Abstract
DNA-restructuring activities of RecQ-family helicases play key roles in genome maintenance. These activities, driven by two tandem RecA-like core domains, are thought to be controlled by accessory DNA-binding elements including the helicase-and-RnaseD-C-terminal (HRDC) domain. The HRDC domain of human Bloom’s syndrome (BLM) helicase was shown to interact with the RecA core, raising the possibility that it may affect the coupling between ATP hydrolysis, translocation along single-stranded (ss)DNA and/or unwinding of double-stranded (ds)DNA. Here, we determined how these activities are affected by the abolition of the ssDNA interaction of the HRDC domain or the deletion of the entire domain in E. coli RecQ helicase. Our data show that the HRDC domain suppresses the rate of DNA-activated ATPase activity in parallel with those of ssDNA translocation and dsDNA unwinding, regardless of the ssDNA binding capability of this domain. The HRDC domain does not affect either the processivity of ssDNA translocation or the tight coupling between the ATPase, translocation and unwinding activities. Thus, the mechanochemical coupling of E. coli RecQ appears to be independent of HRDC-ssDNA and HRDC-RecA core interactions, which may play roles in more specialized functions of the enzyme.
Similar content being viewed by others
Introduction
RecQ-family helicases, which are key players in genome maintenance in both bacteria and eukaryotes, perform ssDNA translocation and dsDNA unwinding driven by ATP hydrolysis1,2,3. What makes RecQ enzymes different from other helicases is their unique capability to unwind various non-canonical DNA structures that arise during DNA replication, recombination and repair1,2,3. E. coli (Ec) RecQ helicase, the prototypic family member, is required for the processing of late replication intermediates and stalled replication forks4,5,6, exerts both pro- and anti-recombination activities7,8 and suppresses illegitimate recombination9. Mutations in three of five human RecQ homologs (BLM, WRN (Werner syndrome helicase) and RecQ4) cause severe syndromes bearing the common feature of loss of genomic integrity, highlighting the essential role of RecQ helicases in genome maintenance1,3.
The capability of RecQ helicases to process complex DNA structures is associated with a conserved domain architecture shared by Ec RecQ, human BLM and WRN enzymes, as well as at least one RecQ homolog in almost all investigated organisms (Fig. 1)10. Two conserved RecA-like domains harbor the ATPase cleft and drive 3′–5′-directed ssDNA translocation activity2,11. In most RecQ helicases, these domains are followed by a zinc-binding domain, which appears essential for the structural stability of the enzymes and contributes to DNA binding12,13,14. Furthermore, many RecQ helicases contain the auxiliary winged-helix (WHD) and HRDC domains2,11. The WHD mediates protein-DNA and protein-protein interactions and has been shown to facilitate DNA unwinding and contribute to substrate recognition10.
RecQ constructs used in this study
(a) Schematic domain map of Ec RecQ constructs used in this study. The motor core comprises two RecA-like domains. ZBD, zinc-binding domain; WHD, winged-helix domain; HRDC, helicase-and-RNase-D-C-terminal domain. RecQY555A harbors the Y555A point mutation in the HRDC domain (black bar). RecQ523 lacks the entire HRDC domain. (b–c) Crystal structures of a HRDC-deleted Ec RecQ construct (b, PDB code 1OYW) and a HRDC-containing human BLM construct (c, PDB code 4O3 M). Domains are colored as in a.
The HRDC domain, which is connected to the WHD via a flexible linker, also contributes to the substrate specificity and DNA binding affinity of RecQ helicases2,11,15,16,17. Despite their similar overall fold, HRDC domains of RecQ helicases show highly variable surface properties and ssDNA binding affinities16,18,19,20,21. For Ec RecQ, atomic structures have been solved separately for the HRDC-deleted helicase core and for the isolated HRDC domain (Fig. 1b)16,22. Atomic structures containing the RecQ helicase core together with the HRDC domain have been solved only for human BLM23 (PDB codes 4O3M, 4CDG, 4CGZ). In these structures the HRDC domain folds back onto the core and interacts with both RecA domains (Fig. 1c). Deletion of the BLM HRDC domain increased the ATPase activity of the enzyme while decreasing the rate of dsDNA unwinding23. These findings led to the proposition that the HRDC-core interaction is required for efficient coupling of the ATPase activity to dsDNA unwinding23.
Previous studies by us and others showed that RecQ and BLM helicases translocate on ssDNA with moderate processivity24,25,26,27. Translocation is tightly coupled to ATP hydrolysis and the enzymes translocate 1- 2 nucleotide (nt) upon hydrolysis of one ATP molecule24,25,26. In addition we found that, similar to the situation in BLM, the Ec RecQ HRDC domain suppresses the ssDNA-activated ATPase activity of the enzyme17. These findings suggest a general role for HRDC domains in supporting coupling between ATPase and translocation/unwinding activities through interdomain interactions. To test this hypothesis, here we performed transient and steady-state kinetic measurements using wild-type (WT), HRDC point mutant and an HRDC deletion mutant Ec RecQ constructs to dissect the mechanism of ATPase-coupled translocation and unwinding and its modulation by HRDC domain interactions. We found that the HRDC domain slows down ssDNA translocation and dsDNA unwinding by suppressing the ATPase activity, but it does not influence the processivity of translocation and the coupling of the ATPase activity to translocation and unwinding. These results suggest that the interaction between the motor core and the HRDC domain is conserved among RecQ helicases and this interaction moderates the rate of ATPase-driven activities, but its effect on mechanochemical coupling may vary among RecQ homologs, according to the differing physiological roles of the enzymes.
Results
Protein constructs
To dissect the role of the HRDC domain of Ec RecQ helicase in the enzymatic mechanism, we used three protein constructs: wild-type protein (RecQWT), a construct in which the previously characterized Y555A point mutation abolishes the ssDNA-binding ability of the HRDC domain (RecQY555A)16 and a construct lacking the entire HRDC domain (RecQ523, comprising amino acids (a.a.) 1-523) (Fig. 1a). Circular dichroism measurements indicated proper folding of all constructs (Supplementary Fig. S1).
ATPase suppression by the HRDC domain does not greatly affect ssDNA translocation processivity or ATPase-translocation coupling
To monitor the ATPase activity of helicase constructs, we followed the kinetics of inorganic phosphate (Pi) generation from ATP in a stopped-flow instrument. The amount of Pi produced was quantified using a fluorescently labeled phosphate binding protein (MDCC-PBP) (Supplementary Fig. S2)24,25,26,28,29. In the absence of DNA, traces of Pi generation were linear for all constructs in the presence of saturating ATP concentration (shown for RecQ523 in Fig. 2a). Steady-state DNA-free ATPase slopes for RecQWT were consistent with previous results determined by other methods25,30. The DNA-free ATPase activity (kbasal) was not influenced by either the Y555A mutation or HRDC deletion (Table 1).
The HRDC domain suppresses RecQ ATPase activity
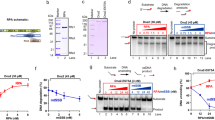
(a) Kinetics of Pi generation from ATP (1 mM) by RecQ523 (20 nM) in the absence of DNA, followed by MDCC-PBP (5 μM) fluorescence in the stopped-flow. (b) Kinetic traces of Pi release by RecQ523 (as in panel a but in the presence of various dT54 concentrations, indicated in nM). Traces were analyzed using Supplementary equation (S1) to determine the steady-state ATPase activity values shown in (c) for RecQWT (solid circles), RecQY555A (open diamonds) and RecQ523 (open circles). Solid lines in (c) show fits using the Hill equation. Determined parameters are listed in Table 1.
As shown previously, ssDNA markedly activates RecQ ATPase activity30,31. To assess the role of the HRDC domain in ssDNA-induced ATPase activation, we rapidly mixed ATP with RecQ constructs pre-mixed with increasing amounts of dT54 and monitored Pi release kinetics. Kinetic traces showed a lag followed by quasi-linear Pi generation for all constructs (shown for RecQ523 in Fig. 2b). The onset of the linear steady-state became faster with increasing DNA concentration with no sign of saturation, suggesting that the lag originated from an enzyme-DNA binding process induced by mixing with ATP (Fig. 2b). Our earlier study showed that ATP binds very rapidly to the enzyme30, ruling out that the lag in Fig. 2b originates from ATP binding. However, we recently showed that the presence of ATP, or its analogs, markedly increases the ssDNA affinity of RecQ17. Therefore, the lag can be explained by an ATP-induced increase in the fraction of ssDNA-bound RecQ molecules occurring upon stopped-flow mixing. This leads to a concomitant increase in the rate of Pi generation until reaching the steady-state.
We determined the apparent rate constant of the ATP-induced DNA binding process (kb) and the steady-state ATPase rate (kss) at each dT54 concentration from the Pi generation transients (Fig. 2b) using the model described in Supplementary equation (S1). Fits to the dT54 concentration dependence of kss using the Hill equation revealed ssDNA-activated kcat values (i.e. maximal kss at saturating dT54 concentration) in line with those determined previously using an NADH-coupled assay (Fig. 2c)17. The analysis indicated small positive cooperativity for the RecQ-ssDNA interaction, which was not systematically altered by the mutations (Hill coefficients were 1.7 ± 0.3, 1.0 ± 0.4 and 1.7 ± 0.3 for RecQWT, RecQY555A and RecQ523, respectively). In agreement with our previous results, kcat was almost identical for RecQWT and RecQY555A, but was 3 fold-higher for RecQ523 (Table 1)17. The apparent dissociation constants for dT54 (Kd,app,dT54), also determined from the dT54 concentration dependence of kss were slightly higher for RecQY555A and RecQ523 compared to RecQWT (Table 1).
The apparent rate constant of ATP-induced DNA binding (kb, determined from lag kinetics using Supplementary equation (S1)) increased linearly with dT54 concentration (Supplementary Fig. S3). This dependence allowed another, independent means of determination of the ssDNA binding Kd values of RecQ constructs during ATPase cycling (Supplementary Fig. S3, Supplementary Table S1). The determined values (Supplementary Table S1) were slightly higher than those determined from kss values (Table 1), but were in line with those determined previously for the RecQWT.AMPPNP (non-hydrolyzable ATP analog) complex17. The mutations caused a slight reduction in the ssDNA affinity of RecQ (Table 1, Supplementary Table S1). In summary, these results suggest that the ssDNA interaction of the HRDC domain has a minor contribution to the overall ssDNA binding affinity of RecQ, but the presence of the HRDC domain—regardless of its ssDNA binding capability—significantly suppresses the ssDNA-activated ATPase activity of the motor core.
To dissect how the HRDC domain affects ssDNA translocation, we monitored the kinetics of Pi generation from ATP hydrolysis during single-round translocation, which was previously shown to be suitable for the determination of translocation processivity and ATPase-translocation coupling (ATP hydrolyzed per nucleotide (nt) traveled)24,25,26,29.
To ensure single-round translocation conditions, we used dextran sulfate (DxSO4) as a DNA-mimicking protein trap25. First we assessed the trapping efficiency of DxSO4 by rapidly mixing RecQ with a pre-mixture of ATP, poly-dT and different concentrations of DxSO4 and monitoring Pi generation using MDCC-PBP in a stopped-flow instrument. Poly-dT was used instead of oligo-dT to minimize DNA end effects. Similar to those recorded in the absence of DxSO4 (Fig. 2b), traces showed a lag followed by linear steady-state Pi generation (Fig. 3a). In the absence of DxSO4, the steady-state rate of Pi generation was 3 times higher for RecQ523 than for RecQWT and RecQY555A (shown for RecQ523 in Fig. 3a), similar to the dT54 experiments of Fig. 2b–c (Table 1). As expected, the steady-state rate markedly decreased with increasing DxSO4 concentration (Fig. 3a). Surprisingly, we found that RecQ523 required 30-fold higher DxSO4 concentration for half-maximal inhibition of the ssDNA-activated steady-state ATPase activity than did RecQWT and RecQY555A (Fig. 3b). This result suggested that the HRDC domain, regardless of its ssDNA-binding capability, is a crucial mediator of DxSO4 trapping of RecQ.
The HRDC domain does not influence the processivity of ssDNA translocation
(a) Kinetics of Pi release observed upon mixing 20 nM RecQ523 with a pre-mixture of ATP (1 mM), poly-dT (25 μM, nt concentration) and various concentrations of DxSO4 (indicated in mg/ml) in the presence of 5 μM MDCC-PBP in the stopped-flow. The control trace was recorded in 6 mg/ml DxSO4 in the absence of poly-dT. The DxSO4 concentration dependence of steady-state ATP hydrolysis rates was used to determine the trapping efficiency (based on Supplementary equation (S2)) values shown in (b). Solid lines are hyperbolic fits to determine the DxSO4 concentrations required for half-maximal trapping (2.3 ± 0.1 μg/ml for RecQWT, 1.8 ± 0.1 μg/ml for RecQY555A and 62 ± 11 μg/ml for RecQ523). Symbols are as in Fig. 2c. (c) Time courses of Pi generation from ATP by RecQ523 during single-round translocation on poly-dT in the presence of 3 μM MDCC-PBP. 20 nM RecQ523 was pre-incubated with poly-dT (25 μM, nt) and mixed with ATP (1 mM) plus different concentrations of DxSO4 (1–8 mg/ml) in the stopped-flow apparatus. Traces consisted of an exponential and a linear phase, corresponding to ATP hydrolysis during translocation along poly-dT and in the DxSO4-bound end state, respectively. (d) DxSO4 concentration dependence of the amplitude of the translocation phase for all RecQ constructs. Symbols are as in Fig. 2c. Solid lines show fits based on Supplementary equation (S3). Determined parameters are listed in Table 1.
We characterized the ssDNA translocation processivity of RecQ constructs by determining the mean number of ATPase cycles during a single translocation run (<nATP>, Table 1). In these experiments, RecQ was preincubated with poly-dT and then rapidly mixed with ATP and different concentrations of DxSO4 in a stopped-flow instrument. The kinetics of Pi generation during ATP hydrolysis was followed by MDCC-PBP. DxSO4 concentrations were chosen from the regime where the trapping efficiency was above 95% (cf. Fig. 3b). Pi generation kinetic traces comprised an exponential phase (characteristic of single-round ssDNA translocation) followed by a slow steady-state that ensued upon dissociation of RecQ from ssDNA after the translocation run (Fig. 3c). Thus, the amplitude of the exponential phase (mol Pi/mol RecQ) equals <nATP>. This amplitude decreased with increasing DxSO4 concentration for all RecQ constructs, indicating that DxSO4 actively facilitates RecQ dissociation from ssDNA during translocation. In the case of RecQ523, the amplitudes showed a shallower DxSO4 concentration dependence than for RecQWT and RecQY555A (Fig. 3d), as expected based on the experiments of Fig. 3a–b. To determine the genuine (DxSO4-free) <n0ATP> value, we extrapolated to zero DxSO4 concentration using Supplementary equation (S3)24,25,29. RecQWT and RecQY555A showed similar <n0ATP> values, whereas this value was slightly higher in RecQ523 (Table 1), reflecting that the HRDC domain does not have a profound effect on the ssDNA translocation processivity of RecQ.
As described earlier, the ATPase-translocation coupling stoichiometry (Ctrans, number of ATP molecules hydrolyzed per nt traveled) can be determined from the ssDNA (oligo-dT) length dependence of Pi generation amplitudes during single-round ssDNA translocation24,25,29. In these experiments, we preincubated RecQ with oligo-dT substrates of different lengths and then rapidly mixed these premixtures with ATP plus DxSO4 in a stopped-flow instrument. Kinetic traces showed exponential and linear phases (Fig. 4a), similar to those observed for poly-dT (Fig. 3c). As expected, the single-round (exponential) Pi generation amplitudes increased with increasing oligo-dT length, showed saturation and their maximal value decreased with increasing DxSO4 concentration (Fig. 4b). Fits to these data using Supplementary equation (S4) revealed a Ctrans value of 1.0 ± 0.1 ATP/nt and an occluded site size (b) of 13 ± 1 nt for RecQ523 along ssDNA, both of which were independent of DxSO4 concentration in the assessed range (2 4 mg/ml) (Table 1). Comparison of these values with those determined earlier for RecQWT under identical or similar conditions revealed that deletion of the HRDC domain does not significantly affect the ATPase-translocation coupling but it decreases the occluded site size (Table 1)25,26.
4 mg/ml) (Table 1). Comparison of these values with those determined earlier for RecQWT under identical or similar conditions revealed that deletion of the HRDC domain does not significantly affect the ATPase-translocation coupling but it decreases the occluded site size (Table 1)25,26.
ATPase-translocation coupling in HRDC-deleted RecQ
(a) Time courses of Pi generation from ATP during translocation of RecQ523 along oligo-dT substrates of different length (dT15 − dT90, lengths in nt are indicated), recorded upon mixing 20 nM RecQ523 plus dTn (1 μM, except for dT15 (2.5 μM) and dT18 (2 μM)) with 1 mM ATP plus 2 mg/ml DxSO4 in the stopped-flow instrument. Traces comprised two phases as in Fig. 3c. (b), Oligo-dT length dependence of Pi generation amplitudes during the first (translocation-coupled) phase. Filled circles, empty circles and triangles represent values obtained in the presence of 2, 3 and 4 mg/ml DxSO4, respectively. Solid lines are fits based on Supplementary equation (S4). Determined parameters are listed in Table 1.
The rate of ATP hydrolysis during translocation along poly-dT (kATP,trans, determined from the initial slopes of the exponential phase in the experiments of Fig. 3c) was similar to those determined for DxSO4-free steady-state ATP hydrolysis for each construct (Fig. 3a, Table 1).
The HRDC domain suppresses the rate of ssDNA translocation in parallel with that of ATP hydrolysis
The above results showed that the HRDC domain does not influence ATPase-translocation coupling (Ctrans), but it suppresses the rate of ATP hydrolysis during translocation (kATP,trans). These findings imply that the rate of ssDNA translocation (ktrans (expressed in nt/s) = kATP,trans/Ctrans) must also be suppressed by the HRDC domain, in parallel with the ATPase rate. Furthermore, we found that <n0ATP> was not greatly affected by the HRDC domain, implying that this domain neither affects the mean processive run length (nt traveled in a single run, <nnt> = <n0ATP>/Ctrans). As <nnt> = ktrans/koff,trans where koff,trans is the rate constant of RecQ dissociation from ssDNA during translocation, one will expect that the latter parameter will be suppressed by the HRDC domain in parallel with the ATPase rate. Thus, the experimental determination of koff,trans provides an independent means of verification of the proposed mechanochemical effects.
The intrinsic (tryptophan, Trp) fluorescence intensity of each ssDNA-bound RecQ construct is markedly lower than that of the DNA-free forms17. This signal is thus suitable for monitoring the kinetics of dissociation of RecQ molecules (i.e., koff,trans) from ssDNA upon completing single-round translocation25. We determined the koff,trans values of RecQ constructs upon rapidly mixing the RecQ-poly-dT complex with excess ATP and varying concentrations of DxSO4 (Fig. 5a). The observed rate constants (kobs) of the transients increased linearly with DxSO4 concentration (Fig. 5b). The slopes and intercepts of the plots, reflecting the DxSO4 sensitivity of the reaction and the DxSO4-free koff,trans value, respectively, were similar for RecQWT and RecQY555A (Fig. 5b, Table 1). Importantly, the DxSO4-free koff,trans of RecQ523 was 3 times higher than that of RecQWT and RecQY555A, providing independent verification of the above mechanochemical considerations.
The HRDC domain suppresses the rate of dissociation from ssDNA during translocation
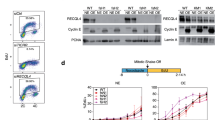
(a) Trp fluorescence stopped-flow time courses monitoring the dissociation of RecQ523 (0.5 μM) from poly-dT (25 μM, nt) during single-round translocation initiated by rapidly mixing the enzyme-DNA complex with 1 mM ATP plus varying concentrations of DxSO4 (bottom to top: 0, 0.5, 3 and 6 mg/ml). Solid lines show single exponential fits. (b) DxSO4 concentration dependence of observed rate constants (kobs) determined for all three RecQ constructs as in (a). Solid lines show linear fits. The y intercept of the plots delineated the trap-free dissociation rate constant from poly-dT during translocation (koff,trans, Table 1). The slopes of the plots, characterizing the DxSO4-sensitivity of ssDNA dissociation, were 2.0 ± 0.3, 1.1 ± 0.1 and 0.04 ± 0.01 s−1(mg/ml)−1 for RecQWT, RecQY555A and RecQ523, respectively. Symbols marked by * indicate kobs values determined in the absence of ATP. Symbols are as in Fig. 2c. AU, arbitrary units.
As expected based on the experiments of Fig. 3, the ssDNA dissociation kobs values of RecQ523 were less sensitive to DxSO4 compared with the other constructs (Fig. 5b, Table 1). In control experiments performed in the absence of ATP, kobs values were several times higher than those in the presence of ATP at the corresponding DxSO4 concentration for all constructs (Fig. 5b). This finding indicates that the Trp fluorescence data obtained in the presence of ATP reliably report dissociation after single-round translocation. Calculated <nnt> values (=ktrans/koff,trans) were slightly higher compared to those determined from MDCC-PBP experiments (<nnt> = <n0ATP>/Ctrans), but were practically identical for RecQWT and RecQ523 in both cases (Table 1).
The HRDC domain does not affect the coupling between ATP hydrolysis and dsDNA unwinding
To determine the coupling between the ATPase and dsDNA unwinding activities, we devised experiments to measure the rate of DNA unwinding and the rate of ATPase activity in the presence of a forked duplex DNA substrate comprising 33 bp dsDNA with two 21-nt arms. First we determined the binding affinity of RecQ constructs to this DNA substrate by fluorescence anisotropy titrations utilizing a fluorescein label placed on the dsDNA-forming 3′-end of one strand (Fig. 6a). Compared to RecQWT, the binding affinity (Kd, forked duplex) was reduced about 4-5-fold in both RecQY555A and RecQ523, reflecting a modest contribution of the HRDC-ssDNA interaction (Table 1).
The HRDC domain modulates forked duplex DNA unwinding but does not affect ATPase-unwinding coupling
(a) Fluorescence anisotropy titration of 10 nM fluorescein-labeled forked duplex DNA substrate (21-nt ssDNA arms + 33-bp dsDNA) with increasing amounts of RecQ constructs. Symbols are as in Fig. 2c. Solid lines show hyperbolic fits. Determined dissociation constants (Kd,forked duplex) are listed in Table 1. (b) Electrophoretogram of a single-turnover unwinding experiment. Preincubation of fluorescein-labeled forked duplex (30 nM) with RecQ523 (100 nM) was followed by rapid mixing with ATP (3 mM) plus excess ssDNA trap strand (3 μM) (final post-mixing concentrations). Reactions were stopped by the addition of EDTA (40 mM) and loading dye at different time points (0–150 s; as in (c)) using a quench-flow instrument or by manual mixing. Amounts of DNA species (forked duplex and ssDNA, depicted by cartoons) labeled with fluorescein (asterisk) were detected by a fluorescence imager. “–“ denotes a 150-s control reaction in which ATP was absent. (c) Single-turnover unwinding kinetics of forked DNA (as in (b)). Symbols are as in Fig. 2c. Solid lines show fits based on Supplementary equation (S5). Determined parameters are listed in Table 1. (d) Steady-state ATPase kcat (DNA-saturated ATPase) values of RecQ constructs (10 nM) in the presence of dT54, poly-dT and forked duplex DNA, determined in a PK-LDH coupled assay. Values obtained for dT54 and poly-dT are in line with those obtained in MDCC-PBP-based ATPase assays (Figs 2c and 3a; Table 1).
To determine dsDNA unwinding rates, we performed rapid kinetic single-turnover unwinding experiments in which we rapidly mixed the RecQ-forked DNA complex with ATP and excess unlabeled ssDNA trap strand in a quenched-flow instrument and monitored the time course of fluorescently-labeled ssDNA generation from forked duplex via gel electrophoresis of reaction products (Fig. 6b). Traces comprised a short (about 0.1-s) initial lag followed by two quasi-exponential rise phases in the case of all constructs (Fig. 6c) and were analyzed based on a previously described n-step kinetic model (Supplementary equation (S5))32. This model assumes that unwinding occurs as a result of n consecutive rate limiting steps that have a uniform rate constant. The model is suitable for the calculation of the macroscopic dsDNA unwinding rate (kunw) (Supplementary equation (S5)). The lag and the rapid unwinding phase of RecQWT was similar to that observed earlier using a Förster resonance energy transfer (FRET)-based assay33. We found that kunw was slightly accelerated in RecQY555A and RecQ523 compared to RecQWT, indicating that the HRDC-ssDNA interaction moderately hinders dsDNA unwinding (Fig. 6c, Table 1).
The unwinding traces of all constructs contained an additional slow exponential phase (apparent after 5 s in Fig. 6c) (Table 1). As discussed earlier for RecBCD and UvrD helicases, this phase may result from a fraction of enzyme molecules bound to DNA non-productively, necessitating a rate-limiting initiation of unwinding32,34. Alternatively, if multiple helicase molecules are initially bound to the 3′-ssDNA overhang, the slow phase may result from the action of one helicase proceeding in the trail of the leading one33,35.
To assess ATPase-dsDNA unwinding coupling, we measured the steady-state ATPase rate of RecQ constructs (kATP,unw) during unwinding of the same forked dsDNA substrate, using a pyruvate kinase-lactate dehydrogenase (PK-LDH) coupled assay (Fig. 6d). Compared to RecQWT, the ATPase activity of RecQY555A was slightly elevated, while RecQ523 had a significantly higher (kATP,unw) value (Fig. 6d, Table 1). The calculated macroscopic ATPase-unwinding coupling stoichiometry (Cunw (expressed as ATP hydrolyzed per bp unwound) = kATP,unw/kunw) was close to 1 ATP/bp for all constructs, suggesting that unwinding is tightly coupled to ATP hydrolysis and is not influenced by the HRDC domain (Table 1). Taken together, these results show similar trends to those found for ssDNA translocation: the HRDC domain hinders the rate of unwinding in parallel with that of the ATPase activity, without influencing the tight coupling between these processes.
Discussion
Although the HRDC domain has generally been considered as an auxiliary ssDNA-binding element, recent crystal structures of human BLM constructs showed that the HRDC domain can fold back onto and interact with both RecA domains, both in the presence and absence of DNA (pdb ids.: 4CGZ, 4CGD, 4O3M) (Fig. 1c)23. These interdomain interactions were proposed to influence the ATPase activity and the coupled ssDNA translocation and dsDNA unwinding by BLM23. Indeed, suppression of DNA-activated ATPase activity by the HRDC domain was observed for various RecQ helicases (Table 2)14,17,23,36. Moreover, in the case of Ec RecQ, ATPase suppression is independent of the ssDNA-binding ability of the HRDC domain, further indicating interdomain interactions (RecQY555A data in Table 1)17.
Importantly, however, the present study shows that the coupling of the RecQ ATPase activity to ssDNA translocation is unaffected by either the HRDC-ssDNA or the HRDC-motor core interactions. Moreover, the processivity of translocation is also unaffected by HRDC deletion in RecQ (Table 1), similar to the lack of effect of WHD-HRDC deletion on ATPase-translocation coupling and processivity in human BLM14.
We also found that the rate of forked duplex DNA unwinding, determined explicitly in transient kinetic experiments (Fig. 6c), is suppressed by the RecQ HRDC domain in parallel with the ATPase activity, indicating that coupling between these processes is not affected by the HRDC domain (Table 1). Interestingly, the DNA unwinding rate of RecQY555A fell between those of RecQWT and RecQ523, indicating that HRDC-ssDNA interactions slightly suppress the rate of unwinding, unlike that of ssDNA translocation (Table 1).
Explicit kinetic rates of dsDNA unwinding have not been measured for HRDC-deletion constructs of other RecQ helicase homologs. Available steady-state unwinding kinetic23 and end-point measurements14,36 are not directly informative of possible changes in mechanochemical coupling, as they can be influenced by more complex features (unwinding processivity, unproductive initiation, reversal during unwinding etc.). Nevertheless, such data reflect the relative unwinding efficiencies of helicase constructs and indicate that the HRDC domain may affect ATPase-driven unwinding of forked duplex DNA by BLM in a way that is different from that in Ec and Deinococcus radiodurans (Dr) RecQ enzymes (Table 2)22,36. BLM constructs lacking the HRDC23 or both the WHD and HRDC domains14 showed decreased efficiency of forked duplex DNA unwinding despite increased ATPase activities (Table 2). Further studies are needed to clarify whether and how the varying properties of HRDC domains of different RecQ homologs contribute to mechanochemical coupling and/or more complex dynamic processes during dsDNA unwinding (see below).
RecQ helicases are thought to translocate along ssDNA and unwind dsDNA via ATP-driven inchworm-type stepping24,25,26. Thus, the interactions and the relative positions of the two RecA-like domains are likely to undergo coordinated changes during the ATP hydrolytic cycle, as reported for various other helicases harboring a similar motor core37,38. Based on the lack of direct polar interactions between the two RecA domains of BLM, it was proposed that the interactions of the HRDC domain with both RecA domains contribute to mechanochemical coupling23. HRDC deletion will therefore increase the flexibility of the RecA core, thereby accelerating the ATPase cycle but possibly decreasing ATPase-unwinding coupling efficiency23. However, other crystal structures (PDB codes 4CDG, 4CGZ) show polar interactions between the RecA domains of ADP-bound BLM, both in the DNA-free and the DNA-bound form. Interestingly, crystal structures of ADP- and DNA-bound ternary complexes of BLM show two different conformations (PDB codes 4CGZ and 4O3M)23. These two structures possibly represent different snapshots of the ATPase cycle, suggesting transient interactions between the two RecA domains and/or between the bound nucleotide and the C-core RecA domain. Similarly, the crystal structure of an Ec RecQ construct lacking the HRDC domain22 shows no direct polar interactions between the RecA domains in the absence of substrates, but such interactions are present in the ATPγS (pre-hydrolytic ATP analog) -bound form of the same construct.
Crystal structures of human RECQ1, a RecQ homolog naturally lacking the HRDC domain, also suggest inter-RecA communication via transient polar interactions39. RECQ1 bound to a dsDNA substrate with a 5-nt 3′-ssDNA overhang exhibits no direct polar inter-RecA interactions, while the C-core RecA domain interacts with DNA39. On the other hand, RECQ1 bound to dsDNA with a 4-nt 3′-ssDNA overhang contains an inter-RecA salt bridge, but shows no C-core RecA-DNA interaction (PDB code 4U7D). These findings reflect that inter-RecA coordination can be modified by interactions with the DNA substrate. Interestingly, the two RecA domains form numerous direct as well as water-mediated polar contacts in ADP-bound DNA-free RECQ139. Together with the markedly (about 10 times) lower DNA-activated ATPase activity of RECQ140 compared to Ec RecQ and human BLM14,41, this finding provides further indication for the inverse relationship between inter-RecA rigidity and ATPase kinetics. Thus, the HRDC-induced suppression of the ATPase activity of various HRDC-containing RecQ helicases (Table 2) is likely brought about by HRDC-mediated coordination of the RecA domains.
Our previous work showed that the similar macroscopic mechanochemical properties of Ec RecQ and human BLM result from different underlying kinetic mechanisms24,30. The steady-state rate of DNA-activated ATP hydrolysis by Ec RecQ is limited by the ATP cleavage step, whereas in BLM a transition between two ADP bound states is rate-limiting24,30. ATP hydrolysis by RecA-type ATPases is thought to be triggered by the so-called “arginine finger” residue that interacts with the γ-phosphate of the bound nucleotide in SF1 helicases42,43. The putative arginine finger located in the C-core RecA domain of different RecQ helicases shows variation in its nucleotide interactions, suggesting its possible role in kinetic tuning. The arginine finger does not interact with the bound nucleotide in DNA-free Ec RecQ·ATPγS22,44 and human RECQ1·ADP complexes39. In contrast, the arginine finger of DNA-bound BLM45 interacts with both phosphate groups of ADP (PDB code 4CGZ). These differences may indicate that the dynamic interaction of the arginine finger with the bound nucleotide during the hydrolytic cycle, which can in turn be influenced by HRDC-RecA core interactions, contributes to limiting the rate of enzymatic activity.
The HRDC domain of RecQ helicases is connected to the rest of the protein through a long and flexible loop, raising the possibility of dynamic interactions with the RecA core of the protein and/or the ssDNA regions of the DNA substrate. Amino acids involved in ssDNA binding by the isolated HRDC domain of BLM18 are buried in crystal structures of HRDC-containing BLM constructs23 due to the interaction with the RecA core, suggesting that the HRDC-ssDNA and HRDC-RecA core interactions are mutually exclusive.
Synthesis of our current findings on Ec RecQ with earlier data on BLM reveals that in both enzymes, depending on the structure of the DNA substrate encountered, the HRDC domain is prone to interact with ssDNA regions outside the DNA segment tracked by the RecA core. We found that the unwinding of forked DNA substrates by Ec RecQ is noticeably slowed by HRDC-ssDNA interactions (cf. RecQWT vs. RecQY555A profiles in Fig. 6c, Table 1), but not by HRDC-RecA core interactions (cf. quasi-identical profiles of RecQY555A and RecQ523 in the same experiment). Consistent with this observation, we found that the deletion of the HRDC domain drastically reduced the DxSO4 sensitivity of ssDNA translocation kinetics by Ec RecQ (Figs 3d and 5b), suggesting that the HRDC domain greatly assists ssDNA-RecQ-DxSO4 ternary complex formation. In parallel with these observations, the HRDC domain of BLM was suggested to bind to ssDNA regions outside the one tracked by the RecA core during unwinding of G4-containing DNA substrates46 and the HRDC domains of Dr RecQ and BLM were found to greatly contribute to recognition and processing of Holliday junction structures36,47.
On the other hand, in the presence of simple ssDNA substrates and absence of trap, the HRDC domain appears to dominantly interact with the RecA core, as inferred from the very small effects of the Y555A point mutation but marked effects of HRDC deletion under these conditions (Fig. 2c; and trap-free dissociation rate constants in Fig. 5b; Table 1). Taken together, available data suggest that the HRDC domain may dynamically switch between ssDNA- and RecA core-interacting modes, thereby fine-tuning DNA-restructuring processes.
Materials and Methods
Reagents
All reagents were from Sigma-Aldrich unless otherwise stated. ATP was from Roche Applied Science. Inorganic phosphate (Pi) standard was from Merck. MDCC (7-diethylamino-3-((((2-maleimidyl)ethyl)amino)carbonyl)coumarin) was from Life Technologies. ε260 = 8400 M−1 cm−1 nt−1 was used for oligo-dT and poly-dT concentration determination. DNA concentrations are expressed as those of oligo- or polynucleotide molecules (as opposed to those of constituent nucleotide units (nt)) unless otherwise stated. Unless indicated otherwise, all experiments were performed at 25 °C in SF50 buffer (50 mM Tris-HCl pH 7.5, 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 50 μg/mL BSA).
Protein expression and purification
RecQ variants were expressed and purified as described in ref. 17. Fluorescently labeled Pi binding protein (MDCC-PBP) was prepared as described in ref. 28.
Circular dichroism (CD) spectroscopic measurements
UV (190–250 nm) CD spectra of proteins were recorded in a Jasco J-715 CD spectropolarimeter in a 1-mm path length cuvette. Measurements were done in SF50 buffer without BSA at 25 °C. Spectra were recorded three times independently and were averaged. Data were corrected with the CD spectra of BSA-free SF50 buffer.
dsDNA substrates
Forked duplex DNA substrates were generated as follows. Equimolar amounts of oligonucleotides fork1 (5′-CGTTGCGCAATTAAGCTCTAAGCCATCCGCAAATTTTTTTTTTTTTT-TTTTTTT-3′) and fork2 (5′-TCCTTTTGATAAGAGGTCATTTTTGCGGATGGTTAGAGCTTAATTG-CGCAACG-Fluorescein) were mixed in a buffer comprising 10 mM Tris-HCl (pH 7.5) and 50 mM NaCl. Samples were heated to 100 °C and were left to cool down to room temperature overnight. The annealed DNA mixtures were purified on a MonoQ anion exchange column using a 0.01–1 M NaCl gradient for elution. Eluted fractions were analyzed by PAGE. Fractions containing the desired DNA structures were desalted by using an Amicon Ultra centrifuge filter (Millipore). DNA substrates were aliquotted and stored at –80 °C. The ssDNA trap strand used in single-round unwinding experiments is the unlabeled version of the fork2 oligonucleotide.
PK-LDH-coupled ATPase measurements
Steady-state ATPase experiments were carried out in SF50 buffer by using a pyruvate kinase-lactate dehydrogenase (PK-LDH) coupled assay (14 U/mL PK, 20 U/mL LDH, 1 mM ATP, 1 mM phosphoenol pyruvate, 200 μM NADH). Time courses of NADH absorbance (ε340 = 6220 M−1cm−1) were followed in a Shimadzu UV-2101PC spectrophotometer.
Stopped-flow measurements
Stopped-flow measurements were carried out in a BioLogic SFM 300 instrument. Post-mixing concentrations are stated. Trp fluorescence was detected through a 320-nm cutoff filter at 280-nm excitation with a 4-nm bandwidth. Pi generation from ATP was followed using MDCC-PBP. The setup for MDCC-PBP calibration and single-round translocation experiments were as described earlier24,25,30.
Fluorescence anisotropy titrations
10 nM of forked duplex substrate was titrated with increasing concentrations of RecQ constructs. Fluorescence anisotropy was measured in a Synergy H4 Hybrid Multi-Mode Microplate Reader (BioTek).
DNA unwinding experiments
In single-turnover unwinding experiments DNA substrates (30 nM; final reaction concentrations stated) were incubated with excess enzyme (100 nM) at 25 °C for 5 minutes in Buffer H (30 mM Tris-HCl pH 7.5, 100 mM KCl, 1 mM DTT, 50 μg/ml BSA, 20 mM creatine phosphate, 20 μg/ml creatine kinase). Reactions were started at 25 °C by mixing the DNA-enzyme complex with Buffer H containing ATP (3 mM), MgCl2 (3 mM) and ssDNA trap strand (3 μM) to provide single-turnover conditions. Reactions were stopped after pre-set reaction times by the addition of EDTA (40 mM final, pH 8.1) and loading dye (10 mM Tris-HCl pH 7.5, 40 mM EDTA, 60% v/v glycerol, 0.075% w/v Orange G, 0.83% w/v SDS). Reaction times shorter than 5 seconds were achieved by using a rapid quench-flow instrument (RQF-3, KinTek). Longer reactions were started and stopped manually. Post-reaction mixtures were incubated at 25 °C for additional 3 min and were held on ice until further processing. Samples were then loaded on 12% non-denaturing polyacrylamide gels in TBE buffer (89 mM Tris-HCl pH 7.5, 89 mM boric acid, 20 mM EDTA). Electrophoresis was carried out at 4 °C. Fluorescently-labeled DNA was detected by using a Typhoon TRIO+Variable Mode Imager (Amersham Biosciences). The intensities of bands corresponding to the DNA substrate and unwinding products were quantified by densitometry (GelQuant Pro software (DNR Bio Imaging Ltd.)).
Data analysis
Means ± SEM values (n = 3) are reported in the paper, unless otherwise specified. Data analysis was performed using OriginLab 8.0 (Microcal corp.).
Additional Information
How to cite this article: Harami, G.M. et al. The HRDC domain of E. coli RecQ helicase controls single-stranded DNA translocation and double-stranded DNA unwinding rates without affecting mechanoenzymatic coupling. Sci. Rep. 5, 11091; doi: 10.1038/srep11091 (2015).
References
Brosh, R. M. DNA helicases involved in DNA repair and their roles in cancer. Nat Rev Cancer. 13, 542–558 (2013).
Vindigni, A., Marino, F. & Gileadi, O. Probing the structural basis of RecQ helicase function. Biophys Chem. 149, 67–77 (2010).
Croteau, D. L., Popuri, V., Opresko, P. L. & Bohr, V. A. Human RecQ Helicases in DNA Repair, Recombination and Replication. Annu Rev Biochem. 83, 519–552 (2014).
Courcelle, J. & Hanawalt, P. C. RecQ and RecJ process blocked replication forks prior to the resumption of replication in UV-irradiated Escherichia coli. Mol Gen Genet. 262, 543–551 (1999).
Hishida, T. et al. Role of the Escherichia coli RecQ DNA helicase in SOS signaling and genome stabilization at stalled replication forks. Genes Dev. 18, 1886–1897 (2004).
Suski, C. & Marians, K. J. Resolution of converging replication forks by RecQ and topoisomerase III. Mol Cell. 30, 779–789 (2008).
Harmon, F. G. & Kowalczykowski, S. C. RecQ helicase, in concert with RecA and SSB proteins, initiates and disrupts DNA recombination. Genes Dev. 12, 1134–1144 (1998).
Handa, N., Morimatsu, K., Lovett, S. T. & Kowalczykowski, S. C. Reconstitution of initial steps of dsDNA break repair by the RecF pathway of E. coli. Genes Dev. 23, 1234–1245 (2009).
Hanada, K. et al. RecQ DNA helicase is a suppressor of illegitimate recombination in Escherichia coli. Proc Natl Acad Sci USA. 94, 3860–3865 (1997).
Harami, G. M., Gyimesi, M. & Kovács, M. From keys to bulldozers: expanding roles for winged helix domains in nucleic-acid-binding proteins. Trends Biochem Sci. 38, 364–371 (2013).
Vindigni, A. & Hickson, I. D. RecQ helicases: multiple structures for multiple functions? HFSP journal. 3, 153–164 (2009).
Guo, R. B., Rigolet, P., Zargarian, L., Fermandjian, S. & Xi, X. G. Structural and functional characterizations reveal the importance of a zinc binding domain in Bloom’s syndrome helicase. Nucleic Acids Res. 33, 3109–3124 (2005).
Liu, J. L., Rigolet, P., Dou, S. X., Wang, P. Y. & Xi, X. G. The zinc finger motif of Escherichia coli RecQ is implicated in both DNA binding and protein folding. J Biol Chem. 279, 42794–42802 (2004).
Gyimesi, M. et al. Complex activities of the human Bloom’s syndrome helicase are encoded in a core region comprising the RecA and Zn-binding domains. Nucleic Acids Res. 40, 3952–3963 (2012).
Bernstein, D. A. & Keck, J. L. Domain mapping of Escherichia coli RecQ defines the roles of conserved N- and C-terminal regions in the RecQ family. Nucleic Acids Res. 31, 2778–2785 (2003).
Bernstein, D. A. & Keck, J. L. Conferring substrate specificity to DNA helicases: role of the RecQ HRDC domain. Structure. 13, 1173–1182 (2005).
Kocsis, Z. S., Sarlós, K., Harami, G. M., Martina, M. & Kovács, M. A Nucleotide-dependent and HRDC Domain-dependent Structural Transition in DNA-bound RecQ Helicase. J Biol Chem. 289, 5938–5949 (2014).
Kim, Y. M. & Choi, B. S. Structure and function of the regulatory HRDC domain from human Bloom syndrome protein. Nucleic Acids Res. 38, 7764–7777 (2010).
Sato, A. et al. Solution structure of the HRDC domain of human Bloom syndrome protein BLM. J Biochem. 148, 517–525 (2010).
Kitano, K., Yoshihara, N. & Hakoshima, T. Crystal structure of the HRDC domain of human Werner syndrome protein, WRN. J Biol Chem. 282, 2717–2728 (2007).
Liu, Z. et al. The three-dimensional structure of the HRDC domain and implications for the Werner and Bloom syndrome proteins. Structure. 7, 1557–1566 (1999).
Bernstein, D. A., Zittel, M. C. & Keck, J. L. High-resolution structure of the E.coli RecQ helicase catalytic core. EMBO J. 22, 4910–4921 (2003).
Swan, M. K. et al. Structure of human Bloom’s syndrome helicase in complex with ADP and duplex DNA. Acta Crystallogr D Biol Crystallogr. 70, 1465–1475 (2014).
Gyimesi, M., Sarlós, K. & Kovács, M. Processive translocation mechanism of the human Bloom’s syndrome helicase along single-stranded DNA. Nucleic Acids Res. 38, 4404–4414 (2010).
Sarlós, K., Gyimesi, M. & Kovács, M. RecQ helicase translocates along single-stranded DNA with a moderate processivity and tight mechanochemical coupling. Proc Natl Acad Sci USA. 109, 9804–9809 (2012).
Rad, B. & Kowalczykowski, S. C. Efficient coupling of ATP hydrolysis to translocation by RecQ helicase. Proc Natl Acad Sci USA. 109, 1443–1448 (2012).
Rad, B. & Kowalczykowski, S. C. Translocation of E. coli RecQ helicase on single-stranded DNA. Biochemistry. 51, 2921–2929 (2012).
Brune, M., Hunter, J. L., Corrie, J. E. & Webb, M. R. Direct, real-time measurement of rapid inorganic phosphate release using a novel fluorescent probe and its application to actomyosin subfragment 1 ATPase. Biochemistry. 33, 8262–8271 (1994).
Gyimesi, M., Sarlós, K., Derényi, I. & Kovács, M. Streamlined determination of processive run length and mechanochemical coupling of nucleic acid motor activities. Nucleic Acids Res. 38, e102 (2010).
Sarlós, K., Gyimesi, M., Kele, Z. & Kovács, M. Mechanism of RecQ helicase mechanoenzymatic coupling reveals that the DNA interactions of the ADP-bound enzyme control translocation run terminations. Nucleic Acids Res. 43, 1090–1097 (2015).
Umezu, K., Nakayama, K. & Nakayama, H. Escherichia coli RecQ protein is a DNA helicase. Proc Natl Acad Sci USA. 87, 5363–5367 (1990).
Ali, J. A. & Lohman, T. M. Kinetic measurement of the step size of DNA unwinding by Escherichia coli UvrD helicase. Science. 275, 377–380 (1997).
Zhang, X. D. et al. Escherichia coli RecQ is a rapid, efficient and monomeric helicase. J Biol Chem. 281, 12655–12663 (2006).
Lucius, A. L., Wong, C. J. & Lohman, T. M. Fluorescence stopped-flow studies of single turnover kinetics of E.coli RecBCD helicase-catalyzed DNA unwinding. J Mol Biol. 339, 731–750 (2004).
Pan, B. Y. et al. Mutual inhibition of RecQ molecules in DNA unwinding. J Biol Chem. 285, 15884–15893 (2010).
Killoran, M. P. & Keck, J. L. Three HRDC domains differentially modulate Deinococcus radiodurans RecQ DNA helicase biochemical activity. J Biol Chem. 281, 12849–12857 (2006).
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell. 97, 75–84 (1999).
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell. 127, 1349–1360 (2006).
Pike, A. C. W. et al. Structure of the human RECQ1 helicase reveals a putative strand-separation pin. Proc Natl Acad Sci USA. 106, 1039–1044 (2009).
Cui, S. et al. Analysis of the unwinding activity of the dimeric RECQ1 helicase in the presence of human replication protein A. Nucleic Acids Res. 32, 2158–2170 (2004).
Gyimesi, M. et al. Visualization of human Bloom’s syndrome helicase molecules bound to homologous recombination intermediates. FASEB journal : official publication of the Federation of American Societies for Experimental Biology. 27, 4954–4964 (2013).
Soultanas, P. & Wigley, D. B. Site-directed mutagenesis reveals roles for conserved amino acid residues in the hexameric DNA helicase DnaB from Bacillus stearothermophilus. Nucleic Acids Res. 30, 4051–4060 (2002).
Ye, J., Osborne, A. R., Groll, M. & Rapoport, T. A. RecA-like motor ATPases–lessons from structures. Biochim Biophys Acta. 1659, 1–18 (2004).
Killoran, M. P. & Keck, J. L. Sit down, relax and unwind: structural insights into RecQ helicase mechanisms. Nucleic Acids Res. 34, 4098–4105 (2006).
Ren, H. et al. The arginine finger of the Bloom syndrome protein: its structural organization and its role in energy coupling. Nucleic Acids Res. 35, 6029–6041 (2007).
Chatterjee, S. et al. Mechanistic insight into the interaction of BLM helicase with intra-strand G-quadruplex structures. Nature communications. 5, 5556 (2014).
Wu, L. et al. The HRDC domain of BLM is required for the dissolution of double Holliday junctions. EMBO J. 24, 2679–2687 (2005).
Acknowledgements
This work was supported by the Human Frontier Science Program (RGY0072/2010 to M.K. and K.C.N.), the “Momentum” Program of the Hungarian Academy of Sciences (LP2011-006/2011 to M.K.) and ELTE KMOP‐4.2.1/B‐10‐2011‐0002 Grant to M.K. This work was supported in part by the Intramural Research Program of the National Heart, Lung and Blood Institute, National Institutes of Health (HL001056-07 to K.C.N.). We are grateful to Dr. Tünde Juhász for assistance with CD measurements and to Dr. Yeonee Seol for discussions.
Author information
Authors and Affiliations
Contributions
G.M.H., K.C.N. and M.K. conceived and designed the work. G.M.H., N.T.N. and M.M. performed experiments. G.M.H. and M.K. analyzed data. G.M.H., K.C.N. and M.K. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Harami, G., Nagy, N., Martina, M. et al. The HRDC domain of E. coli RecQ helicase controls single-stranded DNA translocation and double-stranded DNA unwinding rates without affecting mechanoenzymatic coupling. Sci Rep 5, 11091 (2015). https://doi.org/10.1038/srep11091
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11091
This article is cited by
-
The toposiomerase IIIalpha-RMI1-RMI2 complex orients human Bloom’s syndrome helicase for efficient disruption of D-loops
Nature Communications (2022)
-
The structural and functional characterization of human RecQ4 reveals insights into its helicase mechanism
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.