Abstract
H3K9 methylation is an important histone modification that is correlated with gene transcription repression. The asymmetric H3K9 dimethylation (H3K9me2) pattern between paternal and maternal genomes is generated soon after fertilization. In the present study, we carefully determined the dynamics of H3K9me2 changes in mouse zygotes and investigated the regulatory mechanisms. The results indicated that histone methyltransferase G9a, but not GLP, was involved in the regulation of asymmetric H3K9me2 and G9a was the methyltransferase that induced the appearance of H3K9me2 in the male pronucleus of the zygote treated with cycloheximide. We found that there were two distinct mechanisms that regulate H3K9me2 in the male pronucleus. Before 8 h of in vitro fertilization (IVF), a mechanism exists that inhibits the association of G9a with the H3K9 sites. After 10 h of IVF the inhibition of G9a activity depends on yet unknown novel protein(s) synthesis. The two mechanisms of transfer take place between 8–10 h of IVF and the novel protein failed to inhibit G9a activity in time, resulting in the appearance of a low level de novo H3K9me2 in the male pronucleus.
Similar content being viewed by others
Introduction
Epigenetic alterations are heritable changes that are not encoded by DNA sequences in a cell and such changes can be delivered stably throughout development and cell proliferation. Epigenetic mechanisms are essential for normal development and maintenance of tissue-specific gene expression patterns in mammals1. Thus, a comprehensive understanding of epigenetic mechanisms, their interactions and alterations, has become a high research priority2. The methylation of histones on different lysine sites results in different biological effects3. Generally, H3K9, H3K27 and H4K20 methylations are correlated with gene transcription repression, whereas H3K4 and H3K36 methylations are correlated with gene transcription activation4,5,6,7,8,9. The H3K9 site can have single, double and triple methylation and different degrees of methylation results in different distributions and functions. H3K9 methylation catalyzed by histone-lysine-methyltransferases involves transcriptional silencing and heterochromatin formation. For all model species, the heterochromatin protein HP1 or HP1 homologue show high affinity with the H3K9 methylation10,11.
In mammalian cells, H3K9 methyltransferase enzymes include SUV39H112, SUV39H213,14, Eu-HMTase/GLP15, G9a16, ESET/SETDB117 and RIZ118. SUV39H1, SUV39H2 and ESET are thought to catalyze H3K9 trimethylation (H3K9me3), while G9a and GLP catalyze H3K9 dimethylation (H3K9me2). G9a plays a catalytic role in H3K9me2 activity at euchromatin regions and causes gene expression inhibition16. Another euchromatin area displays histone methyltransferase GLP that plays a role in silencing of E2F- and Myc-responsive genes in quiescent cells15. G9a and GLP form heteromeric complexes and are linked by the zinc finger protein Wiz19,20,21. G9a and GLP play dominant roles in euchromatic histone H3K9 methylation and are essential for early embryogenesis; G9a- and GLP-deficient embryos display severe growth retardation and early lethality19,22.
There are several kinds of epigenetic alterations in mouse zygotes, including asymmetry of DNA methylation and a series of histone modifications between male and female pronuclei23. Therefore, the zygote is an excellent stage to study the epigenetic mechanisms. In this paper, we investigated the alterations and mechanisms of H3K9me2 in mouse zygotes. Previous studies have shown that the asymmetric H3K9me2 pattern between parental genomes is generated soon after fertilization. H3K9me2 showed a very weak or absent methylation signal in the male pronucleus, whereas a distinct methylation signal was detected in the female pronucleus24. Our study carefully examined H3K9me2 levels in different pronuclear stages of zygotes; we found that a low level de novo H3K9me2 occurred in the male pronucleus at 10 h of in vitro fertilization (IVF), while de novo H3K9me2 of zygotes treated with cycloheximide occurred at the same time, suggesting that there might be two distinct mechanisms that regulate the male pronucleus H3K9me2 in mouse zygotes.
Results
Alteration of H3K9me2 in mouse zygotes
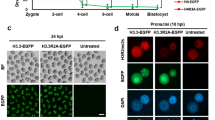
Fertilized eggs were collected at 4 h, 6 h, 8 h, 10 h and 12 h of IVF, respectively. After immunostaining with H3K9me2 antibody, the female pronucleus displayed high H3K9me2 levels in all stages (Fig. 1A). In contrast, the male pronucleus showed no H3K9me2 signal or only low H3K9me2 levels. These results are consistent with previous reports24,25,26,27. However, we noticed that there was a low but distinct H3K9me2 signal in the male pronucleus after 10 h of IVF; in contrast, almost no H3K9me2 signal was observed before 8 h of IVF. When treated with cycloheximide, the male pronucleus showed increased H3K9me2 to a level that was similar to that of the female pronucleus after 10 h of IVF (indicated with white arrow, Fig. 1A).
Alteration of H3K9me2 in mouse zygotes after different treatments.
(A) The H3K9me2 state of zygotes at 4, 6, 8, 10 and 12 h of IVF detected by immunostaining. Control indicates zygotes cultured in normal KSOM medium and CHX indicates zygotes cultured in KSOM medium containing cycloheximide. (B) The H3K9me2 state of zygotes at 12 h of IVF treated with different agents. CHX, Puro, ROS and Aphi indicate zygotes cultured in KSOM medium containing cycloheximide, puromycin, roscovitin and aphidicolin, respectively. Confocal micrographs show the immunostained H3K9me2 (green) and DNA (DAPI, blue) in mouse zygotes. White arrows indicate the increased H3K9me2 of the male pronucleus. Scale bar = 20 μm.
We further investigated the effect of inhibition of protein synthesis, DNA replication and protein kinase on H3K9me2 in mouse zygotes, as shown in (Fig. 1B). Cycloheximide and puromycin are two protein synthesis inhibitors with different mechanisms28. Roscovitine is a purine analog that is a potent and selective inhibitor of cyclin-dependent kinases (CDK)29,30. Aphidicolin is an inhibitor of DNA replication hat specifically inhibits DNA polymerase α31. Fertilized eggs were transferred at 2h of IVF into KSOM medium containing different reagents and collected at 12 h of IVF. Immunostaining revealed that H3K9me2 levels in zygotes treated with cycloheximide and puromycin displayed a clear increase, especially in the male pronucleus (indicated with white arrow, Fig. 1B), suggesting that the asymmetric H3K9 methylation pattern between paternal and maternal genomes disappeared after inhibiting protein synthesis. Meanwhile, the H3K9me2 level of zygotes treated with aphidicolin and roscovitin showed no significant difference compared with the control group.
BIX 01294 inhibited the increase of H3K9me2 in mouse zygotes
The above results showed that the H3K9me2 level of the euchromatin area in the male pronucleus was dramatically increased after protein synthesis inhibition. G9a and GLP were the primary histone methyltransferases with catalytic H3K9me2 activity at euchromatin regions. We first investigated the effect of BIX 01294, a selective inhibitor of G9a and GLP, on H3K9me2 levels in mouse zygotes32,33. The H3K9me2 level of zygotes treated with BIX 01294 showed no significant difference with the control group (Fig. 2A). The H3K9me2 level of the male pronucleus in mouse zygotes treated with both cycloheximide and BIX 01294 did not increase (indicated with red arrow, Fig. 2A). This result indicates that BIX 01294 inhibits the catalytic H3K9me2 activity of histone methyltransferase in mouse zygotes treated with cycloheximide.
The influence of BIX 01294 and G9a or GLP antibody microinjection on H3K9me2.
(A) The H3K9me2 state of zygotes at 12 h of IVF treated with BIX 01294. CHX and BIX indicate zygotes cultured in KSOM medium containing cycloheximide and BIX01294, respectively. BIX+CHX indicates zygotes cultured in KSOM medium containing cycloheximide and BIX01294 together. (B) The H3K9me2 state of zygotes at 12 h of IVF after G9a or GLP antibody microinjection. α -G9a and α –GLP indicates zygotes cultured in normal KSOM medium after antibody microinjection. And α -G9a+CHX and α –GLP+CHX indicates zygotes cultured in KSOM medium containing cycloheximide after antibody microinjection. White arrow indicates the increased H3K9me2 of the male pronucleus and red arrows indicate the low H3K9me2 level of the male pronucleus. Scale bar = 20 μm.
G9a, but not GLP, was involved in the increase of H3K9me2 in mouse zygotes
Next, we investigated the function of G9a and GLP in mouse zygotes treated with cycloheximide, by microinjection of antibody, siRNA and mRNA. G9a or GLP antibody was microinjected into the cytoplasm of fertilized eggs at 2 h of IVF, then cultured for 10 h in KSOM medium containing cycloheximide. Immunostaining showed that asymmetric H3K9me2 disappeared in zygotes microinjected with G9a antibody and treated with cycloheximide (indicated with red arrow, Fig. 2B). But in the GLP antibody microinjection group, the asymmetric H3K9me2 still persisted after treatment with cycloheximide (indicated with white arrow, Fig. 2B). We further confirmed this phenotype by siRNA injection. The interference efficiency of G9a and GLP siRNA were detected by both quantitative real time-PCR and western blot analysis. G9a and GLP siRNA significantly reduced the expression of mRNA and protein, respectively (Fig. 3A,B). Importantly, similar results were obtained with G9a and GLP siRNA microinjection (indicated with red and white arrow, Fig. 3C).
The influence of G9a or GLP siRNA microinjection on H3K9me2.
(A) The mRNA and protein expression of G9a were detected after 24 h of G9a siRNA microinjection (G9a-si). L, the G9a long isoform. S, the G9a short isoform. *P < 0.01. (B) The mRNA and protein expression of GLP were detected after 24 h of GLP siRNA microinjection (GLP-si). Actb (beta-actin) was used as a loading control. *P < 0.01. (C) The H3K9me2 state of zygotes at 12 h of IVF after G9a or GLP siRNA microinjection. NC (negtive control) indicates zygotes microinjected with scrambled control siRNA. NC+CHX, G9a-si+CHX and GLP-si+CHX indicate zygotes cultured in KSOM medium containing cycloheximide after siRNA microinjection. White arrow indicates the increased H3K9me2 of the male pronucleus and red arrow indicates the low H3K9me2 level of the male pronucleus. Scale bar = 20 μm.
We next over-expressed G9a and GLP protein in mouse zygotes by mRNA microinjection. The protein level of G9a and GLP increased significantly at 6 h of microinjection as detected by western blotting (Fig. 4A,B). After G9a over-expression, the asymmetric H3K9me2 disappeared and both pronuclei showed an intensive immunostaining signal (indicated with white arrow, Fig. 4C). However, after GLP over-expression, the asymmetric H3K9me2 in zygotes still persisted (indicated with red arrow, Fig. 4C) although the immunostaining signal of the GLP over-expression group appeared to be more distinct than the control and rabbit Globin over-expression group.
The influence of G9a and GLP over-expression on H3K9me2.
(A) The protein expression of G9a was detected after 6 h of G9a mRNA microinjection (G9a-mi). L, the G9a long isoform. S, the G9a short isoform. (B) The protein expression of GLP was detected after 6 h of GLP mRNA microinjection (GLP-mi). Rabbit Globin mRNA (Globin-mi) was microinjected as control. Actb (beta-actin) was used as a loading control. (C) The H3K9me2 state of zygotes at 12 h of IVF after mRNA microinjection. White arrow indicates the increased H3K9me2 of the male pronucleus and red arrow indicates the low H3K9me2 level of the male pronucleus. Scale bar = 20 μm.
G9a over-expression did not increase H3K9me2 before 8 h of IVF
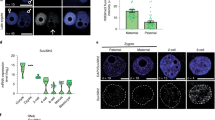
As mentioned above, a different mechanism for asymmetric H3K9me2 was apparent in zygotes before 8 h of IVF, as the H3K9me2 level in the male pronucleus showed no increase after cycloheximide treatment by this time point (Fig. 1A). We then investigated the change of H3K9me2 at the PN 3 stage after G9a mRNA microinjection. After over-expression, G9a protein already increased clearly at 6 h of IVF (Fig. 5A), but the H3K9me2 level of the male pronucleus still showed no increase at 8 h of IVF (Fig. 5B).
G9a and GLP may function independently without WIZ in mouse zygotes.
(A) Immunostaining for the G9a in zygotes at 6 h of IVF after G9a mRNA microinjection. DNA was counterstained with DAPI. (B) The H3K9me2 state of zygotes at 8 h of IVF after G9a mRNA microinjection. Rabbit Globin mRNA (Globin-mi) was microinjected as control. White arrow indicates the increased H3K9me2 of the male pronucleus. Scale bar = 20 μm. (C) The expression of WIZ in GV oocytes, one-cell embryo (1C), two-cell embryo (2C) and testis. Gapdh was used as a loading control. (D). The expression of G9a and GLP in GV oocytes, one-cell embryo (1C), two-cell embryo (2C) and four-cell embryo (4C). Actb (beta-actin) was used as a loading control. L, the G9a long isoform. S, the G9a short isoform.
WIZ showed no expression in mouse zygotes
In mammalian cells, the zinc finger protein WIZ targets G9a and GLP to the chromatin and mediates the G9a/GLP heteromeric complex-dependent H3K9 methylation as well as gene repression34,35. The Wiz/G9a/GLP tri-complex may protect G9a from degradation and Wiz plays a major role in G9a/GLP heterodimer formation20. However, indications are that there may not be any expression of WIZ in zygotes and early embryos in the EST profile of NCBI (UniGene, Mm.274948). Our results also showed that there was almost no expression of WIZ protein in zygotes and early embryos, compared to the expression in testis (Fig. 5C). The expression of G9a and GLP proteins was stable during early embryo development (Fig. 5D).
Discussion
In this study, we investigated the alterations of H3K9me2 in mouse zygotes and the roles of G9a and GLP in these alterations. Our results suggest that there are two distinct mechanisms that regulate H3K9me2 in the male pronucleus of mouse zygotes (Fig. 6). Before 8 h of IVF, H3K9me2 of the male pronucleus remained at a low level and the asymmetric H3K9me2 pattern of paternal and maternal genomes remained after cycloheximide treatment (Fig. 1A), which indicates that this asymmetric H3K9me2 pattern does not depend on new protein synthesis. After over-expression of G9a at 2 h of IVF, the asymmetric H3K9me2 pattern of paternal and maternal genomes also showed no change at 8 h of IVF (Fig. 5B). These results indicate that the asymmetric H3K9me2 was not caused by inhibition of the methyltransferase activity, but it was caused because the methyltransferase was prevented from binding to H3k9 sites by certain unknown protein(s).
After 10 h of IVF, the mechanism of the asymmetric H3K9me2 pattern between paternal and maternal genomes depends on new protein synthesis, as summarized in a previous study24. The two mechanisms of transfer between 8–10 h of IVF and the synthesis of unknown new protein(s) cannot totally inhibit the methyltransferase activity in time. This results in the low level of de novo H3K9me2 appearance in the male pronucleus. In this study, we confirmed the function of new protein synthesis inhibition on H3K9me2 of the male pronucleus by cycloheximide and puromycin treatment (Fig. 1B). Cycloheximide36 and puromycin37 are two different protein synthesis inhibitors and both can cause the disappearance of asymmetric H3K9me2 patterns in mouse zygotes.
Between 8–10 h of IVF the mouse zygotes are in the S phase and DNA replication is in progress. We thus investigated whether DNA replication and cell cycle regulation were involved in the regulation of H3K9me2 in mouse zygotes. Aphidicolin and roscovitin inhibit DNA replication and cyclin-dependent kinase (CDK), respectively. The results of aphidicolin and roscovitin treatment indicate that H3K9me2 of the male pronucleus did not significantly change when either DNA replication or cyclin/cyclin-depended kinase is inhibited (Fig. 1B). This indicates that DNA replication and cell cycle regulation may not be involved in the regulation of H3K9me2 in mouse zygotes.
G9a and GLP form a heteromeric complex to mediate H3K9 methylation as well as gene repression in somatic cells34,35. However, in mouse zygotes G9a and GLP may function independently without WIZ expression (Fig. 5C). Our results indicate that G9a, but not GLP, is involved in the regulation of asymmetric H3K9me2 in mouse zygotes. And G9a is the methyltransferase which induces the appearance of H3K9me2 in the male pronucleus of the zygote treated with cycloheximide. Knock-down by G9a antibody or siRNA microinjection and over-expression of G9a by mRNA microinjection both can influence the H3K9me2 of the male pronucleus (Figs 2B,3 and 4). The results indicate that G9a is involved in the regulation of asymmetric H3K9me2 in mouse zygotes.
Post-translational modification of histone proteins in chromatin and DNA methylation are two major mechanisms involved in epigenetic modifications of genomes, which are regulated by distinct, but coupled, pathways38. G9a/GLP complexes independently mediate H3K9 and DNA methylation to silence transcription21. PGC7 binds histone H3K9me2 to block the activity of the Tet3 methylcytosine oxidase in the maternal genome as well as at certain imprinted loci in the paternal genome39, thereby protecting against conversion of 5 mC to 5 hmC in early embryos40. The parental pronuclei have asymmetric reprogramming capacities and the reprogramming factors reside predominantly in the male pronucleus41. Our results indicate that the alterations of H3K9me2 may play a role in the chromatin remodeling and cell reprogramming.
In the current study, we used mouse zygotes from IVF instead of natural fertilized zygotes for experimental feasibility, as we can determine with greater accuracy the fertilization time by using an IVF system. A previous study showed that there was no difference in the H3K9 methylation pattern in in vivo and IVF mouse zygotes42. Taken together, the study of epigenetic regulation in IVF zygotes may benefit improvement of embryo culture and treatment of female infertility. Understanding the epigenetic regulation mechanisms in mouse zygotes will also benefit research of chromatin remodeling and cell reprogramming and it is important for understanding the mechanism of gene expression regulation and cell differentiation in embryo development.
Materials and Methods
Oocyte collection and culture
All experiments and methods were carried out in accordance with the protocols approved by the Animal Research Committee of the Institute of Zoology, Chinese Academy of Sciences, China. Mice were housed in 12-hour alternating light/dark cycles, with free access to water and food. The GV stage oocytes were isolated from ovaries of 6- to 8-week-old female ICR mice and cultured in M16 medium (Sigma) under paraffin oil at 37 °C, 5% CO2 in air for up to 12 h.
IVF and embryo culture
Spermatozoa were collected from the caudal epididymis of adult ICR males and pre-incubated in human tubal fluid (HTF) medium for 1 h in an atmosphere of 5% CO2, 95% air at 37 °C. Superovulated metaphase II-arrested (M II) oocytes were collected from the ampullae of the oviducts in HTF medium, 14–15 h of hCG administration. The oocytes were inseminated with capacitated spermatozoa. Two hours after insemination, fertilized eggs were washed and cultured in KSOM medium43 and collected at the times indicated. All media were pre-incubated for at least 2 h in an atmosphere of 5% CO2, 95% air at 37 °C. The zygotes were collected at 4, 6, 8, 10 and 12 h of IVF, respectively.
Reagent treatments
Fertilized eggs were transferred into KSOM medium containing cycloheximide (CHX, 15 μg/mL. 01810), puromycin (Puro, 1 mg/mL, P8833), BIX 01294 (BIX, 5 μM, B9311), roscovitin (ROS, 200 μM, R7772) or aphidicolin (Aphi, 3 μg/mL, A0781) and collected at the times indicated. Fertilized eggs cultured in normal KSOM medium were used as control group. All reagents were purchased from Sigma-Aldrich Co. and stored following the product instructions.
Plasmid construction and mRNA synthesis
The full-length G9a (GenBank: NM_145830) and that of GLP (GenBank: NM_001012518.3) CDS were cloned to pcDNA3.1 vector. The plasmids were linearized by NotI and purified by gel extraction kit (Promega). T7 highyield capped RNA tanscription kit and Poly (A) Tailing Kit (Ambion) were used for producing capped and tailed mRNA and then the mRNA was purified with RNeasy cleanup kit (Qiagen). The concentration of G9a and GLP mRNA was determined with a Beckman DU 530 Analyzer and then diluted to a high concentration (1.5 mg/mL) for over-expression.
Microinjection of siRNA, antibodies or mRNA
Microinjection was performed using an Eppendorf microinjector and completed within 1 h. The siRNAs were diluted to 20 μM and microinjected into the cytoplasm of GV stage oocytes. After microinjection, the GV stage oocytes were cultured for 12 h in M16 medium supplemented with 0.2 mM IBMX to maintain oocytes at the GV stage and then transferred into normal M16 medium for 12 h for in vitro maturation. The matured M II oocytes were selected and intracytoplasmic sperm injection (ICSI) was performed; culture followed in KSOM medium containing cycloheximide. The G9a siRNA sequence is 5′-AUACGAAUCACAUCGAUGUGCUUGU-3′, GLP siRNA sequence is 5′-AAAUGCAGCCGCUUGCUCAGCUCCA-3′ and scrambled control siRNA sequence is 5′-UUUCUCCAUACCAUUUCAUCC AUCC-3′ (synthesized by Life Technology). G9a (sc-22877, Santa Cruz) and GLP (sc-68165, Santa Cruz) antibodies or mRNA solution were microinjected into cytoplasm of fertilized eggs at 2 h of IVF. The rabbit Globin mRNA was used as negative control.
Quantitative real time-PCR
Approximately 60 oocytes for each group were used to extract RNA for reverse transcription reactions. Expression level of G9a and GLP was validated by quantitative real-time PCR analysis (Roche 480) according to the manufacturer’s instructions. Primers for G9a (PrimerBank ID 22164772a1), GLP (PrimerBank ID 26352233a1) and beta-actin (actb, PrimerBank ID 6671509a1) were synthesized by Sangon Biotech (Shanghai) Co., Ltd. The experiments were repeated at least 3 times.
Western blot analysis
About 200 mouse oocytes or zygotes or 30 μg testis protein per sample were mixed with SDS sample buffer and boiled for 5 min at 100 °C for SDS-PAGE. Western blotting was performed as described previously44 using the antibody dilution anti-G9a (3306S, Cell Signaling Technology) at 1:1000; anti-GLP (ab41969, Abcam) at 1: 1000; anti-Actb (BS6007, Bioward) at 1: 1000; anti-Gapdh (MB001, Bioward) at 1: 1000. The membranes were subsequently incubated with HRP-conjugated secondary antibodies (1:2000; ZB2301 and ZB2305, Zhongshan Golden Bridge Biotechnology) for 1 h at 37 °C. Protein bands were detected using Thermo Supersignal West Pico chemiluminescent substrate.
Immunofluorescence and confocal microscopy
Zygotes were fixed for 1 h in 3.7% paraformaldehyde in PBS and permeabilized with 0.5% Triton X-100 in PBS for 20 min at room temperature. Then the zygotes were incubated at 4 °C overnight with primary antibodies (H3K9me2 at 1:200, 07–212, Millipore; G9a at 1:200, 3306S, Cell Signaling Technology) and then incubated for 1 h with a secondary AlexaFluor 488-conjugated antibody or AlexaFluor 594-conjugated antibody (1:1000, A11008 and A11012, Life Technology). DNA was stained for 20 min with DAPI (4, 6-diamidino- 2-phenylindole). Fluorescence was detected using a Zeiss LSM780 laser-scanning confocal microscope.
Data analysis
All experiments were repeated at least three times. Statistical analysis was performed using SPSS. Data were expressed as mean ± S.E.M. and P < 0.01 was considered as statistically significant.
Additional Information
How to cite this article: Ma, X. et al. The Dynamics and Regulatory Mechanism of Pronuclear H3k9me2 Asymmetry in Mouse Zygotes. Sci. Rep. 5, 17924; doi: 10.1038/srep17924 (2015).
References
Sharma, S., Kelly, T. K. & Jones, P. A. Epigenetics in cancer. Carcinogenesis 31, 27–36 (2010).
Portela, A. & Esteller, M. Epigenetic modifications and human disease. Nat Biotechnol 28, 1057–1068 (2010).
Kouzarides, T. Histone methylation in transcriptional control. Curr Opin Genet Dev 12, 198–209 (2002).
Volkel, P. & Angrand, P. O. The control of histone lysine methylation in epigenetic regulation. Biochimie 89, 1–20 (2007).
Tamaru, H. et al. Trimethylated lysine 9 of histone H3 is a mark for DNA methylation in Neurospora crassa. Nat Genet 34, 75–79 (2003).
Kirmizis, A. et al. Silencing of human polycomb target genes is associated with methylation of histone H3 Lys 27. Genes Dev 18, 1592–1605 (2004).
Schotta, G. et al. A silencing pathway to induce H3-K9 and H4-K20 trimethylation at constitutive heterochromatin. Genes Dev 18, 1251–1262 (2004).
Krogan, N. J. et al. Methylation of histone H3 by Set2 in Saccharomyces cerevisiae is linked to transcriptional elongation by RNA polymerase II. Mol Cell Biol 23, 4207–4218 (2003).
Nishioka, K. et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev 16, 479–489 (2002).
Nakayama, J., Rice, J. C., Strahl, B. D., Allis, C. D. & Grewal, S. I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001).
Freitag, M., Hickey, P. C., Khlafallah, T. K., Read, N. D. & Selker, E. U. HP1 is essential for DNA methylation in neurospora. Mol Cell 13, 427–434 (2004).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
O’Carroll, D. et al. Isolation and characterization of Suv39h2, a second histone H3 methyltransferase gene that displays testis-specific expression. Mol Cell Biol 20, 9423–9433 (2000).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Ogawa, H., Ishiguro, K., Gaubatz, S., Livingston, D. M. & Nakatani, Y. A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296, 1132–1136 (2002).
Tachibana, M., Sugimoto, K., Fukushima, T. & Shinkai, Y. Set domain-containing protein, G9a, is a novel lysine-preferring mammalian histone methyltransferase with hyperactivity and specific selectivity to lysines 9 and 27 of histone H3. J Biol Chem 276, 25309–25317 (2001).
Schultz, D. C., Ayyanathan, K., Negorev, D., Maul, G. G. & Rauscher, F. J., 3rd . SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16, 919–932 (2002).
Kim, K. C., Geng, L. & Huang, S. Inactivation of a histone methyltransferase by mutations in human cancers. Cancer Res 63, 7619–7623 (2003).
Tachibana, M. et al. Histone methyltransferases G9a and GLP form heteromeric complexes and are both crucial for methylation of euchromatin at H3-K9. Genes Dev 19, 815–826 (2005).
Ueda, J., Tachibana, M., Ikura, T. & Shinkai, Y. Zinc finger protein Wiz links G9a/GLP histone methyltransferases to the co-repressor molecule CtBP. J Biol Chem 281, 20120–20128 (2006).
Tachibana, M., Matsumura, Y., Fukuda, M., Kimura, H. & Shinkai, Y. G9a/GLP complexes independently mediate H3K9 and DNA methylation to silence transcription. EMBO J 27, 2681–2690 (2008).
Tachibana, M. et al. G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev 16, 1779–1791 (2002).
Burton, A. & Torres-Padilla, M. E. Epigenetic reprogramming and development: a unique heterochromatin organization in the preimplantation mouse embryo. Brief Funct Genomics 9, 444–454 (2010).
Liu, H., Kim, J. M. & Aoki, F. Regulation of histone H3 lysine 9 methylation in oocytes and early pre-implantation embryos. Development 131, 2269–2280 (2004).
van der Heijden, G. W. et al. Asymmetry in histone H3 variants and lysine methylation between paternal and maternal chromatin of the early mouse zygote. Mech Dev 122, 1008–1022 (2005).
Lepikhov, K. & Walter, J. Differential dynamics of histone H3 methylation at positions K4 and K9 in the mouse zygote. BMC Dev Biol 4, 12 (2004).
Santos, F., Peters, A. H., Otte, A. P., Reik, W. & Dean, W. Dynamic chromatin modifications characterise the first cell cycle in mouse embryos. Dev Biol 280, 225–236 (2005).
Gilly, M. & Pellegrini, M. Puromycin photoaffinity labels small- and large-subunit proteins at the A site of the Drosophila ribosome. Biochemistry 24, 5781–5786 (1985).
Meijer, L. & Raymond, E. Roscovitine and other purines as kinase inhibitors. From starfish oocytes to clinical trials. Acc Chem Res 36, 417–425 (2003).
Bain, J., McLauchlan, H., Elliott, M. & Cohen, P. The specificities of protein kinase inhibitors: an update. Biochem J 371, 199–204 (2003).
Pedrali-Noy, G. & Spadari, S. Mechanism of inhibition of herpes simplex virus and vaccinia virus DNA polymerases by aphidicolin, a highly specific inhibitor of DNA replication in eucaryotes. J Virol 36, 457–464 (1980).
Chang, Y. et al. Structural basis for G9a-like protein lysine methyltransferase inhibition by BIX-01294. Nat Struct Mol Biol 16, 312–317 (2009).
Kubicek, S. et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell 25, 473–481 (2007).
Shinkai, Y. & Tachibana, M. H3K9 methyltransferase G9a and the related molecule GLP. Genes Dev 25, 781–788 (2011).
Bian, C., Chen, Q. & Yu, X. The zinc finger proteins ZNF644 and WIZ regulate the G9a/GLP complex for gene repression. Elife 4 (2015).
Schneider-Poetsch, T. et al. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nat Chem Biol 6, 209–217 (2010).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc Natl Acad Sci USA 109, 413–418 (2012).
Passaro, D. et al. Epigenetic chromatin modifications in the cortical spreading depression. Brain Res 1329, 1–9 (2010).
Kang, J., Kalantry, S. & Rao, A. PGC7, H3K9me2 and Tet3: regulators of DNA methylation in zygotes. Cell Res 23, 6–9 (2013).
Nakamura, T. et al. PGC7 binds histone H3K9me2 to protect against conversion of 5mC to 5hmC in early embryos. Nature 486, 415–419 (2012).
Liu, W. et al. Asymmetric reprogramming capacity of parental pronuclei in mouse zygotes. Cell Rep 6, 1008–1016 (2014).
Huang, J. C. et al. Comparison of histone modifications in in vivo and in vitro fertilization mouse embryos. Biochem Biophys Res Commun 354, 77–83 (2007).
Lawitts, J. A. & Biggers, J. D. Culture of preimplantation embryos. Methods Enzymol 225, 153–164 (1993).
Zhang, D. et al. Intra-oocyte localization of MAD2 and its relationship with kinetochores, microtubules and chromosomes in rat oocytes during meiosis. Biol Reprod 71, 740–748 (2004).
Acknowledgements
We thank Ying-Chun Ouyang, Yi Hou, Shi-Wen Li and Li-Juan Wang for technical assistance. This study was supported by National Basic Research Program of China (No 2012CB944404, 2011CB944501, 2014CB138502) and the National Natural Science Foundation of China (No. 30930065, 31072028).
Author information
Authors and Affiliations
Contributions
Q.Y.S., H.L.L. and X.S.M. conceived and designed the experiments. X.S.M., S.B.C., X.J.H., F.L., L.Q., X.G.W., T.G.M. and C.C.Z. performed the experiments. H.S. read and modified the manuscript. X.S.M. and Q.Y.S. wrote the paper. All authors analyzed the results and approved the final version of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ma, XS., Chao, SB., Huang, XJ. et al. The Dynamics and Regulatory Mechanism of Pronuclear H3k9me2 Asymmetry in Mouse Zygotes. Sci Rep 5, 17924 (2016). https://doi.org/10.1038/srep17924
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17924
This article is cited by
-
Regulation of paternal 5mC oxidation and H3K9me2 asymmetry by ERK1/2 in mouse zygotes
Cell & Bioscience (2022)
-
PRC2 and EHMT1 regulate H3K27me2 and H3K27me3 establishment across the zygote genome
Nature Communications (2020)
-
Rad9a is involved in chromatin decondensation and post-zygotic embryo development in mice
Cell Death & Differentiation (2019)
-
SIRT1-dependent modulation of methylation and acetylation of histone H3 on lysine 9 (H3K9) in the zygotic pronuclei improves porcine embryo development
Journal of Animal Science and Biotechnology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.