Abstract
Atherosclerosis is the primary cause of cardiovascular events and its molecular mechanism urgently needs to be clarified. In our study, atheromatous plaques (ATH) and macroscopically intact tissue (MIT) sampled from 32 patients were compared and an integrated series of bioinformatic microarray analyses were used to identify altered genes and pathways. Our work showed 816 genes were differentially expressed between ATH and MIT, including 443 that were up-regulated and 373 that were down-regulated in ATH tissues. GO functional-enrichment analysis for differentially expressed genes (DEGs) indicated that genes related to the “immune response” and “muscle contraction” were altered in ATHs. KEGG pathway-enrichment analysis showed that up-regulated DEGs were significantly enriched in the “FcεRI-mediated signaling pathway”, while down-regulated genes were significantly enriched in the “transforming growth factor-β signaling pathway”. Protein-protein interaction network and module analysis demonstrated that VAV1, SYK, LYN and PTPN6 may play critical roles in the network. Additionally, similar observations were seen in a validation study where SYK, LYN and PTPN6 were markedly elevated in ATH. All in all, identification of these genes and pathways not only provides new insights into the pathogenesis of atherosclerosis, but may also aid in the development of prognostic and therapeutic biomarkers for advanced atheroma.
Similar content being viewed by others
Introduction
Cardiovascular diseases are the leading cause of morbidity and mortality worldwide and atherosclerosis is known to be the primary underlying factor responsible for the development of these diseases1. Despite extensive research, the detailed molecular mechanisms underlying the development of atherosclerosis and causing plaque rupture still remain unclear and new findings are urgently needed to complement the current knowledge and to identify new drug targets2. Rapid advances in biological technology, including DNA microarrays, able to detect the expression levels of tens of thousands of genes simultaneously, might help to provide comprehensive insights into the pathogenesis of atherosclerosis.
Gene-expression profiling of atherosclerosis has recently been used to identify genes and pathways relevant to vascular pathophysiology. It has previously been used to analyze altered gene expression in normal and diseased arteries3, establish crucial players in atherosclerotic plaque progression4,5, identify differentially expressed genes (DEGs) by comparing plaques with or without cerebrovascular symptoms6, discover candidate pathways and genes related to atherosclerosis7 and find gene expression changes of atherosclerotic plaques in different vascular beds8. However, some drawbacks are associated with those previous studies. In microarray studies comparing atheroma with normal tissues3,7, differences in the cellular compositions and morphologies of atherosclerotic plaques and normal arteries may result in differential gene expression profiles that simply reflect this variation9. In addition, irregular sample-collection methods existed in some studies3,8 , for example, samples from different sites or sources or small sample sizes, may affect the reliability of studies10. Furthermore, in animal model experiments4,5, a high degree of variability in plaque composition and gene expression between humans and animal models may limit the extension of cDNA array studies on animal material to clinical use11. Features of unstable plaques, such as surface ulceration, rupture, intraplaque hemorrhage and thrombus, may also occur in both asymptomatic and symptomatic patients, which may also confound studies6 that classify samples according to patient symptomatology12. Additionally, the relative lack of systematic bioinformatic analysis of cDNA microarrays in current studies limits the effective exploitation of gene-expression data sets10. Therefore, an integrated bioinformatic analysis based on cDNA microarray studies of human tissues may help to clarify the mechanisms underlying the development and progression of atherosclerosis.
To our knowledge, the variations between different individuals or blood vessels may affect the reliability of studies and it is very difficult to obtain healthy and diseased tissue from the same blood vessel of the same individual in human studies. To overcome the difficulty, we used a gene expression dataset from a previously published study13, comparing atheroma and its surrounding tissues from the same individual to track gene changes with disease progression and validated our findings with similar tissues. Besides, to interpret the biological relevance of these changes in gene expression, the microarray data were analyzed by integrated bioinformatic analysis expanding on traditional microarray analysis methods, namely gene-ontology and pathway analysis, thereby allowing the construction of interaction networks, that might identify novel prognostic markers and therapeutic targets.
Results
Identification of differentially expressed genes
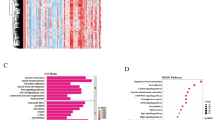
Through our microarray analysis, a total of 816 differentially-expressed genes (DEGs) were identified between atheroma plaque (ATH) and macroscopically-intact tissue (MIT), including that 443 genes were up-regulated and 373 genes were down-regulated (Fig. 1A,B). The greatest fold differential expressions were a six-fold up-regulation of the FABP4 gene (fatty acid-binding protein 4) and a 3.3-fold down-regulation of the CNTN1 gene (contactin 1) in ATH compared with MIT.
Differentially expressed genes were identified between ATH and MIT.
(A) The overlapping gene set dually identified by the SAM and FC method. (B) Volcano plots for all differentially expressed genes in comparison. The dots indicate that up-(red) and down-regulated (blue) DEGs were significant both at false-discovery rate <0.05 and Fold-change >1.5 or <0.667.
Gene ontology and pathway analyses
Two hundred-ninety and 26 GO terms were significantly enriched among the up-regulated and down-regulated genes, respectively. Table 1 shows the ten most overrepresented GO terms for the up-regulated and down-regulated DEGs, including immune-related biological process, such as “cell activation” and “cytokine production” and “muscle system process”. Meanwhile, the KEGG pathway analysis identified 77 and 26 significantly enriched pathways for up-regulated and down-regulated genes, respectively. The ten most overrepresented KEGG pathways for up-regulated and down-regulated DEGs are shown in Table 2, with “B cell receptor signaling pathway” and “TGF-beta signaling pathway” being most significantly enriched.
Protein–protein interaction (PPI) network analysis
We constructed a PPI network to identify more important proteins and biological modules that may play crucial roles in the development of atherosclerosis. To confine the interactions only to those close to the DEGs, only first level interactions between DEGs and their neighbors were selected. There were 3,990 PPI pairs and 2,491 nodes in our constructed PPI network. The degree represents the number of neighboring nodes in the network and changes in the proteins/genes with higher degrees have more effects than changes in those with smaller degrees. SMAD9, LYN, PTPN6, ZBTB16, SYK, PRKCB, SVIL, VAV1, BMPR1B and BTK were located in the more important positions of network with higher degrees of 108, 102, 95, 81, 74, 66, 63, 62, 60 and 56, respectively, indicating those proteins play irreplaceable and critical roles in the maintaining the whole protein interactions in the network (Fig. 2A).
PPI interactions network of DEGs and the disease-relevant module found in the network.
Nodes and links represent human proteins and protein interactions; Nodes represent the encoding genes of proteins; Red color indicates up-regulated genes annotated in the PPI network; Blue color indicates the down-regulated genes annotated in the PPI network; Pink nodes represent the non-DEGs which have an interaction with DEGs (A,B). The disease-relevant module contains the most nodes in CFinder software (B).
CFinder software was used to identify the disease-relevant modules in the PPI network. Figure 2B shows the module containing the most nodes with parameter k = 5. This module contained eight DEGs, including CSF2RB, LCP2, LYN, PLCG2, PTPN6, PTPRC, SYK and VAV1. Jointly using topology of network and module analysis to select candidates would identify genes that have higher significance in the PPI network. Finally, we focused on four genes (SYK, LYN, PTPN6 and VAV1) that were overlapped between the top 10 nodes ranked by degrees and the disease-relevant modules in the PPI network (Table 3).
Verification of differentially expression genes in clinical samples
To confirm and validate the expression of four candidates determined from microarray data analysis in clinical samples, eight fresh sets of ATH and MIT samples were collected from surgery. The mRNA expression of four candidates (VAV1, SYK, LYN and PTPN6) were examined by qRT-PCR in eight sets of atherosclerotic sample (8 ATH and 8 MIT, n = 16). The results from qRT-PCR showed that mRNA level of VAV1 (12.6 ± 5.0), LYN (3.7 ± 1.1), SYK (13.1 ± 4.4) and PTPN6 (11.2 ± 4.9) increased by 12.6, 3.7, 13.1 and 11.2 folds respectively in ATH compared with in MIT (Fig. 3A). The results are consistent with data from microarray analysis although the differences in mRNA level were even higher than the differences determined in the microarray analysis. Moreover, the whole lysates from four sets of atherosclerotic samples (4 ATH and 4 MIT, n = 8) were analyzed by western blot. As Fig. 3B shows, the protein level of LYN (2.2 ± 0.4), SYK (27.3 ± 8.3) and PTPN6 (2.0 ± 0.5) significantly increased by degree of 2.2, 27.3 and 2.0 folds respectively in all ATH samples compared with MIT samples. We noticed that VAV1 was not detected in the atherosclerotic samples at the protein level by current antibody.
Validation of four candidate genes expression determined from microarray data analysis in clinical samples.
(A) VAV1, LYN, SYK and PTPN6 expression was analyzed by qRT- PCR in 8 sets of atherosclerotic sample (8 ATH and 8 MIT, n = 16). *p < 0.05; **p < 0.01. The data is a representative of three independent experiments. (B) whole lysate from atherosclerotic samples (4 ATH and 4 MIT, n = 8) were analyzed by western blot. Top panel, the representative images of three independent experiments; Bottom panel, the quantitative data of the images in western blot by ImageJ software. The full-length blots are presented in Supplementary materials.
Discussion
Microarray studies have great potential to provide novel insights into the pathogenesis of complex diseases. In the present study, we systematically applied integrated bioinformatic methods to mine new candidate players in the process of atherosclerosis and validated our findings in an independent set of samples at both mRNA and protein levels. In our study, we identified a total of 816 genes differentially expressed in ATH compared with MIT, including 443 up-regulated and 373 down-regulated DEGs. GO functional-enrichment analysis of these DEGs showed that genes mainly related to inflammation and immune responses were altered with disease progression. “Cell adhesion”, “proliferation”, “differentiation”, “motility”, “cell death”, “lipid metabolism” and “immune response” have all been reportedly associated with atherosclerosis14 and these processes were also identified in our enrichment results. Interestingly, although we identified similar numbers of up-regulated and down-regulated genes in ATHs, we observed an excess of significant GO categories for up-regulated genes, suggesting that the up-regulated genes are functionally more important in atherosclerosis progression.
Pathway-enrichment results showed an overabundance of immune and inflammatory signals, represented by the “chemokine-signaling pathway”, “natural killer cell-mediated cytotoxicity” and “FcεRI-signaling pathway” in atherosclerosis. Our finding indicates that innate and adaptive immune cells might contribute to the development and progression of atherosclerosis, especially in the advanced stages. Hypercholesterolemia was initially considered to be the major risk factor for atherosclerosis, but recent advances have proven that chronic inflammation and autoimmunity play major roles in the initiation and progression of the disease15, as supported by our pathway-enrichment results. In addition, the “TGF-β signaling”, “calcium signaling” and “osteoclast differentiation” were all significantly enriched among DEGs.
Mast cells are frequently in an activated state and participate in the process of atherosclerosis16. “FcεRI-mediated signaling” in mast cells is initiated by the interaction of an antigen with IgE bound to the extracellular domain of the α-chain of FcεRI. The mast cells activated by crosslinking of the FcεRI via IgE-antigen complexes could release and secrete biogenic amines, cytokines, lipid mediators and proteoglycans, which contribute to inflammatory responses. IgE and FcεRI have been implicated in several aspects of autoimmunity and chronic inflammatory diseases17. LYN, SYK and VAV1, the key modulators in our PPI network and module, are also implicated in this pathway (Figure S1). Our results indicated that this pathway might play a crucial role in the process of atherosclerosis; however, there is currently no proof of a relationship between “FcεRI-mediated signaling” and atherosclerosis. And it is worth noting that calcium signaling is associated with FcεRI -mediataed signaling. “Calcium signaling”18, “TGF-β signaling”19 and “osteoclast differentiation”20 have all been shown to be involved in the process of atherosclerosis. Our pathway-enrichment analysis supported the involvement of some pathways known to be associated with atherosclerosis initiation and progression and also highlighted the FcεRI-mediated signaling pathway, which, to the best of our knowledge, has not previously been reported in association with carotid atheroma.
Our PPI network and further module analysis showed that VAV1, SYK, LYN and PTPN6 overlapped between the top 10 nodes and the disease-relevant module, suggesting that these genes play more crucial roles in the pathogenesis of carotid atheroma. Several studies have indicated important related functions for SYK and VAV1, suggesting that they play significant roles in atherosclerosis. SYK has been reported to be involved in the pathogenesis of atherosclerosis by activating monocyte chemotactic protein-1 expression21. Choi et al.22 found that TLR4/SYK-mediated macrophage responses may contribute to chronic inflammation in human atherosclerosis. Furthermore, the SYK inhibitor fostamatinib attenuated atherogenesis in mice, suggesting that SYK is a potential anti-inflammatory therapeutic target in atherosclerosis23. The validation study also indicated SYK was up-regulated both at the level of mRNA and protein which strengthens the assertion that it might play an important role in atherosclerosis development.
VAV1, a member of the VAV gene family, is expressed exclusively in hematopoietic cells. It is a signal transduction molecule that acts as guanine nucleotide exchange factor for Rac1 and Rho GTPases and also functions as an adaptor platform24. VAV1 impacts on processes that are highly relevant to atherogenesis, such as NADPH oxidase-mediated generation of reactive oxygen species, cell death and leukocyte activation. An in vivo carotid artery thrombosis model showed that genetic deletion of Vav1 and Vav3 together may prevent the development of occlusive thrombi in mice fed a high-fat diet25. Deletion of Vav1 alone led to modest inhibition of oxidized low-density lipoprotein uptake and foam-cell formation, while deletion of both Vav1 and Vav3 led to nearly complete inhibition of oxidized low-density lipoprotein uptake and foam-cell formation, suggesting that Vavs act as a critical regulator in the process of atherogenesis and thus represents a novel therapeutic target26. In our validation study, VAV1 was not detected in the atherosclerotic samples at the protein level by current antibody, which worked well in the experiment of positive control tissues. Another VAV1 antibody was also applied, which still could not dectect the expression of this gene. Though we found similar expression patterns for upregulated VAV1 in both the qPCR and microarray analyses of ATH samples, VAV1 may not play a major role in the progression of atherosclerosis because its expression may be blocked at the translation phase.
LYN encodes a tyrosine protein kinase that is involved in the regulation of mast-cell degranulation. Lyn is the major Src-family kinase regulating glycoprotein VI signaling and its absence caused a delay in activation and a marked reduction in platelet aggregation on collagen in a laser-injury model27. However, an apparently contradictory study showed that Lyn inhibited platelet activation and that Lyn was increasingly inactivated as platelet aggregation progressed28. Miki et al.29 suggested that Lyn plays an important role in the metabolism of serum lipids and could induce the expression of monocyte chemotactic protein-1, which is related to atherosclerosis, during the development of atherosclerotic lesions on high-fat diets. Previous studies reflect the complex roles of LYN and our validation experiments showed that LYN was up-regulated in the atheroma at the levels of mRNA and proteins, which might promote the progression of this disease.
PTPN6 is a member of the protein tyrosine phosphatase family of signaling molecules. It regulates a variety of cellular processes including cell growth, differentiation, mitotic cycle and oncogenic transformation30. Kamata et al.31 suggested that PTPN6 acts as a negative regulator in the development of allergic responses such as allergic asthma. Dubois et al.32 concluded that PTPN6 played a crucial role in the negative modulation of insulin action and clearance in the liver, thus regulating whole-body glucose homeostasis. However, here we found PTPN6 up-regulated in atheroma at both the mRNA and proteins level and for the first time linked PTPN6 to atherosclerosis.
Additionally, to ensure the robustness of our candidate DEGs, we confirmed the expression of our candidate genes in another dataset GSE28829 (Supplementary Figure S2). The analysis revealed that a similar representation of the gene expression patterns of our candidate DEGs was seen in the dataset GSE28829, suggesting that VAV1, SYK, LYN and PTPN6 genes may play an important role in progression of atherosclerosis.
To our knowledge, this is the first integrated bioinformatic analysis comparing gene expression between carotid plaque and macroscopically intact arterial tissue. The large-scale gene expression profile analysis in our study is a significant strength in addition to the fact that paired tissue samples were obtained from the same individual. Our integrated methods are based on pre-specified specific algorithms, established topology information of networks and existing knowledge from databases and literature. The integrated methods have an advantage over traditional, single-analysis microarray approaches and other enrichment-analysis methods, such as DAVID33 and NetGestalt34 and ensure the reliability and accuracy of the results. Key candidates were also validated in Chinese patients, indicating the generalization of molecular mechanisms among different ethnic groups. Meanwhile, our integrated bioinformatic analysis might reveal the relationship between DEGs at molecular interaction and pathway levels, which provided some clues for the deeper mechanism of our candidate DEGs.
The discrepancies of expression between the qRT-PCR and microarray results may have been caused by a sensitivity bias between the two methods, difference in ethnicity, diet and lifestyle between French and Chinese people or by the use of different statistical methods in qRT-PCR and microarray. However, there are also some limitations in our study. First, the study population from the microarray analysis underwent carotid endarterectomy at the university hospital of Lyon, so the gene expression profile may be influenced by their ethnicity, diet and lifestyle. Secondly, the cohort consisted of older subjects that were predominantly male and the majority had hypertension. The generalization of our findings is unknown as the present results are limited to high risk populations with signs of atherosclerosis and severe symptoms. Finally, our work is a reanalysis of previously published dataset and although some previous work and our validation experiments support our gene expression analysis results, the work requires further study to identify the mechanism of action and to assess the relevance of our findings. This work serves as an excellent foundation and reference for further studies to expand on these findings in the future.
Conclusion
This study identified SYK, LYN, PTPN6 and the “FcεRI-mediated signaling pathway” as potential candidate players involved in the pathogenesis of atherosclerosis. These findings enhance our understanding of the molecular mechanisms of this important disease. Further studies, such as gene functional studies, are needed to support the results of our study, with the aim of identifying candidate biomarkers with sufficient predictive power to act as prognostic and therapeutic biomarkers for advanced atheroma.
Methods
Source of data
An existing dataset GSE43292 within the Gene Expression Omnibus database was used for this work and obtained through approved access. The dataset was generated using the Affymetrix Human Gene 1.0 ST Array13. Ethical approval, sample tissue collection and preparation methods and characteristics of study participants were described in a previous report13. In brief, the dataset included 32 from 34 consecutive patients admitted to the university hospital of Lyon in 2009 for carotid endarterectomy. Paired samples were taken from individuals meaning that 64 carotid artery samples were analyzed. The mean age of participants was 70 years (±10 years) and the majority were male and with hypertension, with elevated blood lipid levels and just over one third of the sample were diabetic (Supplementary Table S1)13. Tissue samples were removed during surgery and dissected into two fragments: atheroma plaque tissue (ATH, subsequently identified as mostly stage IV and/or V lesions according to the American Heart Association classification) and macroscopically intact tissue (MIT, almost exclusively composed of stage I and II lesions)13.
Identification of differentially expressed genes
The raw gene dataset obtained from the previous work13 was converted into expression measures and background correction and quantile data normalization were performed using the robust multiarray average algorithm from the Affy package to obtain the expression profile data35. After deleting duplicated probes and averaging the multiple probes values for the same Entrez Genes (the unique integers as identifiers for gene records)36, we finally obtained expression profiles for 19,924 genes in the 64 samples.
Because the differentially expressed genes (DEGs) might have stronger relationship with the development of disease, the significance analysis of microarrays37 and fold-change methods were jointly used to identify DEGs between ATH and MIT.
Functional-enrichment analysis
We integrated GO annotation into the total DEGs by mining for enriched GO terms of proteins using the R-based GO function software packages, which extracts biologically relevant terms from statistically significant GO terms for a disease38,39.
The DEGs were chosen for further analysis of Kyoto Encyclopedia of Gene and Genome (KEGG) enrichment. The SubpathwayMiner is a pathway identification system40 and accurately assessed the pathway structure to locate disease-relevant KEGG pathways41 and subpathways in DEGs relative to the genomic background.
Protein–protein interaction network construction
In the study, we downloaded protein-protein interaction (PPI) data from human protein reference database (Release 9) on the website (http://www.hprd.org/)42. These interactions were derived from literature of experimental validation, including physical interactions and enzymatic reactions found in signal transduction pathways. The PPI data were preprocessed, including removing redundancy and self-loops, resulting in a connected network of 9,618 nodes (unique Entrez IDs) and 39,240 documented interactions. PPI networks are visualized in Cytoscape43 with the nodes representing the proteins/genes and the edges representing interactions between any two proteins/genes.
We constructed the PPI network by mapping the DEGs to the PPI network using the following steps. First, we extracted the nodes and relationships between DEGs and their direct interacting neighbors to confine the interactions only to those close to the DEGs using R software44, with each pair of interacting proteins in two lists of a text file. The DEGs (gene symbols) were listed in a NOA file with different node attribution annotations (down-regulated genes, up-regulated genes) and mapped to the constructed PPI network by the menu of “File-Import-Node Attributes”. Second, the degrees of nodes in the PPI networks were calculated by Network Analysis plugin by the menu of “Plugins- Network Analysis-Analyze network”. In our network, the degree of a node was the number of neighboring nodes in the network and node size was proportional to the degree of the protein. Third, CFinder software45 was used to find disease-related modules based on the Clique Percolation Method46, which is a free software for finding and visualizing overlapping dense groups of nodes in networks. PPI data from a text file was imported using the menu of “File- Open new network-run” with default parameters. The results of CFinder are highly correlated to the value of the parameter k. Larger k values correspond to smaller subgraphs with a higher density of links within them.
Ethics statement and validation study sample collection
The collection of clinical samples was under approval by the Medical Ethics Committee of NanFang Hospital (Number: NFEC-2014-117) and informed consent was obtained from all subjects. The study was carried out in accordance with the standards set by the Declaration of Helsinki and Good Clinical Practice guidelines.
Human carotid atherosclerotic plaques were obtained from patients who underwent endarterectomy at the Vascular Surgery Department of Nanfang hospital of Southern medical university (Guangzhou, China). Patients (n = 8, mean age: 67.3 years, range: 53–80 years) with internal carotid artery stenosis >70% were included. The carotid atheromas were separated as ATH and MIT according to macroscopic observation. Dissected vascular tissues were rapidly frozen in liquid nitrogen and stored at –80 °C. Sample characteristics used for each experiment are shown in supplementary information Table S2.
Quantitative real-time PCR (qRT-PCR)
Total RNA from specimens (n = 16) was isolated using Trizol reagent (TaKaRa Bio Inc, Japan) according to the manufacturer’s instructions. Complementary DNA was synthesized from 1000 ng of total RNA using the PrimeScriptTM RT reagent Kit with gDNA Eraser (TaKaRa Bio Inc, RR047Q) according to the manufacturer’s instruction, including the DNase step. Amplification was performed using SYBR® Premix Ex TaqTM (TaKaRa Bio Inc, RR420A). Quantitative real-time polymerase chain reaction (q-PCR) analysis was performed on Lightcycler96 (Roche Applied Science) according to the manufacturer’s protocol. GAPDH was used as internal control to normalize mRNA levels. All experiments were repeated three times. Primer sequences are listed in Table 4. Analysis was performed by the comparative delta–delta–Ct method47.
Western blotting
Samples (n = 8) were taken from ultra-low temperature freezer and crushed in lysis buffer under liquid nitrogen. After detecting the concentration, proteins were separated on 7% SDS-polyacrylamide gel electrophoresis and transferred onto nitrocellulose membranes. Membranes were blocked with TBS-T(TBS/0.1% Tween-20) containing 5% non-fat dry milk for 1.5 hour at room temperature. Then, they were probed successively with mouse monoclonal anti-VAV1 (Cell Signaling Technology, Danvers, MA, USA), rabbit monoclonal anti-LYN (Abcam, Cambridge, MA, USA), rabbit monoclonal anti-PTPN6 (Abcam, Cambridge, MA, USA) and rabbit monoclonal anti-SYK (Abcam, Cambridge, MA, USA) antibodies at 4 °C overnight. Mouse monoclonal antibodies against GAPDH (Cell Signaling Technology, Danvers, MA, USA) were used as a loading control. Membranes were washed in TBS-T (TBS/0.1% Tween-20) three times for 5 min and probed with an anti-rabbit or -mouse HRP-conjugated secondary antibody in TBS-T with 5% of nonfat dry milk at room temperature for 1.5 h. Protein detection was performed using ECL reagents. Western blot bands were scanned using the ChemiDoc™ XRS Imaging System (BioRad, USA). Western blot bands were quantified using ImageJ software by measuring the band intensity for each group and normalized by GAPDH. The final results are expressed as fold changes by normalizing the data to the control values.
Statistical analyses
Microarray analysis: DEGs for the microarray were identified using the fold-change and significance analysis of microarrays methods, with multiple testing corrections applied using the Benjamini-Hochberg method48. False-discovery rate <0.05 and fold-change >1.5 or <0.667 were set as the cutoff values of DEGs. For the functional enrichment analysis, significantly enriched GO terms in DEGs relative to the genomic background by GO function software packages were identified using the hypergeometric tests with an adjusted p-value <0.01, calculated by the Benjamini-Hochberg method48. Pathway-enrichment analysis was done using the R-based SubpathwayMiner software packages. Significantly enriched pathways were identified using hypergeometric tests and a p-value <0.01 was applied as the cut-off value for statistical significance.
Validation study: The data in this study are shown as the mean ± S.D. For the real time PCR, groups were compared using the Wilcoxon signed-rank test for continuous variables (SPSS 19.0, Chicago, IL) and a 2-sided p value <0.05 was considered statistically significant.
Additional Information
How to cite this article: Nai, W. et al. Identification of novel genes and pathways in carotid atheroma using integrated bioinformatic methods. Sci. Rep. 6, 18764; doi: 10.1038/srep18764 (2016).
Change history
19 August 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Mendis, S., Puska, P. & Norrving, B. Global Atlas on Cardiovascular Disease Prevention and Control. (Geneva: World Health Organization, 2011).
Shalhoub, J. et al. Systems biology of human atherosclerosis. Vasc Endovascular Surg 48, 5–17 (2014).
Hiltunen, M. O. et al. Changes in gene expression in atherosclerotic plaques analyzed using DNA array. Atherosclerosis 165, 23–32 (2002).
Chen, Y. C. et al. A novel mouse model of atherosclerotic plaque instability for drug testing and mechanistic/therapeutic discoveries using gene and microRNA expression profiling. Circ Res 113, 252–265 (2013).
Koizumi, G. et al. Gene expression in the vascular wall of the aortic arch in spontaneously hypertensive hyperlipidemic model rats using DNA microarray analysis. Life Sci 93, 495–502 (2013).
Gertow, K. et al. 12- and 15-lipoxygenases in human carotid atherosclerotic lesions: associations with cerebrovascular symptoms. Atherosclerosis 215, 411–416 (2011).
King, J. Y. et al. Pathway analysis of coronary atherosclerosis. Physiol Genomics 23, 103–118 (2005).
Levula, M. et al. Genes involved in systemic and arterial bed dependent atherosclerosis–Tampere Vascular study. PLoS One 7, e33787 (2012).
Papaspyridonos, M. et al. Novel candidate genes in unstable areas of human atherosclerotic plaques. Arterioscler Thromb Vasc Biol 26, 1837–1844 (2006).
Bijnens, A. P. et al. Genome-wide expression studies of atherosclerosis: critical issues in methodology, analysis, interpretation of transcriptomics data. Arterioscler Thromb Vasc Biol 26, 1226–1235 (2006).
Laguna, J. C. & Alegret, M. Regulation of gene expression in atherosclerosis: insights from microarray studies in monocytes/macrophages. Pharmacogenomics 13, 477–495 (2012).
Golledge, J., Greenhalgh, R. M. & Davies, A. H. The symptomatic carotid plaque. Stroke 31, 774–781 (2000).
Ayari, H. & Bricca, G. Identification of two genes potentially associated in iron-heme homeostasis in human carotid plaque using microarray analysis. J Biosci 38, 311–315 (2013).
Van Assche, T. et al. Transcription profiles of aortic smooth muscle cells from atherosclerosis-prone and -resistant regions in young apolipoprotein E-deficient mice before plaque development. J Vasc Res 48, 31–42 (2011).
Hansson, G. K. & Hermansson, A. The immune system in atherosclerosis. Nat Immunol 12, 204–212 (2011).
Hansson, G. K. & Libby, P. The immune response in atherosclerosis: a double-edged sword. Nat Rev Immunol 6, 508–519 (2006).
Rottem, M. & Mekori, Y. A. Mast cells and autoimmunity. Autoimmun Rev 4, 21–27 (2005).
Mak, S. et al. Differential expression of genes in the calcium-signaling pathway underlies lesion development in the LDb mouse model of atherosclerosis. Atherosclerosis 213, 40–51 (2010).
Mallat, Z. et al. Inhibition of transforming growth factor-beta signaling accelerates atherosclerosis and induces an unstable plaque phenotype in mice. Circ Res 89, 930–934 (2001).
Sinningen, K. et al. Monocytic expression of osteoclast-associated receptor (OSCAR) is induced in atherosclerotic mice and regulated by oxidized low-density lipoprotein in vitro. Biochem Biophys Res Commun 437, 314–318 (2013).
Koo, T. Y. et al. Mycophenolic acid regulates spleen tyrosine kinase to repress tumour necrosis factor-alpha-induced monocyte chemotatic protein-1 production in cultured human aortic endothelial cells. Cell Biol Int 37, 19–28 (2013).
Choi, S. H. et al. polyoxygenated cholesterol ester hydroperoxide activates TLR4 and SYK dependent signaling in macrophages. PLoS One 8, e83145 (2013).
Hilgendorf, I. et al. The oral spleen tyrosine kinase inhibitor fostamatinib attenuates inflammation and atherogenesis in low-density lipoprotein receptor-deficient mice. Arterioscler Thromb Vasc Biol 31, 1991–1999 (2011).
Swat, W. & Fujikawa, K. The Vav family: at the crossroads of signaling pathways. Immunol Res 32, 259–265 (2005).
Chen, K. et al. Vav guanine nucleotide exchange factors link hyperlipidemia and a prothrombotic state. Blood 117, 5744–5750 (2011).
Rahaman, S. O., Swat, W., Febbraio, M. & Silverstein, R. L. Vav family Rho guanine nucleotide exchange factors regulate CD36-mediated macrophage foam cell formation. J Biol Chem 286, 7010–7017 (2011).
Severin, S. et al. Distinct and overlapping functional roles of Src family kinases in mouse platelets. J Thromb Haemost 10, 1631–1645 (2012).
Ming, Z. et al. Lyn and PECAM-1 function as interdependent inhibitors of platelet aggregation. Blood 117, 3903–3906 (2011).
Miki, S. et al. Reduction of atherosclerosis despite hypercholesterolemia in lyn-deficient mice fed a high-fat diet. Genes Cells 6, 37–42 (2001).
Kawakami, T., Xiao, W., Yasudo, H. & Kawakami, Y. Regulation of proliferation, survival, differentiation and activation by the Signaling Platform for SHP-1 phosphatase. Adv Biol Regul 52, 7–15 (2012).
Kamata, T. et al. src homology 2 domain-containing tyrosine phosphatase SHP-1 controls the development of allergic airway inflammation. J Clin Invest 111, 109–119 (2003).
Dubois, M. J. et al. The SHP-1 protein tyrosine phosphatase negatively modulates glucose homeostasis. Nat Med 12, 549–556 (2006).
Huang da, W. et al. DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res 35, W169–175 (2007).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res 41, W77–83 (2013).
Irizarry, R. A. et al. Exploration, normalization and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Maglott, D., Ostell, J., Pruitt, K. D. & Tatusova, T. Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res 39, D52–57 (2011).
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98, 5116–5121 (2001).
Wang, J. et al. GO-function: deriving biologically relevant functions from statistically significant functions. Brief Bioinform 13, 216–227 (2012).
Consortium, G. O. The gene ontology: enhancements for 2011. Nucleic Acids Res 40, D559–D564 (2012).
Li, C. et al. SubpathwayMiner: a software package for flexible identification of pathways. Nucleic Acids Res 37, e131 (2009).
Kotera, M., Hirakawa, M., Tokimatsu, T., Goto, S. & Kanehisa, M. The KEGG databases and tools facilitating omics analysis: latest developments involving human diseases and pharmaceuticals. Methods Mol Biol 802, 19–39 (2012).
Keshava Prasad, T. S. et al. Human Protein Reference Database–2009 update. Nucleic Acids Res 37, D767–772 (2009).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504 (2003).
Team, R. D. C. R: A language and environment for statistical computing. R Foundation for Statistical Computing (2012).
Adamcsek, B., Palla, G., Farkas, I. J., Derenyi, I. & Vicsek, T. CFinder: locating cliques and overlapping modules in biological networks. Bioinformatics 22, 1021–1023 (2006).
Palla, G., Derenyi, I., Farkas, I. & Vicsek, T. Uncovering the overlapping community structure of complex networks in nature and society. Nature 435, 814–818 (2005).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B (Methodological) 57, 289–300 (1995).
Acknowledgements
We sincerely thank Dr. HANÈNE AYARI and Dr. GIAMPIERO BRICCA for our free access to the microarray data of carotid atheroma. This work was supported by Science and Technology Foundation of Guangzhou city (Project Number: 2013J4500040).
Author information
Authors and Affiliations
Contributions
Y.L.Y. and M.D. conceived the research project and supervised study and revised the manuscript. W.Q.N., Z.J.O., Y.F., Y.D. and Y.Y.W. performed bioinformatic analysis and interpretation of microarray data. Y.F., L.L.S. and H.Y.W. collected the specimens from Vascular department. J.B.L. supervised the experiments and distinguished the MIT from ATH tissues. H.Y.W., W.Q.N. and K.W.Z. participated in the validation experiments and performed the statistical analysis. W.Q.N. and D.T drafted the manuscript and supplementary materials. All authors reviewed and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Nai, W., Threapleton, D., Lu, J. et al. Identification of novel genes and pathways in carotid atheroma using integrated bioinformatic methods. Sci Rep 6, 18764 (2016). https://doi.org/10.1038/srep18764
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18764
This article is cited by
-
Identification of immune infiltration-related biomarkers in carotid atherosclerotic plaques
Scientific Reports (2023)
-
Bioinformatics analysis of vascular RNA-seq data revealed hub genes and pathways in a novel Tibetan minipig atherosclerosis model induced by a high fat/cholesterol diet
Lipids in Health and Disease (2020)
-
Analysis of protein-protein interaction network based on transcriptome profiling of ovine granulosa cells identifies candidate genes in cyclic recruitment of ovarian follicles
Journal of Animal Science and Technology (2018)
-
Microarray profiling of miRNA and mRNA expression in Rag2 knockout and wild-type mouse spleens
Scientific Data (2018)
-
A Computational Method of Defining Potential Biomarkers based on Differential Sub-Networks
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.