Abstract
The combinatorial cross-regulation of transcription factors (TFs) plays an important role in cellular identity and function; however, the dynamic regulation of TFs during glioma progression remains largely unknown. Here, we used the genome-wide expression of TFs to construct an extensive human TF network comprising interactions among 513 TFs and to analyse the dynamics of the TF-TF network during glioma progression. We found that the TF regulatory networks share a common architecture and that the topological structures are conserved. Strikingly, despite the conservation of the network architecture, TF regulatory networks are highly grade specific and TF circuitry motifs are dynamically rewired during glioma progression. In addition, the most frequently observed structure in the grade-specific TF networks was the feedforward loop (FFL). We described applications that show how investigating the behaviour of FFLs in glioblastoma can reveal FFLs (such as RARG-NR1I2-CDX2) that are associated with prognosis. We constructed comprehensive TF-TF networks and systematically analysed the circuitry, dynamics and topological principles of the networks during glioma progression, which will further enhance our understanding of the functions of TFs in glioma.
Similar content being viewed by others
Introduction
Glioma is the most common and fatal primary brain tumour; approximately 10,000 new cases of high-grade or malignant glioma occur each year1,2,3. Significant new evidence suggests that the dysregulation of gene expression palys critical roles in the development of glioma4. Sequence-specific transcription factor (TF) binding performs key functions in gene expression control. In particular, a TF can interact with and regulate another TF and such cross-regulation among TFs defines the regulatory subnetworks that underlie complex diseases5,6,7,8. However, despite their central roles in cellular identity and function, both the structure of human TF-TF networks and their critical functions in the progression of glioma are largely undefined.
Gene expression-based studies have provided profiles of thousands of individual transcripts, thus revealing novel genes that are dysregulated, concordant with other clinical features of gliomas, such as histological grade and patient survival9,10,11. In particular, TFs play important roles in the transcriptional networks that regulate gene expression and modify and control cancer phenotypes. Differentially expressed TFs in glioma and their downstream gene targets may be potential therapeutic biomarkers of glioma. For example, TP53, SP1, JUN, STAT3 and SPI1 were identified as crucial TFs involved in the development of glioma12. In addition, some TFs were found to play key roles in specific grades of glioma. The TF YY1 has previously been identified as a regulator of the proneural-specific expression pattern in glioma13. This TF plays an essential role in oligodendrocyte progenitor differentiation. Moreover, E2F1 and E2F3 are both master regulators and promote cell-cycle progression and they were amplified in transgenic mouse models of glioma14,15. These candidate TFs and their downstream target genes may play important roles in the progression of glioma and could be potential biomarkers for clinical treatment.
The acquisition of more genome-wide expression datasets from glioma patients, has demonstrated that the processes of tumorigenesis and progression likely involve the coordinated dysregulation of molecular networks16,17 rather than single genes. Therefore, the abnormal expression of single genes may be better viewed collectively at the molecualr network level18. In addition, in contrast to single gene/protein targets for the treatment of glioma, these networks establish broader, multi-target pathways for glioma therapy19. In contrast to the entire transcription regulatory network, the subnetworks are formed by interactions among the TFs that are expressed in a given cell type and these subnetworks make the cell system nimble and robust20. Although many transcriptional regulations are ignored in simple analyses of TF-TF regulation subnetworks, research has demonstrated that the cross-regulation among TFs defines a core subnetwork that plays important roles in cellular identity and complex functions, such as development, differentiation and complex diseases. Moreover, Shane et al. demonstrated that human TF networks are highly cell specific and are driven by cohorts of TFs, including TFs with previously unrecognized roles in the control of cellular identity5. Although regulatory networks provide a representation of molecular interactions, they appear to undergo dynamic reconfiguration in specific contexts21. Through an analysis of the transcriptional network for the mesenchymal transformation of brain tumours, Califano et al. identified the transcriptional module that controls the expression of the mesenchymal signature22. In addition, the transcriptional regulatory network of proneural glioma determines which genetic alterations are selected during tumour progression4. These results indicate that constructing and anlysing of the glioma-progression-associated TF-TF regulatory network might be necessary to understand the mechanism underlying glioma progression. Unfortunately, studying the factors that influence the malignant progression of glioma is limited by the poor reporting of relevant data regarding the expression of genes that are simultaneously profiled across samples with different grades.
To clarify this issue, we reanalysed the extensive dataset obtained through the mRNA profiling of a 160-patient cohort with different grades of glioma in China that was generated for one of our recent studies5. This data and the TF regulation information were integrated and we constructed TF-TF regulatory networks for each grade. By analysing the grade-specific TF regulatory networks, we found that all three of these TF-TF networks share conserved network structures and a common architecture. However, the constructed TF-TF networks are highly grade selective and the network motifs dynamically rewire during glioma progression. We then found an association between some network motifs, especially feedforward loops (FFLs) and patient survival. These systematic analyses provide novel insight into regulatory mechanisms at the TF level during glioma progression and both the method and predictions that were generated in the current study could serve as important resources for future experimental dissection of TF functions during glioma progression.
Results
Comprehensive mapping of TF networks during glioma progression
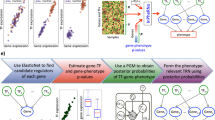
To evaluate the dynamic rewiring of TF circuits during the progression of glioma, we integrated genome-wide gene expression profiles with sequence binding information (Fig. 1A). We used the annotated database (TRANSFAC) of TF-binding motifs to infer TF-TF regulations through Match23. Consequently, we obtained 88,971 regulations among 522 TFs. However, evidence of physical binding does not directly imply transcriptional functional activity of the TFs. Exploring the regulatory relationships among TFs that are specifically co-expressed in the same glioma grade could offer useful information to elucidate the role of TF-TF regulation in glioma progression. Through an analysis of the TF-TF coexpression in each grade, we observed distinct distributions of correlation coefficients for the three grades (Fig. S1). Moreover, the distributions were distinct for different datasets, implying the genetic heterogeneity of different patients.
Construction of comprehensive transcriptional regulatory networks during glioma progression.
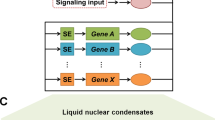
(A) Schematic for the construction of TF regulatory networks during glioma progression. The network nodes represent TFs. Each TF can regulate other TFs and can also be regulated by other TFs; these cross-regulations formed regulatory networks. The sequence-binding information and the genome-wide gene expression were integrated. (B) Grade-specific versus shared regulatory interactions in the TF networks of the three glioma grades. The solid yellow lines represent TF-TF connections that were grade specific and the black lines represent TF connections that occurred in more than two grades. The order of TFs along each circular axis is sorted by their degree in grade II.
Similar to previous studies, we used |Pearson’s coefficient|>0.5 and p < 0.01 to define a link between co-expressed TFs in the regulatory network. To obtain a robust regulatory network of different grades, a regulatory interaction between TFs was only established if correlation criteria were met in two independent datasets. By incorporating physical binding information, we obtained grade-specific TF regulatory networks (Fig. 1B and Fig. S2). Although 88,971 interactions were identified based on sequence binding, only a fraction (0.92%–1.74%) of these potential edges were activated in each grade. Specifically, 1,549 regulations among 342 TFs, 1,143 regulations among 313 TFs and 820 regulations among 299 TFs were obtained for grades II, III and IV, respectively.
Common topological architecture of grade-specific TF-TF regulatory networks
An analysis of the topological features of the TF regulatory networks across each grade revealed some common topological features of the TF-TF network. We found that 48.83%–56.73% of the TFs regulate at least two targets and approximately 61.87%–64.91% of TFs are co-regulated by 2 or more other TFs. Most TFs are connected and form a large connecting subnetwork. These results indicate a complicated combination in terms of both target multiplicity and TF cooperation. We evaluated the out-degree distribution of TF regulators and the in-degree distribution of TF targets and observed a power law and an exponential distribution, respectively. These results indicate that most TFs have few targets and somewhat more regulators. Moreover, the out-degree and in-degree distributions were conserved in the three grades (Fig. 2A). Therefore, similar to many large-scale networks, the grade-specific TF regulatory networks display scale-free characteristics; this result indicates that these networks are not random but are characterized by a core set of structural organizing principles that distinguish them from randomly linked networks.
The TF regulatory networks show conserved structures and motif composition.
(A) The in-degree and out-degree distributions of TFs in three regulatory networks. (B) Network motif enrichment. The relative enrichment or depletion of the 13 possible three-node motifs within the TF-TF regulatory network for each grade is shown. The dashed line represents the distribution of the Z-scores of the sequence-binding network of TFs.
Complex regulatory networks from diverse organisms are mainly formed by a set of simple building blocks termed network motifs24. Network motifs are simple regulatory subnetworks and the topology of a given regulatory network can be reflected by the distribution of the normalized frequencies of network motifs. We therefore analysed the topology of the grade-specific TF regulatory networks. We first extracted the 13 possible three-node motifs and then computed the relative frequency and relative enrichment or depletion of each motif within each grade-specific network. Then, we compared the results of the network for each grade and found that the topologies of the TF networks in the three grades were strikingly similar (Fig. 2B). However, when we recomputed these values solely from the TF binding sites within the promoter regions of each TF without considering the grade-specific coexpression, remarkable similarity of the motif distribution was almost lost (Fig. 2B). This result affirms the criticality of using the grade-specific expression to obtain biologically meaningful conclusions from networks. These results highlight the observation that grade-specific TF-TF networks are highly similar in their structures and organizing principles.
Dynamic rewiring of transcription regulatory networks during glioma progression
Although grade-specific networks share common topological features, we found that most TF-TF interactions are unique to a specific grade (Fig. 3A and Fig. S3). Approximately 26.78% of the edges were common to more than two glioma grades. In addition, we found substantial difference in the degree of TFs between different glioma grades (Fig. 1B). Because network hub nodes have been found to play important roles in many networks, we identified hub TFs in each grade-specific network. Generally, these TF hubs manifest grade-specific behaviours (Fig. 3B). Only ten hubs (such as RUNX1, CEBPB and SOX4) were shared by these three grades. In addition, the TF-TF regulations that are unique to grades typically formed a highly connected subnetwork (Fig. S4), implying that grade-specific regulatory differences are not caused by the actions of a few TFs but rather by TF-TF interactions.
Dynamic network rewiring of TFs during glioma progression.
(A) Network snapshots. Each column corresponds to a TF-TF interaction that occurs in at least one grade network; rows correspond to glioma grade. A purple entry indicates active interactions in the network. (B) The hub TFs in each grade-specific network were defined as those in which the degrees were in the top 10%. This Venn diagram shows the overlap of hub TFs for the three grades. (C) The proportions of TFs in each layer. Blue, master regulators; orange, intermediate regulators and green, low-level regulators. (D) The rewiring of regulators during glioma progression. (E) The hierarchical structure of cancer-associated TF regulatory networks during glioma progression. The TFs are assigned by their layer in each grade.
Because the regulome is directed by networks, it is of particular interest to examine whether the transcription regulatory networks in the progression of glioma harbour a hierarchical structure and if so, how that hierarchy is organized25,26. In addition, TFs in different hierarchical levels are with different roles. For example, Jothi et al. demonstrated that TFs at the top-level in yeast are more likely to be essential and are more conserved27. Next, we used simulated annealing to reveal the organization of TFs in three layers of master-regulators, intermediate regulators and low-level regulators28 (details in methods). The algorithm found a similar proportion of three types of regulators in three grades (Fig. 3C), with approximately 40% of TFs at the top layer of the network in each grade. However, the composition of TFs at each level was dynamically rewired in the progression of glioma (Fig. 3D). In total, 206 TFs were observed in all three grades. Of these TFs in each level, only approximately 30% were retained in the progression of glioma, indicating that the TFs in each level are highly grade specific, even between adjacent grades. The dynamic grade-specific TF usage in each level suggested that the transcription regulatory network changes rapidly between grades. In addition, the TF overlaps in each level between distant grades were lower than those of adjacent grades (Fig. S5), suggesting that the rewiring of TF levels in adjacent grades occurs more frequently than in distant grades.
Next, we focused on the regulatory subnetwork of known cancer TFs and we found dynamic rewiring of the transcription regulatory subnetwork of these TFs during glioma progression. The position of some TFs in the subnetwork changed with glioma progression. For instance, STAT3 has emerged as a key initiator and master regulator of the mesenchymal transformation in malignant gliomas22. We observed that STAT3 was located in the intermediate level in grade II and changed to the top level in grade III (Fig. 3E). This location change may contribute to the progression because the changes corresponds to activation of the STAT3 pathway. Another example is TF-CREB1, which underwent a top-to-intermediate transition during glioma progression. CREB1 plays key roles in glioma29 and CREB1 is essential for glioma cell growth and cell survival in vitro and is critical for gliomagenesis in vivo30. The grade specificity and rapid level transitions of TFs suggest that dynamic rewiring of the regulatory network is an important mechanism for glioma progression.
Dynamic composition of FFLs during glioma progression
Next, to assess whether the conserved network topological architecture (Fig. 2) in each grade was driven by a common core of regulatory interactions, we identified all instances of each three-node network motif within each grade, extracted the constituent TFs and computed how the same TFs were connected for each grade. This comparison revealed that despite the conservation of the overall network architecture among the three grades, specific combinations of TFs comprising individual regulatory motifs underwent substantial remodelling during glioma progression (Fig. 4A,B); this remodeling was greater than that observed among individual edges. Overall, approximately 15% of the combinations of three TFs that were found in either grades II and III or grades III and IV were maintained. For example, FFLs are one of the most important network motifs in biological regulatory networks; biological information is propagated forward from the top TF through the middle TF to the bottom TF, with direct top TF-to-bottom TF reinforcement31. Indeed, only approximately 2.82% of all observed FFLs across the three grades were common (Fig. S6). We next explored the enrichment of three-node circuits in a network that was constructed using edges that were present in two grades (II and III; III and IV). The TF circuits that were conserved between grades were enriched in seven major network motif types, including FFLs, the ‘regulated mutual’ motif and the ‘clique’ motif.
Dynamic network motif rewiring during glioma progression.
(A) Left: frequency with which individual three-node motifs are identically maintained between grades II and III. Middle: percentage of specific three-node motifs that were identically maintained between grades II and III. Right: enrichment of the three-node motif in a network that was constructed using edges that were present in grades II and III. (B) Simialr to A, but for grades III and IV. (C) Frequency with which TFs from FFLs occupy different positions (driver, first passenger and second passenger) for each glioma grade.
In addition, we found that 61 common TFs formed the FFLs in the three studied grades of glioma. However, these TFs also demonstrated grade-specific behaviours. Rather than simply measuring the degree of these TFs, we sought to measure the behaviour of TFs within each glioma grade by determining their topological position within FFLs. For each grade, we computed the relative frequency of FFLs containing a specific TF at each of the three FFL positions (top, middle or bottom). We found that although the positions of some TFs were preseved, some TFs, including some cancer-related TFs, performed different roles during glioma progression (Fig. 4C). For example, ATF3, RUNX1 and CEBPA were likely to locate in the top position of the FFLs in all three grades and RUNX1 preserved high connectivity in all regulatory networks, indicating that this TF may play critical roles in glioma32. Moreover, we observed that some TFs showed dynamic transformation during the glioma progression. For example, HOXB4 was frequently located on the bottom of the FFL in grade II, but tended to locate in the top position in grade III and grade IV. Another example is TF-POU3F2 (Fig. 4C), whose encoded protein is involved in neuronal differentiation and enhances the activation of corticotrophin-releasing-hormone regulated genes. Taken together, these observations highlight the importance of analysing grade-specific regulatory networks to explore the mechanism of glioma progression.
TF FFL biomarkers associated with the survival of glioblastoma (GBM)
We found that the most frequently observed structure in the grade-specific TF networks was the FFL structure. This structure has several important functions in a biological network: detecting persistent signals, filtering noise signals and accelerating response. FFLs play an essential role in the central nervous system. Thus, we next explored whether some FFL motifs were associated with the survival of grade IV-GBM. The grade-IV specific network contains 117 FFLs. We found that the FFL (RARG-NR1I2-CDX2) motif could distinguish the GBM patients with different survival times at a significance level of 0.05 in the training and testing datasets of the TCGA dataset (Fig. 5A,B). The survival-prediction capability of this FFL was validated in the test and CGGA datasets (Fig. 5B,C). In addition, when we used the CGGA dataset to obtain training and test datasets and using the TCGA dataset for independent validation, we also identified this FFL (Fig. S7). RARG plays an important role in the proliferation, metastasis and chemoresistance of cholangiocarcinoma through the simultaneous activation of the Akt/NF-kappaB and Wnt/beta-catenin pathways33. Moreover, the TFs in the FFL could not help to differentiate patients with good prognosis from those with poor prognosis. In the RARG-NR1I2-CDX2 motif, the Cox p-values for these three TFs were 0.84, 0.06 and 0.176 for the TCGA training dataset. We found that the log-rank p-values for this motif in the training and testing of the TCGA dataset were 0.03 and 0.007, respectively. These results suggest that when using a biological network as the searching platform, network-based biomarkers (i.e., FFLs) are expected to be more effective than individual TFs. This result is consistent with previous studies in which network-based biomarkers were more robust and provided valuable insights into the molecular mechanisms of disease34,35.
Network motifs are associated with the survival of grade-IV patients.
(A) The FFL motif (RARG-NR1I2-CDX2) and its functions. (B) Kaplan-Meier survival plots were generated for this motif using the TCGA training dataset. (C) Kaplan-Meier survival plots were generated for this motif using the TCGA test dataset. (D) Kaplan-Meier survival plots were generated for this motif using the CGGA dataset.
Previous studies have shown that patients with IDH mutations have much better survival rates and represent a distinct glioma grade compared with those for wild-type IDH tumours. Next, we determined whether the survival association of this FFL with survival depends on the IDH mutation status; the connection between the prognosis associated FFL and IDH mutation status was assessed by Fisher’s exact test. Here, we found that no distinct difference in the IDH mutation states in the low- and high- risk groups in the training (p = 0.055), test (p = 0.55) and total CGGA datasets (p = 0.49). These results suggest that this FFL may play key roles in glioma progression and may be a potential target for glioma therapy.
Discussion
TF-TF regulations form basic regulatory networks for biological systems and the dysregulation of TF-TF regulatory networks plays critical roles in various types of cancer. However, despite their critical role in human complex diseases, human TF regulatory networks during glioma progression are presently poorly understood. An enduring frustration is the lack of the genome-wide TF expression data for patients with different glioma grades. Here, we reanalyzed the TF expression profiles in 160 Chinese glioma patients generated by one of our recent studies. Integrating this information with the sequence binding information and other two independent gene expression datasets from TCGA and REMBRANDT, we constructed the grade-specific TF regulatory networks for glioma grade II, III and IV. The networks that we constructed here for the three grades provide an extensive resource for analysing the circuitry, dynamics and topological principles of the human TF regulatory network during glioma progression.
Our comparative analysis of transcriptional regulatory networks has revealed several key features of human TF regulation in glioma. First, we found that human transcription regulatory networks of glioma are markedly grade specific, with 47.20%–59.59% of all regulatory interactions only being observed in specific grades. This finding highlights the regulatory diversity during glioma progression and the importance of analysing grade-specific TF-TF regulatory networks. In contrast to the high regulatory diversity between different grades of glioma, we found that the regulatory networks of all grades share a common network topological architecture. In addition, the network motif patterns were conserved during the glioma progression. To explore whether some regulatory interactions might drive the conserved topological architecture during glioma progression, we compared network motifs between adjacent grades. These findings suggest that a distinct set of the network motifs for each grade independently specifies the conserved topological architecture of the TF-TF networks.
Moreover, we found that FFLs were the most observed network motifs in each grade. Through an analysis of the FFLs in GBM, we identified the prognosis-associated FFLs. Some FFLs were independently validated using an independent dataset, providing new potential targets for glioma therapy.
In addition, although grade-IV gliomas are mostly associated with wild-type and grade II- and III are associated with mutated IDH, all of the grades include both cancer types. The glioma grades for cases wild-type are distinct from those of cases with mutated IDH and we further reconstructed both grade-specific and IDH wild-type or IDH mutated networks based on the CGGA dataset. As a result, similar to the TF regulatory network in the main text, the IDH wild-type and IDH mutated grade-specific networks had a relatively low overlap among the three grades (Fig. S8). Grade-II-specific regulations predominanted in IDH mutated samples, whereas grade-IV regulations predominanted in wild-type IDH samples (Table S1). Through further analyses of these regulatory networks, we observed that all six regulatory networks also shared the common network topological architecture of the networks in the main text. In addition, the network motif patterns were conserved during glioma progression. These findings reveal the regulatory diversity during glioma progression and highlight the importance of considering IDH information when analysing grade-specific TF-TF regulatory networks.
Despite the successes of our approach, some additional steps could be used to improve our understanding of the TF regulation during glioma progression. As the number of the datasets of ChIP-Seq36 and DNaseI footprint maps increases, the regulatory networks derived from these datasets may be added to map and analyse grade-specific regulatory networks in cancer. The approach that we utilized in this study does not consider indirect regulatory interactions among TFs that may also affect the expression of a given TF. The identification of such indirect interactions among TFs may provide more details for modelling such interactions. In summary, our current results provided the grade-specific TF networks and a resource to analyse the circuitry, dynamics and topological principles of the human TF regulatory network during glioma progression. Systematic analyses provided novel insight into regulatory mechanisms during glioma progression and both the method and networks that are generated in the current study have the potential to provide novel insight into human TF regulation during glioma progression.
Materials and Methods
Genome-wide gene expression during glioma progression
The genome-wide gene expression profiles of 160 glioma patients were obtained from one of our previous studies18. Briefly, all of these patients were from Beijing Tiantan Hospital. The samples were histologically graded according to the WHO classification of tumours of the nervous systems and included 63 grade-II patients, 33 grade-III patients and 64 GBM patients. Written informed consent was obtained from all donors. Clinical investigations were conducted after approval by the local research ethics committee (Harbin Medical University) and were in accordance with ethical principles. Genome-wide mRNA expression profiles were obtained using an Agilent Whole Human Genome Array. The probes were filtered to remove non-specific probes and probes lacking association with Entrez Gene IDs. In addition, if multiple probes corresponded to a single gene, the expression values of these redundant probes were averaged. The expression values for each gene were background subtracted, quantile-normalized and log-transformed. Additionally, the top 1% low expressed genes were excluded. In total, 515 TFs were expressed in the array. All of the data were deposited in the Chinese Glioma Genome Atlas (CGGA, http://www.cgga.org.cn/), which is a publicly available database that focuses on glioma in China. The sample IDs, IDH mutation status and clinical information of the samples that were used in this study are provided in Table S2.
In addition, two additional sets of gene expression data of different grades of glioma patients were obtained from The Cancer Genome Atlas (TCGA) and The REpository of Molecular BRAin Neoplasia DaTa (REMBRANDT) projects. For the TCGA dataset, whole-genome gene expression profiles were obtained for 78 grade-II, 94 grade-III and 158 grade-IV glioma patients using RNA-seq V2. The level-3 datasets were directly downloaded and then log-transformed. For the REMBRANDT dataset, we obtained the expression as measured by Array Platforms (HG-U133_Plus_2) for 90 grade-II, 68 grade-III and 125 grade-IV patients. Gene expression analysis was performed using packages in Bioconductor37. The Robust Multichip Averaging algorithm (RMA) with quantile normalization was used for the normalization of the Affymetrix microarrays38. Additionally, the top 1% low-expressed genes in these two datasets were also excluded.
Construction of the TF regulatory network
To construct the TF regulatory network for each grade, we first obtained 523 TFs with known motif binding information from TRANSFAC39. We then downloaded the defined promoter region (−2000/+2000bp around the transcription start site, TSS) of these TFs from UCSC Table Browser40. Next, we performed a binding site search using the MatchTM software that is integrated into TRANSFAC Professional. For the purpose of this study, we used a pre-calculated cut-off to minimize false positive matches (minFP) and create a high-quality matrix as described in previous studies41. To restrict the search, we also required a core score of 1.00 and a matrix score of 0.95.
To identify the functional regulations among TFs, we combined both the computational target predictions at the sequence level and the expression relationships between regulators and targets in the context of glioma progression (Fig. 1A). Thus, the Spearman correlation coefficients were calculated for each predicted TF-TF pair in each dataset and the threshold was set to >0.5 or <−0.5 with a corresponding p-value of <0.01. In our study, regulatory interaction between the TFs was established only if the correlation criteria were met in two independent datasets. Finally, after assembling all of the functional TF-TF pairs for each grade, we generated the TF regulatory networks for three grades of glioma. These networks are directed graphs. A directed edge (connection) from a regulatory TF to one of its TF targets exists if their relationship is functionally regulated. To facilitate the analysis of the topological structure of the networks, the weights of all of the edges were set to 1.
Visualization and topological features of the TF regulatory network
TF regulatory network of each grade was visualized using Circos42. Circular plots were formed by two parts and each half was divided into 423 points, with one point representing a TF. Solid lines linking the left and right parts represent TF-TF interactions. The solid yellow lines are TF-TF connections that were specific to the indicated glioma grade. The solid black lines represent TF-TF connections that occurred in more than two grades. The TFs are ordered by their degree in the grade-II network. The in-degree of a TF is defined as the number of regulators that regulated the TF and the out-degree is defined the number of genes that it regulated.
Significance profiles of three-node motifs
The self-interactions of TFs were removed from each grade-specific network and the mfinder software tool was used for network motif analysis24. We focused on the 13 possible network motifs with three TFs (three-node network motifs) in our current study. We calculated a z-score for each network motif and 1000 randomized networks of the same size were used to estimate a null distribution. Then, the z-score was vectorized for every grade and normalized each to unit length to create triad significance profiles as described by Milo et al.24. Similar to the study published by Neph et al.5, if the motif type had a normalized Z-score above 0, it was considered as enriched. Otherwise, if the motif type had a normalized Z-score below 0, it was considered depleted.
Classification of TF regulators based on network topology
All of the TFs in each grade-specifc network were classified into three levels based on their network topology. This process was performed using the method proposed by Snyder et al.28. Briefly, we computed the out-degree (O) and in-degree (I) of each TF and further defined a normalized ‘hierarchy height’ metric, h = (O − I)/(O + I). The ‘master regulators’ are those TFs that regulate many other TFs (h~1), ‘low-level regulators’ are more regulated than regulating (h ~ −1) and ‘intermediate-level regulators’ are both regulated and regulating (h ~ 0). The code that was used for the analysis was downloaded from http://encodenets.gersteinlab.org/enets16.hierarchy_levels.m.
Identification of prognosis associated FFL
To identify the prognosis-associated FFL signature in grade IV-GBM, the specimens in the TCGA dataset were randomly assigned to a training data set or a test dataset. The training and test subsets were required to have approximately the same size (Table S3). According to previous studies43,44, we used the splitting strategy instead of cross-validation because there was no overlap between the training and test sample sets; this lack of overlap was an advantage over cross-validation.
Next, we aimed to identify the FFL signature from the training sets and tested them using the testing datasets. All of the FFLs that were identified in the GBM TF regulatory network were considered candidate signatures. For each FFL, we first used univariate Cox regression analysis to evaluate the association between survival and the expression level of each TF in FFL. Then, a mathematical formula for survival prediction was constructed that considered both the strength and positive or negative association of each TF with survival. The regulatory relationships between the three nodes in FFL motifs were not further considered. Specifically, we assigned a risk score to each patient according to a linear combination of TF expression values as weighted by regression coefficients. The risk score for each patient was calculated as follows:

where  is the Cox regression coefficient of TF i in the training set and
is the Cox regression coefficient of TF i in the training set and  is the expression of TF i in the patient. All of the patients in the training data set were thus dichotomized into high-risk and low-risk groups using the median risk score as the threshold. Then, the coefficient and threshold values that were derived from the training sets were directly applied to the TF expression data of the testing sets to divide the patients in the testing sets into high-risk and low-risk groups. The Kaplan-Meier method was used to estimate the overall survival time for the two groups. The difference in the survival times was compared with the log-rank test. FFLs with p-values less than 0.05 in the training and test datasets were identified as prognostic biomarkers.
is the expression of TF i in the patient. All of the patients in the training data set were thus dichotomized into high-risk and low-risk groups using the median risk score as the threshold. Then, the coefficient and threshold values that were derived from the training sets were directly applied to the TF expression data of the testing sets to divide the patients in the testing sets into high-risk and low-risk groups. The Kaplan-Meier method was used to estimate the overall survival time for the two groups. The difference in the survival times was compared with the log-rank test. FFLs with p-values less than 0.05 in the training and test datasets were identified as prognostic biomarkers.
Validation of the TF FFL prognostic biomarkers in independent datasets
To validate the TF FFL prognostic biomarkers in independent datasets, we first split the TCGA dataset into training and test datasets to identify the FFL biomarkers and then validated these biomarkers in the CGGA dataset. The expression of TFs in the FFL prognostic biomarkers was extracted. We then clustered the gene expression profiles by the k-mean method (k = 2) in the validation dataset. The patients were divided into two groups and the difference in the survival times was compared with the log-rank test. The FFLs with p-values of less than 0.05 were analysed futher.
In addition, the CGGA dataset was also split into training and test datasets to identify the FFL biomarkers, which were then validated using the TCGA datasets. The two subsets of FFL biomarkers were overlapped to obtain the final ones.
Additional Information
How to cite this article: Li, Y. et al. Construction and analysis of dynamic transcription factor regulatory networks in the progression of glioma. Sci. Rep. 5, 15953; doi: 10.1038/srep15953 (2015).
References
Cancer Genome Atlas Research, N. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–8 (2008).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–12 (2008).
Zong, H., Verhaak, R. G. & Canoll, P. The cellular origin for malignant glioma and prospects for clinical advancements. Expert Rev Mol Diagn 12, 383–94 (2012).
Sonabend, A. M. et al. The transcriptional regulatory network of proneural glioma determines the genetic alterations selected during tumor progression. Cancer Res 74, 1440–51 (2014).
Neph, S. et al. Circuitry and dynamics of human transcription factor regulatory networks. Cell 150, 1274–86 (2012).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–9 (2015).
Yosef, N. et al. Dynamic regulatory network controlling TH17 cell differentiation. Nature 496, 461–8 (2013).
Stergachis, A. B. et al. Conservation of trans-acting circuitry during mammalian regulatory evolution. Nature 515, 365–70 (2014).
Jiang, Y. X., Ma, Y. & Cheng, Y. Transcriptome and coexpression network analysis of the human glioma cell line Hs683 exposed to candoxin. J Int Med Res 40, 887–98 (2012).
Weller, M. et al. Molecular predictors of progression-free and overall survival in patients with newly diagnosed glioblastoma: a prospective translational study of the German Glioma Network. J Clin Oncol 27, 5743–50 (2009).
Ivliev, A. E., t Hoen, P. A. & Sergeeva, M. G. Coexpression network analysis identifies transcriptional modules related to proastrocytic differentiation and sprouty signaling in glioma. Cancer Res 70, 10060–70 (2010).
Wei, B. et al. Identification of differentially expressed genes regulated by transcription factors in glioblastomas by bioinformatics analysis. Mol Med Rep 11, 2548–54 (2015).
Baritaki, S. et al. YY1 Over-expression in human brain gliomas and meningiomas correlates with TGF-beta1, IGF-1 and FGF-2 mRNA levels. Cancer Invest 27, 184–92 (2009).
Bhatia, B., Hsieh, M., Kenney, A. M. & Nahle, Z. Mitogenic Sonic hedgehog signaling drives E2F1-dependent lipogenesis in progenitor cells and medulloblastoma. Oncogene 30, 410–22 (2011).
Takahashi, Y., Rayman, J. B. & Dynlacht, B. D. Analysis of promoter binding by the E2F and pRB families in vivo: distinct E2F proteins mediate activation and repression. Genes Dev 14, 804–16 (2000).
Xiao, Y. et al. Identifying dysfunctional miRNA-mRNA regulatory modules by inverse activation, cofunction and high interconnection of target genes: a case study of glioblastoma. Neuro Oncol 15, 818–28 (2013).
Xu, J. et al. MiRNA-miRNA synergistic network: construction via co-regulating functional modules and disease miRNA topological features. Nucleic Acids Res 39, 825–36 (2011).
Li, Y. et al. Comprehensive analysis of the functional microRNA-mRNA regulatory network identifies miRNA signatures associated with glioma malignant progression. Nucleic Acids Res 41, e203 (2013).
Hu, Y. et al. Identifying novel glioma associated pathways based on systems biology level meta-analysis. BMC Syst Biol 7 Suppl 2, S9 (2013).
Bruggeman, F. J., Hornberg, J. J., Boogerd, F. C. & Westerhoff, H. V. Introduction to systems biology. EXS 97, 1–19 (2007).
Califano, A. Rewiring makes the difference. Mol Syst Biol 7, 463 (2011).
Carro, M. S. et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature 463, 318–25 (2010).
Matys, V. et al. TRANSFAC: transcriptional regulation, from patterns to profiles. Nucleic Acids Res 31, 374–8 (2003).
Milo, R. et al. Network motifs: simple building blocks of complex networks. Science 298, 824–7 (2002).
Yu, H. & Gerstein, M. Genomic analysis of the hierarchical structure of regulatory networks. Proc Natl Acad Sci USA 103, 14724–31 (2006).
Cheng, C. et al. Construction and analysis of an integrated regulatory network derived from high-throughput sequencing data. PLoS Comput Biol 7, e1002190 (2011).
Jothi, R. et al. Genomic analysis reveals a tight link between transcription factor dynamics and regulatory network architecture. Mol Syst Biol 5, 294 (2009).
Boyle, A. P. et al. Comparative analysis of regulatory information and circuits across distant species. Nature 512, 453–6 (2014).
Barresi, V. et al. p-CREB expression in human gliomas: potential use in the differential diagnosis between astrocytoma and oligodendroglioma. Hum Pathol 46, 231–8 (2015).
Tan, X. et al. cAMP response element-binding protein promotes gliomagenesis by modulating the expression of oncogenic microRNA-23a. Proc Natl Acad Sci USA 109, 15805–10 (2012).
Murugan, R. Theory on the dynamics of feedforward loops in the transcription factor networks. PLoS One 7, e41027 (2012).
Perry, C. et al. Complex regulation of acetylcholinesterase gene expression in human brain tumors. Oncogene 21, 8428–41 (2002).
Huang, G. L. et al. Oncogenic activity of retinoic acid receptor gamma is exhibited through activation of the Akt/NF-kappaB and Wnt/beta-catenin pathways in cholangiocarcinoma. Mol Cell Biol 33, 3416–25 (2013).
Liu, R., Guo, C. X. & Zhou, H. H. Network-based approach to identify prognostic biomarkers for estrogen receptor-positive breast cancer treatment with tamoxifen. Cancer Biol Ther 16, 317–24 (2015).
Yang, Y. et al. Gene co-expression network analysis reveals common system-level properties of prognostic genes across cancer types. Nat Commun 5, 3231 (2014).
Yang, J. H., Li, J. H., Jiang, S., Zhou, H. & Qu, L. H. ChIPBase: a database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Res 41, D177–87 (2013).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5, R80 (2004).
Irizarry, R. A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31, e15 (2003).
Wingender, E., Dietze, P., Karas, H. & Knuppel, R. TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic Acids Res 24, 238–41 (1996).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res 12, 996–1006 (2002).
Sun, J., Gong, X., Purow, B. & Zhao, Z. Uncovering MicroRNA and transcription factor mediated regulatory networks in glioblastoma. PLoS Comput Biol 8, e1002488 (2012).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res 19, 1639–45 (2009).
Yu, S. L. et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell 13, 48–57 (2008).
Simon, R., Radmacher, M. D., Dobbin, K. & McShane, L. M. Pitfalls in the use of DNA microarray data for diagnostic and prognostic classification. J Natl Cancer Inst 95, 14–8 (2003).
Acknowledgements
This work was supported by the National High Technology Research and Development Program of China [863 Program, Grant No. 2014AA021102], the National Program on Key Basic Research Project [973 Program, Grant No. 2014CB910504], the National Natural Science Foundation of China [Grant Nos 91439117, 61473106, 61203264, 31571331 and 61502126], the China Postdoctoral Science Foundation [Grant No. 2014T70364, 2015M571436 and LBH-Z14134], Natural Science Foundation of Heilongjiang Province [Grant Nos QC2015020], WeihanYu Youth Science Fund Project of Harbin Medical University and Harbin Special Funds of Innovative Talents on Science and Technology Research Project [Grant No. RC2015QN003080].
Author information
Authors and Affiliations
Contributions
X.L. and J.X. participated in the design and coordination of the study. Y.L., T.S. and C.J. acquired and analyzed the data. Y.L., T.S. and C.J. carried out the function analysis. J.B., Z.W. and J.Z. participated in the figures preparation. L.Z., Z.Z. and Y.L. prepared the supplemental files. Y.L., T.S. and J.X. wrote the manuscript. All authors reviewed the manuscript. All authors have read and approved the manuscript and its contents and are aware of responsibilities connected to authorship.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, Y., Shao, T., Jiang, C. et al. Construction and analysis of dynamic transcription factor regulatory networks in the progression of glioma. Sci Rep 5, 15953 (2015). https://doi.org/10.1038/srep15953
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15953
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.