on the degradome signatures followed by secondary structure prediction. Only the miRNA precursor candidates with hairpin-like structures are retained. Then, the sRNA short reads from HTS data sets are mapped onto these candidates to identify the miRNA-miRNA* duplexes. AGO enrichment analysis, an optional filtering step, is provided to extract AGO-associated miRNAs. Finally, miRNA Digger reports the known and novel miRNA-miRNA* duplexes and the corresponding pre-miRNAs.
In more details, miRDigger initially subdivides the genomic sequences into 10,000-nt fragments with 500-nt overlaps. The degradome tags are mapped onto the 10,000-nt fragments to obtain the degradome signatures(see “Extraction of the degradome signatures” in “Materials and Methods”)and their positions on the genome. Two pre-miRNA candidates of (30+L) nt in length will be obtained based on a single potential cleavage locus (Fig. 2). The secondary structures of each pre-miRNA candidates were predicted by using RNAfold. Based on the predictions, miRNA Digger retains the pre-miRNA candidates containing classic stem-loop structures (see “RNAfold-based secondary structure identification” in “Materials and Methods”).
Based on the biogenesis model of miRNAs, cropping of a precursor by Dicer could result in the production of a miRNA and a miRNA*26. And, slight variations at the Dicer recognition sites on miRNA precursors could result in the generation of miRNA-centered isomiR clusters along with miRNA*-centered isomiR* clusters22. In this regard, miRNA Digger retains potential pre-miRNAs containing detectable miRNA-miRNA* duplex (see “Screen of potential pre-miRNAs” in “Materials and Methods”).
It is well known that, miRNAs exert their regulatory function by incorporating into AGO-associated RISCs. However, several recent reports have revealed that a considerable number of AGO-associated miRNA*s have biological regulatory roles20,22,29. Therefore, if the AGO-associated sRNA HTS data sets are available, the optional step “AGO-enrichment analysis” is strongly recommended. Thus, miRNA Digger retains miRNA-miRNA* duplexes containing AGO-enriched miRNA(*)s, which can provide researchers with valuable hints for further investigations.
Finally, miRNA Digger outputs a report containing information about the pre-miRNA-coding loci on the genome (which can be used as their original identity), the degradome signatures loci on pre-miRNAs, the sequences and read counts (in RPM) of miRNAs and miRNA*s, the miRNA-miRNA* duplexes loci within the corresponding pre-miRNAs and the sequences and the secondary structures of the pre-miRNAs. If a miRNA(*) sequence has been registered in miRBase, its name will be output simultaneously.
The miRNA Digger software package is available at http://www.bioinfolab.cn/miRNA_Digger/index.html. Additionally, it carries several auxiliary programs for data pre-processing. The sRNA HTS data sets and degradome sequencing data sets should be pre-treated by using the “HTS Data Process” option (see “pre-treatment of high-throughput sequencing data” in “Materials and Methods”) before they could be used for novel miRNA-miRNA* duplex digging. The downloaded chromosome sequences file should be separated into several files by using the “Chromosome Process” option (each file contains one chromosome sequence in fasta format). The “miRBase Update” option links to the miRBase Database, which can be used for downloading the latest version of registered mature miRNAs and pre-miRNAs of different species. MiRNA Digger is designed for Windows System (Vista, Win7 or advanced versions). As the degradome-based scanning of cleavage signals is a time-consuming task, several weeks might be required for scanning a single chromosome. Thus, in order to reduce the total running time, miRNA Digger is designed to run several algorithms in parallel. Additionally, if the running tasks are unfortunately break off under particular situation, miRNA Digger could continue the unfinished tasks when it is restarted.
Case study: discovery of novel miRNAs from different tissues of Arabidopsis
To estimate the sensitivity of miRNA Digger, it was further employed for miRNA-miRNA* duplexes discovery from eight previous reported sRNA HTS data sets of Arabidopsis. Then all of the predicted novel miRNAs and miRNA*s were further recruited for the identification of miRNA–target interactions.
The sRNA HTS data sets from the flowers (six-week-old wild type plants, GSM707678; AGO1-associated sRNA, GSM707682), the leaves (four-week-old wild type plants, GSM707679; AGO1-associated sRNA, GSM707683), the roots (four-week-old wild type plants, GSM707680; AGO1-associated sRNA, GSM707684) and the seedlings (wild type seedlings, GSM707681; AGO1-associated sRNA, GSM707685) of Arabidopsis were contributed by Wang et al.30. The genomic DNA sequences of Arabidopsis were retrieved from the Arabidopsis Information Resource (TAIR, release 10). Eight degradome data sets (AxIDT_raw, AxIRP_raw, AxSRP_raw, Col_raw, ein5l_raw, TWF_summary and Tx4F_summary) were retrieved from PARE (http://mpss.udel.edu/at_pare/)28. The sRNA HTS and degradome sequencing data sets were pre-treated by using the “HTS Data Process” option and the genomic DNA sequences (chromosome 1 to chromosome 5) were pre-processed by using the “Chromosome Process” option. Then, all of the pre-treated data sets were imported into miRNA Digger. As a result, 76 out of 176, 74 out of 153, 71 out of 132 and 74 out of 183 expressed known miRNA-miRNA* duplexes were identified from the flowers, leaves, roots and seedlings, respectively (Fig. 3 and supplementary table S2). It indicates that a large portion of the expressed miRNA-miRNA* duplexes could be uncovered by miRNA Digger. Notably, although these sRNA HTS data sets have been intensively mined for miRNAs30,31,32 miRNA Digger still discovered 24, 25, 21 and 22 novel miRNA-miRNA* duplexes in flowers, leaves, roots and seedlings, respectively (Fig. 4, supplementary table S3 and supplementary figure S1).
Recovery of expressed known miRNAs from different tissues of Arabidopsis by miRNA Digger.
The total numbers of expressed known miRNA-miRNA* duplexes in different tissues of Arabidopsis were shown in blue columns and the numbers of recovered miRNA-miRNA* duplexes by miRNA Digger were shown in orange yellow columns.
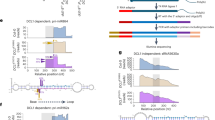
Discovery of novel miRNA from different tissues of Arabidopsis.
The degradome-supported novel miRNA-1/miRNA-1* loci within the corresponding pre-miRNAs were showed in panel A. The miRNA-1 sequences was marked by red line the miRNA-1* was marked by blue line and the degradome signature loci were marked by arrows. The expression level of miRNA-1/miRNA-1* in different tissues were listed in panel B. The degradome sequencing data-based validation of the miRNA-target interactions were showed in panel C, in which two libraries of degradome sequencing data libraries (TWF_summary and Tx4F_summary) were recruited for T-plot profiling (a to f). The IDs of the target transcripts were listed on the top. The y axes measured the normalized reads (in RMP, reads per million) of the degradome signals and the x axes represented the position of the cleavage signals on the target transcripts. The binding sites of the miRNA-1 on its target transcripts were denoted by grey horizontal lines and the dominant cleavage signals were marked by black arrows.
All of the miRNAs and miRNA*s from these newly discovered miRNA-miRNA* duplexes were recruited for target prediction by using miRU algorithm33 and then followed by the degradome-based identification of miRNA–target interactions20,22. As a result, 11 novel miRNA–target interactions involving six novel miRNAs were validated (Table 1 and Supplementary Figure S2).
Among them, miRNA-1 was found to target an RNA processing factor 2 (RPF2) gene (AT1G62670.1), three PPR genes (AT1G62860.1, AT1G62910.1 and AT5G16640.1) and two TPR genes(AT1G62930.1 and AT1G63130.1). The RPF2 protein has been revealed involving in the generation of a distinct 5′ terminus of transcripts of subunit 9 of the NADH DEHYDROGENASE complex (nad9) and RPF2 also determines the efficiency of 5′ end formation of the mRNAs for subunit 3 of the CYTOCHROME C OXIDASE (cox3), in Arabidopsis34. The TRP and PPR proteins are wide-spread on the Arabidopsis and regulate RNA maturation, additionally, the TPR proteins are also considered to function in protein—protein interaction35. miRNA-2 was detected targeting an Argonaute 2 (AGO2) encoding gene, AT1G31280.1, which has been revealed play important role in antibacterial immune responses in Arabidopsis by Jin and his teammates36,37.
miRNA-3 was detected to target an mRNA capping enzyme family protein encoding gene, AT3G09100.1. Eukaryotic mRNA capping enzymes are bifunctional, carrying both RNA triphosphatase (RTPase) and guanylyltransferase (GTase) activities, which is essential for gene expression38,39.
The transposable element gene AT5G33223.1, which contributes to plant gene and genome evolution40, was found to be regulated by miRNA-4. miRNA-5 regulates a MKK9 gene which has been revealed act as a negative regulator of the abiotic stress response in Arabidopsis. miRNA-6 was found to target genes (AT3G54990.1, AT4G36920.1, AT5G04275.1, AT5G60120.1 and AT5G60120.2) involving in plant development, AT3G54990.1 and AT4G36920.1 encoding integrase-type DNA-binding superfamily protein, which has been revealed may regulating the progression of plant developmental cycle41. miR172b (AT5G04275.1) controls the Transition to Autotrophic Development of Arabidopsis42. The targets of early activation tagged (EAT)2, AT5G60120.1 and AT5G60120.2 have been revealed involving in flowering and change of phenotype of Arabidopsis43.
The above results indicated that miRNA Digger could successfully recovered the known miRNA-miRNA* duplexes and the corresponding pre-miRNAs with reasonable sensitivity (52–61%). Secondly, by recruiting HTS data sets, miRNA Digger could perform genomic-wide scan to search for the novel miRNA-miRNA* duplexes. Finally, although the data sets used have already been specifically mined for miRNA22,30, miRNA Digger still predicts 30 novel miRNA-miRNA* duplexes and six of them were identified to play important regulatory functions in Arabidopsis. Therefore, miRNA Digger is a sensitive and reliable tool for genome-wide prediction of novel miRNA-miRNA* duplexes.
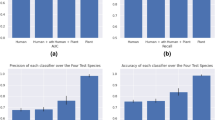
miRDeep is a popular software for miRNA detection based on the biological model of Dicer-mediated processing of the miRNA precursors44. miRPlant is a plant-specific tool for miRNA discovery based on the similar algorithm45. In order to test the functionality of miRNA Digger, we compared all of the miRNAs discovered by miRNA Digger from the flowers of Arabidopsis (Supplementary table S2a and S3a) with those reported by miRDeep and miRPlant with an additional AGO1-enrichment filter step (Supplementary table S4 and S5). As a results, 76, 74 and 54 known miRNA-miRNA* pairs, were recovered by miRNA Digger, miRDeep and miRPlant, respectively. However, only 26 known pairs were recovered by all three tools and 38 known pairs were specifically recovered by miRNA Digger (Fig. 5A). The results indicated that, miRNA Digger presented a different sensitivity and specificity for miRNA detection by relying on the algorithm of finding degradome-supported Dicer cleavage sites. Additionally, 24, 38 and 14 novel miRNA-miRNA* pairs were also discovered by miRNA Digger, miRDeep and miRPlant, respectively. However, no miRNA candidates were discovered by all these three tools (Fig. 5B).
Discussion
Various bioinformatics tools have been developed for systematical detection of novel miRNAs, however, these available tools appeared diverse sensitivity and specificity in miRNA detection among different species9,44. In this study, we proposed a novel algorithm for detection of miRNA-miRNA* duplexes from sRNA HTS data based on the genomic-wide scanning of degradome-supported Dicer/DCL1 cropping sites on genome of animals/plants. Although similar with miRNA Digger that, miRDeep and miRPlant discovers miRNAs based on the processing mechanism of the miRNA precursors by Dicer, they are utterly dependent on the sRNA HTS data sets. However, our approach has the virtue of finding evidence of Dicer-mediated cleavage on miRNA precursors, which reduces the interference of other non-Dicer processed sRNAs and provides a different sensitivity and specificity for miRNA discovery. Therefore, the algorithm of miRNA Digger is a good complement of those previous algorithms with respect to digging miRNA genes from the available sRNA HTS data. Additionally, as it is the first time that the degradome-supported Dicer cleavage sites scanning was reported integrating into the core heart of an algorithm for miRNA detection, we believe that, compared with the previous reported algorithms, miRNA Digger can discover the miRNA-miRNA* duplexes more confidently with this additional criteria.
The sensitivity of miRNA Digger is also determined by the availability and depth of degradome sequencing data and sRNA HTS data. Therefore, miRNA Digger is database-supported and the advances in the field of next-generation sequencing will greatly contribute to the efficiency of miRNA Digger. Additionally, as the application of degradome sequencing technology focused on the validation of miRNA-target interaction previously20,21,22,23,24, the construction and application of miRNA Digger for miRNA mining significantly expanded its application range.
Further improvements of miRNA Digger will be performed in our future work including the enhancement of the rate of degradome-based cleavage signals scanning by integrating with other algorithms to reduce the whole processing time for miRNA discovery. Besides, miRNA Digger is designed only for genome scanning in this study, further improvement will allow the input of expressed sequence tag (EST) for analysis, when no genome sequencing is available.
Conclusion
We developed a novel tool, miRNA Digger, for miRNA detection from different samples of animals and plants, based on the evidence that, the Dicer/DCL1 cropping sites on pre-miRNAs can be well supported by the degradome signatures. miRNA Digger was tested and proved to capable of both recovering the majority of known miRNA-miRNA* duplexes present in heterogeneous samples and reporting novel miRNAs with high confidence, indicating that, miRNA Digger can be widely utilized for novel miRNA discovery and investigation of gene regulation mechanism in both animals and plants.
Materials and Methods
Pre-treatment of high-throughput sequencing data sets
The short sequences containing ‘N’s or with ‘0’ read counts were discarded and then the read count of each short sequence was normalized and measured with reads per million (PRM) as described previously21, to enable the cross-comparison between different libraries.
Extraction of the degradome signatures
The following criteria are applied to extract the degradome-based cleavage signals on genome: 1) for the degradome tags from one library, the averaged read count (in RPM) of all the degradome tags with their 5′-ends mapped to a potential cleavage site should be ‘five times or more’22,46 than that mapped to the regions surrounding this cleavage site; 2) for the tags from the same degradome library, among the degradome tags mapped to a potential cleavage site, the most abundant tag should be among the top 12 most abundant degradome tags that perfectly matched to the corresponding 10000-nt fragment19.
Extraction of pre-miRNA candidates
All of the pre-miRNAs registered in miRBase have been deposited into miRNA Digger, which can be periodically updated. Then, the length of the longest pre-miRNA (the parameter “L”) was calculated for each plant or animal species. When using miRNA Digger, the selection of a species can automatically result in a search for pre-miRNAs by using the default length parameter “L”. Dicer-mediated cleavages probably locate at either end of the miRNA(*) and the length of the mature miRNAs of animals and plants are less than 30 nt25. In this consideration, for the 3′ armed miRNA(*), if a potential cleavage site (degradome-based cleavage signal) locates at the 5′or 3′ end of the miRNA(*), the pre-miRNA candidate will be included within the downstream “L”-nt region plus the 30-nt upstream region. And, for the 5′ armed miRNA(*), if the potential cleavage site locates at the 5′or 3′ end of the miRNA(*), the pre-miRNA candidate will be included within the upstream “L”-nt region plus the 30-nt downstream region. Thus, based on a single potential cleavage locus, two pre-miRNA candidates of (30+L) nt in length will be obtained for further screening.
RNAfold-based secondary structure identification
RNAfold is recruited to predict the thermodynamically optimal secondary structure (minimum free energy secondary structure) of all pre-miRNA candidates47. As we do not know the true 5′ and 3′ ends of pre-miRNA candidates, the unrelated termini of the respective sequences which do not belong to the true pre-miRNA, which might cause the pre-miRNA structure to be thermodynamically suboptimal. Therefore, the pre-miRNA candidates were further employed for base complementarity analysis. As we do not know the precisely position of miRNA/miRNA* within a pre-miRNA candidates, two possible “Stem” structure regions (30nt in length) of a single miRNA precursor candidates around the cleavage locus were analyzed (Fig. 6), if one of them possesses complementarity higher than a setting value (adjustable parameter: the default value is set as 70%), the miRNA precursor candidate was then retained.
The default value of complementarily filtering was set based on the analysis of all “Stem” structure regions (30 nt in length including miRNA-miRNA*) of the known pre-miRNAs of Arabidopsis (miRBase release 21) and we found that, about 95% (404 out of 427) of them possess complementarity higher than 70%. This value can be optimized and adjusted to cart the need of researchers with their interest of miRNAs in different species.
Screen of potential pre-miRNAs
miRNA Digger retains the potential pre-miRNA with detectable miRNA cluster and the corresponding miRNA* cluster. The most abundant miRNA within a miRNA cluster is treated as mature miRNA and the position of its corresponding miRNA* is then determined based on the structurally featured 2-nt 3′ overhangs of the miRNA-miRNA* duplex25. If no such canonical miRNA* is detected within the miRNA* cluster, which might result from fast degradation of miRNA* after Dicer-mediated processing, then the most abundant isomiR* sequence within the miRNA* cluster is treated as the counterpart of mature miRNA. Thus, the filtered pre-miRNA sequences encoding the miRNA and miRNA* sequences, along with 5-nt additional sequence at both 5′ and 3′ ends is obtained.
AGO-enrichment filter
For optional AGO-enrichment filter, the read count of each AGO-associated miRNA should above a setting value (adjustable parameter: the default value is set as 1RPM) and the AGO-enriched miRNA should be several times or more (adjustable parameter: the default parameter is set as 3) abundant than that of the control21.
Compare with miRDeep and miRPlant
The sRNA HTS data sets from the flowers (GSM707678, GSM707682)30 and degradome data sets (AxIDT_raw, AxIRP_raw, AxSRP_raw, Col_raw, ein5l_raw, TWF_summary and Tx4F_summary) (http://mpss.udel.edu/at_pare/)28 were pre-treated by using the “HTS Data Process” and the genomic DNA sequences of Arabidopsis (TAIR, release 10) were pre-treated by using the “Chromosome Process” option. Then, all of them were imported into miRNA Digger for analysis.
The pre-treated sRNA HTS data sets of AGO1-associated sRNA in flowers (GSM707678) and the genomic DNA sequences of Arabidopsis (TAIR, release 10) were imported into miRDeep and miRPlant for analysis. The reported miRNA candidates were further employed for an additional AGO-enrichment filter (select the same enrichment filter parameters as those of miRNA Digger) by using the pre-treated sRNA HTS data set GSM707682 (flower tissues in Col-0) as control.
Additional Information
How to cite this article: Yu, L. et al. miRNA Digger: a comprehensive pipeline for genome-wide novel miRNA mining. Sci. Rep. 6, 18901; doi: 10.1038/srep18901 (2016).
References
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
Alvarez-Garcia, I. & Miska, E. A. MicroRNA functions in animal development and human disease. Development 132, 4653–4662 (2005).
Wilfred, B. R., Wang,W. X. & Nelson,P. T. Energizing miRNA research: A review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol. Genet. Metab. 91, 209–217 (2007).
Tili, E., Michaille, J. J., Costinean, S. & Croce, C. M. MicroRNAs, the immune system and rheumatic disease. Nature Clin. Pract. Rheum. 4, 534–541 (2008).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Boeri, M. et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc. Natl Acad. Sci. U.S.A. 108, 3713–3718 (2011).
Buckley, P. G. et al. Chromosomal and microRNA expression patterns reveal biologically distinct subgroups of 11q- neuroblastoma. Clin. Cancer Res. 16, 2971–2978 (2010).
Lai, C. Y. et al. MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS One 6, e21635 (2011).
Li, Y. et al. Performance comparison and evaluation of software tools for microRNA deep-sequencing data analysis. Nucleic Acids Res. 40, 4298–4305 (2012).
Hendrix, D., Levine, M. & Shi, W. MmetihRodTRAP, a computational method for the systematic identification of miRNAs from high throughput sequencing data. Genome Biology. 11:R39 (2010).
Williamson, V. et al. Detecting miRNAs in deep-sequencing data: a software performance comparison and evaluation. Brief. Bioinform. 14, 36–45 (2013).
Zhang, Z., Jiang, L., Wang, J. & Chen, M. MTide: an integrated tool for the identification of miRNA-target interaction in plants. Bioinformatics. 31, 290–291 (2014).
Wang, W. C. et al. miRExpress: analyzing high-throughput sequencing data for profiling microRNA expression. BMC Bioinformatics. 10, 328 (2009)
Mathelier, A. & Carbone A. MIReNA: finding microRNAs with high accuracy and no learning at genome scale and from deep sequencing data. Bioinformatics 26, 2226–2234 (2010).
Leclercq, M., Diallo, A. B. & Blanchette, M. Computational prediction of the localization of microRNAs within their pre-miRNA. Nucleic Acids Res. 41, 7200–7211 (2013).
He, C. et al. MiRmat: mature microRNA sequence prediction. PLoS One. 7, e51673 (2012).
Pritchard, C. C., Cheng, H. H. & Tewari, M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 13, 358–369 (2012).
Reddy, A. S., Marquez, Y., Kalyna, M. & Barta, A. Complexity of the alternative splicing landscape in plants. Plant Cell 25, 3657–3683 (2013).
Fukunaga, R. et al. Dicer partner proteins tune the length of mature miRNAs in flies and mammals. Cell 151, 533–46 (2012).
German, M. A. et al. Global identification of microRNA-target RNA pairs by parallel analysis of RNA ends. Nat Biotechnol. 26, 941–946 (2008).
Liu, S. et al. StarScan: a web server for scanning small RNA targets from degradome sequencing data. Nucleic Acids Res. 43, W480–W486 (2015).
Shao, C., Chen, M. & Meng, Y. A reversed framework for the identification of microRNA-target pairs in plants. Brief. Bioinform. 14, 293–301 (2013).
Li, Y. F. et al. Transcriptome-wide identification of microRNA targets in rice. Plant J. 62, 742–759 (2010).
Meng, Y., Gou, L., Chen, D., Wu, P. & Chen, M. High-throughput degradome sequencing can be used to gain insights into microRNA precursor metabolism. J Exp Bot. 61, 3833–3837 (2010).
Ma, X., Shao, C., Jin, Y., Wang, H. & Meng, Y. Long non-coding RNA: a novel endogenous source for the generation of Dicer-like 1-dependent small RNAs in Arabidopsis thaliana. RNA Biol. 11, 373–390 (2014).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Min, H. & Yoon, S. Got target? Computational methods for microRNA target prediction and their extension. Exp Mol Med. 42, 233–244 (2010).
Nakano, M. et al. Plant MPSS databases: signature-based transcriptional resources for analyses of mRNA and small RNA. Nucleic Acids Res. 34 (Database issue), D731–735 (2006).
Guo, L. & Chen, F. A challenge for miRNA: multiple isomiRs in miRNAomics. Gene 544, 1–7 (2014).
Wang, H. et al. Deep sequencing of small RNAs specifically associated with Arabidopsis AGO1 and AGO4 uncovers new AGO functions. Plant J. 67, 292–304 (2011).
Meng,Y., Shao, C., Ma, X., Wang, H. & Chen, M. Expression-Based Functional Investigation of the Organ-Specific MicroRNAs in Arabidopsis. PLoS One. 7, e50870 (2012).
Shao, C. et al. Identification of novel microRNA-like-coding sites on the long-stem microRNA precursors in Arabidopsis. Gene 527, 477–483 (2013).
Zhang, Y. miRU: an automated plant miRNA target prediction server, Nucleic. Acids Res. 33(Web Server issue), W701–704 (2005).
Jonietz, C., Forner, J., Hölzle, A., Thuss, S. & Binder, S. RNA PROCESSING FACTOR2 is required for 5′ end processing of nad9 and cox3 mRNAs in mitochondria of Arabidopsis thaliana. Plant Cell 22, 443–453 (2010).
Geddy, R. & Brown, G. G. Genes encoding pentatricopeptide repeat (PPR) proteins are not conserved in location in plant genomes and may be subject to diversifying selection. BMC Genomics. 8, 130 (2007).
Zhang, X. et al. Arabidopsis Argonaute 2 regulates innate immunity via miRNA393(*)-mediated silencing of a Golgi-localized SNARE gene, MEMB12. Mol. Cell 42, 356–366 (2011).
Seo, J. K., Wu, J., Lii, Y., Li, Y. & Jin, H. Contribution of small RNA pathway components in plant immunity. Mol. Plant Microbe. Interact. 26, 617–625 (2013).
Cho, E. J., Takagi, T., Moore, C. R. & Buratowski, S. mRNA capping enzyme is recruited to the transcription complex by phosphorylation of the RNA polymerase II carboxy-terminal domain. Genes Dev. 11, 3319–3326 (1997).
Takagi, T. et al. The Caenorhabditis elegans mRNA 5′-capping enzyme. In vitro and in vivo characterization. J. Biol. Chem. 278, 14174–14184 (2003).
Bennetzen, J. L. Transposable element contributions to plant gene and genome evolution. Plant Mol Biol. 42, 251–269 (2000).
Balaji, S., Babu, M. M., Iyer, L. M. & Aravind, L. Discovery of the principal specific transcription factors of Apicomplexa and their implication for the evolution of the AP2-integrase DNA binding domains. Nucleic. Acids Res. 33, 3994–4006 (2005).
Zou, Y. et al. miR172b controls the transition to autotrophic development inhibited by ABA in Arabidopsis. PLoS One 8, e64770 (2013).
Aukerman, M. J. & Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 15, 2730–41 (2003).
Friedländer, M. R. et al. Discovering microRNAs from deep sequencing data using miRDeep. Nat Biotechnol. 26, 407–415 (2008).
An, J., Lai, J., Sajjanhar, A., Lehman, M. L. & Nelson, C. C. miRPlant: an integrated tool for identification of plant miRNA from RNA sequencing data. BMC Bioinformatics. 15, 275 (2014).
Ma, X., Shao, C., Jin, Y., Wang, H. & Meng, Y. A novel endogenous source for the generation of Dicer-like 1-dependent small RNAs in Arabidopsis thaliana. RNA Biology. 11, 373–390 (2014).
Clote, P., Ponty, Y. & Steyaert, J. M. Expected distance between terminal nucleotides of RNA secondary structures. J. Math. Biol. 65, 581–599 (2012).
Acknowledgements
We would like to thank all the publicly available data sets and the scientists behind them. This work was supported by the National Natural Sciences Foundation of China [31271380], [31371328] and [31571349], Zhejiang Provincial Natural Science Foundation of China [LY15C060004], [LY15C060006] and [LQ16C060001], Scientific Research Fund of Zhejiang Provincial Education Department [Y201533314] and the Research Fund of Huzhou University [201441].
Author information
Authors and Affiliations
Contributions
L.Y. wrote the main manuscript text and prepared all the figures and tables. C.S. constructed the miRNA Digger pipeline. X.Y. created the website for downloading of miRNA Digger package. M.C., Y.M., L.Y. and C.S. conceived and designed the experiments. L.Y. and C.S. analyzed the data. Y.Z. accomplished the comparative analysis of miRDeep and miRPlant. All of the authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yu, L., Shao, C., Ye, X. et al. miRNA Digger: a comprehensive pipeline for genome-wide novel miRNA mining. Sci Rep 6, 18901 (2016). https://doi.org/10.1038/srep18901
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18901
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.