Abstract
Bacterial adhesins play a pivotal role in the tight bacteria-host cells attachment to initiate the downstream processes and bacterial infection of hosts. In this study, we identified a novel adhesin, VpadF in V. parahaemolyticus. Deletion of VpadF in V. parahaemolyticus markedly impaired its attachment and cytotoxicity to epithelial cells, as well as attenuated the virulence in murine model. Biochemical studies revealed that VpadF recognized both fibronectin and fibrinogen. The binding of VpadF to these two host receptors was mainly dependent on the its fifth bacterial immunoglobulin-like group domain and its C-terminal tail. Our finding suggested that VpadF is a major virulence factor of V. parahaemolyticus and a potential good candidate for V. parahaemolyticus infection control for both vaccine development and drug target.
Similar content being viewed by others
Introduction
Vibrio parahaemolyticus is an inhabitant of estuarine, marine and coastal environments. It is commonly free swimming and attaching to aquatic animals, including corals, fish, oyster, sponges, shrimp and zooplankton1,2. V. parahaemolyticus has also been recognized as the causative agent of seafood-related gastroenteritis, wound infections and septicemia3.
To survive in these diverse and challenging environments, V. parahaemolyticus is presumed to evolve rapidly. Molecular profiling studies on environmental and clinical isolates, as well as comparative genomic analysis of pre-pandemic and pandemic strains, reveals that V. parahaemolyticus genomes are highly versatile and the emergence of pandemic strain could be associated with recombination and novel gene acquisition4,5,6,7. For instance, thermostable direct hemolysin (tdh), tdh related hemolysin (trh) and type III secretion system 2 (T3SS2) are commonly found within the pathogenicity island (Vp-PAI) from clinical isolates, suggesting that V. parahaemolyticus strains acquire these virulence determinants by horizontal gene transfer (HGT)8. TDH and TRH directly lead to cytotoxicity and enterotoxicity while T3SS2 is essential for the enteritis, colonization and competition to protists in aquatic environment9,10,11,12. In addition to TDH, TRH and T3SS2, some V. parahaemolyticus strains also acquire other virulence factors, such as VpaH and ZnuA13,14. These findings reinforce the notion that V. parahaemolyticus is a versatile pathogen with its virulence tightly linked to the acquisition of virulence factors through HGT.
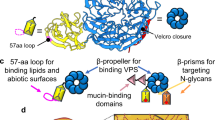
The attachment of pathogens to host cells is the prerequisite for the efficient translocation of effectors that suppress host immune response and/or modulate cellular signaling pathways to aid its infection. Moreover, adhesion subverts host actin cytoskeleton and triggers cellular signaling pathways to recruit downstream signaling proteins to the plasma membrane to facilitate subsequent pathogen invasion. Attachment also ensures pathogens persistence in host niches15,16,17. To date, the multivalent adhesion molecule 7 (MAM7), mannose-sensitive haemagglutinin (MSHA) pilus, enolase, capsular polysaccharide and two type VI secretion systems (T6SSs) have been reported to contribute to cell attachment of V. parahaemolyticus18,19,20,21,22. However, among these adhesive organelles, only mam7 was found to be required for the pathogenesis of V. parahaemolyticus in the worm infection model. MAM7 is a transmembrane protein that consists of a transmembrane motif at the N-terminus and seven mammalian cell entry domains that mediate its adherence to different types of cells by binding to fibronectin (Fn) and phosphatidic acid in the initial stage of infection and subsequent cytotoxicity against 3T3 fibroblasts and RAW264.7 macrophages, but not to HeLa or Caco-2 epithelial cells. Sequence homology search showed that MAM7 is not only present in V. parahaemolyticus, but also a conservative protein possessed by several Gram-negative pathogens, including enteropathogenic Escherichia coli, Vibrio cholerae and Yersinia pseudotuberculosis22. Given that each Gram-negative pathogen usually produces more than one adhesive factor to initiate infection23,24,25 and V. parahaemolyticus frequently acquires virulence factors, it is reasonable to speculate that V. parahaemolyticus has evolved or acquired specific, yet poorly understood mechanism to strengthen host-pathogen interactions. In this study, we identified a novel adhesin gene vp1767, which was referred to as VpadF (V. parahaemolyticus adhesive Factor), contributing to the attachment to and cytotoxicity against epithelial cells. This gene is essential for the lethal effect of V. parahaemolyticus on mice. Most importantly, VpadF bound to cell surface receptors, fibronectin (Fn) and fibrinogen (Fg), which may contribute to its host colonization and pathogenesis.
Methods
Bacterial strains, plasmids and growth conditions
V. parahaemolyticus strains, E. coli strains and plasmids used in this study were listed in ST 1. V. parahaemolyticus strains were cultured in LB medium supplemented with 2.5% sodium chloride (LBS) at 37 °C. Thiosulfate-citrate-bile salts-sucrose agar (TCBS) was used to select V. parahaemolyticus strains.
Construction of gene deletion and complementary strains
The vp1767 gene was deleted from clinical V. parahaemolyticus strain VP3218 by homologous recombination as described previously14. Similar procedures were used to obtain ΔvcrD1 and ΔvcrD1ΔVpadF using specific primers (ST 2). To construct VpadF complementary strain, DNA fragment encoding vp1767 gene with a C-terminal Flag tag and N-terminal ribosome binding site was amplified using primers vp1767com-F and vp1767com-R (ST 2). PCR product was digested and cloned into the same digested pMMB207 to create pMMB207:VpadF. The recombinant plasmid was transformed to E. coli SY327 λpir and then conjugated to V. parahaemolyticus strains with a helper partner E. coli SY327 λpir carrying pPK2013. Transconjugants were selected on TCBS containing 5 μg/ml chloramphenicol to obtain the complementary strains.
Bioinformatics analysis
Protein domain search was performed using PFAM (http://pfam.sanger.ac.uk/search), InterPro (http://www.ebi.ac.uk/interpro) and SMART (http://smart.embl-heidelberg.de). The subcellular location and the transmembrane region were predicated using TMHMM Server v2.0 (http://www.cbs.dtu.dk/services/TMHMM/) and TMpred (http://www.ch.embnet.org/software/TMPRED_form.html). Multiple sequence alignments were achieved using Clustal W2 (http://www.ebi.ac.uk/Tools/msa/clustalw2/). Phylogenetic analysis was performed using MEGA version 5 after multiple alignment of the data via CLUSTAL_X. Distances were obtained using options according to Kimura’s two-parameter model and clustering was performed by using the neighbor-joining method. The topology of the neighbor-joining phylogenetic tree was evaluated by using bootstrap resampling with 1000 replications26.
Fractionation of bacterial cells
Outer membrane proteins of V. parahaemolyticus were obtained as previously described27. Briefly, exponential phase cultures (4 ml) were pelleted, lysed by sonication and then centrifuged at 20,000 g for 2 min. Supernatants were then transferred to fresh tubes and centrifuged again (20,000 g, 30 min, 4 °C). The pellets were re-suspended in 500 μl 1% sodium lauryl sulfate in 10mM HEPES (pH7.4) and incubated at room temperature for 30 min. Outer membrane proteins were obtained after centrifugation at 20,000 g for 30 min (4 °C). The detergent soluble and insoluble fractions of the outer membrane proteins were separated by SDS-PAGE on 11% polyacrylamide gel, transferred to PVDF membrane and subjected to Western Blotting using rabbit α-VpadV polyclonal antibody (1:2000, Pierce).
Expression and purification of recombinant proteins
Different fragments of VpadF were amplified by PCR using the primers listed in ST 2 and cloned into pET28 vector. Recombinant proteins were induced by IPTG and then purified using affinity chromatograph methods as previously described28, dialyzed into PBS buffer and examined by SDS–PAGE. Protein concentrations were measured by comparison with the BSA standards (Amresco).
Solid phase binding assay
One hundred μl each of recombinant fibrinogen (F3879, sigma), full-length fibronectin (F2006, Sigma), HBD (F9911, Sigma) and CBD (F0162, 45-kDa, Sigma) domains of fibronectin at a concentration of 5 μg/ml in 50mM Na2CO3, pH 9.6 was coated onto ELISA plates (IWAKI) at 4 °C overnight. After three-time washes with PBS, the plate was blocked with 1% (w/v) BSA in 50 mM Na2CO3 (pH 9.6) and incubated at room temperature for 1h. The ELISA plate was then washed three times and then incubated with 3xFlag-tagged, recombinant VpadF truncation fragments at room temperature for 1 h. After triple washes, 100 μl mouse α-3xFlag-HRP antibody (diluted 1:1000 0) was added to each well and incubated at room temperature for 1 h. 100 μl TMB (T4444, sigma) was added to each well for color development (5 min, room temperature). After quenching with 100 μl H2SO4 (1 M), absorbance was read at 450 nm. BSA was coated simultaneously as negative control.
Cell attachment and cytotoxicity assays
HeLa and HT-29 cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM, Invitrogen) supplemented with 10% fetal bovine serum (FBS, Invitrogen) at 37 °C in 5% CO2. Attachment assay was carried out as previously described29. Briefly,70–90% confluence cells were infected with freshly prepared V. parahaemolyticus at a multiplicity of infection (MOI) of ~10 CFU/cell. To consider the factor that V. parahaemolyticus may replicate during the experiment, bacterial cell were added to the empty wells of the cell culture plates and incubate for the same time as the binding experiment to determine the final total number of V. parahaemolyticus for the binding experiment. After 1 h infection, cells were washed three times with PBS, lysed with 1% Triton X-100 at 37 °C for 10 min to get the successful attached bacteria. The cell lysates and control bacteria were serially diluted and plated on LBS agar. Attachment rate was calculated by dividing bound bacteria to the total bacterial load.
For cytotoxicity assay, similar conditions were used as mentioned above. The supernatants from infected epithelial cell cultures (50–80% confluence) were collected at specific time points. The amounts of LDH release were determined using CytoTox 96 Non-Radioactive Cytotoxicity kit (Promega) following the manufacturer’s instructions. The LDH activity in the supernatant of uninfected cells was also measured to obtain a spontaneous LDH value. Percent cytotoxicity was calculated with the following formula: (test LDH release-spontaneous release)/(maximal release-spontaneous release).
RT-PCR
Overnight V. parahaemolyticus culture was firstly diluted 1:100 in fresh LBS broth and grown to exponential growth phase (OD600 ≈ 0.5–0.7). Cells were collected and re-suspended in the same volume of DMEM with different concentration of FBS (0, 5% and 10% FBS), respectively. After incubation for 30 min at 37 °C, bacteria were collected. Exponential growth phase culture that grown in LBS was directly collected. RNA was extracted using Trizol (Invitrogen) following the manufacturer’s instructions. Residual DNA was removed from the sample with DNase (Turbo DNase, Ambion). RT-PCR reactions were performed using Superscript one-step RT-PCR system (Invitrogen) following the manufacturer’s instructions. Primer pairs, rtVpadF-F/rtVpadF-R and rtrpoA-F/rtrpoA-R, were used to amplify the target genes, respectively (ST 2).
Murine Infection assay
V. parahaemolyticus strains (108 CFU) were intraperitoneally injected into 6- to 10-week-old C57BL/6 mice (n = 10) as described previously30,31,32 and the survival of mice was measured at the indicated time points. Three independent experiments were performed. The animal experiments were conducted in the National Institute for Communicable Disease Control and Prevention, Chinese Center for Disease Control and Prevention (CDC) following the guidelines and policies approved by China CDC.
Statistical analysis
Statistical analysis was performed using Prism software (version 5.0, GraphPad software). The data were the averages of three independent experiments. A P value of 0.05 or lower was considered significant. For analysis of the murine survival ratio, Kaplan-Meier and log rank tests were performed and P values of <0.01 was considered statistically significant.
Results
VpadF is probably a adhesin gene in V. parahaemolyticus
To identify potential adhesins, we searched the genome sequence of a pandemic strain RIMD2210633. An open-reading frame, vp1767, designated VpadF, which encodes a putative protein with 736 amino acid residues, attracted our attention. VpadF consists of a putative transmembrane domain at the N-terminal, followed by five bacterial immunoglobulin-like tandem repeats (Bigs). Sequence alignment analysis showed that Bigs in VpadF possesses ~30% amino acid identity to the last Big of LigA and LigB, which possess 13 and 12 Bigs, respectively (SF3). LigA and LigB were described as adhesive molecules in Leptospira interrogans33,34,35,36. Bigs are widely present in numerous proteins, especially in surface proteins that involved in pathogen-host cells interactions36,37. It is likely that VpadF is an adhesive factor.
Proteins that possess similar domain configuration to that of VpadF also exist in other bacteria, such as Butyrivibrio fibrisolvens, Gemmatimonas aurantiaca, V. campbellii, V. harveyi, etc (Fig. 1A), suggesting that they share a common ancestor and represent a protein family. Phylogenetic analysis based on the core regions of VpadF and its homologs revealed that VpadF formed a distinct branch related to its homolog from V. cholerae, while it was separated from that of V. harveyi (Fig. 1B). PCR screening of VpadF on environmental and clinical V. parahaemolyticus strains showed that almost all isolates possessed this gene with a few exceptional cases (Table 1). BLAST of the 293 available draft genome sequences of V. parahaemolyticus in GenBank confirmed that about 256 out of 293 (87%) whole genome sequences contained VpadF gene. BLASTN of GenBank non-redundant (NR) database failed to find any high homologues in any bacteria other than V. parahaemolyticus. In addition, the codon adaption index (CAI) and expression level analysis of VpadF were found to be normal compared to other genes in V. parahaemolyticus (Data not shown). These data suggested that VpadF is likely to be an intrinsic gene in V. parahaemolyticus and gets lost in some cases rather than a horizontal gene transferred (HGT) gene.
Schematic representation and phylogenetic analysis of VpadF.
(A) Schematic representation of VpadF and its homologues; (B) Neighbor-joining tree based on five Bigs sequences showing the phylogenetic relationships of VpadF and its homologues. The protein sequences were obtained from either NCBI or UniProt. Bootstrap values (>50%) are shown at branch nodes. Bar, 0.2 difference at the amino acid level.
VpadF localizes to the outer membrane
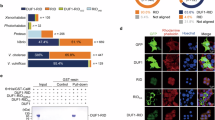
Firstly, we tried to find the appropriate condition for endogenous VpadF expression because some adhesin genes are not able to be expressed in vitro, or just transcribed under specific condition38,39. As shown in Supplementary Figure 1, the transcription of VpadF was stimulated in DMEM media in a FBS-independent manner, while its expression was inhibited in LBS broth. We then analyzed the localization of VpadF by heterogeneous expression in E. coli BL21. After Coomassie blue staining of the SDS-PAGE gel, a ~110 KDa protein band was clearly visible from the outer membrane fractions of E. coli BL21 expression VpadF (Fig. 2A). To further determine the cellular localization of VpadF in V. parahaemolyticus, the outer membrane fractionation of three clinical V. parahaemolyticus strains, VPATCC17802, VP3218 and VP1074, were probed with VpadF antibody generated in our lab. It showed that VpadF was localized at the detergent insoluble fraction of the outer membrane of V. parahaemolyticus (Fig. 2B).
VpadF mediates V. parahaemolyticus infection to epithelial cells
Having confirmed that VpadF is a surface protein, we then assessed the role of VpadF on V. parahaemolyticus attachment to HeLa and HT-29 epithelial cells. A ΔvcrD1 background strain (T3SS1 negative), which showed a much slower cell lysis rate, was constructed as wild type (WT) V. parahaemolyticus caused cell lysis quickly after infection, making it unfeasibl for the attachment assay40. The deletion of VpadF in the ΔvcrD1 strain dramatically decreased its attachment to HeLa and HT-29 epithelial cells (Fig. 3B). The complementary strain, ΔVpadFΔvcrD1:pVpadF, showed ~3 fold higher cell adherence ratio than that of the ΔVpadFΔvcrD1 strain (Fig. 3B), suggesting that VpadF is required for bacterial cell attachment and in trans expression of VpadF increases its cell attachment ratio.
Contribution of VpadF to V. parahaemolyticus host cell attachment and cytotoxicity.
(A) Adhesion of different strains of V. parahaemolyticus including wild type (WT), VpadF deletion mutant (ΔVpadF) and VpadF deletion mutant complementary strain (ΔVpadF:pVpadF) to HeLa and HT-29 epithelial cells. Values represent the mean ± the SE of three independent experiments. Statistical comparisons were performed with a one-way analysis of variance (ANOVA) with Tukey’s multiple comparison tests. **p < 0.01. Cytotoxicity of different strains of V. parahaemolyticus including wild type (WT), VpadF deletion mutant (ΔVpadF) and VpadF deletion mutant complementary strain (ΔVpadF:pVpadF) to HeLa (B) and HT-29 epithelial cells (C). The LDH released from lysed HeLa and HT-29 epithelial cells were measured at specific time points. Values represent the mean + the SE of three independent experiments.
To test whether the host cell adherence mediated by VpadF affects cytotoxicity of V. parahaemolyticus against the epithelium, the lysis of HeLa and HT-29 epithelial cells were measured by monitoring the release of lactate dehydrogenase (LDH) after infection with WT V. parahaemolyticus, ΔVpadF strain (vcrD1 positive) and the complementary strain ΔVpadF:pVpadF. At 3.5 h, ΔVpadF caused a ~50% decreased cytotoxicity compared to that caused by WT and ΔVpadF::pVpadF. After 5.5 h infection, HeLa and HT-29 epithelial cells were nearly completely lysed by WT and the VpadF complementary strains, while ΔVpadF infected cells showed ~30% less lysis (Fig. 3C). This suggested that VpadF plays an important role in epithelial cell infection.
VpadF is responsible for lethality in mice
To explore whether VpadF affects V. parahaemolyticus pathogenesis, mice were infected with WT V. parahaemolyticus VP3218 and its VpadF deletion mutant, respectively. As shown in Fig. 4A–C, mice infected with WT strain displayed only 20% survival, while deletion of VpadF nearly completely abolished the lethality in mice after 48 h infection. After 96 h infection, no further lethal effect was observed in the mice infected by WT V. parahaemolyticus VP3218 or its VpadF deletion mutant (data not shown). This indicated that VpadF mediating V. parahaemolyticus-host interaction is required for its pathogenesis in mice.
Survival rates of murine models infected with V. parahaemolyticus strains.
C57BL/6 mice (n = 10) were infected intraperitoneally with WT or ΔVpadF strains (108 CFU) and the dead mice were recorded at specific time points during a 96 period of time. Only the data for the first 48 h was shown since no further change was observed in these mice for the next 48 h. Data from three independent experiments, (A–C), were shown. Kaplan-Meier and log rank tests were used to analyze the data (P < 0.001).
VpadF binds both fibronectin (Fn) and fibrinogen (Fg)
To gain further insight into the mechanism by which VpadF mediates host cells attachment and thus contributes to the pathogenesis, we set out to identify potential host receptors of VpadF. It was suggested that besides mam7, V. parahaemolyticus produces other adhesins to bind Fn29. Initially, we assessed the Fn binding ability of VpadF. VpadFA, the VpadF truncated fragment that lack of 36 N-terminal amino acid residues was first purified (Fig. 5A and SF2). The specific interaction between VpadFA and Fn was examined using the solid-phase binding assay. As shown in Fig. 5B, VpadFA was able to interact with Fn in a pronounced dose-dependent manner and reached a saturated status, with a Kd (the concentration able to saturate 50% of substrate) of ~200 nM. To evaluate which region in VpadF contributes to its binding property, all five Big repeats, VpadFB1 (the first Big domain), VpadFB2 (the second Big domain), VpadFB3 (the third Big domain), VpadFB4 (the fourth Big domain), VpadFB5 (the fifth Big domain) and the C-terminal end, VpadFC, were individually purified, respectively. Our results demonstrated that VpadFB2, VpadFB5 and VpadFC strongly bound to Fn and VpadFC displayed highest binding affinity. In contrast, VpadFB1, VpadFB3 and VpadFB4 showed weak binding activities to Fn (Fig. 5C).
Characterization of interactions between VpadF and its cell surface receptors.
(A) Schematic representation of VpadF, its truncation domains and fibronectin; VpadFA (residues 37–736), VpadFB1 (residues 98–174), VpadFB2 (residues 184–255), VpadFB3 (residues 272–348), VpadFB4 (residues 356–433), VpadFB5 (residues 442–522) and VpadFC (residues 625–736); (B) Binding of rVpadFA to fibronectin; (C) Binding of various domains of rVpadF, rVpadFB1, B2, B3, B4, B5 and C to fibronectin; (D,E) Binding of rVpadFA, B2, B5 and C to HBD and CBD of fibronectin; (F) Binding of rVpadFA to fibrinogen; (G) Binding of various domains of rVpadF, rVpadFB1, B2, B3, B4, B5 and C to fibrinogen. BSA was used as negative control. Values represent the mean + the SE of three independent experiments.
To further demonstrate which parts of Fn are bound by VpadF, we tested the binding activities of VpadFA, B2, B5 and C to the immobilized N- terminal 30-KDa (heparin binding domain, HBD) and 45-KDa fragments (gelatin and collagen binding domain, CBD) of Fn. As shown in Fig. 5D,E, both HBD and CBD were targeted by VpadFA, B2, B5 and C. VpadFB5 and VpadFC bound HBD and CBD with similar avidities, while higher than those of VpadFB2.
Septicemia caused by V. parahaemolyticus is potential fatal to immunocompromised and liver failure individuals41. In Gram-positive pathogens, septicemia is related to their binding abilities to Fg since Fg is not only rich in human blood but also present in extracellular matrix42,43. Thus, we tested the binding ability of VpadF to Fg. Our results displayed that Fg was bound by VpadFA, with a Kd of ~400 nM (Fig. 5F). Similar to VpadF binding Fn, this protein also interacted with Fg mainly dependent on its C-terminal region, VpadFB5 and VpadFC (Fig. 5G).
Discussion
Initial host cell adhesion is the first important step in bacterial pathogenesis. In the case of V. parahaemolyticus, its infectious processes require the adherence to intestinal epithelial cells, resulting in epithelial cell extrusion, villus disintegration and formation V. parahaemolyticus-filled cavities44. Adhesins play a pivotal role in the tight bacteria-host cells attachment to initiate the downstream processes. In the present study, we discover a novel adhesin gene, VpadF, that plays a significant role on the pathogenesis of V. parahaemolyticus.
VpadF has several features that distinguish it from other previously described adhesive factors, such as MSHA pilus, enolase, capsular polysaccharide, T6SSs and MAM7 in V. parahaemolyticus. Firstly, this protein shows a unique domain organization. Secondly, VpadF is required for the lethal effects of V. parahaemolyticus on mice. Moreover, VpadF is a multifunctional adhesin that is capable of interact with both Fn and Fg. To the best of our knowledge, VpadF is the first reported adhesin that be able to bind Fg in Vibrio species. It was shown that Fg binding proteins have essential roles in pathogenesis by enabling bacteria to penetrate host barriers and spread in tissues43. VpadF is likely to both strengthen the attachment of V. parahaemolyticus to epithelial cells and modulate its spreading in the infected tissues. The molecular mechanism of the interactions between VpadF and host receptors is also different from those of other adhesins in Vibrio. For instance, OmpU from V. vulnificus recognizes RGD tripeptide that localizes the middle part of Fn and MAM7 interact with HBD of Fn29,45. In contrast, VpadF can bind the whole N-terminal fragments, HBD and CBD of Fn.
VpadF also differs from Bigs possessing adhesins in Leptospira interrogans. Biochemical analysis revealed that the fifth Big repeat in VpadF significantly contributes to Fn and Fg binding, while in LigA and LigB, their binding avidities are mainly dependent on the 13th and 12th Big domain, respectively46,47,48. Moreover, unlike in LigA and LigB, only the last Bigs can recognize host components, the 2nd Big repeat in VpadF also exhibited Fn/Fg binding property to some extent. These results implicates the number of Bigs is not the primary determinant of function and individual Big repeat in a single protein could be functional diverged even each Big fold shares some sequence similarity with each other (SF3).
In conclusion, we identified and characterized a novel and essential adhesion factor from V. parahaemolyticus and demonstrated its significant role on host cell attachment, cytotoxicity and pathogenicity. VpadF is a potential good candidate for V. parahaemolyticus infection control for both vaccine development and drug target.
Additional Information
How to cite this article: Liu, M. and Chen, S. A novel adhesive factor contributing to the virulence of Vibrio parahaemolyticus. Sci. Rep. 5, 14449; doi: 10.1038/srep14449 (2015).
References
Baffone, W. et al. Detection of free-living and plankton-bound vibrios in coastal waters of the Adriatic Sea (Italy) and study of their pathogenicity-associated properties. Environmental microbiology 8, 1299–1305 (2006).
Chimetto, L. A. et al. Vibrios dominate as culturable nitrogen-fixing bacteria of the Brazilian coral Mussismilia hispida. Systematic and applied microbiology 31, 312–319 (2008).
Blake, P. A., Weaver, R. E. & Hollis, D. G. Diseases of humans (other than cholera) caused by vibrios. Annu Rev Microbiol 34, 341–367 (1980).
Yan, Y. et al. Extended MLST-based population genetics and phylogeny of Vibrio parahaemolyticus with high levels of recombination. International journal of food microbiology 145, 106–112 (2011).
Theethakaew, C. et al. Genetic relationships of Vibrio parahaemolyticus isolates from clinical, human carrier and environmental sources in Thailand, determined by multilocus sequence analysis. Applied and environmental microbiology 79, 2358–2370 (2013).
Caburlotto, G., Lleo, M. M., Gennari, M., Balboa, S. & Romalde, J. L. The use of multiple typing methods allows a more accurate molecular characterization of Vibrio parahaemolyticus strains isolated from the Italian Adriatic Sea. FEMS microbiology ecology 77, 611–622 (2011).
Gennari, M., Ghidini, V., Caburlotto, G. & Lleo, M. M. Virulence genes and pathogenicity islands in environmental Vibrio strains nonpathogenic to humans. FEMS microbiology ecology 82, 563–573 (2012).
Okada, N. et al. Identification and characterization of a novel type III secretion system in trh-positive Vibrio parahaemolyticus strain TH3996 reveal genetic lineage and diversity of pathogenic machinery beyond the species level. Infection and immunity 77, 904–913 (2009).
Matz, C., Nouri, B., McCarter, L. & Martinez-Urtaza, J. Acquired type III secretion system determines environmental fitness of epidemic Vibrio parahaemolyticus in the interaction with bacterivorous protists. PloS one 6, e20275 (2011).
Shirai, H. et al. Molecular epidemiologic evidence for association of thermostable direct hemolysin (TDH) and TDH-related hemolysin of Vibrio parahaemolyticus with gastroenteritis. Infection and immunity 58, 3568–3573 (1990).
Matsuda, S. et al. Association of Vibrio parahaemolyticus thermostable direct hemolysin with lipid rafts is essential for cytotoxicity but not hemolytic activity. Infection and immunity 78, 603–610 (2010).
Ohnishi, K. et al. Relationship between heat-induced fibrillogenicity and hemolytic activity of thermostable direct hemolysin and a related hemolysin of Vibrio parahaemolyticus. FEMS microbiology letters 318, 10–17 (2011).
Park, K. S., Arita, M., Iida, T. & Honda, T. vpaH, a gene encoding a novel histone-like nucleoid structure-like protein that was possibly horizontally acquired, regulates the biogenesis of lateral flagella in trh-positive Vibrio parahaemolyticus TH3996. Infection and immunity 73, 5754–5761 (2005).
Liu, M., Yan, M., Liu, L. & Chen, S. Characterization of a novel zinc transporter ZnuA acquired by Vibrio parahaemolyticus through horizontal gene transfer. Frontiers in cellular and infection microbiology 3, 61 (2013).
Kline, K. A., Falker, S., Dahlberg, S., Normark, S. & Henriques-Normark, B. Bacterial adhesins in host-microbe interactions. Cell Host Microbe 5, 580–592 (2009).
Pizarro-Cerda, J. & Cossart, P. Bacterial adhesion and entry into host cells. Cell 124, 715–727 (2006).
Carabeo, R. Bacterial subversion of host actin dynamics at the plasma membrane. Cell Microbiol 13, 1460–1469 (2011).
Jiang, W. et al. Vibrio parahaemolyticus enolase is an adhesion-related factor that binds plasminogen and functions as a protective antigen. Appl Microbiol Biotechnol 98, 4937–48 (2014).
Yu, Y. et al. Putative type VI secretion systems of Vibrio parahaemolyticus contribute to adhesion to cultured cell monolayers. Arch Microbiol 194, 827–835 (2012).
Hsieh, Y. C. et al. Study of capsular polysaccharide from Vibrio parahaemolyticus. Infection and immunity 71, 3329–3336 (2003).
O’Boyle, N., Houeix, B., Kilcoyne, M., Joshi, L. & Boyd, A. The MSHA pilus of Vibrio parahaemolyticus has lectin functionality and enables TTSS-mediated pathogenicity. Int J Med Microbiol 303, 563–573 (2013).
Krachler, A. M., Ham, H. & Orth, K. Outer membrane adhesion factor multivalent adhesion molecule 7 initiates host cell binding during infection by gram-negative pathogens. Proc Natl Acad Sci USA 108, 11614–11619 (2011).
Bardiau, M., Szalo, M. & Mainil, J. G. Initial adherence of EPEC, EHEC and VTEC to host cells. Veterinary research 41, 57 (2010).
Mikula, K. M., Kolodziejczyk, R. & Goldman, A. Yersinia infection tools-characterization of structure and function of adhesins. Frontiers in cellular and infection microbiology 2, 169 (2012).
Wagner, C. & Hensel, M. Adhesive mechanisms of Salmonella enterica. Advances in experimental medicine and biology 715, 17–34 (2011).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance and maximum parsimony methods. Molecular biology and evolution 28, 2731–2739 (2011).
Carlone, G. M., Thomas, M. L., Rumschlag, H. S. & Sottnek, F. O. Rapid microprocedure for isolating detergent-insoluble outer membrane proteins from Haemophilus species. J Clin Microbiol 24, 330–332 (1986).
Baldwin, M. R., Bradshaw, M., Johnson, E. A. & Barbieri, J. T. The C-terminus of botulinum neurotoxin type A light chain contributes to solubility, catalysis and stability. Protein Expr Purif 37, 187–195 (2004).
Krachler, A. M. & Orth, K. Functional characterization of the interaction between bacterial adhesin multivalent adhesion molecule 7 (MAM7) protein and its host cell ligands. J Biol Chem 286, 38939–38947 (2011).
Hiyoshi, H., Kodama, T., Iida, T. & Honda, T. Contribution of Vibrio parahaemolyticus virulence factors to cytotoxicity, enterotoxicity and lethality in mice. Infection and immunity 78, 1772–1780 (2010).
Pineyro, P. et al. Development of two animal models to study the function of Vibrio parahaemolyticus type III secretion systems. Infection and immunity 78, 4551–4559 (2010).
Whitaker, W. B., Parent, M. A., Boyd, A., Richards, G. P. & Boyd, E. F. The Vibrio parahaemolyticus ToxRS regulator is required for stress tolerance and colonization in a novel orogastric streptomycin-induced adult murine model. Infection and immunity 80, 1834–1845 (2012).
Choy, H. A. et al. Physiological osmotic induction of Leptospira interrogans adhesion: LigA and LigB bind extracellular matrix proteins and fibrinogen. Infection and immunity 75, 2441–2450 (2007).
Palaniappan, R. U. et al. Cloning and molecular characterization of an immunogenic LigA protein of Leptospira interrogans. Infection and immunity 70, 5924–5930 (2002).
Palaniappan, R. U. et al. Expression of leptospiral immunoglobulin-like protein by Leptospira interrogans and evaluation of its diagnostic potential in a kinetic ELISA. Journal of medical microbiology 53, 975–984 (2004).
Matsunaga, J. et al. Pathogenic Leptospira species express surface-exposed proteins belonging to the bacterial immunoglobulin superfamily. Molecular microbiology 49, 929–945 (2003).
Isberg, R. R., Voorhis, D. L. & Falkow, S. Identification of invasin: a protein that allows enteric bacteria to penetrate cultured mammalian cells. Cell 50, 769–778 (1987).
Dorsey, C. W., Laarakker, M. C., Humphries, A. D., Weening, E. H. & Baumler, A. J. Salmonella enterica serotype Typhimurium MisL is an intestinal colonization factor that binds fibronectin. Molecular microbiology 57, 196–211 (2005).
Seo, H. S. et al. Bacteriophage lysin mediates the binding of streptococcus mitis to human platelets through interaction with fibrinogen. PLoS Pathog 6, e1001047 (2010).
Burdette, D. L., Yarbrough, M. L., Orvedahl, A., Gilpin, C. J. & Orth, K. Vibrio parahaemolyticus orchestrates a multifaceted host cell infection by induction of autophagy, cell rounding and then cell lysis. Proc Natl Acad Sci USA 105, 12497–12502 (2008).
Blake, P. A., Merson, M. H., Weaver, R. E., Hollis, D. G. & Heublein, P. C. Disease caused by a marine Vibrio. Clinical characteristics and epidemiology. The New England journal of medicine 300, 1–5 (1979).
Adams, R. A., Schachtrup, C., Davalos, D., Tsigelny, I. & Akassoglou, K. Fibrinogen signal transduction as a mediator and therapeutic target in inflammation: lessons from multiple sclerosis. Current medicinal chemistry 14, 2925–2936 (2007).
Rivera, J., Vannakambadi, G., Hook, M. & Speziale, P. Fibrinogen-binding proteins of Gram-positive bacteria. Thrombosis and haemostasis 98, 503–511 (2007).
Ritchie, J. M. et al. Inflammation and disintegration of intestinal villi in an experimental model for Vibrio parahaemolyticus-induced diarrhea. PLoS Pathog 8, e1002593 (2012).
Goo, S. Y. et al. Identification of OmpU of Vibrio vulnificus as a fibronectin-binding protein and its role in bacterial pathogenesis. Infection and immunity 74, 5586–5594 (2006).
Lin, Y. P. & Chang, Y. F. A domain of the Leptospira LigB contributes to high affinity binding of fibronectin. Biochemical and biophysical research communications 362, 443–448 (2007).
Lin, Y. P., McDonough, S. P., Sharma, Y. & Chang, Y. F. Leptospira immunoglobulin-like protein B (LigB) binding to the C-terminal fibrinogen alphaC domain inhibits fibrin clot formation, platelet adhesion and aggregation. Molecular microbiology 79, 1063–1076 (2011).
Lin, Y. P., McDonough, S. P., Sharma, Y. & Chang, Y. F. The terminal immunoglobulin-like repeats of LigA and LigB of Leptospira enhance their binding to gelatin binding domain of fibronectin and host cells. PloS one 5, e11301 (2010).
Acknowledgements
We acknowledge Drs. Wenchao Liu and Zhuo Wang for providing HeLa and HT-29 epithelial cells, respectively. We also thank Drs. Hans Wolf-Watz, Douglas R. Call, Kim Orth and Roland Nordfelth for kindly providing E. coli SY327 λpir strain and plasmids pDM4, pMMB207 and pPK2013. This work is supported by Chinese National Key Basic Research and Development (973) Program (2013CB127200) and the Research Fund for the Control of Infectious Diseases from the Food and Health Bureau, the Government of Hong Kong SAR (13121422 to SC).
Author information
Authors and Affiliations
Contributions
M.L. designed and performed experiments, analyzed the data and wrote manuscript; S.C. designed the experiments, analyzed the data, wrote manuscript and coordinated the whole project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Liu, M., Chen, S. A novel adhesive factor contributing to the virulence of Vibrio parahaemolyticus. Sci Rep 5, 14449 (2015). https://doi.org/10.1038/srep14449
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14449
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.