Abstract
To determine how well DNA barcodes from the chloroplast region perform in forest dynamics plots (FDPs) from global CTFS-ForestGEO network, we analyzed DNA barcoding sequences of 1277 plant species from a wide phylogenetic range (3 FDPs in tropics, 5 in subtropics and 5 in temperate zone) and compared the rates of species discrimination (RSD). We quantified RSD by two DNA barcode combinations (rbcL + matK and rbcL + matK + trnH-psbA) using a monophyly-based method (GARLI). We defined two indexes of closely-related taxa (Gm/Gt and S/G ratios) and correlated these ratios with RSD. The combination of rbcL + matK averagely discriminated 88.65%, 83.84% and 72.51% at the local, regional and global scales, respectively. An additional locus trnH-psbA increased RSD by 2.87%, 1.49% and 3.58% correspondingly. RSD varied along a latitudinal gradient and were negatively correlated with ratios of closely-related taxa. Successes of species discrimination generally depend on scales in global FDPs. We suggested that the combination of rbcL + matK + trnH-psbA is currently applicable for DNA barcoding-based phylogenetic studies on forest communities.
Similar content being viewed by others
Introduction
Species discrimination (or identification, delimitation) refers to recognizing biological units based on morphological, ecological or molecular characters, which is one of central topics to biology and particularly important for biodiversity conservation and evolutionary ecology1,2,3,4,5. The tropics and subtropics hold the majority of global biodiversity, containing 30 of the 34 biodiversity hotspots around the world (according to Conservation International up to year 2010)6,7. Theoretically and empirically, incorrect/ambiguous species discrimination could result in inaccurate knowledge about taxonomic classification3, biased estimations on ecological patterns and processes in biological communities8 and ineffective strategy-making for biodiversity conservation5, etc. Previous work has highlighted that most animal taxa can be identified to species level via DNA barcoding technology from a mitochondrial gene cytochrome c oxidase subunit 1 (COI)9,10, which indicates that DNA barcodes can solve the identification problem in biology-related fields. Furthermore, DNA barcoding approach works better but costs less than traditional morphology-based taxonomic practices for species discrimination in vegetation surveys3. Traditional plant taxonomy will encounter with practical difficulties when discriminating samples at early stage (e.g., shoots, seedlings, treelets) and/or only having an un-featured fragment (e.g., broken branches, dried stems, powdered leaves). More problems will arise, especially in the areas with masses of rare, not well-known and potentially threatened plant species11. Therefore, detecting the general performance of plant DNA barcodes in discriminating species and exploring potential factors influencing discrimination rates are of significant importance for researches on biodiversity and ecology.
A two-locus combination of rbcL + matK from the chloroplast region has been approved as a core DNA barcode to identify land plants, which achieves a rate of 72% correct species level identification based on a data set of 550 species representing the major lineages of land plants (including 445 angiosperm, 38 gymnosperm and 67 cryptogam species)12. However, several empirical studies demonstrate that, the trnH-psbA intergenic spacer13,14, ycf1 from the chloroplast region15 and nuclear ribosomal internal transcribed spacer (ITS) could also be complemented as the core DNA barcodes for seed plants16, in consideration of their inherent advantages (containing more evolutionary information) and disadvantages (different to amplify, unavailable to access sequence data for some forest plots at present). To date, most evaluations on DNA barcodes focused on particular taxonomic groups (taxon-level; family or genus) and usually came out a relatively weak discriminatory power5,17,18. In the family Combretaceae (63 species in six genera) in southern Africa, a two-locus core DNA barcode is found to perform poorly but the addition of trnH-psbA to work well, which indicate that evolutionary and biogeographic histories may influence the success of DNA barcoding in discriminating closely-related taxa19. However, for studies in the context of limited geographic forest regions (community-level) where many species from distantly related clades co-occur, it is likely that plant DNA barcoding can achieve a high rate of species discrimination20. Several studies have involved forest communities, but the rates of correct species identification by DNA barcodes are relatively inconsistent. In the two 1-ha plots of an Amazonian tropical forest in French Guiana, a TaxonDNA method achieved a rate of correct plant identification less than 70%, regardless of combinations of eight plant DNA markers tested in 252 juvenile tree species21. Meanwhile, with the aid of a supermatrix phylogeny-building approach, a BLAST method obtained much higher rates of correct species identification, via combinations of a two-locus core DNA barcode and a three-locus DNA barcode (rbcL + matK + trnH-psbA) accordingly; greater than 92%/98% in the 50-ha BCI FDP in Panama14, 89%/93% in the 16-ha Luquillo FDP in Puerto Rico22 and 81%/87% in the 20-ha Dinghushan FDP in China23. Among a data set of 436 plant species (269 genera) in a temperate Canadian Koffler Scientific Reserve, rates of species discrimination were 100% for gymnosperms, bryophytes, lycophytes and monilophytes, compared with 92.7% in angiosperms using monophyly-based method24.
DNA barcoding is a powerful method in detecting new species even in well explored lineages and geographic areas under morphological scrutiny, with the aids of regional and international DNA barcoding libraries25. These remarkable findings inspire us to probe into the significance and effectiveness of plant DNA barcodes when discriminating species in forest communities. As more studies conducted using different methods, results of discrimination should reconcile with one another to move DNA barcoding method forward and investigations on rates of correct species discrimination in diverse forest communities could be applied across multiple ecosystems and forest types. Furthermore, we are eager to detect possibly general patterns of plant DNA barcodes performing on more diverse taxonomic groups and at larger geographic scales worldwide and try to explore potential factors influencing discrimination rates. Our work does not solely focus on issues of taxonomy, but dedicates to provide a new pathway to discriminate plant taxa sampled in forest communities using the universal DNA barcoding method, which will hopefully facilitate species classification and be applied to related research fields, especially in an era of shortage of taxonomists specialized on large-scale floristic studies.
Forest dynamic plots (FDPs) provide a unique opportunity to study large-scale research and multidisciplinary researches on forests including DNA barcoding. DNA barcoding libraries based on FDPs could then provide a non-distorted picture. To the best of our knowledge, the present study comprehensively reported a general performance of the suggested two-locus core DNA barcode of land plants on tree species discrimination and the improvements when adding a complementary DNA barcode (i.e., trnH-psbA spacer region) using a monophyly-based method (i.e., GARLI), utilizing large data sets from 13 CTFS-ForestGEO FDPs ranging from tropics to temperate zones. The present study included three objects. Firstly, we compared rates of species discrimination using two DNA barcode combinations and along a latitudinal gradient. GARLI method was utilized to calculate rates of species discrimination when using DNA barcoding sequence data. Secondly, we have regressed closely-related taxa ratios (S/G and Gm/Gt ratios) against a latitudinal gradient. Two indexes of closely-related taxa were used; specifically, ratio of multiple-species genus/total genera (Gm/Gt ratio: the number of multiple-species genus ( species per genus) to the total number of genera) and species/genus ratio (S/G ratio: the number of species per genus). These ratios were mapped accordingly along a latitudinal gradient. Finally, we correlated ratios of closely related taxa with rates of species discrimination. We predicted that the accuracy of correct species discriminations using plant DNA barcodes was mainly determined by the ratios of closely-related taxa (i.e., S/G ratio and Gm/Gt ratio) in global FDPs.
species per genus) to the total number of genera) and species/genus ratio (S/G ratio: the number of species per genus). These ratios were mapped accordingly along a latitudinal gradient. Finally, we correlated ratios of closely related taxa with rates of species discrimination. We predicted that the accuracy of correct species discriminations using plant DNA barcodes was mainly determined by the ratios of closely-related taxa (i.e., S/G ratio and Gm/Gt ratio) in global FDPs.
Results
Rates of species discrimination
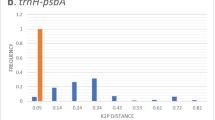
Locally (n = 13), average rates of species discrimination (RSDs) were 88.64 ± 2.12% for rbcL + matK and 91.52 ± 1.44% for rbcL + matK + trnH-psbA; mean ± S.E.) (Fig. 1). For rbcL + matK, RSD was the highest in Wabikon-Lake (100%) and Wytham-Woods (100%) from the temperate zone, but the lowest in Nanjenshan (72.41%) from the subtropics. For rbcL + matK + trnH-psbA, RSD was also the highest in Wabikon-Lake (100%) and Wytham-Woods (100%), but the lowest in BCI (84%) from the tropics. Regionally (n = 3), average RSDs were 83.84 ± 8.11% for rbcL + matK and 83.33 ± 7.35% for rbcL + matK + trnH-psbA. RSD was the highest in the temperate zone (100%) both for rbcL + matK and rbcL + matK + trnH-psbA, but the lowest in the subtropics (74.46% for rbcL + matK and 77.23% for rbcL + matK + trnH-psbA). Globally (among all the species) (n = 1), RSDs were 75.51% for rbcL + matK and 76.09% for rbcL + matK + trnH-psbA.
Improvements of species discrimination by adding trnH-psbA to rbcL + matK combination
An additional DNA barcoding locus (trnH-psbA) had increased rates of species discrimination to various degrees. Specifically, average RSDs increased from 88.65 ± 7.65% to 91.52 ± 5.19% (t = 2.458, P = 0.030, n = 13) locally, from 83.84 ± 8.11% to 85.33 ± 7.35% (t = 1.848, P = 0.206, n = 3) regionally and from 72.51% to 76.09% globally (n = 1). Furthermore, the discrimination of all the plots united without multiple-species genera increased to 84.46% when compared to that with multiple-species genera.
Species discrimination along a latitudinal gradient
Determination coefficients (R2) values were 0.548 (P = 0.019) and 0.715 (P = 0.002) for rbcL + matK and rbcL + matK + trnH-psbA with latitudinal factors under a quadratic model (Fig. 2), while only 0.294 and 0.504 when under a linear model. Correlation coefficients (Pearson’s r) were 0.740 and 0.846 under quadratic models, indicating a relatively high correlation. Rates of species discrimination varied along a latitudinal gradient, which tended to decrease from low to middle latitudes and to increase from middle to high latitudes. Consequently, a quadratic regression model seems to be more appropriate when describing distribution patterns of species discrimination rates along a latitudinal gradient.
A latitudinal gradient of closely-related taxa ratios (S/G and Gm/Gt ratios)
S/G ratios ranged from 1.000 to 1.750 (on average 1.466) among all the 13 plots; specifically from 1.316 to 1.614 (on average 1.466) in tropics, from 1.304 to 1.714 (on average 1.554) in subtropics and from 1.000 to 1.750 (on average 1.378) in temperate zones. S/G ratios in subtropics were higher than in tropics and temperate zones, but no statistical significance could be detected (F = 0.835, P = 0.462). Gm/Gt ratios ranged from 0 to 0.178 (on average 0.095) among all the 13 plots; specifically from 0.076 to 0.100 (on average 0.092) in tropics, from 0.043 to 0.178 (on average 0.123) in subtropics and from 0 to 0.150 (on average 0.069) in temperate zones. Gm/Gt ratios in subtropics were higher than in tropics and temperate zones, but no statistical significance could be detected (F = 1.309, P = 0.313). Determination coefficients (R2) were 0.362 and 0.390 for S/G and Gm/Gt ratios with latitudinal factors under a quadratic regression model, while only 0.142 and 0.134 when under a linear model. Consequently, both S/G ratios and Gm/Gt ratios tended to increase from low to middle latitudes and to decrease from middle to high latitudes. Correlation coefficients (r) were 0.602 and 0.624 for S/G and Gm/Gt ratios with latitudinal factors using a quadratic model, indicating a moderate correlation.
Correlations of closely-related taxa ratios with rates of species discrimination
RSDs tended to be higher in FDPs with lower ratios of closely-related taxa (S/G and Gm/Gt ratios) and to be lower in FDPs with higher ratios. R2 values varied from 0.005 to 0.192 for S/G ratio and from 0.003 to 0.173 for Gm/Gt ratio when using a linear regression model (Fig. 3). Rates of species discrimination were negatively correlated with ratios of closely-related taxa, but no statistical significance was detected (all P > 0.05).
Discussion
A central goal of this work is to assess how well DNA barcodes perform in global FDPs by comparing the success of species correct discrimination. Higher rates of species discrimination in forest communities can improve the resolution of community phylogenies, thus phylogenetically related biodiversity indexes could be more accurate and the mechanisms governing the community assembly across large spatio-temporal scales could be more clearly inferred14,26,27,28,29. Many fields of the natural sciences are declining, especially traditional disciplines like taxonomy30. Taxonomists have become an endangered profession in the era of genomics over the past few decades, but now taxonomy suddenly becomes fashionable again due to revolutionary methods in taxonomy called DNA barcoding. Integrative taxonomy incorporates various types of data and quantitative methods for documenting biodiversity and facilitates the integration of data from different sources31,32. With the development of universal loci, standard working flow, complete taxonomic sampling and rapid species discrimination, DNA barcoding may invigorate traditional disciplines in the context of local, regional and global scales29,33,34. Certainly, genetic data like DNA barcoding sequences should be used with caution, since there is no common distance threshold identified for species delimitation, no single classification technique applied universally for species identification and practical difficulties to discriminate closely-related species adopting genetic distance methods. Hence, we need corroboration from ecology, morphology, geography and molecular information, especially the case in the context of species discovery35,36. The data sets used in the study (13 FDPs from the CTFS-ForestGEO network) at least provide a good opportunity to describe general patterns of tree species discriminations using plant DNA barcodes at the local, regional and global scales and to detect how potential factors (e.g., S/G ratio, Gm/Gt ratio and latitudes) influence rates of species discrimination.
For species discrimination in diverse forest communities, plant DNA barcoding satisfies criterions of simplicity, repeatability, effectiveness and standardization. GARLI method had a relatively high rates of species discrimination (88.64–91.52%) among the 13 FDPs (Fig. 1), suggesting that the monophyly-based method distinguished excellent in terms of species discrimination, though monophyly-based methods using DNA barcodes has been criticized because it assumes that phylogenetic reconstruction is reliable21. Our results showed that rbcL, matK and trnH-psbA could be treated as most proper (though not ideal) genetic regions for plant DNA barcoding studies in forest communities. Besides of monophyly-based method, distance and character-based methods is also proposed to classify species through a unique combination of diagnostic character states from molecular sequences that have been organized hierarchically36,37,38, which may perform well for species identification of populations and genera. None method was found to be the best to discriminate plant species and a feasible option could be “the simplest, the best” in diverse FDPs.
Numerous studies have found that rbcL + matK results in low rates of species discrimination in taxonomic groups (i.e., at the taxon-level), because this combination is limited in its ability to correctly identify species due to low levels of sequence variability in these two chloroplast gene regions. CBOL Plant Working Group has reported a general rate of ~72% successful discrimination rate based on a comprehensive analysis across 550 species representing the major lineages of land plants12. Sometimes, these low rates of species discrimination can be extremely variable; for example, it is only 26% of RSD in Rhododendron with 173 species39, 73% RSD in Taxus covering all species in Eurasia40, depending on the particular taxa examined. However, this two-locus core DNA barcode performs much better in geographically restricted regions (i.e., at the community level), especially in forest dynamics plots. In temperate zone, a national DNA barcode resource covering the native flowering plants and conifers for Wales, provides discrimination rates of 1143 species ranging from 69.4% to 74.9%41. In a Canadian Arctic flora, a distance-based analysis of combined rbcL + matK sequence data discriminates 56% of 490 vascular plant species42, while at the Koffler Scientific Reserve, Canada, the rbcL + matK DNA barcode can result in 93.1% of species resolution in the context of local northern temperate floras using monophyly-based method24, which could be caused by the different calculation methods. In an African rainforest plot in Cameroon, a BLAST analysis discriminates 80% of 272 species with the rbcL + matK combination43. It’s reasonable that rbcL + matK can obtain relatively high rates of species discrimination in diverse FDPs at local scale, which can be largely explained by the presence of a broad taxonomic sampling and relatively few closely related taxa co-occurring in the geographically restricted region20,44.
The strategy combining one locus with slower evolutionary rate plus another one or two loci with faster evolutionary rate, utilizing markers from chloroplast (e.g., rbcLa/rbcLb, matK and trnH-psbA) and/or nuclear genes (e.g., ITS/ITS2), could be used as an effective DNA barcode to discriminate species for land plants (at the taxon or community levels), depending on the phylogenetic information from the study sites and ratios of closely-related taxa. The addition of a non-coding region trnH-psbA to the core DNA barcode increased species resolution by 2.2% (to 95.3% at the Koffler Scientific Reserve, Canada)24, by 4% (to 93% in the 16-ha Luquillo FDP in Puerto Rico)22, by 6% (to 87% in the 20-ha Dinghushan FDP in China and to 98% in the 50-ha BCI FDP in Panama)14,23 and by 8% (to 88% in a 50-ha FDP in Cameroon)43. However, our results found that a complementary DNA barcode (i.e., trnH-psbA) could averagely improve RSDs by 2.9% (to 91.5% using GARLI method) for the 13 FDPs (Fig. 1), especially high in subtropical regions. A meta-analysis of DNA sequences from 3495 species of land plants belonging to 498 genera in 149 families searched from GenBank finds that single trnH-psbA results in a relative low rate of species discrimination (from 35.5% in gymnosperms to 72.2% in mosses) using BLAST method, while trnH-psbA + ITS2 combination performs better or equally well compared with other combinations (e.g., trnH-psbA, matK, rbcL and ITS2 from 586 species belonging to 71 genera and 47 families) in most taxonomic groups studied45. The addition of ITS/ITS2 is more powerful in specific taxonomic groups due to its faster evolutionary rate than trnH-psbA44,46.
Species-genus (S/G) ratio and multiple-species genus to the total genera (Gm/Gt) ratio are sensitive to diversity and may indicate recent speciation in a given area47,48, which is hoped to serve as a potential proxy indicator to describe the phylogenetic relatedness of targeted clades in a biological community. S/G ratios are expected to present a strong latitudinal gradient correlation at the global scale, with high ratios in tropics and low ratios in temperate zones49. Similarly, a significant negative correlation was found between the number of species per genus and the rate of species discrimination per genus in a Canadian Arctic flora42 and a weak relation in a Canadian temperate flora24. Our results partially supported the prediction of latitudinal diversity gradient, but with an exception in subtropical region, which may relate to speciation and range-expansion events. Furthermore, our results found that rates of species discriminations negatively correlated with ratios of closely related taxa (S/G and Gm/Gt ratios) across global CTFS-ForestGEO FDPs (Fig. 3).
Species diversity and congener number decrease with the rising of the latitudinal gradient50,51. Our results showed that a nonlinear trend of species discrimination rates varied along a latitudinal gradient, which tended to decrease from low to middle latitudes and to increase from middle to high latitudes, indicating a turning point in subtropics (Fig. 2). Consequently, a quadratic regression model rather than linear or other types of regression models (with higher coefficients of determination), was probably proper to describe distribution patterns of species discrimination rates along a latitudinal gradient. The turning point in subtropical region was likely attributed to insufficient sampling, though a comprehensively up-to-date dataset of 13 FDPs had been included in the study. Likewise, the turning point could also be explained by particular species composition and geographical characteristics. Subtropical China (including mainland and Taiwan Island) cover diverse vegetation types and endemic plant species52,53,54, which may undergo regional complex climate changes and local environmental factors throughout the last ice-age cycles from individual to ecosystem. The relatively low rates of species discrimination in subtropical FDPs could be caused by cryptic plant diversity and endemism in these refuge areas55. Species assembly in local communities (e.g., species number in a specific genus, or the number of multiple-species genus) is relevant to deterministic (e.g., species traits, interspecies interactions and environmental conditions) and/or stochastic processes (e.g., birth, death, colonization, extinction and speciation)56,57,58.
ITS/ITS2 from nuclear gene region will definitely provide more phylogenetic information for community studies and is also expected to play an important role to discriminate closely-related taxa in taxonomic groups. The combination of trnH-psbA + ITS and rbcL + matK + ITS (ITS2) was suggested as the preferred barcodes for tropical and subtropical tree species identification in natural reserve, respectively44,59. In conclusion, based on the comprehensive consideration among the rates of species discrimination, cost-effectiveness, sequence recovery and alignment and phylogenetic reconstruction, we suggested that a three-locus DNA barcode (rbcL + matK + trnH-psbA) from chloroplast gene region is effective and proper for studies on plant communities, especially in CTFS-ForestGEO FDPs (Table 1).
Methods
Selected forest dynamics plots
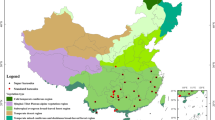
Thirteen FDPs ranging from 2 to 50-ha of CTFS-ForestGEO network (http://www.ctfs.si.edu/) across diverse forests were included in the present study (Supplementary Table S1). These plots span c. 50 degrees of latitude across the globe. All the FPDs were set up according to the standard Smithsonian CTFS protocol60. This work was conducted based on Forestry Standards for “Observation Methodology for Long-term Forest Ecosystem Research” of the People’s Republic of China (LY/T 1952–2011).
DNA barcode libraries
Three standard DNA barcodes from the chloroplast region (i.e., rbcL, matK and trnH-psbA) were used to obtain the sequence data for plant species in each plot61 (Supplementary Table S2). All the 13 FDPs contained 1227 plant species (excluding repeated species), comprising samples with a high-quality sequence for at least one of the three regions. To facilitate comparative analyses, we obtained 17 subsets with sequences for all the three gene regions: 13 individual FDPs with 8 to 226 species at the local scale; tropical, subtropical and temperate regions with 472, 326 and 106 species at the regional scale; and all the FDPs united with 896 species (excluding repeated species) accompanied with DNA barcode sequences from three loci simultaneously at the global scale.
The selected 896 plant species accompanied with DNA sequences from the three loci simultaneously, encompassing 428 genera in 105 families and 37 orders and varying from 8 to 226 species in multiple FDPs. For each forest plot, ten variables were collected, namely latitude, climatic region, hemisphere, plot size, number of genus, number of species, number of multiple-species genus, S/G ratio, Gm/Gt ratio and rates of species discrimination.
Rates of species discrimination
Rates of species discrimination (RSD) for all data sets were obtained using monophyly-based method. The phylogenetic relationship was inferred by maximum likelihood method using GARLI (Genetic Algorithm for Rapid Likelihood Inference) available on CIPRES (http://www.phylo.org/portal2/updateProfile.action) for each barcoding library. The above method allows sequences of three gene regions combined as a matrix, which were not feasible in BLAST method. All DNA sequence data were kept as FASTA format and were aligned as NEXUS format using NCL Converter on CIPRES for subsequent phylogenetic tree reconstruction. The numbers of monophyletic lineages showed how well a phylogenetic tree was reconstructed. More details about GARLI method and analyses can be found in a previous literature61.
Data analyses
To test how much can be improved when adding the third locus trnH-psbA to the core DNA barcode (rbcL + matK) for land plants, two DNA barcode combinations were used for all the analyses at the local, regional and global scales. Oneway ANOVA and Tukey Post Hoc Tests were used to illustrate differences between the discrimination rates. To detect the trend of species discrimination rates along a latitudinal gradient, several regression models were compared. The one with the highest determination coefficient (R2) was selected as the best fit model.
Additional Information
How to cite this article: Pei, N. et al. Closely-related taxa influence woody species discrimination via DNA barcoding: evidence from global forest dynamics plots. Sci. Rep. 5, 15127; doi: 10.1038/srep15127 (2015).
References
Valentini, A., Pompanon, F. & Taberlet, P. DNA barcoding for ecologists. Trends Ecol. Evol. 24, 110–117 (2009).
Dexter, K. G., Pennington, T. D. & Cunningham, C. W. Using DNA to assess errors in tropical tree identifications: How often are ecologists wrong and when does it matter? Ecol. Monogr. 80, 267–286 (2010).
Thomson, K. A. & Newmaste, S. G. Molecular taxonomic tools provide more accurate estimates of species richness at less cost than traditional morphology-based taxonomic practices in a vegetation survey. Biodivers. Conserv. 23, 1411–1424 (2014).
Gomes, A. C. S. et al. Local plant species delimitation in a highly diverse Amazonian forest: do we all see the same species? J. Veg. Sci. 24, 70–79 (2013).
Lahaye, R. et al. DNA barcoding the floras of biodiversity hotspots. Proc. Natl. Acad. Sci. USA 105, 2923–2928 (2008).
Myers, N., Mittermeier, R. A., Mittermeier, C. G., Fonseca, G. A. B. D. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858 (2000).
Pimm, S. L. et al. The biodiversity of species and their rates of extinction, distribution and protection. Science 344, 1246752 (2014).
Swenson, N. G. Phylogenetic resolution and quantifying the phylogenetic diversity and dispersion of communities. PLoS ONE 4, e4390 (2009).
Hebert, P. D. N., Ratnasingham, S. & deWaard, J. R. Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc. Roy. Soc. Lon. Ser. B. 270, S96–S99 (2003).
Ji, Y. et al. Reliable, verifiable and efficient monitoring of biodiversity via metabarcoding. Ecol. Lett. 16, 1245–1257 (2013).
ter Steege, H. et al. Hyperdominance in the Amazonian Tree Flora. Science 342, 1243092 (2013).
CBOL Plant Working Group. A DNA barcode for land plants. Proc. Natl. Acad. Sci. USA 106, 12794–12797 (2009).
Kress, W. J. & Erickson, D. L. A two-locus global DNA barcode for land plants: the coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE 2, e508 (2007).
Kress, W. J. et al. Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama. Proc. Natl. Acad. Sci. USA 106, 18621–18626 (2009).
Dong, W. et al. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 5, 8348 (2015).
China Plant BOL Group et al. Comparative analysis of a large dataset indicates that internal transcribed spacer (ITS) should be incorporated into the core barcode for seed plants. Proc. Natl. Acad. Sci. USA 108, 19641–19646 (2011).
Pang, X. et al. Applying plant DNA barcodes for Rosaceae species identification. Cladistics 26, 1–6 (2010).
Little, D. P. A DNA mini-barcode for land plants. Mol. Ecol. Resour. 14, 437–446 (2014).
Gere, J. et al. Incorporating trnH-psbA to the core DNA barcodes improves significantly species discrimination within southern African Combretaceae. ZooKeys 365, 129–147 (2013).
Chase, M. W. & Fay, M. F. Barcoding of plants and fungi. Science 325, 682–683 (2009).
Gonzalez, M. A. et al. Identification of Amazonian trees with DNA barcodes. PLoS ONE 4, e7483 (2009).
Kress, W. J. et al. Advances in the use of DNA barcodes to build a community phylogeny for tropical trees in a Puerto Rican forest dynamics plot. PLoS ONE 5, e15409 (2010).
Pei, N. Identification of plant species based on DNA barcode technology. Chin. J. Appl. Ecol. 23, 1240–1246 (2012).
Burgess, K. S. et al. Discriminating plant species in a local temperate flora using the rbcL+matK DNA barcode. Methods Ecol. Evol. 2, 333–340 (2011).
Mutanen, M., Kaila, L. & Tabell, J. Wide-ranging barcoding aids discovery of one-third increase of species richness in presumably well-investigated moths. Sci. Rep. 3, 2901 (2013).
Pei, N. et al. Exploring Tree-Habitat Associations in a Chinese Subtropical Forest Plot Using a Molecular Phylogeny Generated from DNA Barcode Loci. PLoS ONE 6, e21273 (2011).
Swenson, N. G. et al. Phylogenetic and Functional Alpha and Beta Diversity in Temperate and Tropical Tree Communities. Ecology 93, S112–S125 (2012).
Joly, S. et al. Ecology in the age of DNA barcoding: the resource, the promise and the challenges ahead. Mol. Ecol. Resour. 14, 221–232 (2014).
Baselga, A. et al. Whole-community DNA barcoding reveals a spatio-temporal continuum of biodiversity at species and genetic levels. Nat. Commun. 4, 1892 (2013).
Tewksbury, J. J. et al. Natural History’s Place in Science and Society. BioScience 64, 300–310 (2014).
Fujita, M. K., Leaché, A. D., Burbrink, F. T., McGuire, J. A. & Moritz, C. Coalescent-based species delimitation in an integrative taxonomy. Trends Ecol. Evol. 27, 480–488 (2012).
Will, K. W., Mishler, B. D. & Wheeler, Q. D. The Perils of DNA Barcoding and the Need for Integrative Taxonomy. Syst. Biol. 54, 844–851 (2005).
Goldstein, P. Z. & DeSalle, R. Integrating DNA barcode data and taxonomic practice: Determination, discovery and description. BioEssays 33, 135–147 (2011).
Papadopoulou, A. et al. Automated DNA-based plant identification for large-scale biodiversity assessment. Mol. Ecol. Resour. 15, 136–152 (2014).
Krishnamurthy, P. K., & Francis, R. A critical review on the utility of DNA barcoding in biodiversity conservation. Biodivers. Conserv. 21, 1901–1919 (2012).
Meier, R., Shiyang, K., Vaidya, G. & Ng, P. K. L. DNA barcoding and taxonomy in Diptera: a tale of high intraspecific variability and low identification success. Syst. Biol. 55, 715–728 (2006).
DeSalle, R., Egan, M. G. & Siddall, M. The unholy trinity: taxonomy, species delimitation and DNA barcoding. Philos. T. R. Soc. B. 360, 1905–1916 (2005).
Saitou, N. & Nei, M. The Neighbor-joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol. Biol. Evol. 4, 406–425 (1987).
Yan, L.-J. et al. DNA barcoding of Rhododendron (Ericaceae), the largest Chinese plant genus in biodiversity hotspots of the Himalaya–Hengduan Mountains. Mol. Ecol. Resour. 15, 932–944 (2015).
Liu, J., Möller, M., Gao, L.-M., Zhang, D.-Q. & Li, D.-Z. DNA barcoding for the discrimination of Eurasian yews (Taxus L., Taxaceae) and the discovery of cryptic species. Mol. Ecol. Resour. 11, 89–100 (2011).
de-Vere, N. et al. DNA Barcoding the Native Flowering Plants and Conifers of Wales. PLoS ONE 7, e37945 (2012).
Saarela, J. M., Sokoloff, P. C., Gillespie, L. J., Consaul, L. L. & Bull, R. D. DNA Barcoding the Canadian Arctic Flora: Core Plastid Barcodes (rbcL + matK) for 490 Vascular Plant Species. PLoS ONE 8, e77982 (2013).
Parmentier, I. et al. How Effective Are DNA Barcodes in the Identification of African Rainforest Trees? PLoS ONE 8, e54921 (2013).
Liu, J. et al. The use of DNA barcoding as a tool for the conservation biogeography of subtropical forests in China. Divers. Distrib. 21, 188–199 (2015).
Pang, X. et al. Utility of the trnH-psbA Intergenic Spacer Region and Its Combinations as Plant DNA Barcodes: A Meta-Analysis. PLoS ONE 7, e48833 (2012).
Gao, T. et al. Evaluating the feasibility of using candidate DNA barcodes in discriminating species of the large Asteraceae family. BMC Evol. Biol. 10, 324 (2010).
Simberloff, D. S. Taxonomic Diversity of Island Biotas. Evolution 24, 23–47 (1970).
Fenner, M., Lee, W. G. & Wilson, J. B. A comparative study of the distribution of genus size in twenty angiosperm floras. Biol. J. Linn. Soc. 62, 225–237 (1997).
Krug, A. Z., Jablonski, D. & Valentine, J. W. Species-genus ratios reflect a global history of diversification and range expansion in marine bivalves. Proc. Roy. Soc. Lon. Ser. B. 275, 1117–1123 (2008).
Janzen, D. H. Herbivores and the number of tree species in tropical forests. Am. Nat. 104, 501–528 (1970).
Leigh, E. G., Jr. et al. Why do some tropical forests have so many species of trees? Biotropica 36, 447–473 (2004).
Hou, H.-Y. Vegetation of China With Reference to Its Geographical Distribution. Ann. Missouri Bot. Gard. 70, 509–549 (1983).
Yu, G. et al. Palaeovegetation of China: a pollen data-based synthesis for the mid-Holocene and last glacial maximum. J. Biogeogr. 27, 635–664 (2000).
Liew, P. M. & Chung, N. J. Vertical Migration of Forests During the Last Glacial Period in Subtropical Taiwan. West. Pac. Earth. Sci. 1, 405–414 (2001).
Qiu, Y.-X., Fu, C.-X. & Comes, H. P. Plant molecular phylogeography in China and adjacent regions: Tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol. Phylogenet. Evol. 59, 225–244 (2011).
Zhou, J. et al. Stochasticity, succession and environmental perturbations in a fluidic ecosystem. Proc. Natl. Acad. Sci. USA 111, E836–E845 (2014).
Hubbell, S. P. The Unified Neutral Theory of Biodiversity and Biogeography. Princeton University Press, (2001).
Chase, J. M. & Myers, J. A. Disentangling the importance of ecological niches from stochastic processes across scales. Philos. T. R. Soc. B. 366, 2351–2363 (2011).
Huang, X.-C., Ci, X.-Q., Conran, J. G. & Li, J. Application of DNA Barcodes in Asian Tropical Trees – A Case Study from Xishuangbanna Nature Reserve, Southwest China. PLoS ONE 10, e0129295 (2015).
Condit, R. Tropical forest census plots. Springer-Verlag and R.G. Landes Company, (1998).
Erickson, D. L. et al. Comparative evolutionary diversity and phylogenetic structure across multiple forest dynamics plots: a mega-phylogeny approach. Front. Genet. 5, 358 (2014).
Acknowledgements
Funding was provided by the Special Fund for Forest Scientific Research in the Public Welfare (20140430105), NSF-China (31200471), the Ministry of Finance of the People’s Republic of China (RITFYWZX201208), China Scholarship Council (201303270006) and CFERN & GENE Award Funds on Ecological Paper. We appreciate J.L., S.G., H.Y., N.Z., Y.F., Y.Z. and X.Y. for useful discussion and thank many assistants of CTFS-ForestGEO FDPs in the field and technicians in the lab. We also thank the anonymous reviewers for their valuable comments on the manuscript.
Author information
Authors and Affiliations
Contributions
W.J.K. and N.P. conceived the idea; W.J.K., D.L.E. and N.P. collected the data; N.P., D.L.E. and J.Z. analyzed the data; N.P., W.J.K. and B.C. wrote the first draft of the manuscript. N.P., D.L.E., B.C., X.G., X.M., N.G.S., J.L.Z., F.A.J., C.L.H., W.Y., Z.H., C.F.H., S.L., N.A.B., J.D.P., J.K.Z., W.J.M., I.C.L., I.F.S., S.J.D., K.M. and W.J.K. contributed to finalizing of the manuscript. S.J.D. is the leader of CTFS-ForestGEO network and K.M. is the leader of CForBio network.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Pei, N., Erickson, D., Chen, B. et al. Closely-related taxa influence woody species discrimination via DNA barcoding: evidence from global forest dynamics plots. Sci Rep 5, 15127 (2015). https://doi.org/10.1038/srep15127
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15127
This article is cited by
-
DNA barcodes in Egyptian olive cultivars (Olea europaea L.) using the rbcL and matK coding sequences
Journal of Crop Science and Biotechnology (2023)
-
DNA barcoding of important fruit tree species of agronomic interest in the genus Garcinia L. from the Western Ghats
Genetic Resources and Crop Evolution (2021)
-
DNA barcoding analysis and phylogenetic relationships of tree species in tropical cloud forests
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.