Abstract
Plant secondary metabolites have been attracting people’s attention for centuries, due to their potentials; however, their production is still difficult and costly. The rich diversity of microbes and microbial genome sequence data provide unprecedented gene resources that enable to develop efficient artificial pathways in microorganisms. Here, by mimicking a natural pathway of plants using microbial genes, a new metabolic route was developed in E. coli for the synthesis of vanillin, the most widely used flavoring agent. A series of factors were systematically investigated for raising production, including efficiency and suitability of genes, gene dosage and culture media. The metabolically engineered strain produced 97.2 mg/L vanillin from l-tyrosine, 19.3 mg/L from glucose, 13.3 mg/L from xylose and 24.7 mg/L from glycerol. These results show that the metabolic route enables production of natural vanillin from low-cost substrates, suggesting that it is a good strategy to mimick natural pathways for artificial pathway design.
Similar content being viewed by others
Introduction
Vanillin (4-hydroxy-3-methoxybenzaldehyde) is one of the most widely used flavoring agents in the world. With an intensely and tenacious creamy vanilla-like odor, it is often used in foods, perfumes, beverages and pharmaceuticals1. As a plant secondary metabolite, natural vanillin is extracted from the seedpods of orchids (Vanilla planifolia, Vanilla tahitensis and Vanilla pompona). The annual worldwide consumption of vanillin exceeds 16,000 tons; however, owing to the slow growth of orchids and the low concentrations of vanillin in these plants (about 2% of the dry weight of cured vanilla beans), only about 0.25% of consumed vanillin originates from vanilla pods2. Most market demand is met by chemical synthesis of vanillin from lignin or fossil hydrocarbons, but the process is not environmentally friendly and it lacks substrate selectivity. Another drawback of chemical synthesis is that the synthetic, “unnatural” vanillin is sold for only $12 per kilogram, which is only 0.3% of the price of natural vanillin3. An alternative is the use of microbial biotechnology to produce vanillin from natural substrates; the resulting product is classified as natural vanillin under European and US food legislation4. Rising demand for natural products has led to the development of various biotechnological approaches for the production of vanillin. Phytochemicals such as ferulic acid are the major substrates used in natural vanillin production methods. Although different microorganisms capable of converting ferulic acid to vanillin have been isolated and studied in the past decade5,6,7, the high price of ferulic acid has limited its application.
Biosynthesis of vanillin from simple carbon sources like glucose is much more attractive because these sources are much cheaper and more readily available (glucose costs less than 30 cents per kilogram)8. Two artificial pathways have been developed for the production of vanillin from simple carbon sources. As shown in Supplementary Fig. 1, Frost et al. designed a pathway in recombinant Escherichia coli for the de novo biosynthesis of vanillic acid from glucose via the shikimic acid pathway; the vanillic acid was then enzymatically reduced to vanillin by aryl aldehyde dehydrogenase in vitro9. This route requires isolated dehydrogenase and costly cofactors and only trace amounts of vanillin can be detected. Hansen et al. demonstrated de novo biosynthesis of vanillin from glucose in Schizosaccharomyces pombe and Saccharomyces cerevisiae by a similar route, but they introduced an aromatic carboxylic acid reductase gene to avoid the extracellular reaction (Supplementary Fig. 1). In S. pombe and S. cerevisiae, vanillin production was 65 and 45 mg/L, respectively10. Brochado et al. expressed a glycosyltransferase in the vanillin-producing S. cerevisiae strain to reduce product toxicity and compared with the previous work, vanillin production was increased approximately fivefold in batch mode through in silico design11. However, the glycosylation step may have reduced the maximum theoretical yield12 and the use of dehydroshikimic acid may limit the aromatic amino acid biosynthesis pathway. Moreover, many alcohol dehydrogenases in yeast can act on vanillin and the precursor protocatechuic aldehyde, leading to loss of product. Recently, Kunjapur et al. used an E. coli with reduced aromatic aldehyde reduction as a platform for aromatic aldehyde biosynthesis. After the introduction of a pathway for the biosynthesis of vanillin, 119 ± 3 mg/L vanillin can be produced from glucose13.
The natural pathway for vanillin production that has evolved in plants could be much more efficient14,15. Mimicking and assembling the natural pathway in E. coli to synthesize vanillin from glucose may overcome the shortcomings of previous works. Based on research into construction of the phenylpropanoid pathway in microorganisms16,17,18,19, a simulated natural pathway including five enzymes was introduced into E. coli to synthesize vanillin from simple carbon sources. Compared with previous pathways, this new metabolic route converted a greater variety of substrates to vanillin, including l-tyrosine and glucose. Xylose (an important pentose component of renewable biomass feedstocks) and glycerol (an inexpensive byproduct of the biodiesel industry) were also used for the production of vanillin. To optimize vanillin production, we investigated the efficiency and suitability of biosynthetic genes from different sources, gene dosage effects and different culture media.
Results
Pathway design and construction
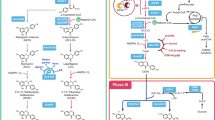
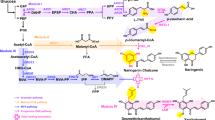
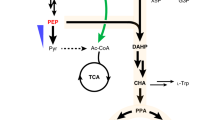
We designed the artificial vanillin biosynthetic pathway shown in Fig. 1. With five enzymes, including tyrosine ammonia-lyase (TAL; EC 4.3.1.23), 4-coumarate 3-hydroxylase (C3H; EC 1.14.14.9), caffeate O-methyltransferase (COMT; EC 2.1.1.68), trans-feruloyl-CoA synthetase (FCS; EC 6.2.1.34) and enoyl-CoA hydratase/aldolase (ECH; EC 4.2.1.17), l-tyrosine could be converted to vanillin. Grafting this artificial pathway into a tyrosine-overproducing E. coli strain would enable production of vanillin from simple carbon sources. To accomplish this, we constructed a single-knockout E. coli strain harboring three plasmids (Fig. 2).
Artificial vanillin biosynthetic pathway constructed in Escherichia coli cells.
The purple part shows the reconstruction of the common aromatic pathway for l-tyrosine overproduction; the global regulatory protein TyrR has been inactivated and four enzymes are overexpressed. As shown in the pale blue part, three enzymes are involved in the artificial pathway from tyrosine to ferulic acid. Black part in the right box shows the three steps from ferulic acid to vanillin, which are catalyzed by two enzymes. TyrR, a regulatory protein; PEPS, phosphoenolpyruvate synthetase; TKT, transketolase; fbr-DAHPS, fbr-3-deoxy-d-arabino-heptulosonate-7-phosphate synthase; fbr-CM/PDH, fbr-chorismate mutase/prephenate dehydrogenase; TAL, tyrosine ammonia-lyase; C3H, 4-coumarate 3-hydroxylase; COMT, caffeate O-methyltransferase; FCS, trans-feruloyl-CoA synthetase; ECH, enoyl-CoA hydratase/aldolase. The orchid was drawn by Jun Ni.
Schematic representation of artificial gene clusters used for vanillin production.
A T7 promoter and ribosomal binding site precede each gene and a T7 terminator is located downstream of each gene cluster. Module one contains the biosynthetic gene cluster used for ferulic acid production. Module two contains the gene cluster used for production of vanillin from ferulic acid. Module three was used for l-tyrosine overproduction. Cmr, chloromycetin resistance; Ampr, ampicillin resistance; Kanr, kanamycin resistance.
Tyrosine ammonia-lyase (TAL) catalyzes non-oxidative deamination of l-tyrosine to 4-coumaric acid and many microbial TALs and their corresponding genes have been reported in recent years20, including the sam8 gene from the actinomycete Saccharothrix espanaensis21 and the RsTAL gene from the photosynthetic bacteriuma Rhodobacter sphaeroides22. 4-Coumarate 3-hydroxylase is a plant-specific cytochrome P450-dependent monooxygenase that converts 4-coumaric acid to caffeic acid by hydroxylation at the 3-position of the benzene ring. Because of the membrane-bound property and the instability of C3H, functional expression of this protein in bacterial systems is very challenging23. Recently, two microbial C3Hs from S. espanaensis and E. coli (encoded by sam5 and hpaBC, respectively) were reported and their functions were characterized21,24. Caffeate O-methyltransferase can convert caffeic acid to ferulic acid25 and the comt gene from Arabidopsis thaliana was codon-optimized and synthesized for this purpose.
The vanillin synthase from Vanilla planifolia can directly convert ferulic acid to vanillin15; however, we failed to express it in E. coli. Instead, the fcs and ech genes (encoding FCS and ECH, respectively) were recently isolated from Streptomyces sp. strain V-1 and their functions were characterized in E. coli26. For conversion of ferulic acid to vanillin, we constructed a plasmid, fcs-ech/pET, bearing both of these genes. The next step was to introduce our artificial biosynthetic pathway into a tyrosine-overproducing strain. An E. coli strain carrying the plasmid tyrAfbr-aroGfbr-tktA-ppsA/pCOLA and a knockout mutation in tyrR fit our requirements27. Inactivation of TyrR-mediated regulation improves the phenotype of aromatic amino acid producers, which leads to an overflow of l-tyrosine biosynthesis. The plasmid expresses phosphoenolpyruvate synthetase (PEPS: ppsA), transketolase (TKT: tktA), feedback-inhibition-resistant (fbr) 3-deoxy-d-arabino-heptulosonate-7-phosphate synthase (fbr-DAHPS: aroGfbr) and fbr-chorismate mutase/prephenate dehydrogenase (fbr-CM/PDH: tyrAfbr). Higher expression of PEPS and TKT can increase carbon flow into the aromatic amino acid biosynthesis pathway and fbr-DAHPS and fbr-CM/PDH can depress inhibition by metabolites.
Thus, our artificial vanillin biosynthetic system, inspired by naturally occurring biosynthetic pathways, comprises an E. coli strain with a knockout mutation, four overexpressed genes and five exogenous genes. With this system, many simple carbon sources, such as l-tyrosine, glucose, xylose and glycerol, may easily convert into vanillin.
Comparison of two recombinant plasmids for ferulic acid production
Previous studies have found that some enzymes involved in the phenylpropanoid pathway are ineffective in recombinant E. coli17,28. Therefore, we compared key biosynthetic genes from different sources to obtain high-yield production of ferulic acid. To evaluate the performance of the genes from gram-positive bacteria (sam8 and sam5) and gram-negative bacteria (RsTAL and hpaBC), sam8/pACYC and tal/pACYC were constructed first and the production of 4-coumaric acid from l-tyrosine by E. coli carrying these plasmids was investigated. In a previous study, a large amount of insoluble TAL was found when a recombinant strain was cultured at 37 °C17; therefore, we cultured the transformed E. coli at 26 °C to reduce the formation of inclusion bodies. The recombinant strains were cultured in Luria-Bertani (LB) medium supplemented with an additional 2 g/L l-tyrosine and 0.2 mM IPTG. After 48 h, high-performance liquid chromatography (HPLC) and liquid chromatography–mass spectrometry (LC–MS) analysis showed that the concentration of 4-coumaric acid reached 875 mg/L and 821 mg/L in cultures of the recombinant strains containing sam8/pACYC and tal/pACYC, respectively (Table 1). There was no significant difference in 4-coumaric acid production between the two recombinant strains. Therefore, to future study the 4-coumaric acid conversion capacity of recombinant strains with genes from different sources, we introduced the sam5 gene into sam8/pACYC and the hpaBC gene into tal/pACYC. The recombinant strains carrying sam8-sam5/pACYC and tal-hpaBC/pACYC were cultured in the same medium and under the same conditions described above and the production of caffeic acid reached 136 mg/L and 44 mg/L, respectively, after 48 h of culture (Table 1). Because actinomycete genes might contain rarely used codons and alterations of mRNA structural elements they are often difficult to express in E. coli29. Nevertheless, the strain with sam8-sam5/pACYC produced 209% more caffeic acid than the strain with tal-hpaBC/pACYC. This may be caused by a side effect of the enzyme encoded by hpaBC, which can catalyze the conversion of tyrosine to l-DOPA and reduce the production of 4-coumaric acid30.
With the addition of the comt gene to the plasmids, the production of ferulic acid reached 156 mg/L (strain VT-1; sam8-sam5-comt/pACYC) and 43 mg/L (strain VT-2; tal-hpaBC-comt/pACYC) after 48 h of culture in the same medium and under the same conditions described above (Fig. 3a). The results indicated that ferulic acid production was more efficient with sam8-sam5-comt/pACYC than with tal-hpaBC-comt/pACYC and plasmid sam8-sam5-comt/pACYC was therefore chosen for further research.
Fermentative production of ferulic acid and vanillin.
The arrow indicates the time of addition of IPTG (0.2 mM). Data are representative of at least three independent experiments and the error bars indicate the standard deviation. (a) For production of ferulic acid, a recombinant strain harboring sam8-sam5-comt/pACYC (red columns) or tal-hpaBC-comt/pACYC (dark cyan columns) was grown in Luria-Bertani (LB) medium with an additional 2 g/L l-tyrosine. Cell growth (lines; colors the same as those for ferulic acid production) is presented as the optical density at 600 nm. (b) Time course of vanillin production from ferulic acid in E. coli VT-3 cultures. (c) Production of vanillin from l-tyrosine using recombinant strains VT-4 and VT-5. (d) Fermentative production of vanillin from glucose. The control strain is E. coli strain K12 harboring sam8-sam5-comt/pACYC and fcs-ech/pET.
The yields of 4-coumaric acid, caffeic acid and ferulic acid were measured by HPLC. These products were also identified using LC–MS and compared with corresponding standards (Fig. 4). During all the experiments mentioned above, no 4-coumaric acid, caffeic acid, or ferulic acid was detected in the culture medium of wild-type E. coli carrying the control (blank) vector.
Mass spectrometry analysis of reaction products.
Electrospray ionization mass spectra were obtained after liquid chromatography–mass spectrometry analysis of the culture medium of recombinant strains and of authentic standards; the corresponding mass spectra are shown in this figure. Negative ion data for the standard compounds were as follows: p-coumaric acid, m/z = 163.0401; caffeic acid, m/z = 179.0350; ferulic acid, m/z = 193.0506; resveratrol, m/z = 227.0714; naringenin, m/z = 271.0612; bisdemethoxycurcumin, m/z = 307.0976 and vanillin, m/z = 151.0415. (a) Sample from recombinant strain harboring sam8/pACYC (upper panel) and authentic 4-coumaric acid standard (lower panel). (b) Sample from recombinant strain harboring sam8-sam5/pACYC (upper panel) and authentic caffeic acid standard (lower panel). (c) Sample from recombinant strain harboring sam8-sam5-comt/pACYC (upper panel) and authentic ferulic acid standard (lower panel). (d) Sample from recombinant strain harboring fcs-ech/pET (upper panel) and authentic vanillin standard (lower panel).
Production of vanillin from tyrosine by recombinant E. coli
Vanillin production from recombinant E. coli harboring fcs-ech/pET (VT-3) was investigated using LB medium containing 1 g/L ferulic acid as substrate and 0.2 mM IPTG as inducer. In HPLC analysis, the retention time of the major product was identical to that of authentic vanillin; we further analyzed the compound by HPLC coupled to MS in the negative-ion mode and observed a molecular ion of m/z 151.04, indicating that this compound was indeed vanillin (Fig. 4). After 36 h of culture at 26 °C, the concentration of vanillin in the medium was 692 mg/L (Fig. 3b) and the corresponding molar conversion rate was 88.3%, which is high compared with previous studies6. This finding suggests that the strain harboring fcs-ech/pET can convert ferulic acid to vanillin with high efficiency and the plasmid is a very good choice for our artificial vanillin biosynthetic system.
Based on these results, we combined the ferulic acid-producing pathway described above with the pathway for conversion of ferulic acid to vanillin. Thus, production of vanillin from tyrosine was carried out with strain VT-4 (recombinant E. coli harboring sam8-sam5-comt/pACYC and fcs-ech/pET). Strain VT-4 was grown in LB medium containing 2 g/L l-tyrosine and 0.2 mM IPTG and 97.2 mg/L vanillin was obtained after 48 h of culture at 26 °C (Fig. 3c). The molar conversion of l-tyrosine to vanillin was 4.8%, which is relatively low. To determine the cause of the low conversion rate, we first examined the degradation of vanillin in the culture medium. Vanillin was added to a culture of strain VT-4 at a concentration close to the production yield. As shown in Supplementary Fig. 2, though vanillin was relatively stable in the culture medium, the degradation of a small amount of vanillin (less than 15% after 24 h) was one of the causes of the low molar conversion rate. To further investigate the low yield of vanillin from tyrosine, we examined the expression levels of the biosynthetic genes (sam8, sam5, comt, fcs and ech) and the amounts of intermediate compounds. Quantitative reverse transcription PCR (RT-PCR) results indicated that all biosynthetic genes were expressed (Fig. 5). The genes in pACYCDuet-1 (sam8, sam5 and comt) had lower expression levels than the genes in pETDuet-1 (fcs and ech) and sam8 and comt had higher expression levels than sam5. l-Tyrosine and intermediate compounds (caffeic acid and ferulic acid) were not detected in the culture medium and about 29.2 mg/L 4-coumaric acid was detected. Besides, a small amount of vanillyl alcohol (about 35.7 mg/L) could be detected in the culture of recombinant strain VT-4. This is in accordance with the results that some aromatic aldehyde reductases in E. coli could convert vanillin to vanillyl alcohol13. Thus, the low conversion rate of tyrosine to vanillin may be due to the endogenous side-reactions and inefficient conversion of 4-coumaric acid to caffeic acid. Plasmid fcs-ech-sam5/pET, with an additional sam5 gene, was constructed to replace plasmid fcs-ech/pET. As shown in Fig. S3, although RT-PCR results indicated that the expression level of sam5 gene was improved in VT-5 (recombinant E. coli harboring sam8-sam5-comt/pACYC and fcs-ech-sam5/pET); the production of vanillin from VT-5 was similar to that from VT-4 (Fig. 3c). Furthermore, a small amount of 4-coumaric acid still remained in the culture of VT-4 (about 21.4 mg/L), indicating that the enzyme activity of C3H may be a critical factor for vanillin production in our system.
Relative transcription levels of vanillin biosynthetic pathway genes.
Total RNA of recombinant strains VT-4 was extracted from glucose-based LB medium 8 h after induction with IPTG and transcription levels were determined by quantitative RT-PCR. All values are relative to the expression level of the sam8 gene, which was set at 1. Dark cyan and purple columns indicate the expression levels of the genes involved in the artificial pathways from tyrosine to ferulic acid and from ferulic acid to vanillin, respectively. Results are presented as the average of three repetitions from independent total RNA samples.
Vanillin production from simple carbon sources
The recombinant E. coli strain containing the plasmid tyrAfbr-aroGfbr-tktA-ppsA/pCOLA and a knockout mutation in tyrR has been tested for its ability to produce tyrosine27. When 10 g/L glucose was used as a carbon source, the engineered E. coli strain produced l-tyrosine in LB medium at a titer of about 1.1 g/L after 24 h and the corresponding molar conversion rate was 10.9%. In contrast, the wild-type E. coli strain produced low quantities of l-tyrosine.
The artificial vanillin biosynthetic pathway from l-tyrosine was constructed in the tyrosine-overproducing E. coli strain by transforming cells with two plasmids (sam8-sam5-comt/pACYC and fcs-ech/pET). Vanillin was produced from 10 g/L glucose at a titer of 19.3 mg/L and the conversion rate was 0.193%. Without glucose, the yield of vanillin was below the detection limit. Moreover, the production of vanillin was quite low in the wild-type E. coli strain containing the biosynthetic pathway from tyrosine to vanillin (Fig. 3d). These results indicate that vanillin was derived from glucose and that metabolic engineering of the aromatic amino acid pathway was necessary for the production of vanillin from a simple carbon source.
To reduce costs, additional experiments were performed with M9 minimal medium and fermentation medium supplemented with 10 g/L glucose. As shown in Fig. 6a, vanillin production of the recombinant E. coli strain in fermentation medium and M9 minimal medium was 15.7 mg/L and 6.4 mg/L, respectively. Compared with LB medium and fermentation medium, vanillin production was much lower in M9 minimal medium; this may have been due to the lower cell density (Fig. 6b), or the M9 minimal medium may have lacked metal cofactors needed for heterologous enzyme activity. Although vanillin production in fermentation medium was slightly lower than production in LB medium, the specific titer (vanillin titer divided by cell density) was roughly 1.5 times higher in fermentation medium (reaching 4.9 mg/L per OD unit) than in LB medium. This was probably the result of more of the carbon source in the LB medium consumed for cell growth.
Optimization of vanillin production from simple carbon sources.
Samples were collected 60 h after induction with IPTG and analyzed by HPLC. Data are representative of three independent experiments. (a) Production of vanillin by engineered strain VT-6 grown in different media with added glucose (blue column), xylose (brown column), or glycerol (green column). (b) Final cell density of engineered strain VT-6 grown in different media. (c) Effect of IPTG concentration on the production of vanillin. (d) Effect of glycerol concentration on the production of vanillin.
Vanillin production from xylose and glycerol was also investigated. When 10 g/L xylose or 10 g/L glycerol was used as carbon source, the engineered E. coli strain produced vanillin in LB medium at a yield of 13.3 mg/L or 24.1 mg/L, respectively (Fig. 6a). Vanillin production was higher with glucose, a better carbon source for E. coli metabolism, than with xylose. On the other hand, the vanillin titer was lower with glucose than with glycerol. This is consistent with previous studies that found that glycerol is a more suitable carbon source than glucose for the production of shikimic acid and other compounds related to the shikimic acid pathway27,31. The recombinant E. coli strain was also cultivated in M9 minimal medium and fermentation medium with xylose or glycerol as substrate. As shown in Fig. 6a, the results were similar to those obtained when glucose was provided as the carbon source: the highest specific titer was achieved in fermentation medium and the lowest production was achieved in M9 minimal medium. These results indicate that glycerol is the best carbon source for vanillin production among the three substrates and fermentation medium is the most cost-effective medium owing to the high specific titer and low cost.
To further optimize the culture conditions for the recombinant E. coli strain, glycerol was chosen as the substrate and fermentation medium was chosen as the preferred production medium. The highest vanillin production (24.7 mg/L) was observed when 0.1 mM IPTG was added to the medium (Fig. 6c). Higher concentrations of IPTG reduced the vanillin titer and only 11.3 mg/L vanillin was detected in the medium when 1 mM IPTG was added; this may have been caused by a metabolic burden on the E. coli host. The “leaky” expression of biosynthetic genes without IPTG induction led to the production of vanillin at a yield of 18.1 mg/L; although a higher yield was achieved with 0.1 mM IPTG, the elimination of IPTG can reduce production costs. As shown in Fig. 6d, vanillin production in medium with 15 g/L, 20 g/L, or 25 g/L glycerol was 24.5 mg/L, 25.3 mg/L and 25.9 mg/L, respectively; vanillin production in medium with 10 g/L glycerol was not significantly different. However, the production of vanillin decreased to 14.3 mg/L when 5 g/L glycerol was used as substrate. These results show that 10 g/L glycerol is sufficient for vanillin production.
Discussion
Synthetic biology is the design and construction of biological devices and systems for useful purposes32. Demand for natural plant metabolites is increasing and synthetic biology has been widely used for the production of these compounds by microorganisms. However, the rational design of feasible pathways is one of the major challenges in this field and artificial pathways have not always performed ideally in the host, often leading to low production33. Furthermore, an artificial pathway may use only one substrate, such as eugenol to vanillin by the biotransformation, which limits its application5. Simulating and assembling natural pathways in the host may counter these problems because natural pathways are the result of evolution over a long period and seem to be more efficient and stable. More importantly, natural synthetic pathways are often connected with basal metabolism34. This connection would enable the production of desired product from various available substrates and the pathways can be easily transplanted to other production platforms. Many plant genes are quite challenging to functionally express in prokaryotic systems; therefore, for successful expression of mimicked natural pathways in microorganisms, microbial genes should be chosen instead of plant genes whenever possible. With the development of genomics and bioinformatics, more and more genes from microorganisms are becoming available for the construction of mimicked pathways.
In this study we succeeded in introducing a mimicked metabolic route for vanillin production into E. coli; with this recombinant strain, tyrosine, glucose, xylose and glycerol can be used as substrate to produce vanillin. This is the first report of vanillin production from tyrosine by microbes and the first attempt to genetically engineer a single recombinant prokaryote for de novo biosynthesis of vanillin. Compared with previously developed artificial pathways for the production of vanillin from simple carbon sources9,10,11, the simulated natural pathway presented here has several advantages. One advantage is that the pathway can use l-tyrosine as substrate, which has less influence on the basal metabolism of the host; it is quite different from the previous artificial vanillin synthetic pathway that limited the biosynthesis of aromatic amino acids. Another advantage is that the simulated pathway can be transplanted to other tyrosine-overproducing strains to improve vanillin production35,36 and most of the genetic modification to improve the yield of tyrosine, such as co-expression of the rate-limiting enzymes shikimate kinase and quinate/shikimate dehydrogenase, also plays a role in increasing the yield of vanillin37. Moreover, expensive precursor chemicals or carbohydrate feedstocks are the main cost of microbial industry; with simulated natural pathways connected to primary metabolism, cheaper and more readily available carbon sources can be used for the production of desired compounds.
Although natural pathway mimicking has many advantages for the production of natural compounds, the yield of vanillin in our system was not very high. Analysis of the low conversion rate of tyrosine to vanillin indicated that it may have been due to the inefficient conversion of 4-coumaric acid to caffeic acid, endogenous side-reactions and, to a lesser extent, the instability of vanillin. In the future, the use of previous strategy to delete aldo-keto reductases and alcohol dehydrogenases may improve the conversion rate13. Furthermore, it will be necessary to look for more efficient enzymes that relate to this biosynthetic route and to C3H in particular. Because the artificial vanillin biosynthetic pathway will consume ATP and NADPH21,26, another strategy for improving the yield of vanillin is to balance NADP+ and ATP. There are various ways to achieve this; for example, to regenerate NADPH, which is the coenzyme of C3H, an NADPH-regenerating enzyme (glucose-6-phosphate dehydrogenase or phosphite dehydrogenase) should be introduced into the system38,39. Many methods of synthetic biology and fermentation technology, such as fed-batch fermentation or the use of bioreactors to increase cell density, may further boost the vanillin titer16. According to our recent research, choosing a suitable promoter is also an effective way to obtain high yields40.
To reduce production costs, we optimized the culture media, the concentration of IPTG and the concentration of substrate. The results will guide the large-scale production of vanillin with our system. Besides glucose, xylose and glycerol, many other inexpensive carbon sources, such as molasses, wheat flour and rice bran, have been found to support microbial growth and enzyme production41. Combining a de novo vanillin biosynthetic pathway with these agro-industrial byproducts will offer a cost-effective process for vanillin production and may have remarkable economic benefits. Lignocellulosic biomass, the most abundant raw material on Earth, contains a lot of reducing sugars and aromatic compounds42. The products of its decomposition, including xylose, ferulic acid and caffeic acid, can be used as substrates for vanillin production with our system. Here we have provided new perspectives and ideas for the simultaneous and effective utilization of complex components.
Recently, more and more plant metabolites were identified and proved with potential bioactivities, their metabolic pathways and key enzymes have also been clarified41. The rich diversity of microbes and explosive growth of microbial genome sequence data provide unprecedented gene resources that enable to mimick efficient pathways in microorganisms. Moreover, the rising demand of high-value plant metabolites will promote the use of this method. In conclusion, we have designed a simulated natural pathway to biosynthesize vanillin, making it possible to use microbes to produce vanillin from inexpensive carbon sources. This is a successful example of mimicking natural pathway for de novo biosynthesis by using cheap carbon sources for the efficient production of valuable plant metabolites.
Methods
Chemicals, bacterial strains and plasmids
All chemicals were purchased from Sigma-Aldrich (St. Louis, MO) unless otherwise specified. Restriction enzymes, ligase (New England Biolabs Inc.) and DNA polymerase (Takara Biochemicals Inc.) were used for cloning and plasmid construction. Oligonucleotides were synthesized by Sangon Biotech Co. (Shanghai, China). The characteristics of the bacterial strains and plasmids used in this study are provided in Table 2. Saccharothrix espanaensis (DSM 44229) and R. sphaeroides (DSM 158) were obtained from DSMZ. E. coli DH5α and E. coli BL21 (DE3) were used for general cloning and expression of biosynthetic genes in feeding experiments, respectively. The l-tyrosine- overproducing strain containing a knockout mutation in tyrR and a plasmid expressing the aroGfbr, tyrAfbr, ppsA and tktA genes was a gift from Professor Minami and was used for shake flask experiments. A pEASY-Blunt cloning vector (Transgen, China) was used for subcloning of genes. Plasmids pACYCDuet-1 and pETDuet-1 were purchased from Novagen (San Diego, CA) and used for gene overexpression in E. coli.
Bacterial cultivation conditions
Streptomyces sp. strain V-1 (CCTCC M 206065) was cultivated at 30 °C in seed medium, which contained 10 g/L glucose, 5 g/L yeast extract, 10 g/L peptone, 5 g/L beef extract and 2 g/L NaCl (pH 7.0), as previously described33. Escherichia coli cells used for gene cloning, plasmid propagation and inoculum preparation were cultured at 37 °C in LB medium supplemented with appropriate antibiotics. The working concentrations of antibiotics were 100 μg/mL for ampicillin, 50 μg/mL for kanamycin and 20 μg/mL for chloromycetin. For production of 4-coumaric acid, caffeic acid, ferulic acid and vanillin from tyrosine, 500 μL of overnight LB culture was inoculated into 50 mL of LB medium with 2 g/L l-tyrosine and grown at 37 °C. After the OD600 reached 0.5–0.6, IPTG was added to the cultures to a final concentration of 0.2 mM and cultures were transferred to a gyratory shaker at 26 °C for 3 days. For de novo biosynthesis of vanillin, 500 μL of overnight LB culture was inoculated into 50 mL of LB, M9, or fermentation medium supplemented with 10 g/L glucose, xylose, or glycerol; the culture conditions were the same as those used for vanillin production from tyrosine. The fermentation medium was modified M9 minimal salt medium containing 1 g NH4Cl, 6 g Na2HPO4, 3 g KH2PO4, 0.5 g NaCl, 2 mmol MgSO4·7H2O, 0.1 mmol CaCl2·2H2O and 0.5 g yeast extract per liter. Trace elements (0.03 mg/L H3BO3, 1 mg/L thiamine, 0.94 mg/L ZnCl2, 0.5 mg/L CoCl2, 0.38 mg/L CuCl2, 1.6 mg/L MnCl2 and 3.6 mg/L FeCl2) were added to the LB and fermentation media. Samples were collected at intervals of 6 or 12 h and analyzed by HPLC and LC–MS.
Heterologous pathway construction and assembly
Genomic DNA was extracted from R. sphaeroides, E. coli, S. espanaensis and Streptomyces sp. V-1 using a bacterial genomic DNA extraction kit (QIAGEN, Hilden, Germany). Lists of primers used in this study can be found in Tables S1 and S2. The genes tal (GenBank accession No. CP000144.2) and sam8 (GenBank accession No. DQ357071) were amplified by high-fidelity PCR from the genomic DNA of R. sphaeroides and S. espanaensis, respectively. The resulting PCR products were cloned into the NcoI and EcoRI sites of pACYCDuet-1, resulting in tal/pACYC and sam8/pACYC. The genes hpaBC (GenBank accession No. CP001509) and sam5 (GenBank accession No. DQ357071) were amplified from the genomic DNA of E. coli and S. espanaensis, respectively and the NdeI–XhoI fragment of the resulting products was cloned into tal/pACYC and sam8/pACYC, generating tal-hpaBC/pACYC and sam8-sam5/pACYC, respectively. The comt gene from A. thaliana (GenBank accession No. NM124796) was codon-optimized, fused with a T7 promoter and synthesized by recombinant PCR43; the primers used for recombinant PCR are listed in Table S2. After digestion with SacI and NotI, comt was cloned into tal-hpaBC/pACYC and sam8-sam5/pACYC to generate tal-hpaBC-comt/pACYC and sam8-sam5-comt/pACYC. The genes fcs (GenBank accession No. KC847405) and ech (GenBank accession No. KC847406) were amplified by PCR from chromosomal DNA of Streptomyces sp. V-1. The ech and fcs genes were ligated into the NcoI and HindIII sites of Multiple Cloning Site (MCS) II and the NdeI and XhoI sites of MCS I of pETDuet-1, respectively, to construct fcs-ech/pET. With the primers HindIII-T7-F and NotI-sam5-R, the sam5 gene with the T7 promoter was amplified from sam8-sam5/pACYC and cloned into fcs-ech/pET, generating fcs-ech-sam5/pET. Gene sequences and orientations were confirmed by nucleotide sequencing after each round of cloning. The maps for plasmids sam8-sam5-comt/pACYC and fcs-ech/pET are shown in Fig. 2. To construct the strains producing 4-coumaric acid, caffeic acid and ferulic acid, E. coli BL21 (DE3) was transformed with sam8/pACYC, sam8-sam5/pACYC and sam8-sam5-comt/pACYC, respectively. To construct the vanillin-producing strains, sam8-sam5-comt/pACYC and fcs-ech/pET (or fcs-ech-sam5/pET) were co-transformed into E. coli BL21 (DE3) or the l-tyrosine-overproducing strain.
Quantitative RT-PCR analysis of synthetic pathway genes
The recombinant E. coli strain harboring sam8-sam5-comt/pACYC and fcs-ech/pET was grown in LB medium at 37 °C and 0.2 mM IPTG was added to the culture when the OD600 reached 0.6. Cells were cultured on a gyratory shaker at 26 °C for 8 h and then collected for total RNA extraction using an RNAprep Pure Cell/Bacteria Kit (Tiangen Biotech Co., Beijing, China). RNA was quantified using a NanoVue spectrophotometer (GE Healthcare Bio-Sciences, Sweden). After removal of genomic and plasmid DNA from RNA preparations using DNase I (Thermo Scientific), a total of 2 μg of RNA was used in reverse transcription reactions with random primers and SuperScript III Reverse Transcriptase (Invitrogen, Shanghai, China). Meanwhile, the RNA sample was used as a temple and the PCR reaction was performed to certify there is no plasmid DNA in the RNA sample. Relative RNA concentrations were determined by quantitative RT-PCR using a 7300 Real-time PCR system with RealMasterMix (SYBR Green) (Tiangen Biotech Co.). Primers were designed using Beacon Designer 8.12 and are listed in Table S1. The amount of mRNA was quantified against a standard curve using the CT value.
HPLC/ESI-MS analysis of cultures
Culture samples containing more than 500 mg/L l-tyrosine were alkalized to a final concentration of 0.25 M KOH and incubated for 30 min at room temperature to dissolve all l-tyrosine. All samples taken from the cultures were centrifuged at 15,000 × g for 10 min and the supernatants were filtered through a 0.2-μm syringe filter. The samples were analyzed by HPLC using an Agilent 1200 series instrument with an Eclipse XDB-C18 column (4.6 × 150 mm) and an Ultimate 3000 Photodiode Array Detector maintained at 25 °C. Vanillyl alcohol was analyzed according to the previous method13. Other compounds were analyzed use the following method. The flow rate was 1 mL/min and the mobile phase consisted of solvent A (0.1% trifluoroacetic acid in water) and solvent B (0.1% trifluoroacetic acid in acetonitrile). The following gradient elution program was used: 0 min, 95% solvent A + 5% solvent B; 8 min, 20% solvent A + 80% solvent B; 10 min, 80% solvent A + 20% solvent B; 11 min, 95% solvent A + 5% solvent B. Production of l-tyrosine was monitored by measuring the absorbance at 280 nm and production of 4-coumaric acid, caffeic acid, ferulic acid and vanillin was monitored by measuring the absorbance at 310 nm. The retention times of the five above-mentioned compounds were 3.4, 5.3, 4.7, 5.5 and 5.7 min, respectively. After HPLC, LC–MS was performed using an Agilent UPLC-TOF-MS system. Compounds were identified and quantified by comparing the observed retention times, peak areas and mass chromatograms with those of the corresponding chemical standards. The data shown in this study were generated from at least three independent experiments and analyzed using Microsoft Office Excel 2007 and IBM SPSS Statistics.
Additional Information
How to cite this article: Ni, J. et al. Mimicking a natural pathway for de novo biosynthesis: natural vanillin production from accessible carbon sources. Sci. Rep. 5, 13670; doi: 10.1038/srep13670 (2015).
References
Priefert, H., Rabenhorst, J. & Steinbuchel, A. Biotechnological production of vanillin. Appl. Microbiol. Biotechnol. 56, 296–314 (2001).
Brochado, A. R. et al. Improved vanillin production in baker’s yeast through in silico design. Microb. Cell Fact. 9, 84–98 (2010).
Walton, N. J., Narbad, A., Faulds, C. B. & Williamson, G. Novel approaches to the biosynthesis of vanillin. Curr. Opin. Biotechnol. 11, 490–496 (2000).
Muheim, A. & Lerch, K. Towards a high-yield bioconversion of ferulic acid to vanillin. Appl. Microbiol. Biotechnol. 51, 456–461 (1999).
Overhage, J., Steinbuchel, A. & Priefert, H. Highly efficient biotransformation of eugenol to ferulic acid and further conversion to vanillin in recombinant strains of Escherichia coli. Appl. Environ. Microbiol. 69, 6569–6576 (2003).
Hua, D. L. et al. Biotransformation of isoeugenol to vanillin by a newly isolated Bacillus pumilusstrain: identification of major metabolites. J. Biotechnol. 130, 463–470 (2007).
Masai, E. et al. Cloning and characterization of the ferulic acid catabolic genes of Sphingomonas paucimobilis SYK-6. Appl. Environ. Microbiol. 68, 4416–4424 (2002).
US Census Bureau. Foreign trade statistics, US Census Bureau, Washington, DC. (2004).
Li, K. & Frost, J. W. Synthesis of vanillin from glucose. J. Am. Chem. Soc. 120, 10545–10546 (1998).
Hansen, E. H. et al. De novo biosynthesis of vanillin in Fission Yeast (Schizosaccharomyces pombe) and Baker’s Yeast (Saccharomyces cerevisiae). Appl. Environ. Microbiol. 75, 2765–2774 (2009).
Brochado, A. R. et al. Improved vanillin production in baker’s yeast through in silico design. Microb. Cell Fact. 9, 84–98 (2010).
Kaur, B. & Chakraborty, D. Biotechnological and molecular approaches for vanillin production: a review. Appl. Biochem. Biotechnol. 169, 1353–1372 (2013).
Kunjapur, A. M., Tarasova, Y. & Prather, K. L. Synthesis and accumulation of aromatic aldehydes in an engineered strain of Escherichia coli. J. Am. Chem. Soc. 136, 11644–11654 (2014).
Dixon, R. A. et al. The phenylpropanoid pathway and plant defence, a genomics perspective. Mol. Plant Pathol. 3, 371–390 (2002).
Gallage, N. J. et al. Vanillin formation from ferulic acid in Vanilla planifolia is catalysed by a single enzyme. Nat. Commun. 5, 4037 (2014).
Zhang, H. R. & Stephanopoulos, G. Engineering E. coli for caffeic acid biosynthesis from renewable sugars. Appl. Microbiol. Biotechnol. 97, 3333–3341 (2013).
Kang, S. Y. et al. Artificial biosynthesis of phenylpropanoic acids in a tyrosine overproducing Escherichia coli strain. Microb. Cell Fact. 11, 153–161 (2012).
Huang, Q., Lin, Y. H. & Yan, Y. J. Caffeic acid production enhancement by engineering a phenylalanine over-producing Escherichia coli strain. Biotechnol. Bioeng. 110, 3188–3196 (2013).
Santos, C. N., Koffas, M. & Stephanopoulos, G. Optimization of a heterologous pathway for the production of flavonoids from glucose. Metab. Eng. 13, 392–400 (2011).
Kyndt, J. A., Meyer, T. E., Cusanovich, M. A. & Beeumen, Van. J. J. Characterization of a bacterial tyrosine ammonia lyase, a biosynthetic enzyme for the photoactive yellow protein. FEBS Lett. 512, 240–244 (2002).
Berner, M. et al. Genes and enzymes involved in caffeic acid biosynthesis in the actinomycete Saccharothrix espanaensis. J. Bacteriol. 188, 2666–2673 (2006).
Watts, K. T., Lee, P. C. & Schmidt-Dannert, C. Exploring recombinant flavonoid biosynthesis in metabolically engineered Escherichia coli. Chembiochem 5, 500–507 (2004).
Kim, Y. H. et al. Gene engineering, purification, crystallization and preliminary X-ray diffraction of cytochrome P450 p-coumarate-3-hydroxylase (C3H), the Arabidopsis membrane protein. Protein Expres. Purif. 79, 149–155 (2011).
Lin, Y. & Yan, Y. Biosynthesis of caffeic acid in Escherichia coli using its endogenous hydroxylase complex. Microb. Cell Fact. 11, 42–50 (2012).
Choi, O. et al. Biosynthesis of plant-specific phenylpropanoids by construction of an artificial biosynthetic pathway in Escherichia coli. J. Ind. Microbiol. Biotechnol. 38, 1657–1665 (2011).
Yang, W. W. et al. Characterization of two Streptomyces enzymes that convert ferulic acid to vanillin. PLoS One, 8, e67339. (2013).
Nakagawa, A. et al. A bacterial platform for fermentative production of plant alkaloids. Nat. Commun. 2, 326–333 (2011).
Xue, Y., Zhang, Y., Grace, S. & He, Q. Functional expression of an Arabidopsis p450 enzyme, p-coumarate-3-hydroxylase, in the cyanobacterium Synechocystis PCC 6803 for the biosynthesis of caffeic acid. J. Appl. Phycol. 26, 219–226 (2014).
Kim, S. & Lee, S. B. Rare codon clusters at 5′-end influence heterologous expression of archaeal gene in Escherichia coli. Protein Expres. Purif. 50, 49–57 (2006).
Muñoz, A. J. et al. Metabolic engineering of Escherichia coli for improving L-3, 4-dihydroxyphenylalanine (L-DOPA) synthesis from glucose. J. Ind. Microbiol Biot. 38, 1845–1852 (2011).
Ahn, J. O. et al. Exploring the effects of carbon sources on the metabolic capacity for shikimic acid production in Escherichia coli using in silico metabolic predictions. J. Microbiol. Biotechnol. 18, 1773–1784 (2008).
Serrano, L. Synthetic Biology: Promises and Challenges. Mol. Syst. Biol. 3, 158–162 (2007).
Andrianantoandro, E., Basu, S., Karig, D. K. & Weiss, R. Synthetic biology: new engineering rules for an emerging discipline. Mol. Syst. Biol. 2, 1–14 (2006).
Goossens, A. et al. A functional genomics approach toward the understanding of secondary metabolism in plant cells. Proc. Natl. Acad. Sci. USA 100, 8595–8600 (2003).
Ikeda, M. & Katsumata, R. Metabolic engineering to produce tyrosine or phenylalanine in a tryptophan-producing Corynebacterium glutamicum strain. Appl. Environ. Microbiol. 58, 781–785 (1992).
Bongaerts, J., Krämer, M., Müller, U., Raeven, L. & Wubbolts, M. Metabolic engineering for microbial production of aromatic amino acids and derived compounds. Metab. Eng. 3, 289–300 (2001).
Lütke-Eversloh, T. & Stephanopoulos, G. Combinatorial pathway analysis for improved l-tyrosine production in Escherichia coli: Identification of enzymatic bottlenecks by systematic gene overexpression. Metab. Eng. 10, 69–77 (2008).
Lee, W. H., Park, J. B., Park, K., Kim, M. D. & Seo, J. H. Enhanced production of ε-caprolactone by overexpression of NADPH-regenerating glucose 6-phosphate dehydrogenase in recombinant Escherichia coli harboring cyclohexanone monooxygenase gene. Appl. Microbiol. Biotechnol. 76, 329–338 (2007).
Johannes, T. W., Woodyer, R. D. & Zhao, H. Efficient regeneration of NADPH using an engineered phosphite dehydrogenase. Biotechnol. Bioeng. 96, 18–26 (2007).
Xu, Y. Q. et al. Systematic metabolic engineering of Escherichia coli for high-yield production of fuel bio-chemical 2,3-butanediol. Metab. Eng. 23, 22–23 (2014).
Mehta, V. J., Thumar, J. T. & Singh, S. P. Production of alkaline protease from an alkaliphilic actinomycete. Bioresour. Technol. 97, 1650–1654 (2006).
Binder, J. B. & Raines, R. T. Simple chemical transformation of lignocellulosic biomass into furans for fuels and chemicals. J. Am. Chem. Soc. 131, 1979–1985 (2009).
Hoover, D. M. & Lubkowski, J. DNA Works: an automated method for designing oligonucleotides for PCR-based gene synthesis. Nucleic Acids Res. 30, e43 (2002).
Acknowledgements
We thank Professor Hiromichi Minami for providing the tyrosine-overproducing E. coli strain. The authors also acknowledge the previous financial support on this study from Shanghai Apple Flavor & Fragrance (China).
Author information
Authors and Affiliations
Contributions
J.N. and P.X. conceived and designed the project and experiments. J.N., H.D. and F.T. performed the experiments. J.N. and F.T. analyzed the data. J.N., F.T. and P.X. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ni, J., Tao, F., Du, H. et al. Mimicking a natural pathway for de novo biosynthesis: natural vanillin production from accessible carbon sources. Sci Rep 5, 13670 (2015). https://doi.org/10.1038/srep13670
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13670
This article is cited by
-
Minimal aromatic aldehyde reduction (MARE) yeast platform for engineering vanillin production
Biotechnology for Biofuels and Bioproducts (2024)
-
Strategies for improving the production of bio-based vanillin
Microbial Cell Factories (2023)
-
Sustainable biosynthetic pathways to value-added bioproducts from hydroxycinnamic acids
Applied Microbiology and Biotechnology (2023)
-
Biosynthesis of eriodictyol from tyrosine by Corynebacterium glutamicum
Microbial Cell Factories (2022)
-
Biosynthesis of vanillin by different microorganisms: a review
World Journal of Microbiology and Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.