Abstract
Identification of mutants with impairments in auxin biosynthesis and dynamics by forward genetic screening is hindered by the complexity, redundancy and necessity of the pathways involved. Furthermore, although a few auxin-deficient mutants have been recently identified by screening for altered responses to shade, ethylene, N-1-naphthylphthalamic acid (NPA) or cytokinin (CK), there is still a lack of robust markers for systematically isolating such mutants. We hypothesized that a potentially suitable phenotypic marker is root curling induced by CK, as observed in the auxin biosynthesis mutant CK-induced root curling 1 / tryptophan aminotransferase of Arabidopsis 1 (ckrc1/taa1). Phenotypic observations, genetic analyses and biochemical complementation tests of Arabidopsis seedlings displaying the trait in large-scale genetic screens showed that it can facilitate isolation of mutants with perturbations in auxin biosynthesis, transport and signaling. However, unlike transport/signaling mutants, the curled (or wavy) root phenotypes of auxin-deficient mutants were significantly induced by CKs and could be rescued by exogenous auxins. Mutants allelic to several known auxin biosynthesis mutants were re-isolated, but several new classes of auxin-deficient mutants were also isolated. The findings show that CK-induced root curling provides an effective marker for discovering genes involved in auxin biosynthesis or homeostasis.
Similar content being viewed by others
Introduction
Auxin plays key roles in the regulation of plant growth and development. The predominant natural form is indole-3-acetic acid (IAA). However, various other small organic acids, both natural (e.g. 4Cl-IAA and phenylacetic acid) and synthetic (e.g. 2,4-dichlorophenoxyacetic acid and 1-naphthaleneacetic acid; 2,4-D and NAA, respectively) also exhibit auxin-like activities1.
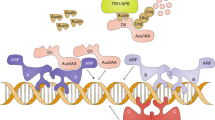
In plants, IAA is produced by multiple highly connected biosynthetic pathways2,3. It is then transported between cells by various efflux and influx carrier proteins that create so-called auxin maxima and concentration gradients, which play important regulatory roles in embryogenesis and organogenesis4. In planta, auxin signaling is initiated by auxin binding to receptors such as TRANSPORT INHIBITOR RESPONSE1/AUXIN SIGNALING F-BOX (TIR1/AFB) receptors5, AUXIN BINDING PROTEIN 1 (ABP1)6,7 and the Arabidopsis cell cycle F-box protein S-phase kinase-associated protein 2A (SKP2A)8. The best characterized TIR1/AFB-Aux/IAA co-receptor system regulates auxin-dependent transcription within the nucleus and auxin binding enables SKP2A to mediate degradation of transcription factors that repress cell division, such as E2-promoter binding factor C (E2FC) and E2F dimerization partner B (DPB)8. Despite the obvious auxin binding activity of ABP1 and its reported involvement in the so-called fast, nontranscriptional cellular responses and some developmental processes6,7,9, recent work with abp1 null mutants suggests that ABP1 is not essential for Arabidopsis development under normal conditions10.
Auxin biosynthesis is less well understood than auxin transport and signaling, but includes several tryptophan (Trp)-dependent and -independent pathways11. Four Trp-dependent pathways have been proposed, named according to the major putative intermediates: the indole-3-pyruvic acid (IPA), indole-3-acetamide (IAM), tryptamine (TAM) and indole-3-acetaldoxime (IAOx) pathways3. Only the IPA pathway has been completely characterized at genetic and biochemical levels to date. It involves two reactions that convert L-Trp to IAA, sequentially catalyzed by TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS 1 (TAA1) and YUCCA (YUC) flavin monooxygenases12,13,14. The genetic bases, potential intermediates and significance for IAA biosynthesis (at least in Arabidopsis) of the Trp-independent pathway(s) are all uncertain15.
The multiple interconnected biosynthesis pathways and their high genetic redundancy complicate efforts to detect auxin-deficient mutants (which would facilitate dissection of the pathways) via direct genetic screening. A further hindrance is the lack of a reliable phenotypic marker for screening auxin-deficient mutants. Some genes involved in auxin biosynthesis have been identified by isolating mutants with high auxin contents16,17,18,19 or characterizing mutants isolated in studies that initially focused on indirectly associated phenomena13,14,20,21,22. However, only a few auxin-deficient mutants have been detected by such indirect screens2,14 and there is still a lack of a systematic forward genetic screening procedure for isolating auxin-deficient mutants3,23.
We hypothesized that a potentially useful marker may be root curling induced by cytokinin (CK), as recently described in the auxin biosynthesis mutant CK-induced root curling 1 / tryptophan aminotransferase of Arabidopsis 1 (ckrc1/taa1), arising from auxin deficiencies in the root tip20. To test the possibility that this trait could be used to isolate more auxin-deficient mutants, we screened large collections of Arabidopsis T-DNA insertion lines, obtained from the Arabidopsis Biological Resource Center (ABRC), to isolate ckrc-like mutants. The advantage of Agrobacterium T-DNA over classical mutagens is that the plant sequences flanking the insertion site can be isolated easily. This simplifies the identification of genes corresponding to interesting mutants. A number of pools of T-DNA insertion lines are now available with the coverage of the whole genome24,25. Our results show that the trait can be effectively applied to isolate mutants with perturbations in auxin biosynthesis, transport and signaling. Furthermore, mutants with auxin-deficiencies can be rescued by exogenous auxins. Mutants carrying mutations in several previously identified biosynthesis genes were re-isolated and some previously unknown auxin-deficient mutants with significantly reduced endogenous auxin levels were also isolated. The results clearly suggest that the screening system is effective for isolating auxin-deficient mutants and hence discovering genes that participate in auxin biosynthesis or dynamics.
Results
Forward genetic screen for ckrc mutants.
To date few auxin-deficient mutants have been directly isolated by forward genetic screening14, partly because of a lack of suitable markers. However, we have previously shown that CK up-regulates local auxin biosynthesis in root tips but inhibits polar auxin transport (PAT) and exogenous CKs induce strong root curling in the auxin biosynthesis mutant ckrc1/taa120. To test the hypothesis of CK-induced root curling (ckrc) could be used as a marker for detecting other auxin-deficient mutants, we first applied CK to two other Arabidopsis mutants, weak ethylene insensitive 2 (wei2) and wei7. These mutants have reduced auxin levels due to mutations in genes encoding α- and β-subunits, respectively, of anthranilate synthase, a rate-limiting L-Trp biosynthetic enzyme22. We measured their Degrees of Curling (DC) to compare them with those of WT and ckrc1 plants. As expected, when grown vertically on Murashige and Skoog (MS) medium in the presence of 0.1 μM trans-Zeatin (tZ), their primary roots curled, although the phenotype of wei2 roots was relatively weak (Fig. 1a,b).
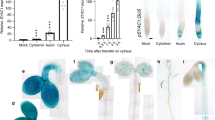
Comparison of 7 DAG root phenotypes between WT and indicated auxin mutants.
(a) Effect of tZ on indicated mutants. (b) The Degree of root Curling (DC) was calculated by dividing the distance between the two ends of seedlings’ roots (L0) by the length of their roots (L). DC of roots of WT seedlings and indicated mutants in the presence and absence of 0.1 μM tZ. Presented data are means ± SD (n = 30). Asterisks indicate statistically significant differences between Mock- and 0.1 μM tZ-treated seedlings according to t-tests (*, ** and *** correspond to P-values of 0.05 > p > 0.01, 0.01 > p > 0.001 and p < 0.001, respectively).
We also tested auxin transport mutants auxin resistant 1 (aux1) and pin-formed 2 (pin2), which have impairments in auxin influx and efflux transport, respectively and both reportedly have defective root gravitropic responses26,27,28,29. In contrast to the auxin-deficient mutants ckrc1, wei2 and wei7, the strong curling of roots of aux1 and pin2 mutants did not depend on the presence of tZ, thus displaying a constitutive root curling (crc) phenotype (Fig. 1a & Table 1). However, roots of pin1, pin3, pin4 and pin7 auxin efflux mutants showed no obvious curling on either tZ-containing or basal MS medium (Fig. 1a). The auxin signaling mutant tir1, defective in an auxin receptor, displayed weak wavy root rather than a curling root phenotype, which was not significantly affected by 0.1 μM tZ (Fig. 1a). Together, these results suggest that the ckrc phenotype differs from those of auxin transport and signaling mutants and may allow large-scale genetic screens for auxin-deficient mutants, which have not been previously reported3.
To optimize screening conditions, ckrc1, wei2 and wei7 mutant seeds were germinated and grown on MS plates with tZ in a range of concentrations and then their root phenotypes were scored 7 days after germination (DAG). Root curling was most distinct at 0.1–1 μM tZ and became weak or masked by growth inhibition at higher tZ concentrations (Supplementary Fig. S1). We chose 0.1 μM tZ as the general screening concentration (for some mutants with weak responses <0.1 μM tZ was better, see below). In its presence, mutants with either curling or wavy root phenotypes were isolated by a two-round screening procedure (putative mutants detected in the first round were confirmed by the second) (Supplementary Fig. S2 & Table 2). In an initial large-scale application of this procedure to two collections of Arabidopsis lines, CS76502/4/6/8 (40,000 lines)24 and CS31100 (62,000 lines)25, we isolated 53 mutants with curled or wavy roots. A further 96 mutants were subsequently isolated after screening the other nine stocks listed in Table 2. In the following sections we present results of genetic and phenotypic characterizations of the first set of 53 isolated mutants. Similar investigations of the other 96 mutants are ongoing and some significant results obtained from their analyses are also noted below.
When grown on MS without tZ, the curling root phenotype of some of the isolated mutants was maintained. These were designated group I, constitutive root curling (crc), mutants. In most of the others the trait was either significantly weaker or disappeared. These were designated group II, CK-induced root curling (ckrc), mutants. A third group consisted of two CK-induced root waving (ckrw) mutants, ckrw1 and ckrw2, whose phenotypes were most distinct at 0.005 and 0.01 μM tZ concentrations, respectively (Fig. 2). On MS without tZ, the ckrw1 phenotype became weak (displaying a longer ‘wavelength’ with a lower ‘frequency’); while the ckrw2 phenotype disappeared (Table 1).
Biochemical complementation tests of the 7 DAG curled/wavy root mutants and WT controls.
Seedlings were grown on MS plates with and without indicated combinations of tZ, auxins and L-Trp. Presented data obtained in the DC analysis are means ± SD (n = 30). Different letters indicate significant differences at P < 0.05, according to ANOVA followed by Tukey’s multiple comparison tests.
Genetic analysis of the progenies produced from backcrosses of these isolated mutants to their respective WTs revealed that WT complemented all of them in the F1 generation and showed 1:3 segregation ratios in the F2 generation, indicating that they carried mutations in single recessive genes (Supplementary Table S1).
Biochemical complementation and allelism tests
The curled root phenotype of auxin-deficient mutants can be rescued by exogenous auxins20 and both influx and efflux transport mutants display differential sensitivities to different kinds of auxins30,31. Moreover, L-Trp can rescue auxin-deficient mutants that are defective in L-Trp biosynthesis22 (Table 1), but not true auxin-biosynthesis mutants, i.e. mutants with perturbations downstream of L-Trp production20. To primarily distinguish between these possibilities, we performed biochemical complementation and genetic allelic tests.
Mutant seedlings were grown on vertical MS/tZ plates with selected auxins or L-Trp and their root phenotypes were observed. The results showed that the group I crc mutants could be divided into two subgroups: one only rescued by NAA and the other only by 2,4-D, which are typical traits of the influx carrier mutant aux1 and the efflux carrier mutant pin2, respectively (Fig. 2 & Table 1). Allelism tests revealed that their mutations are indeed allelic to aux1 or pin2 (Table 1 & Supplementary Table S1).
Unlike group I, the group II ckrc mutants can be rescued by all three of the auxins used, as previously reported20, although often only partly by 2, 4-D. However, they could be divided into two classes by their responses to L-Trp (Table 1). Two currently known auxin-deficient mutants, wei2 and wei7, can be rescued by L-Trp22, but not ckrc1/taa1 (Fig. 2)13,20. Thus, we performed allelism tests by genetic crossing with these three known mutants. The results showed that some group II mutants are allelic to ckrc1, wei2 or wei7 mutants (Table 1). However, one group II mutant is not allelic to any of these three known mutants and was renamed ckrc2 (Fig. 2 & Table 1). Three such group II mutants were also identified among the 96 subsequently isolated mutants (Supplementary Table S2) and gene cloning confirmed that they carry mutations in genes that have not been identified as mutated in any previously characterized auxin-deficient mutants.
The wavy-root mutants ckrw1 and ckrw2 showed different responses in the biochemical complementation tests. While all three auxins could rescue ckrw2, like the group II mutants, only 2, 4-D had rescuing effects on ckrw1 mutants (Fig. 2 & Table 1). Allelism tests showed that these two mutants are not allelic to taa1, wei2, wei7, aux1 or pin2, suggesting that they represent new classes of auxin mutants.
IAA can stimulate root growth and restore root gravitropic responses of ckrc2, ckrw1 and ckrw2 at low concentrations
As shown above, exogenous auxins rescued phenotypes of ckrc2 and ckrw2 mutants in the biochemical complementation tests (Table 1), corroborating their putative auxin-deficiency. Roots of auxin-deficient mutants usually display positive growth responses to exogenous IAA at low concentrations, in contrast to negative responses of WT plants and weak gravitropic responses20. In further tests reported here, we found that the aux1 auxin influx carrier mutant and tir1 auxin signaling mutant were significantly less sensitive than WT to IAA in a root growth inhibition assay, while responses of the pin2 auxin efflux carrier mutant were more similar to WT (Fig. 3a). Like the ckrc1 mutant20, at low concentration IAA stimulates rather than inhibits the root growth of ckrc2, ckrw1 and ckrw2 mutants (Fig. 3b,c). Accordingly, 0.01 μM IAA also rescued the defective or weak gravitropic responses of ckrc2 and ckrw2 mutants (Fig. 3e,g). In contrast, exogenous IAA did not restore gravitropism in aux1 and pin2 mutants (Fig. 3d). Paradoxically, 0.01 μM IAA stimulated ckrw1 root growth (Fig. 3c) and partially rescued its gravitropic response (Fig. 3f), but had little effect on its wavy-root phenotype (Fig. 2).
Effects of exogenous IAA on WT seedlings and indicated auxin-mutants.
(a–c) Primary root elongation measured on 7 DAG; means ± SD (n > 20). Different letters indicate significant differences at P < 0.05 according to ANOVA followed by Tukey’s multiple comparison tests. (d–g) Exogenous IAA can restore root gravitropism in ckrc or ckrw mutants. Approximately 80 seedlings were assessed per genotype or treatment.
ckrc2, ckrw1 and ckrw2 respond normally to IAA and tZ
To see if auxin or CK signaling was affected in ckrc2, ckrw1 and ckrw2 mutants, we analyzed their expression of two auxin (IAA1/2) and two CK (ARR5/15) primary response genes after treatment with these hormones by real-time quantitative PCR (qRT-PCR). As shown in Fig. 4, although the gene response strengths differed among them, none of the mutants showed significantly weaker responses than WT plants, implying that they were not defective in auxin or CK signaling.
Induction of expression of auxin/CK primary response genes by exogenous IAA
(a, b) or tZ (c, d) treatments in WT and indicated mutants. Transcript levels in treated seedlings relative to untreated controls, normalized to ACT-8 mRNA levels. Presented data are means ± SD (n = 4). Different letters indicate significant differences at P < 0.05 according to ANOVA followed by Tukey’s multiple comparison tests.
Endogenous auxin contents are low in ckrc and ckrw mutants
As three tested mutants showed traits associated with auxin-deficiency, such as weak gravitropic responses and positive root growth responses to IAA at low concentrations, we directly measured their endogenous contents of free IAA and its metabolites by LC-MRM-MS32. Like the auxin-biosynthesis mutant ckrc1/taa1, significant overall reductions in levels of both free IAA and most of its metabolites (oxIAA, IAGlu and IAAsp) were observed, especially in ckrc2 mutants (Fig. 5a), suggesting that they are probably true auxin deficient mutants. Reduced levels of free IAA were also detected in the subsequently isolated ckrc3, ckrc4 and ckrc5 mutants (Fig. 5b).
Endogenous levels of IAA and its metabolites.
The data were measured in 7-d-old mutant and respective WT seedlings; means ± SD (n = 4). (a) Different letters indicate significant differences at P < 0.05 according to ANOVA followed by Tukey’s multiple comparison tests. (b) Asterisks indicate statistically significant differences between the mutants and WT according to t-tests (*, ** and *** correspond to P-values of 0.05 > p > 0.01, 0.01 > p > 001 and p < 0.001, respectively). oxIAA, 2-oxindole-3-acetic acid; IAGlu, IAA-glutamate; IAAsp, IAA-aspartate.
Flanking sequences and/or genetic mapping
In order to identify the mutated genes, we used Tail-PCR to amplify the T-DNA flanking sequences or high-throughput sequencing to identify the inserted genes. Overall, 1-5 T-DNA independent insertions were identified per mutant, in accordance with the frequent presence of more than one insertion in T-DNA transformants.
Three T-DNA insertions were identified in ckrw1, but only one (in AT5G49665, WAVY GROWTH 3, WAV333), showed genetic linkage to this mutant and genetic crossing confirmed that it was allelic to wav3.
The T-DNA insertions in ckrc2 and ckrc3 were not genetically linked to their curled root phenotypes. However, map-based cloning located the ckrc2 mutation in the region between the T5F17 and F16A16 markers in chromosome 4 and the ckrc3 mutation between MCK7 and MZN1 in chromosome 5. AT4G28720 (YUC834) and AT5G58450 (TRANSCURVATA2, TCU235) genes are located in these regions and allelism tests confirmed that ckrc2 and ckrc3 are allelic to yuc8 and tcu2, respectively.
In the ckrc5 mutant, no T-DNA insertion was detected either, but genomic DNA sequencing revealed a deletion in the gene AT3G02260 (TIR3/BIG36) and allelism tests confirmed that it is allelic to tir3/big.
Discussion
We have established an effective genetic screening protocol for isolating auxin-deficient mutants by using CK-induced root curling (ckrc) or root waving (ckrw) as a phenotypic marker. Substantial numbers of mutants have been identified using the protocol. Most are allelic to known mutants with perturbations in auxin transport genes (aux1 or pin2 in group I crc mutants) or biosynthesis (taa1/ckrc1 and the two L-Trp biosynthetic mutants wei2 and wei7). However, six (ckrc2, ckrw1, ckrw2, ckrc3, ckrc4 and ckrc5) are not allelic to any known auxin-deficient mutants in previous forward genetic screens (Tables 1 & Supplementary Table S2). Notably, all of the isolated mutants that are allelic to known biosynthetic mutants are group II ckrc mutants (Tables 1 & Supplementary Table S2). Thus, this group appears to consist of (or is enriched in) auxin-deficient mutants with perturbations in auxin biosynthesis or homeostasis. Notably, no allele of known CK or other hormonal mutants was isolated in our screen, suggesting that the curled/wavy root phenotype is specifically related to mutations affecting auxin-mediated processes. This is similar to tir mutations, which reportedly affect auxin signaling (tir1/5), transport (tir3) or biosynthesis (tir2/7)21,37, but different from wei and sav mutations, which include perturbations in the ethylene receptor ERS (wei4), the EIN3-related transcription factor gene EIL1 (wei5)38, a C-22 hydroxylase involved in brassinosteroid biosynthesis (sav1) and a β-tubulin isoform (sav2)14.
In the reported large-scale genetic screen we isolated auxin-deficient mutants that are not allelic to any previously detected in forward genetic screens. They were not defective in auxin/CK signaling responses (Fig. 4), but displayed some typical auxin-deficiency traits, such as weak gravitropic responses (Fig. 3e–g), positive root growth responses to auxin at low concentrations (Fig. 3c) and reduced levels of endogenous IAA and its metabolites (Fig. 5). This is consistent with recent observations of overall reductions in levels of IAA, oxIAA, IAAsp and IAGlu in auxin-biosynthesis mutants (and increases in auxin over-production mutants)32,39.
Our genetic mapping and allelism tests showed that ckrc2 is allelic to yuc8. The YUC flavin monooxygenases were initially proposed to catalyze conversion of tryptamine to N-hydroxytryptamine, but are now believed to convert IPA to IAA12,40. Strong functional redundancy has been found among YUCs34,41 and no single yuc mutant has been previously detected in forward genetic screens. Thus, the successful isolation of ckrc2/yuc8 reveals the utility of the ckrc marker for isolating auxin-deficient mutants. Moreover, the CK-associated root phenotypes of ckrc2 mutants suggest that YUC8 participates in the mediation of root responses to CK. Hence, YUC8 may play a more important general signaling role than previously suspected, as recent reverse genetic evidence indicates that it also mediates jasmonic acid (JA) signaling and hypocotyl growth responses to temperature changes42,43.
The ckrw1 and ckrw2 mutants showed phenotypic deviation (wavy rather than curling roots) from the typical group II ckrc mutants, but their characteristic traits could still be induced or intensified by CK. Our genetic and molecular analyses indicate that ckrw1 is an allele of WAV3, which encodes a RING-containing protein with E3 ubiquitin ligase activity in vitro33. The inability of IAA fully rescue ckrw1 in biochemical complementation tests, despite their reduced endogenous IAA levels (Fig. 5), suggests that this mutant is probably not simply auxin-deficient. Auxin signaling or transport may be also affected in it, as suggested by Sakai et al. (2012)33.
Identification of ckrc3 and ckrc5 as alleles of tcu235 and tir3/big36,37, respectively and their auxin-deficiencies revealed in our screen suggests that these two genes participate in auxin homeostasis. The TCU2 gene encodes the auxiliary subunit of the NatB N-α-acetyltransferase complex, which is required for N-α-terminal acetylation of proteins in Arabidopsis. Mutation of TCU2 causes pleiotropic perturbations in leaf, flower and seed development35. The auxin-deficiency of the ckrc3/tcu2 mutant revealed in our study implies that NatB-mediated N-α-terminal acetylation is required for the maintenance of proper auxin levels.
Previous investigations indicate that TIR3/BIG participates in positioning of auxin efflux carriers at the plasma membrane and thus in polar auxin transport36,37,44. Hence, its mutation may affect diverse hormone and light responses45. Its precise role in PAT is still obscure, but our biochemical complementation results and auxin measurements suggest that BIG may also be involved in auxin homeostasis. It should be noted that similar features, such as low DR5-GUS activities in auxin biosynthesis sites at root tips or young cotyledon/leaf margins have also been reported46,47,48. Moreover, auxin itself can affect the cellular location of its carrier proteins6,44 and reduced basipetal auxin transport has been reported in auxin biosynthesis mutant ckrc120.
Similarly to auxin-deficient mutants previously detected in forward genetic screens, the ckrc/ckrw mutants we isolated are weak, in the sense that they have reduced auxin levels and their mutations have no severe growth or developmental effects. They were able to complete their life cycle and retained fertility under normal growth conditions. More serious perturbations, such as defects in embryogenesis and radically abnormal development with infertility, have only been reported in double or multiple mutants of auxin biosynthesis genes13, clearly suggesting strong functional redundancy.
The characteristic traits of the mutants isolated from the activation-tagged T-DNA lines25 in this study were recessive, indicating that they were caused by loss- rather than gain-of-gene-function mutations. The high frequencies of aux1/ckrc1/pin2 mutants may be explained by their relatively strong phenotypes (making them easy to identify in screens). Mutants with weaker phenotypes are inevitably more difficult to detect, but may be distinguishable under subtle changes in growth conditions. Thus, with further modifications of the selection conditions (such as variations in CK concentration and/or temperature) and screening more mutants, including sets of chemically/physically induced mutants, we may approach saturated (genomic scale) screening for auxin-deficient mutants.
Methods
Plant material and growth conditions
Collections of Arabidopsis thaliana T-DNA insertion lines (Table 2) were purchased from the Arabidopsis Biological Resource Center (ABRC) (http://abrc.osu.edu/), then germinated and cultivated, as previously described20, at 25 °C with a 16 h light / 8 h dark cycle. For growth analyses, seedlings were grown on vertical plates of MS medium solidified with 1.0% w/v agar (hereafter MS plates) supplemented with 10 g/L sucrose.
Arabidopsis thaliana accessions Col-0, 2, 6, 7, C24 and Ws-2 were used as WT. Control wei2 (wei2-1/DR5::GUS) (N16397), wei7 (wei7-2/DR5::GUS) (N16436), aux1 (aux1-7/DR5::GUS) (N16704), pin2 (eir1-1/DR5::GUS) (N16706), pin1 (N5220), pin3 (N9363), pin4 (N9368), pin7 (N9365) and tir1-1 (N3798) mutants were purchased from the Nottingham Arabidopsis Stock Centre (NASC).
Mutant screening and genetic analysis
For the mutant screening, seeds from the T-DNA insertion lines were surface-sterilized and germinated on MS plates containing 0.1 μM tZ (Sigma, http://www.sigmaaldrich.com). Putative mutant seedlings (M1) with curled/wavy roots were identified 10–14 days later. They were planted in soil to harvest seeds (M2) for the second round of screening (Supplementary Fig. S2) and the confirmed mutants were subjected to further genetic analysis.
For genetic analysis, mutants were backcrossed to their respective WTs and the F1 phenotypes were observed. Genetic segregation ratios were calculated from observations of F2 generations.
For genetic allelic tests, the known and newly isolated mutants were reciprocally crossed and the phenotypes of the resulting F1 plants were determined.
Phenotypic characterization
The Degree of root Curling (DC) was calculated by dividing the distance between the two ends of seedlings’ roots (L0) by the length of their roots (L). For biochemical complementation assays, mutant seedlings were grown on vertical MS/tZ plates with various auxins or L-Trp (Sangon, http://www.sangon.com), then their root phenotypes were observed.
For root growth inhibition assays, seedlings were germinated and grown on vertical MS plates supplemented with IAA (Sigma) at a selected range of concentrations for 7 days then their root elongation was measured20. All presented data for these assays are means obtained from three separate experiments, each with at least 20 seedlings.
For root gravitropic assays, seeds were germinated and grown on vertical MS plates at 25 °C. Five days later, the seedlings were transferred to fresh media containing no hormones, IAA, tZ or both at concentrations indicated in Fig. 3d–g. Three hours later, the plates were rotated through 90° and their root bending angle was measured after further incubation for sufficiently long to generate a measurable bend (>12 h, depending on the root growth rates. Approximately 80 seedlings were assessed per genotype or treatment.) The frequencies (%) of root growth direction at intervals of 15° are represented by the lengths of the bars.
Gene cloning and sequence analysis
Tail-PCR was used to clone the T-DNA flanking sequences in the isolated mutants49. All PCR products were electrophoretically separated in 1% agarose gel and the expected TAIL-3 products were purified and sequenced. DNA sequences were aligned with blastn (http://www.ncbi.nlm.nih.gov/BLAST/) and Tair10 (http://www.arabidopsis.org/Blast/) programs.
For the putative new mutants (see Results), T-DNA flanking sequences were also detected by high-throughput sequencing (at ShangHai Biotechnology Corporation, China, http://www.shbiotech.org/ and at Hangzhou Guhe Information and Technology Co., Ltd, China, http://www.guheinfo.com/). If no linkage was found between any detected T-DNA inserts and a mutant’s phenotype, map-based cloning was performed using F2 populations generated by backcrossing the mutants to their respective WTs, as previously described50.
RNA preparation and expression analysis
Arabidopsis seedlings were immediately frozen in liquid nitrogen and stored at −80 °C. RNA was isolated using Trizol (Sangon) and reverse-transcribed using a reverse transcription kit (DRR047A) (Takara, http://www.takara-bio.com/). Quantitative RT-PCR was performed using a Bio-Rad CFX96TM Real-time System (Bio-Rad, http://www.bio-rad.com) and Power SYBR green chemistry (DRR081A) (Takara), with primers listed in Supplementary Appendix S1.
CK- and auxin-inducible gene expression was analyzed in 7-day-old seedlings grown on MS medium, as previously described20, using treatments consisting of exposure to 10 μM tZ for 30 min51 or 20 μM IAA for 1.5 h52 in liquid MS medium. There were four independent biological replicates per treatment.
Endogenous auxin measurement
Endogenous levels of free IAA and its metabolites were measured using previously described LC-MS/MS methods32. Briefly, samples (20 mg fresh weight) of 10-day-old Arabidopsis seedlings were collected, extracted in ice-cold 50 mM sodium phosphate buffer (pH 7) and purified by SPE on hydrophilic-lipophilic balanced reversed-phase sorbent columns (Oasis® HLB, 1 cc/30 mg, Waters). To each extract, 5 pmol of 13C6-IAA, 13C6-oxIAA, 13C6-IAAsp and 13C6-IAGlu were added as internal standards to validate the quantification. All samples were then evaporated at 37 °C to dryness in vacuo. Purified samples were analyzed by the LC-MS/MS system consisting of an ACQUITY UPLC® System (Waters) and Xevo™ TQ-S (Waters) triple quadrupole mass spectrometer. Quantification was obtained using a multiple reaction monitoring (MRM) mode of selected precursor ions and the appropriate product ion32. The linear range spanned at least five orders of magnitude with a correlation coefficient of 0.9989–0.9998. Four independent biological replicates of each mutant were used in these analyses.
Additional Information
How to cite this article: Wu, L. et al. Forward genetic screen for auxin-deficient mutants by cytokinin. Sci. Rep. 5, 11923; doi: 10.1038/srep11923 (2015).
References
Korasick, D. A., Enders, T. A. & Strader, L. C. Auxin biosynthesis and storage forms. J Exp Bot 64, 2541–2555 (2013).
Woodward, A. W. & Bartel, B. Auxin: regulation, action and interaction. Ann Bot 95, 707–735 (2005).
Zhao, Y. Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61, 49–64 (2010).
Leyser, O. Auxin distribution and plant pattern formation: how many angels can dance on the point of PIN? Cell 121, 819–822 (2005).
Chapman, E. J. & Estelle, M. Mechanism of auxin-regulated gene expression in plants. Annu Rev Genet 43, 265–285 (2009).
Robert, S. et al. ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in Arabidopsis. Cell 143, 111–121 (2010).
Xu, T. et al. Cell surface- and rho GTPase-based auxin signaling controls cellular interdigitation in Arabidopsis. Cell 143, 99–110 (2010).
Jurado, S. et al. The Arabidopsis cell cycle F-box protein SKP2A binds to auxin. Plant Cell 22, 3891–3904 (2010).
Grones, P. et al. Auxin-binding pocket of ABP1 is crucial for its gain-of-function cellular and developmental roles. J Exp Bot 10.1093/jxb/erv177 (2015).
Gao, Y. et al. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc Natl Acad Sci USA 112, 2275–2280 (2015).
Tivendale, N. D., Ross, J. J. & Cohen, J. D. The shifting paradigms of auxin biosynthesis. Trends Plant Sci 19, 44–51 (2014).
Dai, X. et al. The biochemical mechanism of auxin biosynthesis by an arabidopsis YUCCA flavin-containing monooxygenase. J Biol Chem 288, 1448–1457 (2013).
Stepanova, A. N. et al. TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 133, 177–191 (2008).
Tao, Y. et al. Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 133, 164–176 (2008).
Müller, A. & Weiler, E. W. Indolic constituents and indole-3-acetic acid biosynthesis in the wild-type and a tryptophan auxotroph mutant of Arabidopsis thaliana. Planta 211, 855–863 (2000).
Boerjan, W. et al. Superroot, a recessive mutation in Arabidopsis, confers auxin overproduction. Plant Cell 7, 1405–1419 (1995).
Delarue, M., Prinsen, E., Onckelen, H. V., Caboche, M. & Bellini, C. Sur2 mutations of Arabidopsis thaliana define a new locus involved in the control of auxin homeostasis. Plant J 14, 603–611 (1998).
Barlier, I. et al. The SUR2 gene of Arabidopsis thaliana encodes the cytochrome P450 CYP83B1, a modulator of auxin homeostasis. Proc Natl Acad Sci USA 97, 14819–14824 (2000).
Zhao, Y. et al. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291, 306–309 (2001).
Zhou, Z. Y. et al. Functional characterization of the CKRC1/TAA1 gene and dissection of hormonal actions in the Arabidopsis root. Plant J 66, 516–527 (2011).
Yamada, M., Greenham, K., Prigge, M. J., Jensen, P. J. & Estelle, M. The TRANSPORT INHIBITOR RESPONSE2 gene is required for auxin synthesis and diverse aspects of plant development. Plant Physiol 151, 168–179 (2009).
Stepanova, A. N., Hoyt, J. M., Hamilton, A. A. & Alonso, J. M. A Link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17, 2230–2242 (2005).
Ljung, K. et al. Biosynthesis, conjugation, catabolism and homeostasis of indole-3-acetic acid in Arabidopsis thaliana. Plant Mol Biol 49, 249–272 (2002).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Weigel, D. et al. Activation tagging in Arabidopsis. Plant Physiol 122, 1003–1013 (2000).
Marchant, A. et al. AUX1 regulates root gravitropism in Arabidopsis by facilitating auxin uptake within root apical tissues. EMBO J 18, 2066–2073 (1999).
Chen, R. et al. The Arabidopsis thaliana AGRAVITROPIC 1 gene encodes a component of the polar-auxin-transport efflux carrier. Proc Natl Acad Sci USA 95, 15112–15117 (1998).
Luschnig, C., Gaxiola, R. A., Grisafi, P. & Fink, G. R. EIR1, a root-specific protein involved in auxin transport, is required for gravitropism in Arabidopsis thaliana. Genes Dev 12, 2175–2187 (1998).
Müller, A. et al. AtPIN2 defines a locus of Arabidopsis for root gravitropism control. EMBO J 17, 6903–6911 (1998).
Delbarre, A., Muller, P., Imhoff, V. & Guern, J. Comparison of mechanisms controlling uptake and accumulation of 2,4-dichlorophenoxy acetic acid, naphthalene-1-acetic acid and indole-3-acetic acid in suspension-cultured tobacco cells. Planta 198, 532–541 (1996).
Yamamoto, M. & Yamamoto, K. T. Differential effects of 1-naphthaleneacetic acid, indole-3-acetic acid and 2,4-dichlorophenoxyacetic acid on the gravitropic response of roots in an auxin-resistant mutant of arabidopsis, aux1. Plant Cell Physiol 39, 660–664 (1998).
Novák, O. et al. Tissue-specific profiling of the Arabidopsis thaliana auxin metabolome. Plant J 72, 523–536 (2012).
Sakai, T. et al. The wavy growth 3 E3 ligase family controls the gravitropic response in Arabidopsis roots. Plant J 70, 303–314 (2012).
Cheng, Y., Dai, X. & Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev 20, 1790–1799 (2006).
Ferrandez-Ayela, A. et al. Mutation of an Arabidopsis NatB N-alpha-terminal acetylation complex component causes pleiotropic developmental defects. PLoS One 8, e80697 (2013).
Gil, P. et al. BIG: a calossin-like protein required for polar auxin transport in Arabidopsis. Genes Dev 15, 1985–1997 (2001).
Ruegger, M. et al. Reduced naphthylphthalamic acid binding in the tir3 mutant of Arabidopsis is associated with a reduction in polar auxin transport and diverse morphological defects. Plant Cell 9, 745–757 (1997).
Alonso, J. M. et al. Five components of the ethylene-response pathway identified in a screen for weak ethylene-insensitive mutants in Arabidopsis. Proc Natl Acad Sci USA 100, 2992–2997 (2003).
Stepanova, A. N. et al. The Arabidopsis YUCCA1 flavin monooxygenase functions in the indole-3-pyruvic acid branch of auxin biosynthesis. Plant Cell 23, 3961–3973 (2011).
Kriechbaumer, V., Wang, P., Hawes, C. & Abell, B. M. Alternative splicing of the auxin biosynthesis gene YUCCA4 determines its subcellular compartmentation. Plant J 70, 292–302 (2012).
Cheng, Y., Dai, X. & Zhao, Y. Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 19, 2430–2439 (2007).
Sun, J., Qi, L., Li, Y., Chu, J. & Li, C. PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating arabidopsis hypocotyl growth. PLoS Genet 8, e1002594 (2012).
Hentrich, M. et al. The jasmonic acid signaling pathway is linked to auxin homeostasis through the modulation of YUCCA8 and YUCCA9 gene expression. Plant J 74, 626–637 (2013).
Paciorek, T. et al. Auxin inhibits endocytosis and promotes its own efflux from cells. Nature 435, 1251–1256 (2005).
Kanyuka, K. et al. Mutations in the huge Arabidopsis gene BIG affect a range of hormone and light responses. Plant J 35, 57–70 (2003).
Guo, X., Lu, W., Ma, Y., Qin, Q. & Hou, S. The BIG gene is required for auxin-mediated organ growth in Arabidopsis. Planta 237, 1135–1147 (2013).
Lopez-Bucio, J. et al. An auxin transport independent pathway is involved in phosphate stress-induced root architectural alterations in Arabidopsis. Identification of BIG as a mediator of auxin in pericycle cell activation. Plant Physiol 137, 681–691 (2005).
Yamaguchi, N. et al. CRM1/BIG-mediated auxin action regulates Arabidopsis inflorescence development. Plant Cell Physiol 48, 1275–1290 (2007).
Liu, Y. G., Mitsukawa, N., Oosumi, T. & Whittier, R. F. Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J 8, 457–463 (1995).
Weigel, D. & Glazebrook, J. Arabidopsis : a laboratory manual (ed. Weigel, D. & Glazebrook, J. ) 155–170 (Cold Spring Harbor Laboratory Press, 2002).
Laxmi, A., Paul, L. K., Raychaudhuri, A., Peters, J. L. & Khurana, J. P. Arabidopsis cytokinin-resistant mutant, cnr1, displays altered auxin responses and sugar sensitivity. Plant Mol Biol 62, 409–425 (2006).
Tian, Q., Uhlir, N. J. & Reed, J. W. Arabidopsis SHY2/IAA3 inhibits auxin-regulated gene expression. Plant Cell 14, 301–319 (2002).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31030045) and the Czech Ministry of Education, through the National Program for Sustainability I (LO1204).
Author information
Authors and Affiliations
Contributions
G.Q.G. designed and performed experiments. L.W., D.W.D. and P.L. performed major parts of the research. L.W. analyzed data, tested statistics and coordinated the figures. P.A., M.S. and O.N. quantified endogenous IAA levels. L.W., P.L., D.W.D., L.W., M.W., C.K.L., S.D.W., L.Z. and T.Z.Z participated in mutant screening. G.Q.G. and L.W. wrote the article. All authors analyzed and discussed the data and the article.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wu, L., Luo, P., Di, DW. et al. Forward genetic screen for auxin-deficient mutants by cytokinin. Sci Rep 5, 11923 (2015). https://doi.org/10.1038/srep11923
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11923
This article is cited by
-
Elucidating the ethylene response and tolerance in non-mutagenized and mutagenized snapdragon (Antirrhinum majus L.) lines using 1-aminocyclopropane-1-carboxylic acid (ACC)
Plant Growth Regulation (2023)
-
Significance of NatB-mediated N-terminal acetylation of auxin biosynthetic enzymes in maintaining auxin homeostasis in Arabidopsis thaliana
Communications Biology (2022)
-
Function of histone H2B monoubiquitination in transcriptional regulation of auxin biosynthesis in Arabidopsis
Communications Biology (2021)
-
Tomato auxin biosynthesis/signaling is reprogrammed by the geminivirus to enhance its pathogenicity
Planta (2020)
-
Functional roles of Arabidopsis CKRC2/YUCCA8 gene and the involvement of PIF4 in the regulation of auxin biosynthesis by cytokinin
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.