Abstract
Cancer types are commonly classified by histopathology and more recently through molecular characteristics such as gene expression, mutations, copy number variations and epigenetic alterations. These molecular characterizations have led to the proposal of prognostic biomarkers for many cancer types. Nevertheless, most of these biomarkers have been proposed for a specific cancer type or even specific subtypes. Although more challenging, it is useful to identify biomarkers that can be applied for multiple types of cancer. Here, we have used a network-based exploration approach to identify a multi-cancer gene expression biomarker highly connected by ESR1, PRKACA, LRP1, JUN and SMAD2 that can be predictive of clinical outcome in 12 types of cancer from The Cancer Genome Atlas (TCGA) repository. The gene signature of this biomarker is highly supported by cancer literature, biological terms and prognostic power in other cancer types. Additionally, the signature does not seem to be highly associated with specific mutations or copy number alterations. Comparisons with cancer-type specific and other multi-cancer biomarkers in TCGA and other datasets showed that the performance of the proposed multi-cancer biomarker is superior, making the proposed approach and multi-cancer biomarker potentially useful in research and clinical settings.
Similar content being viewed by others
Introduction
Cancer is typically classified by tissue-specific scores such as the Gleason score in prostate cancer1, the Dukes or Astler-Coller in colon cancer2, or Figo in cervical cancer3. These have been generalized by TNM staging4. More recently, high-throughput technologies have generated unprecedented molecular characterizations of cancer types, such as the genomic portrayals provided by The Cancer Genome Atlas (TCGA) research network5,6,7. Several cancer types have been divided into subtypes using TCGA data about gene expression8, mutations9, copy number alterations10, microRNA expression11, pseudogenes12, or even biological processes such as inflammation13. Nevertheless, specific subtypes across cancer types seem to share common gene expression properties such as correlations14,15, stromal and immune signatures16, or mesenchymal signatures17. Clinically, better or alternative methods to identify cancer risk groups are always needed.
Although point mutations and chromosomal alterations are currently the subject of active research, a large reservoir of public research has been devoted to the study of gene expression18,19, which is the most broadly studied class of molecular data for cancer so far. More importantly, gene expression is a consequence of cumulative genetic and epigenetic alterations. With the goal of clinically stratifying samples into risk groups, several gene expression biomarkers have been proposed for a large variety of cancer types20,21,22,23,24,25.
However, most biomarkers have been identified and designed for a specific type of cancer. Moreover, some biomarkers can be applied only to specific subtypes; for example, biomarkers exist specifically for grade 2 in colon cancer26 and estrogen receptor-positive and lymph node-negative in breast cancer27. Many proposed gene signatures may not even be considered for clinical use because they could not be reliably validated in other cohorts28. In some cases, a lack of agreement has also been reported among gene expression signatures obtained for the same type of cancer29. In addition, it was recently shown that biomarkers identified for only one cancer type perform modestly or poorly even when clinical data are considered30.
Gene expression biomarkers that can be applied for a broad range of cancers could be highly useful in research and clinical settings. In clinics, such biomarkers may serve as a standard assessment for facilitating the interpretation and broad application of laboratory test results, simplifying laboratory protocols and reducing costs. In research, these biomarkers may help to elucidate broadly observed biological mechanisms and possible drug targets. Nevertheless, gene expression biomarkers that can be applied to more than one cancer type are scarce. Most studies exploit specific properties to identify multi-cancer signatures. For example, signatures have been identified from metastasis-specific solid tumors31, or found to be associated with chromosomal instability32, therapy-failures33, proliferation signatures34, subsequent cancers35 and embryonic stem-cell like gene expression36.
Distinct algorithms and strategies have been used to identify biomarkers for more than 10 years. These methods include variable selection by shrinkage37,38, penalization39,40, clustering41, differential expression42,43, or simply by selection of the top-ranked genes using a univariate Cox score21, among many others44. Most of these methods nevertheless do not consider a priori biological information to identify gene signatures. The use of biological information adds a layer of validation and prioritization45 that can be exploited for biomarker discovery. Common approaches that consider biological information use networks such as protein-protein interactions (PPI) or gene ontologies, which drives the search for modules or terms that could function as gene signatures. For example, a set of significant subnetwork biomarkers to classify breast cancer metastasis was identified by performing a greedy search starting from seed genes and then adding neighbor genes46. These subnetworks were then compared with a null distribution of random subnetworks. Similarly, this algorithm was adapted for a web server that provides network-based biomarkers for survival data47. A network module-based approach applied a Markov clustering algorithm to the correlation of the PPI matrix identifying modules associated with patient survival48. An algorithm similar to that used by Google for ranking web pages has been proposed to order genes according to their association with survival outcomes49. Modularity has also been suggested as an indicator of breast cancer prognosis as determined by an algorithm to find intramodular highly co-expressed and highly interconnected “hub” genes and intermodular hub genes with low co-expression50. Moreover, gene ontology has also been used to identify metastasis network modules combining highly predictive gene ontology sets51. To the best of our knowledge, these network-based approaches have not been tailored to produce network-based multi-cancer biomarkers.
Here we describe a network-based approach that explores, in parallel, gene-to-gene connections in multiple cancer datasets while maximizing the overall association of the subnetwork with clinical outcomes. We implemented this network-based algorithm using, as a proof of concept, the Human Protein Reference Database (HPRD)52, 12 TCGA cancer types53 and a composite Cox-based model54. In these training datasets, the results showed that a gene signature of 41 genes was capable of predicting risk groups across cancer types with high precision. Analysis of a large collection of clinical outcome cancer datasets that included cancers types reported by several authors and many cancer types that were not included in the training datasets validated these results. The predictive power of the biomarker was higher than that of clinical information alone and improved when combined. Our results suggest that it is possible to identify general, compact and biologically driven gene expression biomarkers for multiple cancer types.
Material and Methods
Datasets
We used data from 12 cancer types that belong to the TCGA pan-cancer project repository53 accessed in January 2013, compiled in 11 datasets (http://nature.com/tcga). Detailed lists of datasets, genes and samples used are shown in Table 1 and Supplementary Table 1. Level 3 data were used. Only gene symbols present in all cancer types were used. Microarray data (Agilent and Affymetrix) were transformed using quantile normalization. RNA-Seq data were Log2 transformed and quantile normalized.
Biological networks
We used the protein-protein interaction (PPI) network from the Human Protein Reference Database (HPRD)52 accessed in March 2013. The network covered 9,465 genes and 37,080 interactions. Only genes found in both the TCGA datasets and the network were used.
Performance of network modules
We used gene expression values from each cancer type fitting a Cox model to measure the level of association of a given gene signature. For biomarkers specific to a cancer-type, the negative logarithm of the log-rank test (NLLRT) was used to assess and drive the network-based search. For the multi-cancer biomarker, we used the NLLRT of a reference cancer-type minus the range of the NLLRT from the remaining cancer-types. Subtracting the range of values gave preference to less variable signatures, helping to avoid over-fitting to specific cancer types. We used GBM as the reference cancer-type because its performance was the lowest across the cancer type-specific runs. Nevertheless, we also used other cancer types as reference and compared the results. To assess the overall performance of the prediction after biomarker identification, we used the concordance index (C-index) measure, which is similar to the area under the receiving operating curve55. C-index values close to 0.5 are referred to as random risk predictions whereas C-index values close to 1 are interpreted as nearly perfect risk predictions. To represent the performance of biomarkers graphically, we split the samples by the median of the prognostic index to designate low- and high- risk groups. The prognostic index is the linear component of the exponential function in the Cox model.
Network clinical association (NCA) algorithm
Figure 1 shows a graphical representation of the NCA algorithm. The algorithm proceeds in cycles, starting with the determination of the performance of an isolated gene (a seed gene) across all datasets. Then, a module growth cycle is performed, in which all connected genes are explored, one gene at a time, generating as many grown modules as connections. In the exploration, the performance of the module is evaluated using the NLLRT value described above. Afterwards, only the top 5% of the modules whose NLLRT value improved after the addition of new connections are considered for the next growth cycle. The procedure continues until no improvement is observed. The algorithm starts by using each gene as a seed. This algorithm functions as a type of hill-climbing algorithm. Scripts or executables of this algorithm are available from the corresponding author.
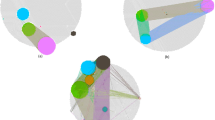
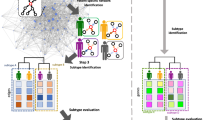
Schematic representation of the network clinical association algorithm (NCA).
Starting from a single seed gene (black), the first cycle generates modules that include the seed gene and each of the connected genes (blue). The 6 modules of 2 genes are then evaluated by their goodness of fit in a Cox survival model. Only those grown modules that improve (filled blue circles) the evaluation are considered for the next grow cycle. Only a proportion of the best improved modules are further explored in the next cycle (represented by a percentage of the distribution of all modules, shown in green, evaluated in the 4th cycle). This procedure continues until no improvement is observed. The NCA algorithm was run for each cancer type and for all cancer datasets (multi-NCA).
Validation analysis
To determine the significance of the C-index values, we generated a null distribution composed of 10,000 random models of 41 genes for the TCGA datasets we used. To assess the C-index prediction of the biomarkers in datasets other than TCGA, we used SurvExpress56, which provides evaluations of gene lists across cancer types. For this, we used normalized datasets that included overall survival times (without considering recurrence, metastases, or relapse) and only those studies containing more than 30 samples. For replicated genes, we selected the highest expressed probe. Analyses were performed in R (http://cran.r-project.org/). For biological validation, we used MSigDB and DAVID57,58 to determine which biological terms were associated with the biomarker gene lists. We also compared the C-index values of our multi-cancer biomarker with those of other multi-cancer biomarkers reported in the literature. For model comparisons including clinical features (e. g. cancer staging), we used the “other factors” option in SurvExpress.
Results
Identification of biomarkers for specific cancer types
We first executed the NCA algorithm (Fig. 1) for each of the 11 cancer datasets. We focused on the network modules with the highest performance value. The results shown in Table 2 suggest that, in general, several network modules existed for each cancer type, from 84 for LUSC to 10,303 for BLCA. Most cancer types generated modules with about 9 genes, ranging in size from 4 for KIRC to 14 for LUSC. To generate a biomarker that is representative of all modules for a specific cancer type, we used the genes that most frequently occurred in modules (around 41 genes for comparisons with the multi-cancer biomarker). The list of genes obtained is provided in Supplementary Table 2. Comparisons of the genes used for these biomarkers across cancer types showed that the pairwise gene overlap was low (ranging from 0 to 5, see Fig. 2A). Although the specific genes used for each biomarker were clearly different, indicating that the biomarkers are cancer type-specific, the prediction across cancer types was surprisingly satisfactory; the average C-index values were higher than 0.75 (Fig. 2B and Supplementary Table 3). Almost all cancer type-specific biomarkers showed C-index values higher than 0.70 for about 8 cancer types (Fig. 2C). We observed consistent C-index values within each cancer type almost irrespective of the network-based biomarker (Fig. 2D). For instance, all biomarkers had a C-index value about 0.97 for BLCA and 0.95 for COADREAD but about 0.65 for OV and 0.62 for GBM. Nevertheless, a random signature analysis indicated that only 14 of the 121 C-index values (11.5%) were significant, mainly those of cancer type-specific biomarkers within the same cancer type dataset (excluding BLCA and COADREAD, see Fig. 2A and Supplementary Table 3).
Comparison of biomarkers generated by the network clinical association (NCA) algorithm.
Panel A shows the number of genes that were included in any two biomarkers. Underlined numbers represent the number of genes per biomarker. Red indicates high overlaps and blue indicates no overlap. The “Sum” row shows the total number of overlaps with other biomarkers while the “Unique” row shows the number of unique genes that overlap. Panel B shows the C-index evaluation of NCA biomarkers (rows) across cancer datasets (columns). Underlined numbers represent the biomarkers evaluated within the cancer dataset. Red indicates high values within the cancer dataset (column) and blue indicates low values. Boldface and framed values represent significant predictions using 10,000 random models of the same length. The “Average” row shows the average C-index per cancer type and the “Average” column shows the mean C-index per biomarker. Panel C shows the NCA biomarkers (horizontal) evaluated in all datasets using C-index (vertical axis). The mean is shown as a horizontal line. Panel D shows cancer types (horizontal) evaluated with all biomarkers using C-index (vertical axis).
Identification of a multi-cancer biomarker
To generate a broadly predictive biomarker, we used the NCA algorithm and considered the 11 datasets in the same run. We estimated a composite performance score based on the individual performance of all cancer types. We maximized the overall performance by taking the NLLRT of a reference cancer type (glioblastoma) and subtracting the range of NLLRT values of the other cancer types. In this way, genes generating large deviances for specific cancer types were avoided in favor of the inclusion of genes that improved the prediction in many cancer types. Two very similar modules consisting of 44 genes were identified (Table 2). Only 6 genes were not present in both modules (JDP2, KIF5B, NTRK3, MMP13, TGFB1 and TGFBRAP1). Therefore, we used the genes present in both modules as the overall multi-cancer biomarker.
The identified network biomarker was composed of 41 genes highly connected by ESR1, PRKACA, LRP1, JUN and SMAD2 (Fig. 3A). This gene signature was able to discriminate between low- and high-risk groups efficiently in the 11 cancer datasets (Fig. 3B and Table 3) through the statistical association of specific genes (Fig. 3C). The log-rank test and the Cox model fitting were highly significant across cancer types (Table 3). The average C-index value across cancer types was 0.81 ranging from 0.65 to 1. Eight of these 11 predictions were significant according to a randomization analysis (Fig. 2B and Supplementary Table 3). The highest C-index predictions were observed for BLCA and COADREAD, whereas the lowest C-index predictions were observed for GBM and OV.
The multi-NCA biomarker identified when all databases were combined.
Panel A shows the genes and network identified. The connections correspond to data from the PPI database used. The most connected genes were PRKACA, ESR1, LRP1, SMAD2 and JUN. Panel B shows the risk group prediction (splitting the prognostic index by the median) of the multi-NCA biomarker across cancer datasets. Panel C depict the color-coded differential expression of genes between risk groups. Darker red indicates more significant differences. The scales were estimated in -Log10 of the t test p value. Only p values <0.01 are highlighted. Darker purple indicates more significant hazard ratio associations within the Cox model. The scales were expressed in -Log10 of the Z p value. Only p values <0.05 are highlighted. Panel D shows, in the top, the curated biological terms and pathways associated with the genes composing the biomarker. The associations of genes with specific cancers based on the literature are shown at the bottom.
In a comparison of the predicted low- and high-risk groups (splitting the prognostic index by the median), we observed several genes differentially expressed across cancer types, except in BLCA (Fig. 3C and Table 3). Apart from LMO4 and DDX5, the other 39 genes were differentially expressed between risk groups in two or more cancer types. LMO4 was not differentially expressed in any cancer but was significantly associated with GBM, LUAD and LUSC according to the Cox model. DDX5 was highly differentially expressed in LUAD and associated with three cancer types according to the Cox model. Similarly, 36 genes were associated with the Cox model for two cancer types or more. Surprisingly, ESR1 was not associated with Cox models but was differentially expressed in two cancer types and served as a hub for connecting 10 genes.
An overrepresentation analysis of the 41 genes using MSigDB57 and DAVID58 revealed important biological associations across pathways, transcriptional control, gene ontologies and other biological terms (Fig. 3D). Some of these pathways are well known to be associated with cancer, such as the MAPK59, LKB160, ERα61 and NGF62 pathways. Some genes were highly associated with transcription factors such as SP163, gene ontologies such as signaling and other biological terms such as immune system, copy number gains in cancer and MIR-18 targets. In addition, at least 36 of the 41 genes have been associated in the literature with one or more cancer types (Fig. 3D).
These findings support the utility of determining which genes predicted specific cancer types and suggest that the signature we generated is robust across cancer subtypes.
Comparison of the multi-cancer and cancer type-specific biomarkers
It has been proposed that molecular processes may be similar across cancer types14,15,64. Consequently, a biomarker of clinical outcomes in a specific cancer type may be a good biomarker in a different cancer type. Therefore, we compared our 12 biomarkers to identify similarities. In terms of gene content (Supplementary Table 2), the multi-cancer (multi-NCA) biomarker was not particularly similar to the cancer type-specific biomarkers (Fig. 2A). Indeed, the biomarker most similar to others was OV, which contained 17 genes (29 occurrences) that overlapped with other biomarkers out of the 418 unique genes. This similarity was considerably higher than that of the multi-NCA, which had 9 genes (17 occurrences) overlapping and that of the most specific biomarker, GBM, which had only 4 genes (5 occurrences) in common with the other biomarkers.
A comparison of the average C-index values across biomarkers and cancer types showed that the multi-NCA biomarker was, overall, the best (average C-index = 0.81) but it was closely followed by OV and BRCA (average C-index of 0.80 and 0.79 respectively; Fig. 2). The C-index of the multi-NCA biomarker was almost always better than those of the cancer type-specific biomarkers (Fig. 2D). Nevertheless, in each cancer type, the C-index was higher using the cancer type-specific biomarker than using the multi-NCA biomarker (by 0.047 on average). Despite this, an analysis of 10,000 random biomarkers showed that 8 of 11 C-index predictions of the multi-NCA biomarker were significant (Fig. 2B and Supplementary Table 3) whereas the C-indexes in most cancer type-specific biomarkers were significant only in one or two cancer types (OV in three and marginally in two more). In terms of prediction power per cancer type, the BLCA and COADREAD average C-index values were, by far, the highest (both 0.98). In contrast, the C-indexes for GBM and OV were the lowest (0.62 and 0.65 respectively).
Comparisons with clinical features
Although biomarkers can be a useful clinical tool to predict outcomes, some of the generated biomarkers may not actually be useful in clinical practice if the gene signature does not add predictive power beyond that of the usual clinical features30. To assess this, we determined the C-index of the multi-NCA biomarker and the available clinical features per cancer type. The Supplementary Figure 1 shows that the multi-NCA biomarker adds between 0.04 and 0.30 of prediction power over clinical features alone. In contrast, the clinical features add only between 0 and 0.075 over the biomarker alone. Overall, in most cancer types (except KIRC) the biomarker makes better predictions than clinical features alone. These results suggest that the multi-NCA signature adds a considerable level of predictive power to clinical features.
We also determined whether the multi-NCA signature was sensitive to stratifications using cancer features. For this, we used the widely used cancer staging system for each cancer type to compare the performance of C-indexes across cancer stages. As shown in Supplementary Figure 2, C-index values varied somewhat across stages, perhaps influenced by the number of TCGA samples available per stage. Of note, in BRCA stage IIIA, in KIRC stage III and IV, OV stage IIIB and UCEC stage IIIC, the C-indexes were lower than 0.05 relative to the overall C-indexes for corresponding cancer types (in these cases the estimation considered more than 20 samples). Nevertheless, the C-index value is still acceptable for most stages. This stratification provides an estimation of the response of markers across a wide spectrum of subtypes.
External comparison and validation of biomarkers
For external validation of the multi-NCA biomarker, we compared the C-index with other 5 multi-cancer biomarkers proposed by other authors32,65,66 representing signatures of chromosome instability (CIN70)32, multiple cancer-related pathways (poised gene cassette, PGC)65, mesenchymal transition (MES)66, mitotic chromosomal instability (CIN)66 and lymphocyte infiltration (LYM)66. The 41 genes in the multi-NCA biomarker did not overlap with any of the genes in CIN70, PGC, MES, CIN and LYM (Supplementary Table 2). The average C-index for the LYM biomarker was 0.796, just below that of our multi-NCA biomarker, which was 0.809 (Supplementary Table 3). The C-index for LYM was nevertheless significant in only 3 TCGA datasets compared with 8 datasets for the multi-NCA biomarker, suggesting that our multi-cancer biomarker is superior to the LYM biomarker.
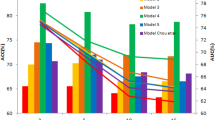
To evaluate the prediction accuracy of the biomarkers in cancer data other than TCGA, we used SurvExpress56 to analyze the multi-NCA and the cancer type-specific biomarkers we generated and the multi-cancer biomarkers generated by other-authors. We used 122 cancer datasets containing 19,105 samples spanning about 20 types of tissues (Supplementary Table 4). These datasets covered cancer types not used to develop the NCA-based biomarkers such as cancer of the bone, esophagus, eye, liver, prostate, pancreas and skin, as well as medulloblastomas and astrocytomas and others. We performed two analyses, the first averaging all 122 datasets and the second normalizing the average per tissue. The second analysis was more important because some tissues have been more studied than others such as lung, ovary, breast, brain and colon. In addition, some cohorts are reported in various datasets. The results showed that our multi-NCA biomarker was one of the top biomarkers evaluated; it was the most accurate in the per-tissue analysis and close to the most accurate in all datasets (Fig. 4 and Supplementary Table 4). Compared with other multi-cancer biomarkers, our multi-NCA signature was more accurate in the per tissue analysis than the CIN, CIN70, PGC, LYM and MES signatures. Among these, the MES was the best in the per-tissue analysis while LYM was first considering all datasets.
Comparison of multi-cancer module evaluation functions
The results reported here represented by the multi-NCA biomarker were obtained using GBM as the reference cancer type minus the range of all other cancer types examined. We also explored the performance of the network-based marker generation using other functions and other cancer types as reference. We first tested the obvious average function, followed by the average minus the range. As demonstrated in Supplementary Figure 4, using only the average function generated the poorest performance, which was improved by subtracting the range but still lower than using GBM as the reference minus the range. Then, we tested the other three cancer types used as references: LUAD, OV and BRCA. Interestingly, using LUAD as the reference generated a lower performance than that of GBM in all cancer types, whereas using OV generated almost the same overall performance as GBM. Surprisingly, using BRCA as the reference resulted in a better performance than that of GBM in 7 cancer types (only LUAD showed a decrease; the overall increase in performance was 0.025).
Discussion
We used NCA, a network-based algorithm, to identify biomarkers highly predictive of survival outcomes in cancer. We first identified biomarkers for specific cancers and then identified a multi-cancer biomarker for 12 cancer types. Interestingly, the gene content varied greatly across biomarkers but the performance was similar when evaluated in each cancer type (Fig. 2D). These results suggest that C-index values are more dependent on cancer type than on gene content of the biomarker. Consequently, survival outcomes may be more difficult to predict in some cancer types than in others. For instance, survival was easier to predict in BLCA and COADREAD than in OV and GBM. This is also supported by the fact that C-index values close to 1 for BLCA and COADREAD were not significant since random markers also showed high C-index values while C-indexes of 0.66 for OV and 0.65 in GBM were highly significant compared with random markers.
The OV-NCA biomarker was the second most accurate biomarker across cancer types (Fig. 2C) even though it was developed using the ovarian serous cystadenocarcinoma dataset only. A comparison of the OV biomarker (Supplementary Figure 3) with the multi-NCA biomarker (Fig. 3) showed that, surprisingly, the ovarian biomarker had more connections than the multi-NCA biomarker. However, the number of differentially expressed genes, the Cox model statistics and the biological terms associated with the signature were more appropriate in the multi-NCA biomarker than in the OV biomarker. The multi-NCA was able to significantly predict survival outcomes in 5 more cancer types than the OV biomarker (Fig. 2B) and it was more accurate in the per-tissue analysis (Fig. 4) than the OV biomarker. These findings indicate that the multi-NCA biomarker was more suitable for multi-cancer predictions than the OV biomarker. Nevertheless, it would be interesting to explore why the OV biomarker was highly predictive of outcomes across cancers. Although ovarian cancer was hard to predict, glioblastoma was even harder but the GBM biomarker was less accurate than the OV biomarker (Figs 2C and 4), so it cannot be easily linked to prediction difficultness. Ovarian serous cystadenocarcinoma can be divided into various subtypes defined by immunoreactive, mesenchymal, proliferative and differentiated characteristics6. These characteristics represent universal tumorigenic processes and are observed in other types of cancer as well6. This heterogeneity is reflected in the relatively high number of individuals (578) included in the TCGA ovarian dataset16, although a similar number of samples was included in glioblastoma and invasive breast carcinoma (Table 1). In addition, the five genes (JUN, PRKACA, SMAD2, ESR1 and BCL3) shared by the OV and multi-NCA biomarkers form a small network module and are recognized as cancer-related genes. Further analysis is needed to explore the reasons for the apparently high inter-cancer accuracy of the OV biomarker.
None of the C-index values in BLCA or in COADREAD were significant in the random model test even though the C-index values reached 1 because 46% and 12% of the random models respectively were equally predictive. Moreover, in BLCA, none of the genes were differentially expressed between risk groups. Although the low number of samples could influence these results (only 54 samples in BLCA and 151 in COADREAD), confirmed results in larger cohorts would imply that many predictive signatures may exist. In our study, the multi-NCA did not depend on the number of samples per cancer type but in the NLLRT of each cancer type. Thus, the selection of the best signature was imposed by other cancer types rather than by BLCA and COADREAD. This may explain why none of the genes was significant in BLCA. Nevertheless, these findings do not necessarily indicate that these genes in BLCA are not important. For instance, high expression of CALR has been associated with high risk in bladder cancer67. HRAS gains have been found in bladder cancer cell lines and have been related to urothelial tumorigenesis68. ITGA4 is part of a methylation gene set used for the detection of bladder cancer69. In COADREAD, AKT isoforms (including AKT1) are associated with high expression of CD133 and CD44 (cancer stem cell markers) and radiation resistance in colon cancer cells70. High expression of DDX5 (previously known as p68) is related to the transition from polyp to adenoma and then to adenocarcinoma71. High levels of DUT protein expression are predictive for tumor resistance to chemotherapy in colorectal cancer72. Finally, up-regulation of JUN is related to the invasiveness of colorectal cancer cells. These findings clearly indicate that the biomarker genes are biologically related to BLCA and COADREAD.
The performance comparisons of the multi-NCA with clinical features suggest that the multi-NCA signature adds predictive power to clinical features. Nevertheless, these comparisons also showed that the predictive power of the multi-NCA biomarker might vary across cancer stages. This may indicate that the biomarker is somehow influenced by the high representation of specific cancer subtypes in the TCGA studies. For example, the results in BRCA were highly influenced by stage II samples, which accounted for more than 50% of total samples, whereas stage IV samples represented only 3% of samples. Other cancer types showed similar staging biases. Inclusion of more samples (as is happening with the TCGA and the International Cancer Genome Consortium datasets) and prefiltering of data to balance stage representation may be good strategies to improve the identification of multi-cancer biomarkers.
The C-index value of our multi-NCA biomarker was higher than that of other previously reported multi-cancer biomarkers, but not substantially. The C-index values of MES and CIN70 were just below that of our multi-NCA biomarker. Some of the other multi-cancer biomarkers however use more genes for the prediction (Supplementary Table 2). Still, these comparisons highlight the fact that our multi-NCA biomarker is highly competitive among the others reported.
The network-based strategy that we used emphasizes the fact that using biological information coupled with gene selection is a powerful strategy to generate biomarkers; this conclusion is consistent with results from other studies46,47,48,49,50,51. However, the network-based strategy that we used is different from other approaches in various ways (Supplementary Table 5). First, we directly evaluated a Cox model that is capable of identifying combinatorial features more robustly than univariate-oriented approaches47, classifiers46,49,51 or components48. Second, unlike in other algorithms, we did not prefilter genes to decrease the complexity of the exploration47. Third, we used population-dependent selection of the most improved models allowing us to explore more combinations than would be possible using other algorithms46,47. Finally, to generate a multi-cancer biomarker, we expanded the Cox evaluation to multiple datasets by subtracting the range of all NLLRT values from the NLLRT value of a reference cancer.
We used the HPRD protein-protein interaction network in our approach. In principle, however, the NCA approach can be applied to other biological networks such BioGrid73, iRefWeb74, STRING75 and to genetic regulatory networks such as MotEvo76 and the conserved transcription factor binding sites track in UCSC (https://genome.ucsc.edu). The NCA algorithm is not limited to gene expression data or to survival analysis as the response variable. The exploration of diverse biological networks, genomic data and response variables may lead to the identification of better or alternative multi-cancer biomarkers.
The identification of novel or alternative multi-cancer biomarkers is also valuable because such biomarkers can represent different biological phenomena that may help to elucidate specific cancer features. For example CIN70 was identified from chromosome instability32, MES from mesenchymal transition66 and LYM from lymphocyte infiltration66. Our multi-NCA biomarker represents a protein-network-based biomarker. In this context, our multi-NCA biomarker does not share genes with other multi-cancer markers and shares only 5 genes with the OV biomarker also identified here.
We tested diverse module evaluation functions in which we varied the reference cancer type. We observed that the biomarkers found and their performance depended on this evaluation. These results have deep implications: the choice of the module-growth function is critical, the function used can be improved and the approach can generate alternative markers. It would be interesting to explore other functions combined with more cancer types.
Recent results have explored the correlation between gene expression and genomic changes such as copy number alterations77. In this context, the predictive power of the multi-NCA biomarker appeared to be specific for gene expression because mutations and copy number alterations were not highly related (Supplementary Table 6). The search for mutation signatures associated with clinical outcomes is starting30. Given the sparseness of the mutational spectrum across cancers, it is difficult to realize that a general mutation signature could be found. It would be exciting to see whether approaches like our proposal are capable of providing interesting solutions.
The identification of multi-cancer biomarkers may lead to proposals of novel diagnostic tools and therapeutic schemes. In this context, using DGIdb78 we observed that 22 of the 41 genes of the multi-cancer biomarker were known drug targets (Supplementary Table 7). Thus, our approach may also shed light on which targets can be assayed in future experiments.
Additional Information
How to cite this article: Martinez-Ledesma, E. et al. Identification of a multi-cancer gene expression biomarker for cancer clinical outcomes using a network-based algorithm. Sci. Rep. 5, 11966; doi: 10.1038/srep11966 (2015).
References
Helpap, B. & Egevad, L. Modified Gleason grading. An updated review. Histol Histopathol 24, 661–666 (2009).
Astler, V. B. & Coller, F. A. The prognostic significance of direct extension of carcinoma of the colon and rectum. Ann Surg 139, 846–852 (1954).
Mutch, D. M., Berger, A., Mansourian, R., Rytz, A. & Roberts, M. A. Microarray data analysis: a practical approach for selecting differentially expressed genes. Genome Biol 2, PREPRINT0009 (2001).
Edge, S. B. & Compton, C. C. The American Joint Committee on Cancer: the 7th edition of the AJCC cancer staging manual and the future of TNM. Ann Surg Oncol 17, 1471–1474, 10.1245/s10434-010-0985-4 (2010).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477, 10.1016/j.cell.2013.09.034 (2013).
Network, C. G. A. R. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615, 10.1038/nature10166 (2011).
Network, C. G. A. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70, 10.1038/nature11412 (2012).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR and NF1. Cancer Cell 17, 98–110, 10.1016/j.ccr.2009.12.020 (2010).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421, 10.1038/nature12477 (2013).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat Genet 45, 1134–1140, 10.1038/ng.2760 (2013).
Hamilton, M. P. et al. Identification of a pan-cancer oncogenic microRNA superfamily anchored by a central core seed motif. Nat Commun 4, 2730, 10.1038/ncomms3730 (2013).
Han, L. et al. The Pan-Cancer analysis of pseudogene expression reveals biologically and clinically relevant tumour subtypes. Nat Commun 5, 3963, 10.1038/ncomms4963 (2014).
Yu, X. et al. The pan-cancer analysis of gene expression patterns in the context of inflammation. Mol Biosyst 10, 2270–2276, 10.1039/c4mb00258j (2014).
Martinez, E. et al. Comparison of gene expression patterns across 12 tumor types identifies a cancer supercluster characterized by TP53 mutations and cell cycle defects. Oncogene, 10.1038/onc.2014.216 (2014).
Hoadley, K. A. et al. Multiplatform Analysis of 12 Cancer Types Reveals Molecular Classification within and across Tissues of Origin. Cell 158, 929–944, 10.1016/j.cell.2014.06.049 (2014).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nature Communications 4, 11, 10.1038/ncomms3612 (2013).
Byers, L. A. et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin Cancer Res 19, 279–290, 10.1158/1078-0432.ccr-12-1558 (2013).
Rung, J. & Brazma, A. Reuse of public genome-wide gene expression data. Nat Rev Genet 14, 89–99, 10.1038/nrg3394 (2013).
Baker, M. Gene data to hit milestone. Nature News 487, 282, doi:10.1038/487282a (2012).
Finak, G. et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat Med 14, 518–527, 10.1038/nm1764 (2008).
Li, Z. et al. Identification of a 24-gene prognostic signature that improves the European LeukemiaNet risk classification of acute myeloid leukemia: an international collaborative study. J Clin Oncol 31, 1172–1181, 10.1200/jco.2012.44.3184 (2013).
Chen, R. et al. A meta-analysis of lung cancer gene expression identifies PTK7 as a survival gene in lung adenocarcinoma. Cancer Res 74, 2892–2902, 10.1158/0008-5472.can-13-2775 (2014).
Peng, Z. et al. An expression signature at diagnosis to estimate prostate cancer patients’ overall survival. Prostate Cancer Prostatic Dis 17, 81–90, 10.1038/pcan.2013.57 (2014).
Verhaak, R. G. et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J Clin Invest 123, 517–525, 10.1172/jci65833 (2013).
Chibon, F. et al. Validated prediction of clinical outcome in sarcomas and multiple types of cancer on the basis of a gene expression signature related to genome complexity. Nat Med 16, 781–787, 10.1038/nm.2174 (2010).
Maak, M. et al. Independent validation of a prognostic genomic signature (ColoPrint) for patients with stage II colon cancer. Ann Surg 257, 1053–1058, 10.1097/SLA.0b013e31827c1180 (2013).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351, 2817–2826, 10.1056/NEJMoa041588 (2004).
Venet, D., Dumont, J. E. & Detours, V. Most random gene expression signatures are significantly associated with breast cancer outcome. PLoS Comput Biol 7, e1002240, 10.1371/journal.pcbi.1002240 (2011).
Ein-Dor, L., Zuk, O. & Domany, E. Thousands of samples are needed to generate a robust gene list for predicting outcome in cancer. Proc Natl Acad Sci USA 103, 5923–5928, 10.1073/pnas.0601231103 (2006).
Yuan, Y. et al. Assessing the clinical utility of cancer genomic and proteomic data across tumor types. Nat Biotechnol 32, 644–652, 10.1038/nbt.2940 (2014).
Daves, M. H., Hilsenbeck, S. G., Lau, C. C. & Man, T. K. Meta-analysis of multiple microarray datasets reveals a common gene signature of metastasis in solid tumors. BMC Med Genomics 4, 56, 10.1186/1755-8794-4-56 (2011).
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N. & Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet 38, 1043–1048, 10.1038/ng1861 (2006).
Glinsky, G. V., Berezovska, O. & Glinskii, A. B. Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest 115, 1503–1521, 10.1172/jci23412 (2005).
Starmans, M. H. et al. Robust prognostic value of a knowledge-based proliferation signature across large patient microarray studies spanning different cancer types. Br J Cancer 99, 1884–1890, 10.1038/sj.bjc.6604746 (2008).
Wan, Y. W., Qian, Y., Rathnagiriswaran, S., Castranova, V. & Guo, N. L. A breast cancer prognostic signature predicts clinical outcomes in multiple tumor types. Oncol Rep 24, 489–494 (2010).
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 40, 499–507, 10.1038/ng.127 (2008).
Tibshirani, R., Hastie, T., Narasimhan, B. & Chu, G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci USA. Jan 20; 99, 6567–6572 (2002).
Tibshirani, R. The lasso method for variable selection in the Cox model. Stat Med 16, 385–395 (1997).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. Journal of the Royal Statistical Society: Series B (Statistical Methodology) 67, 301–320, 10.1111/j.1467-9868.2005.00503.x (2005).
Yoshihara, K. et al. High-risk ovarian cancer based on 126-gene expression signature is uniquely characterized by downregulation of antigen presentation pathway. Clin Cancer Res 18, 1374–1385, 10.1158/1078-0432.ccr-11-2725 (2012).
Park, Y. Y. et al. Development and validation of a prognostic gene-expression signature for lung adenocarcinoma. PLoS One 7, e44225, 10.1371/journal.pone.0044225 (2012).
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA. Jan 20; 98, 5116–5121 (2001).
Stratford, J. K. et al. A six-gene signature predicts survival of patients with localized pancreatic ductal adenocarcinoma. PLoS Med 7, e1000307, 10.1371/journal.pmed.1000307 (2010).
Saeys, Y., Inza, I. & Larranaga, P. A review of feature selection techniques in bioinformatics. Bioinformatics 23, 2507–2517 (2007).
Moreau, Y. & Tranchevent, L. C. Computational tools for prioritizing candidate genes: boosting disease gene discovery. Nat Rev Genet 13, 523–536, 10.1038/nrg3253 (2012).
Chuang, H. Y., Lee, E., Liu, Y. T., Lee, D. & Ideker, T. Network-based classification of breast cancer metastasis. Mol Syst Biol 3, 140, 10.1038/msb4100180 (2007).
Li, J., Roebuck, P., Grunewald, S. & Liang, H. SurvNet: a web server for identifying network-based biomarkers that most correlate with patient survival data. Nucleic Acids Res 40, W123–126, 10.1093/nar/gks386 (2012).
Wu, G. & Stein, L. A network module-based method for identifying cancer prognostic signatures. Genome Biol 13, R112, 10.1186/gb-2012-13-12-r112 (2012).
Winter, C. et al. Google goes cancer: improving outcome prediction for cancer patients by network-based ranking of marker genes. PLoS Comput Biol 8, e1002511, 10.1371/journal.pcbi.1002511 (2012).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol 27, 199–204, 10.1038/nbt.1522 (2009).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat Commun 1, 34, 10.1038/ncomms1033 (2010).
Keshava Prasad, T. S. et al. Human Protein Reference Database--2009 update. Nucleic Acids Res 37, D767–772, 10.1093/nar/gkn892 (2009).
Weinstein, J. N. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet 45, 1113–1120, 10.1038/ng.2764 (2013).
Bradburn, M. J., Clark, T. G., Love, S. B. & Altman, D. G. Survival analysis part II: multivariate data analysis--an introduction to concepts and methods. Br J Cancer 89, 431–436, 10.1038/sj.bjc.6601119 (2003).
Harrell, F. E., Lee, K. L. & Mark, D. B. Multivariable Prognostic Models: Issues In Developing Models, Evaluating Assumptions And Adequacy, And Measuring And Reducing Errors. Statistics in Medicine 15, 361–387 (1996).
Aguirre-Gamboa, R. et al. SurvExpress: An Online Biomarker Validation Tool and Database for Cancer Gene Expression Data Using Survival Analysis. PLoS One 8, e74250, 10.1371/journal.pone.0074250 (2013).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America 102, 15545–15550 (2005).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57, 10.1038/nprot.2008.211 (2009).
Wagner, E. F. & Nebreda, A. R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat Rev Cancer 9, 537–549, 10.1038/nrc2694 (2009).
Hardie, D. G. & Alessi, D. R. LKB1 and AMPK and the cancer-metabolism link - ten years after. BMC Biol 11, 36, 10.1186/1741-7007-11-36 (2013).
Deroo, B. J. & Korach, K. S. Estrogen receptors and human disease. J Clin Invest 116, 561–570, 10.1172/jci27987 (2006).
Molloy, N. H., Read, D. E. & Gorman, A. M. Nerve growth factor in cancer cell death and survival. Cancers (Basel) 3, 510–530, 10.3390/cancers3010510 (2011).
Li, L. & Davie, J. R. The role of Sp1 and Sp3 in normal and cancer cell biology. Ann Anat 192, 275–283, 10.1016/j.aanat.2010.07.010 (2010).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674, 10.1016/j.cell.2011.02.013 (2011).
Yu, K. et al. A precisely regulated gene expression cassette potently modulates metastasis and survival in multiple solid cancers. PLoS Genet 4, e1000129, 10.1371/journal.pgen.1000129 (2008).
Cheng, W. Y., Ou Yang, T. H. & Anastassiou, D. Biomolecular events in cancer revealed by attractor metagenes. PLoS Comput Biol 9, e1002920, 10.1371/journal.pcbi.1002920 (2013).
Chao, M. P. et al. Calreticulin is the dominant pro-phagocytic signal on multiple human cancers and is counterbalanced by CD47. Sci Transl Med 2, 63ra94, 10.1126/scitranslmed.3001375 (2010).
Pinto-Leite, R. et al. Genomic characterization of three urinary bladder cancer cell lines: understanding genomic types of urinary bladder cancer. Tumour Biol 35, 4599–4617, 10.1007/s13277-013-1604-3 (2014).
Yu, J. et al. A novel set of DNA methylation markers in urine sediments for sensitive/specific detection of bladder cancer. Clin Cancer Res 13, 7296–7304, 10.1158/1078-0432.ccr-07-0861 (2007).
Sahlberg, S. H., Spiegelberg, D., Glimelius, B., Stenerlow, B. & Nestor, M. Evaluation of cancer stem cell markers CD133, CD44, CD24: association with AKT isoforms and radiation resistance in colon cancer cells. PLoS One 9, e94621, 10.1371/journal.pone.0094621 (2014).
Shin, S., Rossow, K. L., Grande, J. P. & Janknecht, R. Involvement of RNA helicases p68 and p72 in colon cancer. Cancer Res 67, 7572–7578, 10.1158/0008-5472.can-06-4652 (2007).
Ladner, R. D. et al. dUTP nucleotidohydrolase isoform expression in normal and neoplastic tissues: association with survival and response to 5-fluorouracil in colorectal cancer. Cancer Res 60, 3493–3503 (2000).
Chatr-Aryamontri, A. et al. The BioGRID interaction database: 2013 update. Nucleic Acids Res 41, D816–823, 10.1093/nar/gks1158 (2013).
Turinsky, A. L., Razick, S., Turner, B., Donaldson, I. M. & Wodak, S. J. Navigating the global protein-protein interaction landscape using iRefWeb. Methods Mol Biol 1091, 315–331, 10.1007/978-1-62703-691-7_22 (2014).
Franceschini, A. et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41, D808–815, 10.1093/nar/gks1094 (2013).
Arnold, P., Erb, I., Pachkov, M., Molina, N. & van Nimwegen, E. MotEvo: integrated Bayesian probabilistic methods for inferring regulatory sites and motifs on multiple alignments of DNA sequences. Bioinformatics 28, 487–494, 10.1093/bioinformatics/btr695 (2012).
Ojesina, A. I. et al. Landscape of genomic alterations in cervical carcinomas. Nature 506, 371-+, 10.1038/nature12881 (2014).
Griffith, M. et al. DGIdb: mining the druggable genome. Nat Methods 10, 1209–1210, 10.1038/nmeth.2689 (2013).
Pandey, S. N., Dixit, M., Choudhuri, G. & Mittal, B. Lipoprotein receptor associated protein (LRPAP1) insertion/deletion polymorphism: association with gallbladder cancer susceptibility. Int J Gastrointest Cancer 37, 124–128, 10.1007/s12029-007-9002-y (2006).
Chin, Y. R., Yuan, X., Balk, S. P. & Toker, A. PTEN-deficient tumors depend on AKT2 for maintenance and survival. Cancer Discov 4, 942–955, 10.1158/2159-8290.cd-13-0873 (2014).
Wakefield, A. et al. Bcl3 selectively promotes metastasis of ERBB2-driven mammary tumors. Cancer Res 73, 745–755, 10.1158/0008-5472.can-12-1321 (2013).
Bandini, S. et al. Early onset and enhanced growth of autochthonous mammary carcinomas in C3-deficient Her2/neu transgenic mice. Oncoimmunology 2, e26137, 10.4161/onci.26137 (2013).
Chien, W. et al. Expression of connective tissue growth factor (CTGF/CCN2) in breast cancer cells is associated with increased migration and angiogenesis. Int J Oncol 38, 1741–1747, 10.3892/ijo.2011.985 (2011).
Mazurek, A. et al. DDX5 regulates DNA replication and is required for cell proliferation in a subset of breast cancer cells. Cancer Discov 2, 812–825, 10.1158/2159-8290.cd-12-0116 (2012).
Holst, F. et al. Estrogen receptor alpha (ESR1) gene amplification is frequent in breast cancer. Nat Genet 39, 655–660, 10.1038/ng2006 (2007).
Mange, A. et al. Serum autoantibody signature of ductal carcinoma in situ progression to invasive breast cancer. Clin Cancer Res 18, 1992–2000, 10.1158/1078-0432.ccr-11-2527 (2012).
Langer, S. et al. Jun and Fos family protein expression in human breast cancer: correlation of protein expression and clinicopathological parameters. Eur J Gynaecol Oncol 27, 345–352 (2006).
Sum, E. Y. et al. Overexpression of LMO4 induces mammary hyperplasia, promotes cell invasion and is a predictor of poor outcome in breast cancer. Proc Natl Acad Sci USA 102, 7659–7664, 10.1073/pnas.0502990102 (2005).
Bauer, J. A. et al. Identification of markers of taxane sensitivity using proteomic and genomic analyses of breast tumors from patients receiving neoadjuvant paclitaxel and radiation. Clin Cancer Res 16, 681–690, 10.1158/1078-0432.ccr-09-1091 (2010).
Natarajan, T. G. et al. Epigenetic regulator MLL2 shows altered expression in cancer cell lines and tumors from human breast and colon. Cancer Cell Int 10, 13, 10.1186/1475-2867-10-13 (2010).
Rizki, A. et al. A human breast cell model of preinvasive to invasive transition. Cancer Res 68, 1378–1387, 10.1158/0008-5472.can-07-2225 (2008).
Moody, S. E. et al. PRKACA mediates resistance to HER2-targeted therapy in breast cancer cells and restores anti-apoptotic signaling. Oncogene 0, 10.1038/onc.2014.153 (2014).
Stratford, A. L. et al. Targeting p90 ribosomal S6 kinase eliminates tumor-initiating cells by inactivating Y-box binding protein-1 in triple-negative breast cancers. Stem Cells 30, 1338–1348, 10.1002/stem.1128 (2012).
Reinholz, M. M. et al. Differential gene expression of TGF beta inducible early gene (TIEG), Smad7, Smad2 and Bard1 in normal and malignant breast tissue. Breast Cancer Res Treat 86, 75–88, 10.1023/B:BREA.0000032926.74216.7d (2004).
Slyskova, J. et al. Functional, genetic and epigenetic aspects of base and nucleotide excision repair in colorectal carcinomas. Clin Cancer Res 18, 5878–5887, 10.1158/1078-0432.ccr-12-1380 (2012).
Zhang, J. T. et al. Downregulation of CFTR promotes epithelial-to-mesenchymal transition and is associated with poor prognosis of breast cancer. Biochim Biophys Acta 1833, 2961–2969, 10.1016/j.bbamcr.2013.07.021 (2013).
Ahmed, D. et al. A tissue-based comparative effectiveness analysis of biomarkers for early detection of colorectal tumors. Clin Transl Gastroenterol 3, e27, 10.1038/ctg.2012.21 (2012).
Bae, J. A. et al. An unconventional KITENIN/ErbB4-mediated downstream signal of EGF upregulates c-Jun and the invasiveness of colorectal cancer cells. Clin Cancer Res 20, 4115–4128, 10.1158/1078-0432.ccr-13-2863 (2014).
Nagy, N. et al. Galectin-8 expression decreases in cancer compared with normal and dysplastic human colon tissue and acts significantly on human colon cancer cell migration as a suppressor. Gut 50, 392–401 (2002).
Slattery, M. L., Lundgreen, A., Herrick, J. S. & Wolff, R. K. Genetic variation in RPS6KA1, RPS6KA2, RPS6KB1, RPS6KB2 and PDK1 and risk of colon or rectal cancer. Mutat Res 706, 13–20, 10.1016/j.mrfmmm.2010.10.005 (2011).
Grabowski, P. et al. Heterogeneous expression of neuroendocrine marker proteins in human undifferentiated carcinoma of the colon and rectum. Ann N Y Acad Sci 1014, 270–274 (2004).
Spisak, S. et al. Applicability of antibody and mRNA expression microarrays for identifying diagnostic and progression markers of early and late stage colorectal cancer. Dis Markers 28, 1–14, 10.3233/dma-2010-0677 (2010).
Maldonado, V. & Melendez-Zajgla, J. Role of Bcl-3 in solid tumors. Mol Cancer 10, 152, 10.1186/1476-4598-10-152 (2011).
Weber, R. G., Rieger, J., Naumann, U., Lichter, P. & Weller, M. Chromosomal imbalances associated with response to chemotherapy and cytotoxic cytokines in human malignant glioma cell lines. Int J Cancer 91, 213–218 (2001).
Romao, L. F. et al. Connective tissue growth factor (CTGF/CCN2) is negatively regulated during neuron-glioblastoma interaction. PLoS One 8, e55605, 10.1371/journal.pone.0055605 (2013).
Nicolasjilwan, M. et al. Addition of MR imaging features and genetic biomarkers strengthens glioblastoma survival prediction in TCGA patients. J Neuroradiol, 10.1016/j.neurad.2014.02.006 (2014).
Russo, A. & O’Bryan, J. P. Intersectin 1 is required for neuroblastoma tumorigenesis. Oncogene 31, 4828–4834, 10.1038/onc.2011.643 (2012).
Qiang, L. et al. Isolation and characterization of cancer stem like cells in human glioblastoma cell lines. Cancer Lett 279, 13–21, 10.1016/j.canlet.2009.01.016 (2009).
Beghini, A. et al. The neural progenitor-restricted isoform of the MARK4 gene in 19q13.2 is upregulated in human gliomas and overexpressed in a subset of glioblastoma cell lines. Oncogene 22, 2581–2591 (2003).
Chang, C. C. et al. Connective tissue growth factor activates pluripotency genes and mesenchymal-epithelial transition in head and neck cancer cells. Cancer Res 73, 4147–4157, 10.1158/0008-5472.can-12-4085 (2013).
Rampias, T. et al. RAS/PI3K crosstalk and cetuximab resistance in head and neck squamous cell carcinoma. Clin Cancer Res 20, 2933–2946, 10.1158/1078-0432.ccr-13-2721 (2014).
Menendez, S. T. et al. Frequent aberrant expression of the human ether a go-go (hEAG1) potassium channel in head and neck cancer: pathobiological mechanisms and clinical implications. J Mol Med (Berl) 90, 1173–1184, 10.1007/s00109-012-0893-0 (2012).
Muro-Cacho, C. A., Rosario-Ortiz, K., Livingston, S. & Munoz-Antonia, T. Defective transforming growth factor beta signaling pathway in head and neck squamous cell carcinoma as evidenced by the lack of expression of activated Smad2. Clin Cancer Res 7, 1618–1626 (2001).
Tan, W. et al. Role of inflammatory related gene expression in clear cell renal cell carcinoma development and clinical outcomes. J Urol 186, 2071–2077, 10.1016/j.juro.2011.06.049 (2011).
Penzvalto, Z. et al. Identifying resistance mechanisms against five tyrosine kinase inhibitors targeting the ERBB/RAS pathway in 45 cancer cell lines. PLoS One 8, e59503, 10.1371/journal.pone.0059503 (2013).
Spitz, M. R. et al. Variants in inflammation genes are implicated in risk of lung cancer in never smokers exposed to second-hand smoke. Cancer Discov 1, 420–429, 10.1158/2159-8290.cd-11-0080 (2011).
Chang, C. C. et al. Connective tissue growth factor and its role in lung adenocarcinoma invasion and metastasis. J Natl Cancer Inst 96, 364–375 (2004).
Lu, Y. et al. A gene expression signature predicts survival of patients with stage I non-small cell lung cancer. PLoS Med 3, e467, 10.1371/journal.pmed.0030467 (2006).
Karachaliou, N. et al. BRCA1, LMO4 and CtIP mRNA expression in erlotinib-treated non-small-cell lung cancer patients with EGFR mutations. J Thorac Oncol 8, 295–300, 10.1097/JTO.0b013e31827db621 (2013).
Meng, H. et al. Stromal LRP1 in lung adenocarcinoma predicts clinical outcome. Clin Cancer Res 17, 2426–2433, 10.1158/1078-0432.ccr-10-2385 (2011).
Kobayashi, K. et al. Identification of genes whose expression is upregulated in lung adenocarcinoma cells in comparison with type II alveolar cells and bronchiolar epithelial cells in vivo. Oncogene 23, 3089–3096, 10.1038/sj.onc.1207433 (2004).
Jin, Y. et al. NEDD9 promotes lung cancer metastasis through epithelial-mesenchymal transition. Int J Cancer 134, 2294–2304, 10.1002/ijc.28568 (2014).
Lara, R. et al. An siRNA screen identifies RSK1 as a key modulator of lung cancer metastasis. Oncogene 30, 3513–3521, 10.1038/onc.2011.61 (2011).
Szymanowska-Narloch, A. et al. Molecular profiles of non-small cell lung cancers in cigarette smoking and never-smoking patients. Adv Med Sci 58, 196–206, 10.2478/ams-2013-0025 (2013).
Rolle, C. E. et al. Combined MET inhibition and topoisomerase I inhibition block cell growth of small cell lung cancer. Mol Cancer Ther 13, 576–584, 10.1158/1535-7163.mct-13-0109 (2014).
Kikuchi, R. et al. Promoter hypermethylation contributes to frequent inactivation of a putative conditional tumor suppressor gene connective tissue growth factor in ovarian cancer. Cancer Res 67, 7095–7105, 10.1158/0008-5472.can-06-4567 (2007).
Darb-Esfahani, S. et al. Estrogen receptor 1 mRNA is a prognostic factor in ovarian carcinoma: determination by kinetic PCR in formalin-fixed paraffin-embedded tissue. Endocr Relat Cancer 16, 1229–1239, 10.1677/erc-08-0338 (2009).
Neyns, B. et al. Expression of the jun family of genes in human ovarian cancer and normal ovarian surface epithelium. Oncogene 12, 1247–1257 (1996).
Wang, H. et al. NEDD9 overexpression is associated with the progression of and an unfavorable prognosis in epithelial ovarian cancer. Hum Pathol 45, 401–408, 10.1016/j.humpath.2013.10.005 (2014).
Isaksson, H. S., Sorbe, B. & Nilsson, T. K. Whole genome expression profiling of blood cells in ovarian cancer patients -Prognostic impact of the CYP1B1, MTSS1, NCALD and NOP14. Oncotarget 5, 4040–4049 (2014).
Wang, Y. et al. Ras-induced epigenetic inactivation of the RRAD (Ras-related associated with diabetes) gene promotes glucose uptake in a human ovarian cancer model. J Biol Chem 289, 14225–14238, 10.1074/jbc.M113.527671 (2014).
Davies, S. et al. Effects of bevacizumab in mouse model of endometrial cancer: Defining the molecular basis for resistance. Oncol Rep 25, 855–862, 10.3892/or.2011.1147 (2011).
Li, L. & Guan, K. L. Microtubule-associated protein/microtubule affinity-regulating kinase 4 (MARK4) is a negative regulator of the mammalian target of rapamycin complex 1 (mTORC1). J Biol Chem 288, 703–708, 10.1074/jbc.C112.396903 (2013).
Pressinotti, N. C. et al. Differential expression of apoptotic genes PDIA3 and MAP3K5 distinguishes between low- and high-risk prostate cancer. Mol Cancer 8, 130, 10.1186/1476-4598-8-130 (2009).
Ito, R. et al. Usefulness of tyrosine hydroxylase mRNA for diagnosis and detection of minimal residual disease in neuroblastoma. Biol Pharm Bull 27, 315–318 (2004).
Acknowledgements
We thank CONACyT and Tecnológico de Monterrey CAT220 for funds. We also thank Erica Goodoff, Guy Cardineau, Sean Scott and Raquel Cuevas for writing editions. The results shown here are in whole or part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/.
Author information
Authors and Affiliations
Contributions
E.M. and V.T. designed the study and analyzed the data. R.G.W.V. contributed data, materials, analysis tools and discussions. E.M. and V.T. wrote the manuscript with contributions from R.G.W.V. All authors commented and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Martinez-Ledesma, E., Verhaak, R. & Treviño, V. Identification of a multi-cancer gene expression biomarker for cancer clinical outcomes using a network-based algorithm. Sci Rep 5, 11966 (2015). https://doi.org/10.1038/srep11966
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11966
This article is cited by
-
TFRC, associated with hypoxia and immune, is a prognostic factor and potential therapeutic target for bladder cancer
European Journal of Medical Research (2024)
-
HBS–STACK: hierarchical biomarker selection and stacked ensemble model for biomarker identification and cancer prediction on multi-omics
Neural Computing and Applications (2024)
-
Debiased inference for heterogeneous subpopulations in a high-dimensional logistic regression model
Scientific Reports (2023)
-
Host-response transcriptional biomarkers accurately discriminate bacterial and viral infections of global relevance
Scientific Reports (2023)
-
A novel focal adhesion-related risk model predicts prognosis of bladder cancer —— a bioinformatic study based on TCGA and GEO database
BMC Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.