Abstract
Invertebrates can acquire functional genes via horizontal gene transfer (HGT) from bacteria but fishes are not known to do so. We provide the first reliable evidence of one HGT event from marine bacteria to fishes. The HGT appears to have occurred after emergence of the teleosts. The transferred gene is expressed and regulated developmentally. Its successful integration and expression may change the genetic and metabolic repertoire of fishes. In addition, this gene contains conserved domains and similar tertiary structures in fishes and their putative donor bacteria. Thus, it may function similarly in both groups. Evolutionary analyses indicate that it evolved under purifying selection, further indicating its conserved function. We document the first likely case of HGT of functional gene from prokaryote to fishes. This discovery certifies that HGT can influence vertebrate evolution.
Similar content being viewed by others
Introduction
Horizontal gene transfer (HGT), also termed lateral gene transfer, is the transfer of DNA between organisms outside of traditional reproduction. It often results in adaptive gains of novel genes and traits1,2. Large numbers of HGTs have been identified in prokaryotes and such events constitute an important evolutionary force that modulates the evolution of prokaryote genomes3,4,5,6,7. This is particularly evident when the acquisition confers a selective advantage to the host, such as antibiotic resistance8.
Previously, HGT was thought to play only a minor role in the evolution of animal genomes because heritable changes must enter the germline and separation of germline cells from the soma expectedly leads to low frequencies of HGT. The rapid expansion of newly sequenced animal genomes allows for the identification of additional examples of HGT, which may, in fact, be a far more important source of novel gene evolution than originally thought2,9. Genes of prokaryotic, fungal or botanical origin exist in bdelloid rotifers10,11, tunicates12, jelly fish13, starlet sea anemones14, nematodes15,16 and a variety of insects17,18,19. HGT can contribute to the adaptive potential of the recipient and some lower animals well illustrate this. For example, numerous HGT events have occurred during the evolution of plant parasitic nematodes and such transfers likely allowed the free-living ancestral nematodes to exploit plants as a new ecological niche15,16. HGT of carotenoid biosynthetic genes from fungi has endowed aphids with the ability to produce red body colour, based on which they can vary from between green and red to avoid predators and parasites20. In most cases, acquired genes seem to offer novel functions. Subsequent rounds of gene duplication may expand such functional gains1.
Little well-documented evidence supports cross-kingdom HGTs of encoding sequences to vertebrates. Although the original analysis of the human genome identified a considerable number of protein-coding genes of possible bacterial origin21, further study found them to be phylogenetic artefacts22,23. Evidence for the HGT of glyoxylate cycle enzymes, which may have been acquired by non-placental vertebrates, was also controversial, because the presumptive origin of the gene was uncertain and gene-loss can explain the occurrence24,25. Although HGT has been postulated to occur in primates, flies and nematodes26, it remains unknown in early vertebrates, such as fishes.
Herein, we show that fishes possess one gene transferred from marine bacteria. Gene expression evidence provides strong support of its functionality. These results provide the first credible evidence for functional gene transfer from bacteria to fishes.
Results
One gene in fishes derived from marine bacteria
Using annotated protein sequences of Danio rerio as input queries, a series of stringent filters served to identify promising HGTs (Supplementary Figure 1). After comparison with the proteins of other animals in RefSeq and non-redundant (NR) databases, 2766 fish-specific proteins were retained for subsequent analysis. Sixteen of these protein sequences had high similarity to non-animals and horizontally transferred genes may have encoded these proteins. Subsequent phylogenetic analyses identified one protein with RefSeq ID XP_001335286.1 located on chromosome 16 in D. rerio as being potentially obtained via HGT. Next, we used either BLASTP or TBLASTN27 to compare this protein sequence to proteins, genomes and expressed sequences in the following databases: NR, RefSeq, Ensembl, UCSC, Expressed sequence tags (EST), Transcriptome Shotgun Assembly (TSA) and others. The results further verified its patchy phylogenetic distribution. Outside of fishes the gene had few homologues in animals, yet it was widespread in bacteria and some fungi (Fig. 1). Among non-animals, it had highest similarity with marine bacteria (Supplementary Table 1). Phylogenetic analysis clustered it with bacteria with strong support (BPP >95%; ML and NJ BS >90%) (Fig. 2). It was more likely that the transfer occurred from bacteria to fish than vice versa, assuming parsimony. A physical map of gene XP_001335286.1 in D. rerio documented its presence (Fig. 3a). Another gene with Refseq ID NP_001018555.1 identified initially as presumptive horizontally transferred and with top hits in non-animals, clusters with cyanobacteria well (Supplementary Figure 2). However, its phyletic positioning contrasts with HGT from cyanobacteria to eukaryotes and is more indicative of HGT in the opposite direction, for cyanobacteria embedded within a larger group of eukaryotes and did not cluster with other bacteria species. Herein we mainly study HGT in fishes, so we only focused on XP_001335286.1 in subsequent analyses.
Species-tree for representative eukaryotes and the distributions of XP_001335286.1 homologs among the taxa.
Representative eukaryotic clades were selected from among many to simplify depiction of the distributions of the homologs. Bacteria was used as the outgroup. Genes that were most evenly sampled across major clades were used to construct the phylogeny with ML methods. Red triangles indicate species that possess XP_001335286.1 homologs.
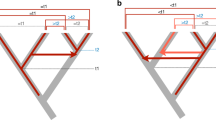
Physical maps of XP_001335286.1 in Danio rerio and its homologs distribution in fishes.
(a) Physical maps of XP_001335286.1 in D. rerio. Arrows denote gene orientation. (b) Phylogeny indicates the general relationship of these fishes based on the NCBI Taxonomy information. Red triangles indicate species that possess XP_001335286.1 homologs.
To exclude the possibility of contamination by bacterial DNA, we confirmed the occurrence of the transferred XP_001335286.1 on multiple fish genomes. We also confirmed that native sequences were physically linked to putative horizontally transferred genes28. The transferred gene contained introns with splice-sites confirmed by RNA-seq data in fishes (Fig. 3a). This also precluded bacterial contamination. Finally, available RNA-seq data supported its expression in transcriptomes (Fig. 3a). Taken together, these analyses provided credible evidence that this horizontally transferred gene was integrated into the genome of fishes and did not represent a bacterial contamination.
Antiquity of the transfer event
Transferred XP_001335286.1 had homologs in most of the sequenced fish genomes. It had greater than 70% similarities with other fishes and these similarities were statistically highly significant with expectation values (E-values) ranging from 3e−144 to 0 (Supplementary Table 1). The levels of significance indicated homology. Their greater similarity among fishes than to bacteria ruled out the possibility of multiple independent HGTs (Supplementary Table 1). Finally, its homologs in fishes located in the same clade adjacent to the potential bacterial donor, further confirming it had a single, ancient transfer (Fig. 2).
Homologs of XP_001335286.1 were not detected in the genomes of basal fishes (Callorhinchus milii and Lepisosteus oculatus). All fishes with this gene belonged to Clupeocephala (Fig. 3b). Elopomorpha (tarpons, bonefishes, eels and relatives), Osteoglossomorpha (bony tongues) and Clupeocephala (all remaining teleosts), constitute the major extant teleosts of early origin29. Because few genomes were sequenced from the two former groups, we used TBLASTN to compare the protein against their available ESTs, TSAs and RNA-seq data, homologs were not found. This failure suggested the transfer likely occurred after emergence of Clupeocephala.
Conserved functions and developmentally regulated expression of HGTs
The gene transferred to fishes was an uncharacterized protein in the RefSeq database. We investigated its possible functions using bioinformatics because no prior analyses or experiments existed. Its bacterial homologs annotated as choline kinase and phosphotransferase. Pfam and SMART analyses revealed it contained the catalytic domain PKc_like super family (cl21453), also named EcKinase and Ecdysteroid kinase (cl17738). This constituted a catalytic domain of the Protein Kinase superfamily (amino acids 27–267) (Fig. 4a). Fishes and bacteria exhibited conserved protein sequence motifs in their domain regions. This conservation suggested that the protein functioned similarly in bacteria and fishes. Although no data document the functional roles that the protein plays, it appeared to be involved in the production of enzymes needed by fishes.
Features and expression of transferred gene in Danio rerio.
(a) Domains of transferred gene coding protein XP_001335286.1. (b) Tertiary structures of transferred gene modelled by SWISS- MODEL server. (c) Expression levels of transferred gene in D. rerio along with developmental stages. Normalized levels of expression were estimated by count of mapped reads to this gene per kilobase of sequence length per million library reads (RPKM). (d) GC content distribution of transferred gene. Red area indicates the density distribution of GC content for D. rerio mRNA. Blue and green lines indicate the GC content of transferred gene in D. rerio and putative donor, respectively.
We explored the cellular locations of the transferred protein using SignalP30, TargetP31, TMHMM32 and WoLFPSORT33. It had no signal peptides or transmembrane helices. WoLFPSORT predicted a cytoplasmic occurrence for it. Structure analyses by the SWISS-MODEL indicated that it had similar structures in fishes and the putative donors (Fig. 4b). Thus, it possibly played similar roles in fishes and bacterial donors.
We used D. rerio as a representative fish to investigate expression profiles of the transferred gene in different developmental stages. Gene XP_001335286.1 was expressed in most stages, except for oocyte and the 1-cell stage. Overall, it was highly expressed in sperm and its level of expression along increased with development up to the 1k-cell stage, indicating probable involvement in development (Fig. 4c).
Due to the absence of experimentation, we could not describe the specific function of this transferred gene. Regardless, its expression in several development stages suggested it still had important functions.
Amelioration and purifying selection of HGTs
Amelioration, the process by which the DNA composition of the acquired gene becomes homogenized to match the composition of the recipient genome34, appeared to have occurred. The GC content of transferred gene was most similar to that of the fishes’ genomes and distinct from the most probable putative donor bacterial genomes (Fig. 4d). All four indices of codon usage bias showed similar patterns of GC3s and GC content (Supplementary Figure 3). Compositional analyses showed other fishes were similar to D. rerio for it.
Likelihood ratio tests can identify strong selection in the evolution of genes35. Whereas positive selection often associates with new functions, purifying selection serves to maintain conserved functions. Global nonsynonymous (Ka)/synonymous (Ks) value of omega for XP_001335286.1 was 0.08845. Values of less than 1 have generally indicated purifying selection on gene sequences. Site-models failed to find signals of positive selection in the homologs as did branch and site models. Overall, our results suggested that purifying selection was the main driving force in the evolution of horizontally transferred XP_001335286.1. This supported the idea that this gene has conserved functions.
Discussion
Our study is the first indication of HGT from bacteria to fishes. The phyletic distribution of the gene indicates one event following emergence of the Clupeocephala. Most new eukaryotic genes and traits originate from gene duplications where mutations subsequently modify one copy to achieve neofunctionalisation1. In comparison, HGT appears to be rare and to play an insignificant role in eukaryotes, especially in complex higher vertebrates1,2. The absence of a well-documented cross-kingdom HGT in vertebrates is not surprising. No mechanism explains the transfer of intact genes from non-animals to vertebrates. Such transfer may be difficult because vertebrate genes are packaged within organelles (principally the nucleus) and, therefore, they are less accessible than genes of bacteria1,36,37. Although HGT may occur in vertebrates26, gene loss, which cannot be ruled out completely, may also explain the observations.
Our analyses involving multiple databases and three tree-building methods to identify possible HGT from prokaryotes to fishes. Many fish genomes contain this horizontally transferred gene. A previous study reported an intriguing case of intra-domain HGT in fishes; a subset of antifreeze proteins (AFPs) appear to have evolved once from a lectin-like protein and then spread by HGT between distantly related species of fishes38,39. Notwithstanding, the marine environment provides fishes an opportunity to obtain genes from environmental bacteria due to their sympatry. The urochordate Ciona intestinalis apparently acquired a cellulose synthase gene from Cyanobacteria40 and similar HGTs may also have occurred in fishes or ancestral vertebrates that have a close evolutionary relationship to C. intestinalis. Many aquatic organisms display HGT events11,12,41,42,43 and a high frequency of HGT occurs in the oceans44. Thus, HGT is not unexpected in fishes.
Of note, another protein within fishes, Refseq ID NP_001018555.1, its phylogenetic tree indicated one possible transfer event from vertebrates to cyanobacteria. Transfer of genes from bacteria to vertebrates is reported uncommonly, while report of functional gene transfer from vertebrates to cyanobacteria is especially rare. NP_001018555.1 may be one of the typical cases of this.
As an alternative explanation to bacteria–fish HGT, gene-loss may explain the pattern. However, this scenario requires multiple wide-scale losses in invertebrates and in vertebrates. Clearly, the origin of this gene does not date to early stages of the origin of metazoans, such as various groups of invertebrates. Further, the gene-loss hypothesis cannot explain the remarkable sequence identity between fishes and their candidate donors. It cannot explain the clustering of transferred genes with bacteria on well-supported nodes in the gene-tree. Therefore, the most plausible explanation for presence of XP_001335286 homologs in fishes is HGT from bacteria.
Though the absence of functional studies for this transfer gene hinders functional annotation, our results indicate that it contains conserved domains and similar structures in recipient fishes and their putative bacterial donors. Its expression patterns suggest involvement in development.
Analyses of the recipient horizontally transferred genes can indicate amelioration and adaptation45. Amelioration of a transferred gene may relate to efficient functioning in the recipient’s genome. For functional efficiency, a transferred gene needs to adjust its nucleotide compositions to match those of the new genome. The extent of amelioration also coincides with the antiquity of a transfer because fine tuning to the genomic background of the recipient requires much time. In fishes, the gene appears to have experienced purifying selection, which indicates its conserved functions. Ka/Ks values relate to protein function such that low values indicate prior purifying selection and possible functional constraints46.
Overall, our results provide the first probable evidence of HGT from bacteria to fishes. The transfer involves the acquisition of functions. This HGT may make significant contribution to the evolution of fishes. Further investigations will lead to a more complete understanding of the biological functions of this gene.
Methods
Data collection and database construction
We retrieved 43138 RefSeq proteins of the zebrafish, Danio rerio and their genomic data from the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/). All protein sequences in RefSeq and non-redundant (nr) database (last accessed August 25, 2014), containing proteins of a wide diversity of eukaryote and prokaryote taxa, were also downloaded from NCBI. The corresponding taxonomy was retrieved from http://www.ncbi.nlm.nih.gov/taxonomy. In addition to D. rerio, sequence data were also downloaded for the following fishes: Oreochromis niloticus, Poecilia reticulate, Neolamprologus brichardi, Pundamilia nyererei, Maylandia zebra, Poecilia formosa, Haplochromis burtoni, Stegastes partitus, Xiphophorus maculates, Oncorhynchus mykiss, Cynoglossus semilaevis, Takifugu rubripes, Tetraodon nigroviridis, Astyanax mexicanus, Lepisosteus oculatus, Oryzias latipes, Callorhinchus milii, Larimichthys crocea, Cyprinus carpio, Lethenteron camtschaticum. Data for Gasterosteus aculeatus, Gadus morhua, Petromyzon marinus, Xenopus tropicalis and Latimeria chalumnae were download from Ensembl (http://www.ensembl.org/) and data for Xenopus laevis were download form Xenopus database (http://www.xenbase.org/). RNA-Seq data of the African lungfish (Protopterus annectens) were retrieved from the Sequence Read Archive of NCBI under accessions SRR505721–SRR505726. Transcriptome assemblies and putative ORF predictions were performed with Trinity47. We used TBLASTN to compare candidate HGTs to Reference of Genome sequences, Expressed Sequence Tag (EST) and Transcriptome Shotgun Assembly (TSA) databases in NCBI to obtain the possible homologous genes27.
We constructed two local proteome databases for the integrated proteins. One database contained 1233189 proteins of animals and the other one was comprised of 10605769 proteins excluding animals.
Screening for candidate HGTs
We used a comprehensive bioinformatic pipeline involving multiple steps to identify HGT candidates in fishes (Supplementary Figure 1). The genome of D. rerio was used to represent fishes due to its high level of integrity and accuracy, which was much better than those of other fishes. Therefore, we mainly focused on proteins of D. rerio to detect candidate HGT events and then searched for such genes in other fishes.
BLAST searching was employed for the initial discovery analysis of putative HGT events27. Firstly, each protein sequence in D. rerio was compared with other animal sequences in the local database using BLASTP. The results were filtered based on an E-value threshold of 1e−10 and a continuous overlap threshold of 33% with the query protein. Because we aimed to detect HGT events in fishes, proteins with homologs in fishes only were selected as possible candidate HGTs45. Next, these candidates were compared to the non-animal database via BLASTP with the same threshold as above. Homologs present in non-animal species were subjected to further phylogenetic analyses to validate the reliability of HGT origins (see below). Putative horizontally transferred genes in D. rerio were compared with other fish proteins in the local database using BLASTP and with their genomes using TBLASTN27.
To eliminate potential DNA contamination, we retained candidate horizontally transferred genes present in many fishes because different kinds of fish constituted independent samples and, therefore, contamination was unlikely to involve identical genes. Further, phylogenies were generated to detect the origins and transmissions of genes adjacent to each candidate gene in the genome48. Finally, considering that bacteria lack introns, bacterial contamination was ruled when candidate horizontally transferred genes had introns.
Validation of horizontal transfer
Phylogenetic analyses were employed to validate the histories of putative horizontally transferred genes. The clustering of a fish gene with a candidate donor group was assumed to support HGT from a particular donor to fish. Homologs of fish horizontally transferred genes were searched in non-redundant databases by BLASTP (E-value = 1e−10). Next, the homologous protein sets were submitted to multiple alignment using ClustalW249. Unrooted tree-building used three approaches: Bayesian inference (BI), maximum likelihood (ML) and distance-based neighbour-joining (NJ). Prottest v3.050 was used to estimate the most optimal models of amino acid substitution. WAG + I + G was the most optimal model for XP_001335286.1. BI analyses were conducted using MrBayes v3.1.251. For each HGT, searches were run using four MCMC chains for one million generations with sampling every 100th tree. Searching was stopped when the average standard deviation of split sequences approached zero. The initial 2500 trees were discarded as burn-in and a 50% majority-rule consensus tree was then constructed to obtain Bayesian posterior probabilities (BPP) for the nodes and those with BPP values greater than 95% were considered to be strongly supported. ML phylogenies were constructed by Phyml v3.152. The branch support values were generated by performing 1,000 bootstrap (BS) pseudoreplicates. NJ trees were obtained using MEGA6.053. The evolutionary distances for NJ construction were computed using the Poisson correction method. Nodal support was obtained by generating 1,000 BS pseudoreplicates. BS values greater than 90% were assumed to indicate strong nodal support. Resulting trees were visualized and edited using FigTree v1.3.1. To the phylogenetic analyses of eukaryotes, 150 genes that were most evenly sampled across major eukaryote clades used in previous study were download54 and ML phylogenies were constructed by Phyml v3.152.
Putative functional assignments
We performed COG, GO and KEGG analyses for each HGT gene to infer its function. A putative HGT protein-coding gene was subjected to SignalP30, TargetP31 and TMHMM32 analyses to identify and predict its possible secretion signals and/or transmembrane domains. Secretory proteins were required to have a signal peptide (SignalP) predicted as secreted (TargetP) and no transmembrane helices (TMHMM). We used WoLFPSORT33 to infer cellular locations. Protein domain predictions were made using Pfam55 and SMART (Simple Modular Architecture Research Tool; http://smart.embl-heidelberg.de/) analyses56. Conserved motif analysis was performed by MEME and Weblogo using default settings57,58. Motifs with p-values less than 1e−10 were considered to be significant. The tertiary structures of HGT-coding proteins were determined using the SWISS-MODEL web server (http://swissmodel.expasy.org/)59.
Gene expression
RNA-Seq data for sperm, oocyte, 1-cell, 4-cell, 16-cell, 64-cell, 128-cell, 256-cell, 1,000-cell and sphere of D. rerio were downloaded from the Gene Expression Omnibus (GEO) repository of NCBI under accession GSE44075. Raw RNA-Seq reads for each sample were stripped of their adaptor sequences with Cutadapt (http://code.google.com/p/cutadapt/) and we removed low quality bases by using Trimmomatic60. Reads that were less than 20 nt in length or that contained an ambiguous nucleotide were discarded. The remaining reads were aligned to D. rerio RefSeq genome Zv9 using TopHat2 while allowing for up to two mismatches61,62. Uniquely mapped reads with mapping qualities larger than or equal to 20 were retained for subsequent analyses. The number of reads mapped to each gene in each RNA-Seq sample was counted using the HTSeq python package (http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html) with the ‘union’ overlap resolution mode and -stranded = no. For each sample, RPKM was computed as the number of reads that mapped per kilobase of exon model per million mapped reads for each gene63.
GC content and codon usage
In general, GC content and patterns of codon usage in horizontally transferred genes have been used to assess the extent of amelioration. Transferred genes were deemed to have already gone through amelioration if their signature bore greater similarity to the genome of the host than the putative donor.
GC content and codon usage bias indices, including CAI (codon adaptation index), CBI (codon bias index), FOP (frequency of optimal codon) and ENC (effective number of codons), were determined for the protein-coding sequences of putative horizontally transferred genes as well as all genes of fishes and the putative donor using CodonW (http://codonw.sourceforge.net/). Non-animal species with the highest sequence identity and occurrence on the same branch in horizontally transferred gene tree was deemed as the putative donor. All coding sequences of donor were downloaded from NCBI. We compared the indices of horizontally transferred gene to the average indices of protein-coding genes of fishes and donor.
Molecular evolution
Selection analyses identified pressures operating on horizontally transferred genes. Selection was assessed using the ML method of Codeml in PAML v4.735. PAL2NAL64 was used to convert protein sequence alignments into the corresponding codon-based DNA alignments. First, an analysis under the one-ratio model (M0) was performed to estimate a global omega (ω) value (dN/dS ratio) across the phylogeny. Global selective pressures were assessed using site models M1a, M2a, M8 and M8a. Evolutionary rates of particular branches of interest (ω1) versus the background ratio (ω0) were computed using the branch model (model = 2). Selective pressures operating on subsets of sites of these branches were calculated using the branch-site models A and A1. The significance of change of ω and evidence of positive selection was assessed using the likelihood ratio test. Positive sites were identified using Bayes Empirical Bayes (BEB)65.
Additional Information
How to cite this article: Sun, B.-F. et al. Horizontal functional gene transfer from bacteria to fishes. Sci. Rep. 5, 18676; doi: 10.1038/srep18676 (2015).
References
Schonknecht, G., Weber, A. P. & Lercher, M. J. Horizontal gene acquisitions by eukaryotes as drivers of adaptive evolution. Bio Essays 36, 9–20 (2014).
Hotopp, J. C. D. Horizontal gene transfer between bacteria and animals. Trends Genet 27, 157–163 (2011).
Boto, L. Horizontal gene transfer in evolution: facts and challenges. Proc Biol Sci 277, 819–827 (2010).
Syvanen, M. Evolutionary implications of horizontal gene transfer. Annu Rev Genet 46, 341–358 (2012).
Pal, C., Papp, B. & Lercher, M. J. Adaptive evolution of bacterial metabolic networks by horizontal gene transfer. Nat Genet 37, 1372–1375 (2005).
Popa, O., Hazkani-Covo, E., Landan, G., Martin, W. & Dagan, T. Directed networks reveal genomic barriers and DNA repair bypasses to lateral gene transfer among prokaryotes. Genome Res 21, 599–609 (2011).
Treangen, T. J. & Rocha, E. P. Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS Genet 7, e1001284 (2011).
Levy, S. B. & Marshall, B. Antibacterial resistance worldwide: causes, challenges and responses. Nat Med 10, S122–129 (2004).
Hotopp, J. C. D. et al. Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317, 1753–1756 (2007).
Boschetti, C. et al. Biochemical diversification through foreign gene expression in bdelloid rotifers. PLoS Genet 8, e1003035 (2012).
Gladyshev, E. A., Meselson, M. & Arkhipova, I. R. Massive horizontal gene transfer in bdelloid rotifers. Science 320, 1210–1213 (2008).
Ni, T. et al. Ancient gene transfer from algae to animals: mechanisms and evolutionary significance. BMC Evol Biol 12, 83 (2012).
Denker, E., Bapteste, E., Le Guyader, H., Manuel, M. & Rabet, N. Horizontal gene transfer and the evolution of cnidarian stinging cells. Curr Biol 18, R858–859 (2008).
Starcevic, A. et al. Enzymes of the shikimic acid pathway encoded in the genome of a basal metazoan, Nematostella vectensis, have microbial origins. Proc Natl Acad Sci USA 105, 2533–2537 (2008).
Danchin, E. G. et al. Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc Natl Acad Sci USA 107, 17651–17656 (2010).
Danchin, E. G. & Rosso, M. N. Lateral gene transfers have polished animal genomes: lessons from nematodes. Front Cell Infect Microbiol 2, 27 (2012).
Acuna, R. et al. Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee. Proc Natl Acad Sci USA 109, 4197–4202 (2012).
Keeling, C. I. et al. Draft genome of the mountain pine beetle, Dendroctonus ponderosae Hopkins, a major forest pest. Genome Biol 14, R27 (2013).
Sun, B. F. et al. Multiple ancient horizontal gene transfers and duplications in lepidopteran species. Insect Mol Biol 22, 72–87 (2013).
Fukatsu, T. Evolution. A fungal past to insect color. Science 328, 574–575 (2010).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Salzberg, S. L., White, O., Peterson, J. & Eisen, J. A. Microbial genes in the human genome: Lateral transfer or gene loss? Science 292, 1903–1906 (2001).
Huerta-Cepas, J., Dopazo, H., Dopazo, J. & Gabaldon, T. The human phylome. Genome Biol 8, R109 (2007).
Kondrashov, F. A., Koonin, E. V., Morgunov, I. G., Finogenova, T. V. & Kondrashova, M. N. Evolution of glyoxylate cycle enzymes in Metazoa: evidence of multiple horizontal transfer events and pseudogene formation. Biol Direct 1, 31 (2006).
Boto, L. Horizontal gene transfer in the acquisition of novel traits by metazoans. Proc Biol Sci 281, 20132450 (2014).
Crisp, A., Boschetti, C., Perry, M., Tunnacliffe, A. & Micklem, G. Expression of multiple horizontally acquired genes is a hallmark of both vertebrate and invertebrate genomes. Genome Biol 16, 50 (2015).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402 (1997).
Richards, T. A. et al. Horizontal gene transfer facilitated the evolution of plant parasitic mechanisms in the oomycetes. Proc Natl Acad Sci USA 108, 15258–15263 (2011).
Chen, J. N., Lopez, J. A., Lavoue, S., Miya, M. & Chen, W. J. Phylogeny of the Elopomorpha (Teleostei): evidence from six nuclear and mitochondrial markers. Mol Phylogenet Evol 70, 152–161 (2014).
Bendtsen, J. D., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340, 783–795 (2004).
Emanuelsson, O., Brunak, S., von Heijne, G. & Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2, 953–971 (2007).
Sonnhammer, E. L., von Heijne, G. & Krogh, A. A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6, 175–182 (1998).
Horton, P. et al. WoLF PSORT: protein localization predictor. Nucleic Acids Res 35, W585–587 (2007).
Lawrence, J. G. & Ochman, H. Amelioration of bacterial genomes: rates of change and exchange. J Mol Evol 44, 383–397 (1997).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24, 1586–1591 (2007).
Taylor, J. S. & Raes, J. Duplication and divergence: the evolution of new genes and old ideas. Annu Rev Genet 38, 615–643 (2004).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nat Rev Genet 11, 97–108 (2010).
Graham, L. A., Lougheed, S. C., Ewart, K. V. & Davies, P. L. Lateral transfer of a lectin-like antifreeze protein gene in fishes. PLoS One 3, e2616 (2008).
Keeling, P. J. Functional and ecological impacts of horizontal gene transfer in eukaryotes. Curr Opin Genet Dev 19, 613–619 (2009).
Nakashima, K., Yamada, L., Satou, Y., Azuma, J. & Satoh, N. The evolutionary origin of animal cellulose synthase. Dev Genes Evol 214, 81–88 (2004).
Yuan, J. B. et al. Horizontally transferred genes in the genome of Pacific white shrimp, Litopenaeus vannamei. BMC Evol Biol 13, 165 (2013).
Jackson, D. J., Macis, L., Reitner, J. & Worheide, G. A horizontal gene transfer supported the evolution of an early metazoan biomineralization strategy. BMC Evol Biol 11, 238 (2011).
Dubuc, T. Q., Traylor-Knowles, N. & Martindale, M. Q. Initiating a regenerative response, cellular and molecular features of wound healing in the cnidarian Nematostella vectensis. BMC Biol 12, 24 (2014).
McDaniel, L. D. et al. High frequency of horizontal gene transfer in the oceans. Science 330, 50 (2010).
Marcet-Houben, M. & Gabaldon, T. Acquisition of prokaryotic genes by fungal genomes. Trends Genet 26, 5–8 (2010).
Wu, D. D. & Zhang, Y. P. Eukaryotic origin of a metabolic pathway in virus by horizontal gene transfer. Genomics 98, 367–369 (2011).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29, 644–652 (2011).
Sun, G. L., Yang, Z. F., Kosch, T., Summers, K. & Huang, J. L. Evidence for acquisition of virulence effectors in pathogenic chytrids. BMC Evol Biol 11, 195 (2011).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22, 4673–4680 (1994).
Abascal, F., Zardoya, R. & Posada, D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59, 307–321 (2010).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30, 2725–2729 (2013).
Katz, L. A. & Grant, J. R. Taxon-rich phylogenomic analyses resolve the eukaryotic tree of life and reveal the power of subsampling by sites. Syst Biol 64, 406–415 (2015).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res 32, D138–141 (2004).
Letunic, I. et al. SMART 5: domains in the context of genomes and networks. Nucleic Acids Res 34, D257–260 (2006).
Grundy, W. N., Bailey, T. L., Elkan, C. P. & Baker, M. E. Meta-MEME: motif-based hidden Markov models of protein families. Comput Appl Biosci 13, 397–406 (1997).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res 14, 1188–1190 (2004).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36 (2013).
Mortazavi, A., Williams, B. A., Mccue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5, 621–628 (2008).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34, W609–W612 (2006).
Yang, Z., Wong, W. S. & Nielsen, R. Bayes empirical bayes inference of amino acid sites under positive selection. Mol Biol Evol 22, 1107–1118 (2005).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (NSFC grant no. 31210103912, 31172072), partially by a grant (No. O529YX5105) from the Key Laboratory of the Zoological Systematics and Evolution of the Chinese Academy of Sciences, by the National Science Fund for Fostering Talents in Basic Research (NSFC-J1210002) and by Program of Ministry of Science and Technology of the Republic of China (2012FY111100).
Author information
Authors and Affiliations
Contributions
D.W.H. conceived and designed the study. B.F.S., T.L., J.X., L.Y.J., L.L. and P.Z. performed the analyses and wrote the manuscript. R.W.M. guided phylogenetic assessments and along with S.H. contributed to manuscript writing. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sun, BF., Li, T., Xiao, JH. et al. Horizontal functional gene transfer from bacteria to fishes. Sci Rep 5, 18676 (2015). https://doi.org/10.1038/srep18676
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18676
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.