Abstract
The centromere/kinetochore interaction is responsible for the pairing and segregation of replicated chromosomes in eukaryotes. Centromere DNA is portrayed as scarcely conserved, repetitive in nature, quickly evolving and protein-binding competent. Among primates, the major class of centromeric DNA is the pancentromeric α-satellite, made of arrays of 171 bp monomers, repeated in a head-to-tail pattern. α-satellite sequences can either form tandem heterogeneous monomeric arrays or assemble in higher-order repeats (HORs). Gorilla centromere DNA has barely been characterized and data are mainly based on hybridizations of human alphoid sequences. We isolated and finely characterized gorilla α-satellite sequences and revealed relevant structure and chromosomal distribution similarities with other great apes as well as gorilla-specific features, such as the uniquely octameric structure of the suprachromosomal family-2 (SF2). We demonstrated for the first time the orthologous localization of alphoid suprachromosomal families-1 and −2 (SF1 and SF2) between human and gorilla in contrast to chimpanzee centromeres. Finally, the discovery of a new 189 bp monomer type in gorilla centromeres unravels clues to the role of the centromere protein B, paving the way to solve the significance of the centromere DNA’s essential repetitive nature in association with its function and the peculiar evolution of the α-satellite sequence.
Similar content being viewed by others
Introduction
In multicellular eukaryotes the pairing and segregation of replicated chromosomes in mitosis and meiosis is essential to guarantee the complete and correct chromosomal complement in daughter cells. This process is mediated by the synchronized work of the spindle apparatus that connects to chromosomes through a proteinaceous bridge called a kinetochore1. The chromosomal counterpart of this molecular link is represented by the centromere. Throughout the mammalian orders, as well as more generally among all higher eukaryotes, the centromeric chromatin has been initially and consistently described by the incorporation of a histone H3 variant, the centromere protein A (CENP-A)2,3 and the recruiting of the centromere protein B (CENP-B)4.
Focusing on the primary sequence of the centromeric DNA, its low conservation as well as its rapid and distinctive evolution are copiously described. Nonetheless, it is generally acknowledged that there is an outstanding maintenance of features, such as its repetitive nature, structure and protein-binding competence5. The centromeric DNA is AT-rich and highly repetitive in almost every kind of plant and animal studied to date (with the exception of the holocentric centromeres of C. elegans)6,7. This constant co-occurrence between the centromeric functionality and repetitive DNA has been interpreted as evidence for a functional role of the satellite DNA, helping the assembly of the kinetochore and allowing a perfect timing at chromosome segregation.
Among primates, the major class of centromeric DNA is the pancentromeric α-satellite, composed of long stretches of 171 bp monomers, tandemly repeated in a head-to-tail pattern that extends for ~250 kbp up to ~5 Mbp per chromosome8,9,10. These sequences have been identified throughout the primate order, including great apes, Old World and New World monkeys11,12,13,14, with the exception of the suborder Strepsirhini15. α-satellite DNA is the most abundant repetitive DNA in all primate species studied, making up to 3–5% of each chromosome10,16.
The α-satellite can be variously classified, depending on its (i) primary sequence, (ii) multimeric structure and (iii) localization with respect to the centromere.
(i) Sequence analyses revealed that all existing primate types of alphoid monomers are most likely to be derived from only two ancient types of units and are therefore designed as “A” or “B” monomers17. (ii) α-satellite sequences can either form tandem heterogeneous monomeric arrays (10–40% divergence between individual monomers) or organize multimeric structures assembled in a period known as higher-order repeats (HORs)18,19,20,21,22,23. HOR units may be composed of two to over 30 monomers and are tandemly repeated several hundreds to thousands times per single centromere13,24,25. (iii) Where investigated, multimeric arrays have been found to contribute to the bulk of the centromeric chromatin, bordered by a monomeric α-satellite that acts as a junction to the pericentromeric regions19,20,26.
In HORs, monomers within a period differ greatly in sequence, while monomers standing at corresponding positions of different periods are virtually identical (<2% sequence divergence)20. The presence of alphoid HORs has been reported throughout the superfamily Hominoidea and is the evolutionary result of sequence homogenization created by molecular drive mechanisms, such as amplification, unequal crossing-over and gene conversion27,28. In particular, homogenization shows three patterns: local homogenization in tandem, intrachromosomal homogenization patterns that are regional but not in tandem and interchromosomal or interarray patterns29.
Among the centromeric proteins that have been characterized, there are two main proteins directly binding the alphoid DNA: pJα and CENP-B. These proteins recognize the 17 bp pJα-motif (CTAPyGGTGPuAAAAGGAA) and CENP-B box (PyTTCGTTGGAAPuCGGGA) within A- and B-type monomers, respectively17. Modern great ape centromere organization emerged from ancestral A-type monomers mixed with more recent B-type monomers17. Like other tandem satellite families, the α-DNA evolves through molecular drive by mechanisms such as unequal crossing-over, gene conversion and transposition16,30,31. Originally, unequal crossover occurred between two similar monomers, creating tandem duplications (ancestral centromeric repeats of monomeres, ACRMs). Subsequent unequal crossovers were able to expand tandem arrays until a subset of ACRMs was multimerized into higher-order α-satellites thus creating several distinct HOR alphoid DNA families. HOR refers then to a structure in which multiple copies of the fundamental repeat units appear periodically19.
Due to the extremely high sequence identity of higher-order α-satellite monomers, unequal crossovers are much more likely to happen in HORs rather than in monomeric α-satellites causing different rates of evolution between them. Indeed, less conservation and higher intraspecies homogenization of orthologous HOR units than orthologous monomeric α-satellites have been described in closely related species32,33.
Based on monomer composition, structure and distribution, HOR DNA in great apes shapes different suprachromosomal families (SFs). In humans, for example, α-satellite sequences have been grouped into five SFs: SF1-5, each characterized by its own specific chromosomal distribution17. The five human SFs were initially revealed by restriction site periodicity and then identified by sequence-based phylogenetic analyse19,34,35. SF1, SF2 and SF5 are dimeric, i.e., contain two different monomers (A- and B-type) alternating regularly, as in SF1 and SF2, or are irregularly assembled, as in SF536,37,38; SF3 is pentameric, i.e., formed by five different types of monomers (two A-type and three B-type)13; SF4 is monomeric, i.e., shaped by arrays of equally related monomers35. SF1-3 are organized in HORs and often designated as “new families” compared to the ancestral SF4 and SF5 and compose the centromeric region of all human chromosomes except the Y chromosome.
Conserved CENP-B boxes are located mostly in dimeric SFs and are regularly distributed every other monomer39,40. It has also been hypothesized that CENP-B dimers have a key role in the assembly of the centromeric chromatin by juxtaposing CENP-B boxes in α-satellite arrays40,41,42.
Since several studies have coupled the centromere function to higher-order rather than monomeric α-satellite in humans, the fairly recent creation of the “new families” may look intriguing and somehow counterintuitive33,39,43. Despite having been deeply characterized in humans, α-satellite structure and organization knowledge in primates has been primarily based on hybridizations of human alphoid sequences and restriction patterns. Among great apes, chimpanzee contains human-like HORs44, while orangutan mostly displays a basic monomeric organization, with HORs being very rare45. Nevertheless, the monomer primary sequence has been maintained similar enough to allow all human SFs to cross-hybridize with orangutan chromosomes at low stringency conditions.
Information on gorilla centromeric DNA is particularly meager and no details on its evolutionary history have been previously reported46. Here, we analyze the centromeric DNA in gorilla with several different approaches to achieve a broad display of organization and evolutionary history of the gorilla-specific α-satellite. Parallelisms with human alphoid sequences as well as gorilla-specific distinctive traits of the alphoid DNA were found.
Results
Our investigation of gorilla centromeres was first achieved by isolating gorilla (GGO) centromeric DNA using human alphoid DNA sequence similarities47,48,49 and subsequently collecting GGO bacterial artificial chromosome (BAC) α-satellite clones and long gorilla centromeric sequences from online databases (whole-genome shotgun sequence, WGSS). Long sequences, such as BAC clones and WGSS, were essential to characterize the alphoid DNA structure and organization.
We isolated the GGO α-satellite by PCR amplification of gorilla genomic DNA using α-27/α-30 primers obtained from the most conserved regions of human alphoid consensus50,51. We then used the amplicons as probes in fluorescence in situ hybridization (FISH) experiments on gorilla metaphase chromosomes. The centromeric region of each chromosome was highlighted and no distribution or intensity differences were observed under either low or high stringency conditions. We cloned the amplicons and tested the clones by FISH on gorilla metaphase spreads; two main hybridization patterns were found (Groups 1 and 2, Table S1, Supplementary Note-Section 1), revealing two specific groups of centromeric sequences as main component of gorilla centromeres. A third subset of plasmids hybridized to chromosomes belonging to both Groups 1 and 2 (Group 3) and, lastly, there were pericentromeric clones hybridizing to only one pair of chromosomes (Group 4) (Table S1, Fig. 1). We also found “exceptional” clones that hybridized to all the chromosomes of one of the two main Groups (1 or 2) plus one single chromosome of the other group: Group 1 plus chromosome XVII or Group 2 plus chromosome V (Table S1 and Supplementary Note-Section 1).
To fully characterize the organization of centromeres in the gorilla genome, we collected gorilla α-satellites containing long-insert BAC clones (n = 41) and tested them by both FISH and in vitro enzymatic restriction. BAC FISH results on GGO metaphases were concordant to the previous results showing the same two main groups of patterns. Restriction patterns revealed that BACs from Group 1 were composed of dimeric sequences, while BACs belonging to Group 2 had a more complex and heterogeneous organization (Table S1, Supplementary Note-Section 2).
The same approach, from short-insert clones to long fully sequenced arrays, was used to perform α-satellite analysis by sequence. A subset of plasmids representative of the whole pool was sequenced, generating about 38 kbp of gorilla-specific alphoid DNA. We obtained 57 sequences ranging from 168 to 1536 bp in length (GenBank accession numbers JQ685164.1-JQ685171.1, JQ685175.1-JQ685179.1, JQ685186.1-JQ685223.1), containing up to eight α-satellite monomers per clone and 171 total monomers. Of the 57 sequences, 39 contained more than one monomer, thus being more informative to study the organization of alphoid arrays. Further, we retrieved and analyzed 66 WGSS containing α-satellites, 1293 to 26,052 bp in length, composed of 2351 monomers, making up 436 kbp of total sequence (Supplementary Note-Section 3).
We extracted 216 17-bp sequences composing one of the two centromeric protein-recognition domain (PRD) from plasmid inserts: 129/216 PRDs were pJα motif-positive, 89 possessed the CENP-B box and 7 monomers had accumulated too many substitutions to be confidently grouped (Supplementary Note-Section 1). We annotated the “core” bases required for binding: 53/129 (41.1%) monomers containing the pJα-motif had the “core” pJα sequence required for protein binding completely conserved17 and 14/53 (26.4%) showed a perfect conservation of the entire 17 bp motif. Furthermore, 44/89 (49.4%) monomers containing the CENP-B box were positive for the binding “core”, i.e., contained all nine essential positions as described by Masumoto et al.52 and 32/44 (72.7%) showed a perfectly intact CENP-B box.
In our entire collection of gorilla alphoid monomers (2521 = 171 from plasmids + 2351 from WGSS), there were 104 extra-long monomeric units (~189 bp), all containing the same insertion. The insertion interrupted the CENP-B box at the 13th position and duplicated 22 bp upstream (the unfinished CENP-B box plus nine more bases) creating a new complete and fully conserved CENP-B box. A 4 bp deletion downstream finally resulted in an 18 bp insertion (Fig. 2). All of these longer monomers, when derived from plasmid inserts, belonged to Group 2.
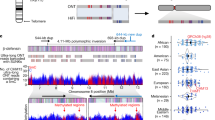
We multi-aligned all the gorilla alphoid monomers (2521) and constructed a phylogenetic tree. A- and B-type monomers were clearly separated in the two main branches (Fig. 3). We derived a general gorilla α-satellite consensus sequence from the multi-alignment and two gorilla-specific A- and B-type consensus sequences (Table 1).
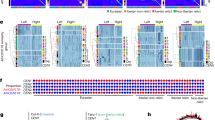
Left panel. Phylogenetic tree built by ClustalW showing all 2521 gorilla alphoid monomers extracted from WGSS plus our plasmid clones. Right panel. Venn diagram of the chromosomal hybridization pattern of the gorilla HOR alphoid suprachromosomal families in great apes according to data both in this work and in the literature. Colors and SF distributions are displayed as follows. Blue: SF1; Red: SF2; Green: SF3; Black: others.
As reported in human17, a CENP-B box can sporadically be found in A-type monomers and likewise the pJα motif in B-type monomers. Therefore, we calculated the p-distances between each GGO alphoid monomer extracted from plasmid inserts and the two GGO A- and B-type consensuses: 98.9% (92/93) of the A-type alphoid units contained the pJα motif and 84.4% (65/77) of the B-type monomers contained the CENP-B box domain (see Methods section). In addition, we found no evidence of a perfect conservation of the essential position of CENP-B box in the type-A monomers nor the “core” of pJα motif perfectly conserved in the type-B monomers. As a sign of regularity and precise organization, 21/39 sequences containing more than one monomer showed a perfect alternance of CENP-B box and pJα motif monomers (Table S2, Supplementary Note-Section 3).
WGSS and plasmid sequences were examined for higher-order periodicities by BLAST comparisons, Tandem Repeats Finder (TRF) and dot plot. 31/66 WGSS and 5/46 plasmid sequences displayed higher order organizations (dimeric, tetrameric, pentameric or octameric) (Table S3, Supplementary Figure S1, Supplementary Note-Section 3); the rest of the sequences were made up of monomeric arrays.
The analysis of the phylogenetic tree pointed out that dimeric sequences were composed of monomers from only two clusters of the tree (blue) while the octameric sequences were from three branches (red) (Fig. 3). Sequences with tetrameric or pentameric patterns (green branches) did not form a clearly separated group on the phylogenetic tree, but rather they mixed with monomeric arrays (black branches) (Tables S3, S4, Fig. 3).
The three groups of sequences were accounted for as representative of gorilla alphoid SFs: SF1, SF2 and SF3 (blue, red and green branches in Fig. 3, respectively; Tables S3 to S5). Each group of sequences was individually studied and gorilla SF-specific substitutions were found (Supplementary Note-Section 3).
We can infer that SF1 (16 WGSS and 7 plasmid clones, Table S4) is dimeric and composed of two regularly alternating types of monomers (A-type and B-type); both presented conserved essential positions in the PRD (Table S3, Supplementary Note-Section 3). SF2 (10 WGSS and 11 plasmid clones, Table S4) is formed by eight different kinds of monomers: three A-type and five B-type monomers. All but two of the three A-type monomers showed a total conservation of the essential positions of the PRD. Moreover, two out of the eight monomers composing SF2 contained the 22 bp insertion previously described and in four analyzed WGSS the insertion was further duplicated (CABD02196940.1, CABD02378856.1, CABD02378856.) or triplicated (CABD02196967.1) (Fig. 4, Table S3, Supplementary Note-Section 3). SF3 (5 WGSS, 1 plasmid, Table S5) is organized as a pentamer (three B-type and two A-type monomers); their PRDs are all conserved in the essential positions except for one of the two A-type monomers and one of the three B-type monomers (Table S3, Supplementary Note-Section 3). Sequences were assigned to SFs exclusively in the case of concordant results between the different analyses and when unequivocally composed of monomers distributed in one of the three groups of clusters of Fig. 4 (Table S4).
Examples, for each SF, of the succession of monomer types obtained by p-distance matrices analysis with both the 16 gorilla monomer units (panels on the right, first lane for each clone) and the 12 human consensus sequences (panels on the right, second lane for each clone) (see Methods section).
Gorilla consensus names have been assigned based on p-distances to human consensus monomers (e.g. the five gorilla type-B consensuses belonging to SF2 are all named “D1” as the human type-B SF2 consensus and further specified as .0 to .2 because of the corresponding growing divercence). SF-specific phylogenetic trees and substitutions of the PRD are also shown (panels on the left). Each position was considered unambiguous if more than 50% of monomers had the same nucleotide at that position. The ambiguous positions were designated as n. V is A/C/G; B is C/G/T; M is A/C; R is A/G; W is A/T; S is C/G; Y is C/T; K is G/T; F is -/A; I is -/C; J is -/G.
Furthermore, each monomer was more similar to its own family consensus than to the consensus sequences of all other monomeric types, thus supporting subfamily classification (Supplementary Note-Section 3). The remaining monomers from 35 WGSS and 27 plasmid sequences not belonging to SF1-3 often intermingled with the SF3 units in the phylogenetic tree (black branches, Fig. 3) or, rather, composed the more ancestral cluster of sequences at the boundary between A- and B-type units. Phylogenetic trees using these sequences showed no clear clusterization of monomers (Supplementary Figure S2) and all were arranged in monomers by TRF and dot plot (Table S3). Indeed, although we can roughly distinguish A- and B-type monomers, they did not form any other HOR family as they showed a totally irregular organization (Fig. 3, Supplementary Figure S1).
We generated 16 new gorilla alphoid consensus sequences, specific for the three different SFs (two monomers for SF1; eight monomers for SF2; five monomers for SF3) and one for the monomeric arrays (gM1) (Table 1). By performing multiple sequence alignment and building p-distance matrices between the 2521 gorilla alphoid monomers and the 12 human alphoid consensus sequences, we revealed that gorilla SF1, SF2 and SF3 parallel human SF1, SF2 and SF3 (Table 2, Supplementary Note-Section 3) (see Methods section). Gorilla A-type monomers gJ1 (SF1) always correspond to human monomer type J1, while the gorilla B-type monomers of the same family have significantly diverged from the human counterpart (J2) (Fig. 4, Supplementary Note-Section 3). Conversely, the SF2 monomer composition revealed an extensive conservation of the B-type monomers. Gorilla SF3 is the less homogeneous SF; the five monomers very rarely form long pentameric stretches (tetramers are more common) (Supplementary Note-Section 3M).
With the correspondence between gorilla and human alphoid SFs ascertained, we gained more informative insight into the evolution of these sequences by hybridizing all the gorilla BAC clones previously described plus plasmids representative of all subgroups (1, 2, 3 and 4) on human (HSA) metaphases (79); then, an exemplificative subset (16) was hybridized on chimpanzee (PTR) and orangutan (PPY). In HSA, all centromeres hybridized with gorilla SF1 and SF2 probes were homologous to the gorilla SF1 and SF2 chromosomes, respectively. However, human chromosomes 4, 8 and 20 did not show this concordancy; the centromeres of these three chromosomes harboring human SF2 sequences showed signals using gorilla SF2 clones while in gorilla they hybridized with clones containing SF1 arrays, thus showing a different organization of these centromeres in human and gorilla. Moreover, due to the evolutionary translocation, gorilla centromeres V and XVII contain human centromeres 17 and 553,54. Despite the translocation, we observed signals on chromosome V using gorilla SF1 and chromosome XVII using SF2 clones in both gorilla and human. The only plasmidic clone containing SF3 sequences showed signals on chromosomes 17 and 11 (Tables 3 and S6). Furthermore, the “exceptional” clones hybridized randomly to human centromeres without maintaining the SF specificity (and hence showing a hybridization pattern similar to the gorilla clones from Group 3). Lastly, centromeric signals on human chromosomes 6 and 10 were not detected with any of the probes used (Fig. 3).
Gorilla SF1 sequences mostly mapped on PTR non-orthologously, while they did not hybridize at all on PPY chromosomes in either high or low stringency conditions. Gorilla SF2 clones, instead, highlighted all PPY chromosomes; although on PTR chromosomes very few signals were detected, mainly on chromosomes 9, 11 and 13 (XI, IX and IIq, respectively). The gorilla SF3 clone hybridized on IIp, XVII and X in PTR and (more faintly) in PPY (and on PTR I, VI and XI and PPY XXII) (Tables 3 and S7, Fig. 3).
Discussion
The centromeric α-satellite DNA in primates is typically composed of tandem repeats of a highly divergent 171 bp monomer repeat unit, with pairwise sequence identities of 60–80% within and between chromosomal subsets20.
At present, the only data available about the α-satellite in gorilla come from comparative hybridization experiments, in which traditionally human alphoid probes have been used. The most detailed study in this species has uniquely shown the extensive conservation of the X chromosome satellites46.
In this work we have explored the centromere DNA in gorilla, investigating more than 8 Mbp of α-satellite, representing roughly the 9% of the total centromeric DNA sequence available for this species.
Our data indicate that the gorilla α-satellite is organized in at least three different HOR subfamilies. The SF1 and SF3 mantain the human organization, while SF2 is represented by an unusual octameric HOR which is totally absent in humans and is likely present on several chromosomes in gorilla (Fig. 3). Gorilla HOR SFs, like in human19,20, show very high sequence similarities between adjacent multimeric units—up to 94% in SF1 and SF3 and up to 99% in SF2 HORs—thus displaying SF2 arrays in gorilla as more incisively homogenized.
FISH results revealed differences in extension and relative localization of HOR versus monomeric arrays. Gorilla SF1, SF2 and SF3 probes gave more intense and centromerically located signals than probes composed of monomeric arrays (Supplementary Figure S3). These data demonstrated that in gorilla, as in human, SF4 and SF5 arrays, are less abundant and localize at the borders of centromeres opposite to SF1-3 that instead create the bulk of centromeres19,20,26. Moreover, our data showed that gorilla centromeric sequence organization, as in human, is quite complex containing more than one type of array48.
In vitro studies proved homodimer CENP-B binds by each amino terminus to a CENP-B box sequence55,56. This creates a complex that contains two CENP-B polypeptides and two DNA molecules57, suggesting that the role of CENP-B in vivo might be able to assemble the higher-order structure of centromere satellite DNA arrays by juxtaposing pairs of CENP-B box sequences. Subsequently, CENP-B might have a role in the establishment of heterochromatin, thereby facilitating cohesion of sister chromatids around the centromere. Although the centromeric DNA is not conserved among species, the CENP-B–CENP-B box interaction is significantly conserved among mammals and may play an important role in the establishment of the specific structure of the kinetochore40. We have analyzed the long-range distribution of the protein-binding sites through α-satellite DNA in gorilla chromosomes and found both very high conservation of the CENP-B box essential positions and regular alternance between monomers. In particular, CENP-B- and pJα-binding monomers alternated perfectly in SF1, while showing B-B(*)-B*-A-B-A-B-A and B-B-B-A-A in SF2 and SF3, respectively (Fig. 4, for details).
Moreover, the frequency of conserved CENP-B boxes or pJα motifs was much lower in monomeric arrays than in HORs, as previously demonstrated in human21. Subsequently, diagnostic mutations specific to each of the SFs were detected, revealing a very high degree of homogenization (Supplementary Note-Section 3).
Hybridization patterns of gorilla centromeric DNA, together with previously published data and sequence analyses, let us conclude that SF1 sequences emerged in the GGO-PTR-HSA common ancestor. Indeed, there is no presence in orangutan47. Their localization in GGO, HSA and PTR indicates that they were created on chromosomes III, VII, X, XII and XIX and were subjected to extensive relocalizations in PTR (Fig. 3).
Clones made of gorilla SF2 sequences hybridized to all orangutan chromosomes, except for chromosomes VI and XII, all acrocentric chromosomes in gorilla and human and very few chimpanzee centromeres on chromosomes IIq, IX and XI. Human SF2 sequences, used as probes, were previously found on chromosomes IIq58 and IX48 in chimpanzee and on chromosome XVII in gorilla59, while pancentromeric hybridization had been concordantly observed in orangutan47. Gorilla SF3 sequences scarcely hybridized to orangutan metaphases and signals were detected in chimpanzee on chromosomes XI, XVII and X, in addition to few others, consistent with prior reports48.
The positive hybridizations obtained in PPY with gorilla SF2 and SF3 probes, might suggest the presence of these specific sequences in this species; nevertheless, since the high similarity of SF2 and especially SF3 arrays to the more ancient SF4 and SF5 monomers, our results might rather be produced by cross-hybridizitions with SF4 and SF5. Hence, before tracing evolutionary hypothesis involving SF2 and SF3 sequences, more focused studies are needed.
The conservation of both the sequence and location of SF2 between human and gorilla chromosomes was previously published47; however, our data prove for the first time that SF1 sequences are also present on homologous chromosomes in these two species, thus giving evidence of a higher similarity of alphoid DNA between human and gorilla than between human and chimpanzee. The only exceptions to the conservation of centromere-specificity between human and gorilla are the centromeres of the chromosomes IV, VIII and XX; all of these chromosomes underwent pericentric inversions during their evolutive history in great apes and this could most probably have affected the centromere evolution60,61,62,63.
Furthermore, human hybridization patterns show the exclusivity of either SF1 or SF2 sequences on human chromosomes, while in gorilla there are exceptions to this rule in regards to chromosomes V and XVII, which we found to contain sequences from both SF1 and SF2. An exchange of centromeric sequences might have happened following the translocation event that created the human-gorilla V and XVII chromosomes53,54.
We described two out of the eight gorilla SF2 monomer types containing a highly conserved insertion (sometimes present two or three times). This extra 22 bp sequence breaks the CENP-B box, duplicates 22 bp upstream and creates a new complete CENP-B box. These “scars” from the creation of “altered monomers” are similar to the mechanism proposed by Alexandrov et al. to explain the evolution of the S4-S5-S3 alphoid trimers in Chiropotes and Pithecia: they arose from an unequal crossover between two different monomers of an S3-S4 dimer that were shifted by 25 bp. As a result, S5 monomers are chimeric and contain a 25 bp duplication19. Both of these events gain interest in light of the assessed very high similarity between the CENP-B and the pogo and Tigger proteins coded by the pogo superfamily of transposable elements64. The human CENP-B and the transposases of the pogo type have highly similar primary sequences, a domain responsible for the coordination of the DNA cleavage during transposition and a DNA binding domain65.
The mechanism of transposition of these elements requires the approach of a pair of terminal inverted repeats (TIRs) to a protein-DNA complex and a following endonucleolytic cleavage and strand transfer. The elements Tigger1 and Tigger2 have an open reading frame similar to CENP-B and the TIRs motifs for Tigger2 are highly similar to the CENP-B box. It is then suggested that CENP-B derived from a transposase. This allowed us to speculate that the functional role of CENP-B at the primary constriction could not strictly be related to the centromere function or kinetochore assembly but rather to the modulation of evolution of the alphoid DNA by inducing recombination hotspots in the case where CENP-B had retained some transposase-associated activities. If this is the case, CENP-B would create nicks 10–20 bp upstream of the CENP-B box after having aligned two of them due to the dimerization of two CENP-Bs.
The role of a CENP-B-mediated transposition in the evolution of the centromere satellite could also possibly complement gene conversion and unequal exchange in solving the conundrum caused by evidence for both recombination and crossover suppression at centromeres31. Indeed, recombination events in α-satellites happen with a higher frequency 10–20 bp upstream of juxtaposed CENP-B boxes, which is actually possible when the CENP-B dimerizes64,66,67,68.
Our data provide new details into the organization and evolution of the centromere DNA in primates, giving direct evidence for the existence of higher-order alphoid SFs in gorilla centromere sequences. We show conserved distribution of SF1 and SF2 between human and gorilla and highlight an astonishing change specific to the gorilla lineage in the organization of SF2 sequences while maintaining a very high degree of sequence conservation. Indeed, SF2 in gorilla is uniquely octameric and periodically includes highly conserved 189 bp monomers. The discovery of these monomers allowed us to propose a role of CENP-B in centromere activity, closely related to the evolution of these sequences. Our model links the complexity of the centromeric sequence to its function and it traces future directions to study centromere functional properties.
Methods
Cell lines
Metaphase spreads and interphase nuclei were prepared from lymphoblastoid or fibroblast cell lines of Pan troglodytes (PTR) PTR5 (Bianka, Budapest zoo), Gorilla gorilla (GGO) (GGO5 ECACC CB1620) and Pongo pygmaeus (PPY) (Sinjo, Hamburg zoo). Human (HSA) metaphase spreads were prepared from Phytohemagglutinin-stimulated peripheral lymphocytes of normal donors by standard procedures. All metaphase spreads were obtained from female individuals.
FISH and Image Analysis
FISH experiments were performed using BAC clones and plasmids directly labeled by nick-translation with Cy3-dUTP as previously described69 with minor modifications. Hybridization was performed at 37 °C in 2x sodium chloride sodium citrate (SSC), 50% (v/v) formamide, 10% (w/v) dextran sulfate, 3 μg C0t-1 DNA and 3 mg sonicated salmon sperm DNA, in a volume of 10 μl. Post hybridization washing was at high stringency (60 °C in 0.1X SSC, three times) or at low stringency (37 °C in 2X SSC, 50% formamide, three times and 42 °C in 2X SSC, three times). Nuclei and chromosome metaphases were DAPI-stained. Digital images were obtained using a Leica epifluorescence microscope equipped with a cooled CCD camera. Fluorescence signals detected with Cy3 filters and chromosomes and nuclei images detected with DAPI filter were recorded separately as grayscale images. Pseudocoloring and merging of images were performed using Adobe Photoshop software.
Polymerase Chain Reaction (PCR) Labeling
DNA probes were directly labeled with Cy3-dUTP by PCR labeling; 200 ng of labeled probe was used for the FISH experiments. The use of PCR labeling avoids the possible contamination from genomic DNA by nick translation labeling of PCR products. PCR labeling was carried out in a final volume of 20 μl that contained 100 ng PCR product, 2.5 μl reaction buffer 10X, 2 μl MgCl2 50 mM, 0.5 μl each primer 10 μM, 0.5 μl dACG 2 mM, 2.5 Cy5-dUTP 1 mM, 5 μl BSA 2% and 0.3 μl Taq polymerase 5 U/μl.
Library screening
Library-hybridization was carried out according to the protocol available at CHORI BACPAC resources (http://bacpac.chori.org/highdensity.htm). The CH255 segment 1 represents a ~7.0-fold clone coverage library (http://bacpac.chori.org).
Alpha PCR
Gorilla genomic DNA were obtained from gorilla lymphoblastoid cell lines by standard methods70. α27 (CATCACAAAGAAGTTTCTGAGAATGCTTC) and α30 (TGCATTCAACTCACAGAGTTGAACCTTCC) primers were used to amplify genomic DNA by Polymerase Chain Reactions. They were obtained from the most conserved regions of human alphoid consensus50,51.
The PCR was performed as previously described14: 2 min initial denaturation at 94 °C, followed by 10 cycles of: 95 °C for 15s, 60 °C for 30s and 72 °C for 1 min; followed by 20 cycles of 94 °C for 15s, 58 °C for 30s and 72 °C for 1 min (20s more each cycle). Final extension was at 72 °C for 7 min (and then at hold 12 °C).
The reaction mixture consisted of 5 μl dNTPs (10X), 0.5 μl each primer (10 μM), 0.3 μl Platinum Taq DNA polymerase (5 U/μl), 1.5 μl MgCl2 (50 mM), 5 μl reaction buffer (Invitrogen) (10X), 3 μl of DNA template (50 ng/μl) and water up to 25 μl.
PCR products were analyzed by 1% agarose gel electrophoresis. They were labeled and used as a probe for FISH experiments on GGO metaphase spreads.
Cloning
PCR products were cloned in pCR-XL-TOPO using the standard protocol Topo cloning XL PCR kit (Invitrogen).
Southern Blot Analysis
Genomic DNAs from gorilla lymphoblastoid cell lines were prepared by standard procedures70. Endonuclease digestions were performed using a 4-fold excess of enzyme under the conditions suggested by the suppliers. Gel electrophoresis was performed in 1X tris-acetate (1X TAE 540 mM Tris-acetate, 1mMethylenediaminetetraacetic acid, EDTA). Genomic DNAs were run in a 0.8% agarose gel for 16–18 h, denatured and DNA transferred to Hybond membrane (Amersham), using as transfer buffer NaOH/NaCl (sodium chloride NaOH 0.25 M, sodium cloride NaCl 1.5 mM).
Clone inserts (50 ng) were labeled with 32P-dATP (3,000 Ci/mmol; Amersham) by using random oligomer priming. Filters were exposed and developed using storm imaging system.
Sequence and phylogenetic analyses
FASTA-formatted sequences were obtained corresponding to each gorilla α-satellite monomer and each sequence was analyzed by NCBI Blast2Sequences tool (http://blast.ncbi.nlm.nih.gov/bl2seq/wblast2.cgi) (BlastN program), aligning each first monomer with the entire sequence using default parameters (1 as reward for a match, −2 as penalty for a mismatch and 5 and 2 as open and extension gap penalties). Gorilla α-satellite sequences (WGSS) were searched by using plasmid sequences as queries in a BLAST search on Gorilla gorilla (taxid: 9593) NCBI databases. High amount of sequences was available thank to the recent gorilla genome sequencing project71. Multiple sequence alignments and consensus sequence extractions were performed using Clustal W72. Phylogenetic analyses were conducted in MEGA6 (Molecular Evolutionary Genetic Analysis, version 6.06)73. The evolutionary history was inferred using the Neighbor-Joining method74. The optimal tree with the sum of branch length = 77.65354306 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. A bootstrap test with 500 replicates and pairwise deletion parameters was conducted to evaluate the statistical significance of each node (Supplementary Figure S4). The evolutionary distances were computed using the p-distance method and are in the units of the number of base differences per site. The analysis involved 2521 nucleotide sequences. All ambiguous positions were removed for each sequence pair. There were a total of 299, 193, 259 and 174, positions in the final dataset for the entire collection of monomers, for SF1, SF2 and SF3 sequences, respectively.
Additional Information
Accession codes: GenBank accession numbers JQ685164.1-JQ685171.1, JQ685175.1-JQ685179.1, JQ685186.1-JQ685223.1.
How to cite this article: Catacchio, C. R. et al. Organization and evolution of Gorilla centromeric DNA from old strategies to new approaches. Sci. Rep. 5, 14189; doi: 10.1038/srep14189 (2015).
References
Rieder, C. L. & Salmon, E. D. Motile kinetochores and polar ejection forces dictate chromosome position on the vertebrate mitotic spindle. J Cell Biol 124, 223–233 (1994).
Sullivan, K. F., Hechenberger, M. & Masri, K. Human CENP-A contains a histone H3 related histone fold domain that is required for targeting to the centromere. J Cell Biol 127, 581–592 (1994).
Sullivan, B. A. & Karpen, G. H. Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat Struct Mol Biol 11, 1076–1083, 10.1038/nsmb845 (2004).
Masumoto, H., Masukata, H., Muro, Y., Nozaki, N. & Okazaki, T. A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J Cell Biol 109, 1963–1973 (1989).
Choo, K. H. Domain organization at the centromere and neocentromere. Dev Cell 1, 165–177 (2001).
Pluta, A. F., Mackay, A. M., Ainsztein, A. M., Goldberg, I. G. & Earnshaw, W. C. The centromere: hub of chromosomal activities. Science 270, 1591–1594 (1995).
Choo, K. H. in The Centromere (Oxford Univ. Press, New York) (1997).
Maio, J. J. DNA strand reassociation and polyribonucleotide binding in the African green monkey, Cercopithecus aethiops. Journal of molecular biology 56, 579–595 (1971).
Manuelidis, L. Chromosomal localization of complex and simple repeated human DNAs. Chromosoma 66, 23–32 (1978).
Vissel, B. & Choo, K. H. Human alpha satellite DNA–consensus sequence and conserved regions. Nucleic Acids Res 15, 6751–6752 (1987).
Alves, G., Seuanez, H. N. & Fanning, T. Alpha satellite DNA in neotropical primates (Platyrrhini). Chromosoma 103, 262–267 (1994).
Musich, P. R., Brown, F. L. & Maio, J. J. Highly repetitive component alpha and related alphoid DNAs in man and monkeys. Chromosoma 80, 331–348 (1980).
Willard, H. F. & Waye, J. S. Chromosome-specific subsets of human alpha satellite DNA: analysis of sequence divergence within and between chromosomal subsets and evidence for an ancestral pentameric repeat. J Mol Evol 25, 207–214 (1987).
Cellamare, A. et al. New insights into centromere organization and evolution from the white-cheeked gibbon and marmoset. Mol Biol Evol 26, 1889–1900, 10.1093/molbev/msp101 (2009).
Lee, H. R., Hayden, K. E. & Willard, H. F. Organization and molecular evolution of CENP-A–associated satellite DNA families in a basal primate genome. Genome Biol Evol 3, 1136–1149, 10.1093/gbe/evr083 (2011).
Waye, J. S. & Willard, H. F. Nucleotide sequence heterogeneity of alpha satellite repetitive DNA: a survey of alphoid sequences from different human chromosomes. Nucleic Acids Res 15, 7549–7569 (1987).
Romanova, L. Y. et al. Evidence for selection in evolution of alpha satellite DNA: the central role of CENP-B/pJ alpha binding region. J Mol Biol 261, 334–340, 10.1006/jmbi.1996.0466 (1996).
Warburton, P. E., Haaf, T., Gosden, J., Lawson, D. & Willard, H. F. Characterization of a chromosome-specific chimpanzee alpha satellite subset: evolutionary relationship to subsets on human chromosomes. Genomics 33, 220–228, 10.1006/geno.1996.0187 (1996).
Alexandrov, I., Kazakov, A., Tumeneva, I., Shepelev, V. & Yurov, Y. Alpha-satellite DNA of primates: old and new families. Chromosoma 110, 253–266 (2001).
Rudd, M. K. & Willard, H. F. Analysis of the centromeric regions of the human genome assembly. Trends Genet 20, 529–533, 10.1016/j.tig.2004.08.008 (2004).
Rosandic, M. et al. CENP-B box and pJalpha sequence distribution in human alpha satellite higher-order repeats (HOR). Chromosome Res 14, 735–753, doi: 10.1007/s10577-006-1078-x (2006).
Paar, V., Basar, I., Rosandic, M. & Gluncic, M. Consensus higher order repeats and frequency of string distributions in human genome. Curr Genomics 8, 93–111 (2007).
Rosandic, M., Gluncic, M., Paar, V. & Basar, I. The role of alphoid higher order repeats (HORs) in the centromere folding. J Theor Biol 254, 555–560, 10.1016/j.jtbi.2008.06.012 (2008).
Wevrick, R. & Willard, H. F. Long-range organization of tandem arrays of alpha satellite DNA at the centromeres of human chromosomes: high-frequency array-length polymorphism and meiotic stability. Proc Natl Acad Sci USA 86, 9394–9398 (1989).
Oakey, R. & Tyler-Smith, C. Y chromosome DNA haplotyping suggests that most European and Asian men are descended from one of two males. Genomics 7, 325–330 (1990).
Warburton, P. E. & Willard, H. F. Genomic analysis of sequence variation in tandemly repeated DNA. Evidence for localized homogeneous sequence domains within arrays of alpha-satellite DNA. Journal of molecular biology 216, 3–16 (1990).
Terada, S., Hirai, Y., Hirai, H. & Koga, A. Higher-order repeat structure in alpha satellite DNA is an attribute of hominoids rather than hominids. J Hum Genet 58, 752–754, 10.1038/jhg.2013.87 (2013).
Schindelhauer, D. & Schwarz, T. Evidence for a fast, intrachromosomal conversion mechanism from mapping of nucleotide variants within a homogeneous alpha-satellite DNA array. Genome Res. 12, 1815–1826 (2002).
Hayden, K. E. et al. Sequences associated with centromere competency in the human genome. Mol Cell Biol 33, 763–772, 10.1128/MCB.01198-12 (2013).
Dover, G. A., Strachan, T., Coen, E. S. & Brown, S. D. Molecular drive. Science 218, 1069 (1982).
Alkan, C., Eichler, E. E., Bailey, J. A., Sahinalp, S. C. & Tuzun, E. The role of unequal crossover in alpha-satellite DNA evolution: a computational analysis. J Comput Biol 11, 933–944 (2004).
Rudd, M. K., Wray, G. A. & Willard, H. F. The evolutionary dynamics of alpha-satellite. Genome Res 16, 88–96, 10.1101/gr.3810906 (2006).
Schueler, M. G., Higgins, A. W., Rudd, M. K., Gustashaw, K. & Willard, H. F. Genomic and genetic definition of a functional human centromere. Science 294, 109–115, 10.1126/science.1065042 (2001).
Iurov Iu, B., Mitkevich, S. P. & Aleksandrov, I. A. [Molecular cytogenetic research on the polymorphism of segments of the constitutive heterochromatin in human chromosomes]. Genetika 24, 356–365 (1988).
Alexandrov, I. A. et al. Definition of a new alpha satellite suprachromosomal family characterized by monomeric organization. Nucleic Acids Res 21, 2209–2215 (1993).
Wu, J. C. & Manuelidis, L. Sequence definition and organization of a human repeated DNA. J Mol Biol 142, 363–386 (1980).
Jorgensen, A. L., Bostock, C. J. & Bak, A. L. Chromosome-specific subfamilies within human alphoid repetitive DNA. J Mol Biol 187, 185–196 (1986).
Thompson, J. D., Sylvester, J. E., Gonzalez, I. L., Costanzi, C. C. & Gillespie, D. Definition of a second dimeric subfamily of human alpha satellite DNA. Nucleic Acids Res 17, 2769–2782 (1989).
Ikeno, M. et al. Construction of YAC-based mammalian artificial chromosomes. Nat Biotechnol 16, 431–439, 10.1038/nbt0598-431 (1998).
Yoda, K. et al. Centromere protein B of African green monkey cells: gene structure, cellular expression and centromeric localization. Mol Cell Biol 16, 5169–5177 (1996).
Muro, Y. et al. Centromere protein B assembles human centromeric alpha-satellite DNA at the 17-bp sequence, CENP-B box. J Cell Biol 116, 585–596 (1992).
Yoda, K., Kitagawa, K., Masumoto, H., Muro, Y. & Okazaki, T. A human centromere protein, CENP-B, has a DNA binding domain containing four potential alpha helices at the NH2 terminus, which is separable from dimerizing activity. J Cell Biol 119, 1413–1427 (1992).
Harrington, J. J., Van Bokkelen, G., Mays, R. W., Gustashaw, K. & Willard, H. F. Formation of de novo centromeres and construction of first-generation human artificial microchromosomes. Nat Genet 15, 345–355, 10.1038/ng0497-345 (1997).
Alkan, C. et al. Organization and evolution of primate centromeric DNA from whole-genome shotgun sequence data. PLoS computational biology 3, 1807–1818 (2007).
Haaf, T. & Willard, H. F. Chromosome-specific alpha-satellite DNA from the centromere of chimpanzee chromosome 4. Chromosoma 106, 226–232 (1997).
Durfy, S. J. & Willard, H. F. Concerted evolution of primate alpha satellite DNA. Evidence for an ancestral sequence shared by gorilla and human X chromosome alpha satellite. J Mol Biol 216, 555–566, doi: 10.1016/0022-2836(90)90383-W (1990).
Baldini, A. et al. Comparative mapping of a gorilla-derived alpha satellite DNA clone on great ape and human chromosomes. Chromosoma 101, 109–114 (1991).
Archidiacono, N. et al. Comparative mapping of human alphoid sequences in great apes using fluorescence in situ hybridization. Genomics 25, 477–484 (1995).
Samonte, R. V., Ramesh, K. H. & Verma, R. S. Comparative mapping of human alphoid satellite DNA repeat sequences in the great apes. Genetica 101, 97–104 (1997).
Waye, J. S. & Willard, H. F. Structure, organization and sequence of alpha satellite DNA from human chromosome 17: evidence for evolution by unequal crossing-over and an ancestral pentamer repeat shared with the human X chromosome. Mol Cell Biol 6, 3156–3165 (1986).
Choo, K. H., Vissel, B., Nagy, A., Earle, E. & Kalitsis, P. A survey of the genomic distribution of alpha satellite DNA on all the human chromosomes and derivation of a new consensus sequence. Nucleic Acids Res 19, 1179–1182 (1991).
Masumoto, H. et al. Properties of CENP-B and its target sequence in a satellite DNA (Springer-Verlag, 1993).
Yunis, J. J. & Prakash, O. The origin of man: a chromosomal pictorial legacy. Science 215, 1525–1530 (1982).
Stanyon, R. et al. Molecular and classical cytogenetic analyses demonstrate an apomorphic reciprocal chromosomal translocation in Gorilla gorilla. Am J Phys Anthropol 88, 245–250, 10.1002/ajpa.1330880210 (1992).
Murphy, W. J. et al. Dynamics of mammalian chromosome evolution inferred from multispecies comparative maps. Science 309, 613–617, 10.1126/science.1111387 (2005).
Wade, C. M. et al. Genome sequence, comparative analysis and population genetics of the domestic horse. Science 326, 865–867, 10.1126/science.1178158 (2009).
Reynolds, A. E., McCarroll, R. M., Newlon, C. S. & Fangman, W. L. Time of replication of ARS elements along yeast chromosome III. Mol Cell Biol 9, 4488–4494 (1989).
D'Aiuto, L., Antonacci, R., Marzella, R., Archidiacono, N. & Rocchi, M. Cloning and comparative mapping of a human chromosome 4-specific alpha satellite DNA sequence. Genomics 18, 230–235, 10.1006/geno.1993.1460 (1993).
Antonacci, R., Rocchi, M., Archidiacono, N. & Baldini, A. Ordered mapping of three alpha satellite DNA subsets on human chromosome 22. Chromosome Res 3, 124–127 (1995).
Marzella, R. et al. Molecular cytogenetic resources for chromosome 4 and comparative analysis of phylogenetic chromosome IV in great apes. Genomics 63, 307–313, 10.1006/geno.1999.6092 (2000).
Ventura, M. et al. Gorilla genome structural variation reveals evolutionary parallelisms with chimpanzee. Genome Res 21, 1640–1649, 10.1101/gr.124461.111 (2011).
Misceo, D. et al. Evolutionary history of chromosome 20. Mol Biol Evol 22, 360–366, 10.1093/molbev/msi021 (2005).
Stanyon, R. et al. Primate chromosome evolution: ancestral karyotypes, marker order and neocentromeres. Chromosome Res 16, 17–39, doi: 10.1007/s10577-007-1209-z (2008).
Smit, A. F. & Riggs, A. D. Tiggers and DNA transposon fossils in the human genome. Proc Natl Acad Sci USA 93, 1443–1448 (1996).
d'Alencon, E. et al. Characterization of a CENP-B homolog in the holocentric Lepidoptera Spodoptera frugiperda. Gene 485, 91–101, 10.1016/j.gene.2011.06.007 (2011).
Kipling, D. & Warburton, P. E. Centromeres, CENP-B and Tigger too. Trends Genet 13, 141–145 (1997).
Yoda, K., Ando, S., Okuda, A., Kikuchi, A. & Okazaki, T. In vitro assembly of the CENP-B/alpha-satellite DNA/core histone complex: CENP-B causes nucleosome positioning. Genes Cells 3, 533–548 (1998).
Tanaka, Y. et al. Crystal structure of the CENP-B protein-DNA complex: the DNA-binding domains of CENP-B induce kinks in the CENP-B box DNA. Embo J 20, 6612–6618, 10.1093/emboj/20.23.6612 (2001).
Lichter, P. et al. High-resolution mapping of human chromosome 11 by in situ hybridization with cosmid clones. Science 247, 64–69 (1990).
Maniatis, T, F. E. & Sambrook, J. Molecular Cloning: A Laboratory Manual. New York (NY): Cold Spring Harbor Laboratory, Cold Spring Harbor (1982).
Scally, A. et al. Insights into hominid evolution from the gorilla genome sequence. Nature 483, 169–175, 10.1038/nature10842 (2012).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22, 4673–4680 (1994).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30, 2725–2729, 10.1093/molbev/mst197 (2013).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4, 406–425 (1987).
Acknowledgements
We thank T. Brown for manuscript editing. This research was funded by Futuro in Ricerca 2010 RBFR103CE3
Author information
Authors and Affiliations
Contributions
M.V. and C.R.C. conceived and designed the experiments. C.R.C., R.R. and G.C. performed molecular biology experiments. R.R. and C.R.C. carried out sequence analysis.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Catacchio, C., Ragone, R., Chiatante, G. et al. Organization and evolution of Gorilla centromeric DNA from old strategies to new approaches. Sci Rep 5, 14189 (2015). https://doi.org/10.1038/srep14189
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14189
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.