Abstract
Single molecule localization microscopy (SMLM) techniques allow for sub-diffraction imaging with spatial resolutions better than 10 nm reported. Much has been discussed relating to different variations of SMLM and all-inclusive microscopes can now be purchased, removing the need for in-house software or hardware development. However, little discussion has occurred examining the reliability and quality of the images being produced, as well as the potential for overlooked preparative artifacts. As a result of the up to an order-of-magnitude improvement in spatial resolution, substantially more detail is observed, including changes in distribution and ultrastructure caused by the many steps required to fix, permeabilize and stain a sample. Here we systematically investigate many of these steps including different fixatives, fixative concentration, permeabilization concentration and timing, antibody concentration and buffering. We present three well-optimized fixation protocols for staining microtubules, mitochondria and actin in a mammalian cell line and then discuss various artifacts in relation to images obtained from samples prepared using the protocols. The potential for such errors to go undetected in SMLM images and the complications in defining a ‘good’ image using previous parameters applied to confocal microscopy are also discussed.
Similar content being viewed by others
Introduction
The recent widespread uptake of new super resolution techniques has revolutionized and invigorated molecular and cellular biology. Methods based on the detection and localization of single molecules such as (f)PALM ((Fluorescence1) Photoactivated localization microscopy2) and (d)STORM ((Direct3) Stochastic optical reconstruction microscopy4) have achieved the most superior gains in spatial resolution with live cell measurements reportedly achieving ~30 nm spatial resolution5 and fixed cell measurements achieving ~10 nm spatial resolution6. These techniques take advantage of the ability to fit the emission pattern of a single emitting molecule's point spread function giving a precise localization coordinate. Whereas conventional widefield or confocal fluorescence microscopy views all fluorophores in an emissive state at once, localization techniques achieve temporal separation through one of several (photo)physical or (photo)chemical methods (For a recent comprehensive review see Klein et al.7).
Extensive protocol and methods papers have been published for PALM, fPALM8, STORM4 and dSTORM9, as well as 3D STORM6,10 and live cell dSTORM11,12. These papers, however, focus on microscope set up and image acquisition and processing providing extensive detail on these matters, but typically provide only outlines of common epifluorescence sample protocols. In an effort to demonstrate the importance of and stimulate further discussion regarding, sample preparation in SMLM, we share here, optimized fixation and staining protocols for single color, 2D SMLM of microtubules (MTs), mitochondria (MC) and actin in COS-7 cells. More importantly, we detail the optimization process via the systematic investigation of the images produced by varying the fixative used, fixation time, temperature and concentration, permeabilization time and concentration, use of blocking buffers and quenching of glutaraldehyde-related autofluorescence. Overall glutaraldehyde is demonstrated to be the best fixative for structure-preservation, though we also detail highly reproducible paraformaldehyde and methanol protocols for MT staining. The images rendered by our optimized protocols are comparable in localization precision and spatial resolution to the very best SMLM images in the literature and surpass many of those images published of un-optimized samples.
A potentially underappreciated consequence of any gain in resolution is that previously satisfactory protocols (e.g. for confocal imaging) may be insufficient for SMLM. This is because any artifacts present on a distance scale smaller than that previously imaged have not been visible in fluorescence imaging, due to the overlapping of point spread functions and the resultant blur. In the case of SMLM, cellular structure must be preserved much more accurately because sub-diffraction deviations will be resolved to the same extent as ‘real’ sub-diffraction detail.
There is not much discussion in the literature about the processes by which sample preparation should be optimized for SMLM and even less research has been conducted into the effects of suboptimal fixation and/or staining on the resulting SMLM image. A handful of publications have endeavoured to highlight some common artifacts seen in SMLM images10,13,14, however, those discussed are more often due to acquisition error. Developing a robust understanding and appreciation of the causes and appearance of artifacts in SMLM images is of paramount importance because oftentimes, these artifacts will not be easily identified. This is particularly true in SMLM involving unknown or dispersed target molecules. A stringent degree of benchmarking is therefore needed to assure that cells imaged using SMLM truly reflect the biologically native state as closely as possible. To do this, targets such as microtubules (MTs) and nuclear pore complexes (NPCs) are ideal as has been demonstrated by extensive publication of these structures imaged using SMLM10,15,16. In addition to well-defined sub-diffraction width, MTs show decreased SM signal in the center of the two-dimensional projection of their three-dimensional cylindrical structure17 and NPCs possess an eight-fold symmetry that can be distinctly resolved15. These features allow for comprehensive and rigorous assessment of the performance of the experiment including preservation of the cells.
Therefore, along with our discussion of optimized protocols for SMLM, images of sub-optimally prepared samples are discussed in relation to our understanding of super-resolution image artifacts as well as the underlying chemistry of fixation. Particular attention is paid to artifact structures observed only in the super-resolution images and not in the epifluorescence images. We draw attention to the inherent artifactual nature of fixed samples as recently highlighted18 and demonstrate the importance of widespread and enhanced understanding of the limitations of SMLM.
Results and Discussion
Optimized Protocols for Paraformaldehyde, Glutaraldehyde and Methanol Fixation
The main outcome of the research presented here is the optimization of several protocols using common fixatives for SMLM. Moreover, the crucial steps in the fixation process are pinpointed and discussed in relation to corresponding SMLM images containing artifacts. For all of these fixation protocols to progress well the chambered coverglass must be removed from the incubator and placed immediately on a pre-warmed 37°C surface with the fixation steps proceeding without delay.
Paraformaldehyde (PFA) fixation
The majority of the culture medium is aspirated using a glass pipette placed carefully into a corner of each chamber. Simultaneously, 500 μl of 37°C 3.7% PFA is added down the side of the chamber using a second glass pipette. This PFA should be freshly prepared from frozen 10% PFA (filtered after thawing and adjusted to pH 7.4) by adding 2X Phosphate buffered saline (PBS) and then the equivalent milli-Q water to make an isotonic solution. This is again pH adjusted using 1 M KOH or HCl to achieve a pH of 7.4.
The 3.7% PFA solution is then left on the cells for 12 minutes with occasional gentle agitation before aspiration using a glass pipette, again positioned in the same corner of each chamber to minimize perturbation damage. 500 μl of PBS is added concurrent to the removal of the fixative solution. This PBS wash is aspirated after approximately 30 seconds and replaced with fresh PBS that is removed after another 60 seconds. Three more PBS washes are administered lasting approximately 5, 10 and 15 minutes each. To permeabilize the cells Triton X-100 in isotonic PBS at concentrations ranging from 0.1–1% for 10 minutes is used. Following permeabilization, cells are again washed in PBS, once for 30 seconds and then twice for 5 minutes.
Blocking can be achieved using bovine serum albumin (BSA) in PBS at concentrations between 1 and 10%. Immunostaining is then done in BSA/PBS at the same concentration as the blocking step with the antibodies used at dilutions ranging 1:100–1:2000 primary antibody and 1:200–1:4000 secondary antibody depending on the antibodies themselves. Three washes of five minutes in PBS follow each staining step.
Gluteraldehyde (GA) fixation
For glutaraldehyde fixation, a previously optimized protocol used for electron microscopy (EM) and SMLM experiments that yielded excellent preservation of the cytoskeletal network6,19,20 was slightly modified. Unlike published SMLM methods using PFA or methanol, which are generally identical to the protocols used for confocal samples, this particular GA protocol was specifically modified for ultrastructure preservation and SMLM measurements and therefore required minimal adjustments. Instead it afforded us the opportunity to investigate what changes to SMLM images resulted from slight deviations from the established, well-developed protocol.
For GA fixation, the culture medium is aspirated and a primary fixative of 0.4% glutaraldehyde and 0.25% Triton X100 in PBS is added simultaneously as detailed in the PFA fixation protocol. After 90 seconds this primary fixative solution is removed, the cells are washed once quickly with 37°C PBS and a second solution of 3% GA in PBS applied. This solution is removed after 15 minutes. Both the 0.4% and 3% glutaraldehyde are made from frozen 10% glutaraldehyde stock, 2X cytoskeleton stabilizing buffer (CSB)21 (1 M NaCl, 100 mM PIPES, 30 mM MgCl2, 10 mM EGTA and 10 mM sucrose) and Milli-Q water to form an isotonic solution, pH 6.2.
The cells are washed following fixation as in the PFA protocol prior to addition of a reductant to quench auto-fluorescence from remaining free aldehyde groups. The 500 μl aliquots of reductant are either NaBH4 (0.1–1% w/v) or glycine (0.3 M) in PBS solution and are left for 15 minutes. Cells are then washed extensively with two quick washes (30, 60 seconds) and then three longer washes (5, 10 and 15 minutes). Blocking and staining steps are carried out as outlined in the PFA protocol.
Methanol(MeOH) fixation
One hour prior to fixation, the medium of cells for methanol fixation is replaced with pre-warmed medium that does not contain FBS. Cells are then returned to the incubator. After one hour the medium is aspirated and the cells are washed once quickly with PBS (37°C). The PBS is then aspirated and 500 μl of −20°C methanol is added to each well. The cells are left in a freezer at −20°C for five minutes and then the methanol is removed and the cells washed with PBS as detailed in the PFA protocol. Cells are then blocked and stained as outlined in the PFA protocol.
Development of well optimized protocols yield exemplary SMLM images of microtubules, mitochondria and actin
In the development of the optimized protocols we focussed primarily on microtubules (MTs) and mitochondria (MC) because they have very well defined structures. MTs are polymeric and expected to measure approximately 60–65 nm in width when immunostained and imaged with a localization precision of <10 nm22. The structure of MC networks is also well documented using SMLM methods10 and by targeting the Tom20 protein in the import receptor on the outer membrane we expect to see 40–50 nm clustering as previously detected using STimulated Emission Depletion (STED)23. By knowing the structure of our targets prior to SMLM dSTORM measurements we were able to more readily identify deviation from the biologically native state in response to changes in fixation and staining protocols.
As such, the three optimized protocols detailed above are a result of careful, systematic consideration of many of the different facets and steps of sample preparation and were initially developed specifically for MT fixation. Conclusively, all three fixatives—paraformaldehyde (PFA), glutaraldehyde (GA) and methanol—when used in accordance with the protocols outlined, give exemplary SMLM images of MTs (Figure 1). The PFA fixation protocol also worked well for MC networks but actin microstructure was lost; conversely the GA fixation protocol worked well for actin but MC appeared susceptible to shrinkage as has been observed before24. Overall, methanol gave unsatisfactory results for both MC and actin stains as expected based on previous literature and the precipitory action of methanol on the MC membrane25,26,27.
Optimized protocols using paraformaldehyde, glutaraldehyde and methanol preserve ultrastructure for SMLM imaging of mitochondria, actin and microtubules.
(A–C) COS-7 cells fixed using the optimized 3.7% paraformaldehyde protocol and stained using Alexa Fluor 647 in conjunction with (A) anti-Tom20, a protein component of the import receptor on the outer membrane of mitochondria, (B) phalloidin which strongly associates with filamentous actin and (C) anti-α/β-tubulin, the unit protein that polymerizes into microtubules. (D–F) Similarly stained mitochondria (D), actin (E) and microtubules (F) in cells fixed using the optimized 3% glutaraldehyde protocol. (G–I) Similarly stained mitochondria (G), actin (H) and microtubules (I) in cells fixed using the optimized −20°C methanol protocol. All images show sub-diffraction structure with the exception of G–H that show substantial damage. Scale bar: 1 μm.
Figure 1 shows these results with highly magnified sections of reconstructed dSTORM images of COS-7 cells stained to show MTs, the MC membrane and actin. These images are representative of images taken of cells routinely fixed following the detailed protocols. The top row (a–c) are images of cells fixed using the optimized PFA protocol, the middle row (d–f) depicts cells fixed using the GA protocol and the bottom row (g–i) shows cells fixed using the methanol protocol. The PFA and GA protocols used to obtain the images in Figure 1c and 1f included a pre-extraction step (Full protocol parameters can be accessed in SI Methods 1).
The images of PFA and GA fixed cells stained for all three target proteins (Figure 1a–f) show sub-diffraction structural detail whereas the methanol protocol yields very poor images of mitochondria and actin (Figure 1g–h) while maintaining good MT structure (Figure 1i). As each image is of a 2 × 2 μm area, the sub-diffraction detail present in these images can immediately and clearly be observed. In the case of the Tom20 stains (1a and 1d), the boundaries of the MC are more clearly delineated in the PFA fixed cell than in the GA fixed cell and similarly, a lower level of background/non-specific stain is observed when PFA is used. The previously reported23 clustering of the import receptor unit on the outer MC membrane is also visible. This clustering is clear evidence of the successful application of SMLM to elucidate sub-diffraction structure as epifluorescence images of identically prepared Tom20 stained cells yield highly homogeneous membrane visualization with no evidence of the underlying clustering.
The successful actin stains (Fig 1b and e) were somewhat more difficult to obtain and cytoskeleton-stabilizing buffers (CSBs) were found to make a significant difference to the preservation of the structure during GA fixation and have accordingly been used throughout. Any perturbation to the architecture of the actin was found to significantly impact the SMLM images. This is because the distribution of actin within cells is far denser, sometimes taking up to 20% of the protein mass in muscle cells and having less defined and more complicated ultrastructure compared to the MT or MC network28. Singular microfilaments of actin are only 4–6 nm in diameter, with intermediate filaments ~10 nm wide29. Larger bundles, known as stress fibres consist of tens of microfilaments and can measure up to 300 nm wide30 with filaments of various widths often distributed <10 nm from neighbouring filaments31. In epifluorescence and confocal imaging only the large stress fibres can be imaged and identified over the homogeneous signal of actin throughout the rest of the cell. Using the optimized PFA protocol (Fig 1b) we were unable to identify structural features beyond this, although the stress fibres themselves were resolved with much enhanced acuity demonstrating cross sections regularly smaller than those limited by diffraction (50–70 nm).
Using GA (Fig 1e) much of the substructure could be observed with fibres detected at less than 25 nm wide and resolved well from microfilaments less than 30 nm away. In both cases, the average localization precision of the Alexa Fluor conjugated F(ab’)2 fragments was calculated to be approximately 13 nm and so the lack of clear structure in Fig 1b is strong evidence that the limitation to spatial resolution is not the measurement or equipment but rather the sample itself.
Under optimized conditions all three fixatives could produce very high quality MT images (Fig 1c,f,i) with good cross section widths (<60 nm in all three examples) and very little background/non-specific stain. This low background stain was achieved by adding a pre-extraction step for both the PFA and GA fixations that removed the cytosol (SI Methods 1) and with it, many of the potential sites for non-specific attachment. It should also be noted that for good continuous structures in the methanol fixed cells, a four-fold increase in antibody concentration was necessary as well as the previously mentioned careful modulation of the temperature of the fixative. It is important to appreciate that conventional background fluorescence does not manifest directly in a reconstructed SMLM image because only single, bright fluorophore PSFs are fitted to Gaussians and localized using rapidSTORM. Instead, any background fluorescence that is present as homogeneous signal, lowers the localization precision of the measurement and can then worsen the spatial resolution of the resulting image (See SI Figure 1). It is fair to assume then that any localized molecule in these images is an Alexa Fluor and the question becomes whether the fluorophore is specific to the target molecule or not.
Sub-optimal initial application of the fixative to cells causes sub-diffraction artifacts
Following optimization of the protocols, we investigated the many artifacts observed in SMLM images of sub-optimally fixed MTs and MC. Worth mentioning but without need for extensive discussion, we observed that temperature, fixative concentration, fixation time, the use of blocking buffers and the importance of quenching after glutaraldehyde fixation, were all parameters that need to be optimized for each new cell type and target protein. These parameters have previously been discussed in the literature extensively14,18,32 and artifacts associated with them were often observed in both the epifluorescence and SMLM images.
Interestingly, the steps within our optimized protocols and, indeed, in all trialled protocols, that were observed to produce the most sub-diffraction scale damage were those involved in the initial application of the fixative. This included both the type of fixative used and the technique with which it was introduced into the cells. Examples of the regularly observed sub-diffraction artifacts that resulted are shown in Figure 2. Contrary to widespread practice, we observed best structure preservation when no initial washing step was conducted for the PFA and GA fixations. Application of PBS even at 37°C for 60 seconds caused changes in the clustering distribution on the MC membrane and introduced short unstained/damaged tracts on the MTs like those shown in Fig 2e. Similarly, any degree of dehydration prior to fixation caused similar artifacts to arise and because of the chamber walls, our samples were particularly susceptible to this if care was not taken. For these reasons we have specified that culture medium be removed from one side of the chamber while 37°C PFA or GA fixative is added simultaneously down the opposite chamber wall. This appears to increase the amount of non-specific aggregated stain on the coverglass outside of the cells but allows very good preservation of the MT, MC and actin substructure.
SMLM microtubule images of sub-optimally fixed cells reveal sub-diffraction artifacts not observable in epifluorescence images.
(A–C) Epifluorescence images of COS-7 cells stained for tubulin using Alexa Fluor 647 after fixation with PFA (A) for a shorter period than optimal, (B) at a lower concentration than optimal and (C) for a longer period than optimal These images are not readily identified as having fixation artifacts. (D–F) Corresponding SMLM images of sub-areas from A–C show clear sub-diffraction scale damage to the microtubule architecture with some areas well preserved and continuously stained in D–E (white arrows) and other long stretches of filament missing or damaged beyond antibody-epitope recognition. Scale bars A–C: 5 μm, D: 1 μm, E–F: 500 nm.
Often extra-cellular protein was observed to precipitate onto the glass or cells during methanol fixation and was not easily removed by extensive washing following fixation For this reason we developed a protocol via which FBS was removed from the cell culture environment prior to fixing and then a single warm wash was applied. It is possible that some of the artifacts seemingly omnipresent in our methanol fixed cells and discussed below are due to the inclusion of these extra steps although precipitating fixatives have previously been identified as causing a significant loss of overall cellular structure27.
Temperature, blocking buffer, quenching, washing and tonicity all affect SMLM images
In the case of the temperature of the applied fixative, the literature values vary from −20°C to room temperature for methanol33 and from 4°C to 45°C for cross linking fixatives34,35,36, depending on the target. Clearly optimization of fixation temperature was required and various temperatures were tested and incorrect temperatures observed often to result in artifacts like those depicted in Figure 2. However for MT, MC and actin, 37°C PFA or GA and −20°C methanol were found reproducibly to yield the best structural preservation. Similarly, the concentrations of fixative and permeabilizer listed in the protocols yielded the best SMLM images when compared with other tested concentrations.
We developed washing steps that increased in duration because this was observed to aid in the removal of the majority of the contaminant in the first 2–3 steps and then any remaining strongly-bound contaminant was removed over the final 10–15 minute washes. The tonicity of the solutions used was also found to be very important in preserving sub-diffraction structure and slightly hyper- or hypotonic solutions resulted in significant changes to the MT architecture and MC size. This was particularly true if the solutions used prior to, or during the fixation and permeabilization steps were not isotonic. This is why we have stressed the use of 2X PBS and CSB to calculate isotonic solutions throughout.
As reported previously14 blocking buffers were used to lower the degree of antibody binding to non-target proteins. However, the antibodies used were found to have very high specificity for our target proteins with there being only small decreases in non-specific binding upon use of a 2–10% blocking buffer. This slight decrease in non-specific dye localizations was only observed when all subsequent steps including antibody incubations and washes were also conducted in the same concentration of BSA blocking buffer. Finally, quenching of samples with either NaBH4 or glycine following GA fixation6 was observed to drastically increase the signal to noise of single Alexa Fluor 647 dyes thus improving overall localization precision. Overall various concentrations of permeabilizing agent (Triton X-100, 0.1–1%), blocking buffer (BSA, 2–10%) and quencher (NaBH4, 0.1–1%) were all observed to produce acceptable images with only slight changes in quality and structure. Finally, in optimization of the fixation protocols it was observed that the use of cytoskeleton stabilizing buffers aided in preservation of both MT and actin architecture but did not affect the images of MCs when used in conjunction with GA.
Over or under exposure of cells to fixative chemicals cause sub-diffraction artifacts
Fixative concentration and incubation time were found to be key to preserving sub-diffraction structure. Figure 2 depicts epifluorescence and super-resolution images of MTs in cells that were fixed in GA for less time than optimal (Fig 2a and d), at a lower concentration than optimal (Fig 2b and e) and for longer than optimal (Fig 2c and f) (Full protocols in SI Methods 2). These types of sub-diffraction artifacts were also routinely encountered when different temperatures or pre-fixation washing steps were used. Moreover, the severity of these artifacts was found to vary depending on several parameters. As such they were often seen and represent any number of biochemical changes to the MTs themselves.
Importantly, the epifluorescence images (Fig 2a–c) are not immediately recognisable as having fixation artifacts and would often be considered fit for publication, especially at low levels of magnification such as in images of multiple cells. For comparison epifluorescence images of cells fixed according to the optimized protocols are provided (SI Figure 1a and e) and show similar structures to those in Figure 2a–c. Conversely the damage to the MT structure is readily identifiable in the corresponding super-resolution images (bottom, 2d–f). Figure 2a–b, d–e depict cells that we hypothesize have not been fixed sufficiently so that the MTs themselves have not been immobilized sufficiently and stabilized in place to avoid damage from subsequent steps. This under-fixation occurs when the biochemical changes necessary for successful fixation are not completed to the required degree prior to removal of the fixative solution. Some areas of the MT architecture are well preserved with continuous, well-stained MTs observed over >500 nm (Fig 2d, white arrow) while in other areas, filaments seems to have either been removed or the epitopes damaged over similar length scales (Fig 2d red arrow). Lower concentrations of fixative or less time for incubation were seen to exacerbate this problem as shown in Fig 2e in which long continuous well-stained MTs (white arrow) are observable with gaps in which blur indicates some disruption to the structure or possible movement during image acquisition (red arrow). Even lower concentrations/shorter incubation times were observed to result in large areas of cells being removed during subsequent washing steps.
Conversely, over-fixing occurred when the primary directive of fixation—to secure the native biological structure in place—was achieved but then further chemical modifications were made. In this case it appears that the MTs have been damaged by the GA in some places but not others. More extensive over-fixation (either by leaving the fixative on longer or at a higher concentration) was observed to erode MT structure completely and, in extreme cases, destroy entire cells. These images clearly demonstrate that the first few minutes of any fixation protocol are the most important in preserving sub-diffraction structure.
Without optimization established protocols often result in artifactual SMLM images
Figure 3 further investigates sub-diffraction artifacts by directly comparing four common protocols used for preparing cells for MT staining and confocal imaging (Full protocols in SI Methods 3). Figure 3a shows a cell fixed using methanol kept at −20°C throughout the fixation. Some epitope damage is observed since there are discontinuous filaments present, (white arrow) despite the use of increased antibody concentration for methanol fixed stains. Minimal background/nonspecific stain is observed without the use of pre-extraction or blocking steps. Figure 3b shows a cell fixed in methanol at −20°C but allowed to sit in ambient conditions for the 20 minutes of fixation, thus raising the temperature of the methanol. There is some indication of non-native curvature in these MTs (blue arrow) as well as structure and clustering in the non-specific dyes localized near the filaments (red arrow). This clustering of the non-specific stain is not observed under any of the other trialled or optimized fixation methods. Figure 3c shows MTs fixed using room temperature, 4% PFA: while MT filaments can be observed in this image and the average cross section of these single MTs is comparable with optimized protocols (~60–65 nm), the image suffers from some discontinuousness in the MT structures as well as increased background/non-specific stain. Finally, Figure 3d shows MTs in a cell fixed using a pre-extraction step and 3% GA in CSB, demonstrating well-conserved architecture and a low level of background/non-specific stain. While methanol is an often-used fixative for MT structure preservation, these results show that GA is preferable and invite further investigation into the biochemical causes for the differences between methanol and GA fixed cells. They are also an excellent illustration of a previously sufficient protocol yielding unexpected artifacts at a sub-diffraction level.
SMLM images of microtubules prepared using standard protocols but with careful initial preparation and application of the fixative solutions show some sub-diffraction artifacts but preserve much of the filamentous architecture.
(A–D) COS-7 cells stained for tubulin using Alexa Fluor 647 after fixation with (A) -20°C methanol following a PBS wash, (B) −20°C methanol allowed to equilibrate to room temperature during fixation, (C) room temperature 4% paraformaldehyde and (D) 3% glutaraldehyde following pre-extraction using 0.3% Triton X-100. White arrows indicate discontinuousness of filaments, blue arrows indicate abnormal curvature of the filaments, red arrows indicate structured but non-filamentous localizations. Scale bar: 1 μm.
Both antibody concentration and permeabilization techniques affect the distribution of fluorophores within the sample
Like the MTs in Fig 1i, Fig 4a shows a cell fixed in −20°C methanol. In this cell almost no non-specific stain is detected despite a high antibody concentration used. Similarly, cells pre-extracted with 0.3% Triton X-100 in growth medium have a very low amount of non-specific stain (Fig 4b) (SI Methods 4). Permeabilization of the cells in a pre-fixation step with a low concentration of GA (0.3% with 0.3% Triton X-100) was observed to leave some cytosolic non-specific stain (Fig 4c) while permeabilizing after fixation (2% GA, followed by 0.3% Triton X-100) yielded the highest amount of non-specific localizations (Fig 4d). In all images the MTs themselves have been stained well with continuous filaments intact and yielding average cross-sections of 55–65 nm. These observations led us to hypothesize that the localized fluorophores not associated with polymeric tubulin were associated with cytosolic dimeric tubulin and were not non-specific in the traditional sense. If they were truly non-specific we would expect to see non-polymeric localizations even with the cytosol removed.
The timing of permeabilization of cells significantly affects the distribution of fluorophores within the cell, resulting in differences in the ‘cleanliness’ of the image as well as the degree to which the image is representative of the biologically native structure and distribution.
(A–D) COS-7 cells stained for tubulin using Alexa Fluor 647 after (A) fixation with −20°C methanol, (B) pre-extraction with 0.3% Triton X-100 in HEPES buffer before 3% glutaraldehyde fixation, (C) simultaneous permeabilization and fixation (0.3% glutaraldehyde with 0.3% Triton X-100) and (D) fixation with 3% glutaraldehyde prior to permeabilization with 0.3% Triton X-100. Scale bar: 1 μm.
Images supporting this hypothesis are shown in Figure 5 which depicts changes observed in response to different antibody concentrations applied to MTs. As mentioned above, homogeneous background fluorescence does not result in localizations therefore this represents only the Alexa Fluor dyes. These cells were all fixed according to the optimized GA protocol and then stained over the dilution range of 1:50 to 1:2000 primary and secondary antibodies. In all cases the antibody solutions were administered in 200 μl aliquots with a primary incubation time of three hours and a secondary incubation time of one hour. Figure 5a shows heavily overstained MTs with an increased average MT width (91 nm) as well as a very high level of background/non-specific stain. A magnitude less antibody results in the MTs depicted in Fig 5b. These MTs have average widths of 58 nm and a significantly lowered amount of non-specific stain and depict the optimal staining density for SMLM. An image of stained MTs resulting from further dilution of the antibodies is shown in Fig 5c depicting an even lower number of non-specific localizations but also suffering from very discontinuous MT filaments. A control stain in which the primary antibody was not used showed almost no localizations demonstrating that the non-specific stain observed in Fig 5 is not simply non-specific secondary antibody. This suggests that these are not ‘traditional’ non-specific fluorophore attachment but are something specific, most likely dimeric tubulin. Previous SMLM publications featuring MTs have not identified the non-polymeric localizations seen in their published images as dimeric tubulin. Nor has it been discussed that by pre- or simultaneously extracting the cytosol from cells the resulting images appear ‘cleaner’ with less non-filamentous localizations and that this effect is achieved with significant alteration of the biology of the sample, i.e. by removing the cytosol and with it, many ‘real’ tubulin localizations. Diffraction limited confocal or epifluorescence images of MTs in a cell would also detect the dimeric tubulin but as a low-intensity homogeneous background that is easily considered noise below the signal of the MTs themselves.
Varying antibody concentration affects apparent microtubule width, non-filamentous stain and filament continuity—all of which can affect spatial resolution.
(A–C) COS-7 cells fixed using the optimized glutaraldehyde protocol and then stained for tubulin using mouse anti-β-tubulin and Alexa Fluor 647 conjugated rabbit-anti-mouse. (A) Both primary and secondary antibodies were diluted 1:50, (B) 1:500 and (C) 1:2000. Scale bar: 1 μm.
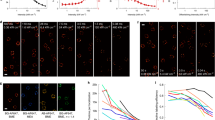
Varying fixative and antibody concentration alters the apparent clustering of Tom20 on the mitochondrial membrane
Our observations of significant changes in fluorophore distribution and apparent structure in response to permeabilization and antibody dilution prompted us to investigate the impact that similar changes in preparation would have on MC structure. As in Figure 1, the mitochondrial membrane import receptor was imaged by staining the Tom20 subunit after fixation with various protocols. In Figure 6 the first column shows cells fixed using the optimized PFA protocol, the second column shows cells fixed using a mixed PFA/GA (3%/0.5%) (SI Methods 6) protocol and the right column shows cells fixed according to the optimized GA protocol. The three rows show cells stained with a relatively high concentration of primary and secondary antibody (1:50, Top, Fig 6a–c), a mid-range concentration of antibody (1:500, Fig 6d–f) and a low concentration of antibody (1:2000, Fig 6g–i). Cells in the top and middle rows have not been quenched or blocked whereas those depicted in the bottom row (6g–i) were quenched using 1% NaBH4 and blocked in 3% BSA. Differences in clustering size and distribution are immediately apparent across antibody concentrations as well as fixatives; indeed, upon visual examination differences in all nine protocols can be seen. This is further demonstrated with basic cluster analysis which was performed by scanning the dSTORM images for clustered fluorophores forming features larger than 2 × 2 10 nm pixels (400 nm2) and smaller than 30 × 30 10 nm pixels (90,000 nm2) and then measuring the size of these clusters. Extensive preprocessing of the data was not necessary, nor was the definition of regions of interest, in order to acquire reasonable estimates of average cluster size. This was found to range from 2020–4330 nm2 corresponding to a range of average diameters (assuming circular arrangements) from 51–74 nm across all fixation and immunostaining conditions tested. The PFA/GA fixed preparation with 1:500 dilution of antibodies showing the smallest clusters (Figure 6e) and the 3.7% PFA fixation with 1:50 dilution of antibodies showing the largest clusters (Figure 6a) (Results summarized SI Table 1). In some cases, this analysis returned clusters larger than previous work which estimated them to be 30–40 nm in diameter. This is a consequence of both the enlargement of the Tom20 clusters by antibodies and fluorophores and the simplicity of the analysis performed. It should not be taken as truly quantitative due to the minimal preprocessing, however the large overall variation in mean size is in good agreement with visual examination of the images and confirms the role of fixation artifacts in the final images. In Figure 6 it can be seen that PFA fixation gives clear clustering at all three antibody concentrations but cluster size decreases as antibody concentration decreases. The mixed PFA/GA fixation yields what appears to be a lower degree of clustering with a more homogeneous distribution of localized dyes irrespective of antibody concentration. GA fixation shows a different pattern of clustering once more as well as some indication of shrinkage of the overall MC structure. This demonstration that slight changes in sample preparation—all of which yielded essentially visually identical epifluorescence images—can alter the sub-diffraction distributions and structures in relation to the known native state23 is extremely important because in many cases the sub-diffraction structure is unknown prior to SMLM. In the case of the Tom20 subunit of the import receptor, previous work using a PFA-based protocol similar to that depicted in Fig. 6d) has detected and quantified this clustering23. Without the reference point of this previously conducted work it would be more difficult to substantiate any claim that one protocol was more biologically accurate compared to the others. The fact that nine different levels of clustering were observed from nine variations of the protocol once again stresses the importance of determining which resembles the true biological native state most closely and to identify the causes of any artifacts.
Varying fixative and antibody concentration affects the apparent clustering distribution of import receptors on the mitochondrial membrane.
(A–I) COS-7 cells stained using Alexa Fluor 647 primary/secondary antibodies against the Tom20 protein subunit of the import receptor on the outer mitochondria membrane. Cells were stained with a high (A–C, 1:50), mid-range (D–F, 1:500) or low (G–I, 1:2000) concentration of primary and secondary antibodies. Cells were fixed using the optimized paraformaldehyde protocol (A, D, G), a mixed 3% paraformaldehyde, 0.5% glutaraldehyde protocol (B, E, H) or the optimized glutaraldehyde protocol (C, F, I). Scale bar: 1 μm.
General implications for SMLM
By focussing on fixation and staining protocols for SMLM we have gained significant appreciation for the importance of speed, hydration and minimal perturbation, particularly in the first steps of fixation, as well as for the necessity of adequate protocol optimization. The importance of carefully made solutions with correct tonicity and concentrations of fixative are also demonstrated. Significantly, all of the SMLM images we have shown here have been of structured targets with well-understood architecture due to extensive previous fluorescence and EM work. In contrast, much of the current and ongoing research using SMLM requires imaging of unknown structures and distributions of biomolecules that have not previously been extensively studied. For such research, the protocols established here are presented as a starting point, in particular the fully optimized double GA and 3.7% PFA protocols, which were found to present good images of all three targets. Following fixation with new or modified protocols, cells should be co-stained to investigate the sub-diffraction preservation of known structures (MTs, MCs, actin, etc.) as this will provide insight into the overall structural preservation of the cell. It is unavoidable that significant work will be required to optimize fixation protocols for new experiments in such a way that the structures and distributions of interest are maintained. However, both efficiency and robustness are maximised by starting with an established protocol and known structures and proceeding to probe in a systematic fashion, the impact of changing the main variables (fixative and fixative concentration, permeabilizer and permeabilizer concentration) on the SMLM images of both known and unknown molecule distributions. Extensive further work is necessary to establish the complicated biochemical effects of fixation on cells and will allow better predictions of fixation damage and artifacts. Live-cell SMLM will also undoubtedly assist in detecting, identifying and minimizing fixation artifacts but at present remains very challenging.
Here, by considering well-known structures, we have been readily able to identify unexpected localizations within the images simply with direct observation. In the case of the MTs, breaks in the filaments were observed as well as non-native curvature after methanol fixation and dyes that were non-specific for the MT filaments but most likely legitimate localizations of dimeric tubulin. In the case of MC, various clustering distributions were seen across different antibody concentrations and fixation protocols and without a known clustering behaviour23, no single result would be identifiable as correct. This demonstrates that careful consideration is required for interpretation of all SMLM images and in SMLM studies of distributions of single proteins in the cell, or of co-localization, a significant amount of benchmarking and protocol development is necessary before the images can be reasonably believed to be a reliable depiction of the native state of the sample. It is also worth noting that while direct observation was sufficient to detect differences in the samples presented here, future development of analytical tools for objective, quantitative appraisal of SMLM images will be crucial for the success of many potential applications.
The identification of fluorophores, such as those attached to dimeric tubulin, which would usually be thresholded out as noise also highlights a key difference between confocal and SMLM images. Specifically, SMLM images are not a true optical rendering rather a coordinate map of single molecule localizations; a distinction which must be continually appreciated when evaluating and interpreting SMLM ‘images’. Moreover, the parameters previously used to judge a confocal fluorescence image—namely the level of signal-to-noise, background and non-specific stain—do not readily apply to SMLM images and thus judging an image ‘good’ is complicated. Another interesting point that became apparent during our measurements is that even working with well-known structures, such as MC and MTs, the SMLM images that resemble the expected images, as well as those with highest signal or the lowest level of background stain, are not necessarily the most biologically relevant.
Ample literature exists detailing the various SMLM configurations4,6,9,37,38,39,40,41,42,43,44 as well as the commercial systems available. Conversely, sample fixation and immunostaining for SMLM has not been well discussed with sample preparation methodologies often published without enough information to be reproduced and with no explanation of optimization or potential artifacts. Our measurements highlight the ability of dSTORM to image sub-diffraction cellular features at a spatial resolution significantly improved (up to ten-fold) on conventional microscopy. We also stress that this improved resolution also brings a similar increase in the apparent severity of artifacts within the image. The many sub-diffraction artifacts we have encountered raise questions over results obtained from samples that were not fixed under optimal conditions and will hopefully be cause for more explicit detailing of the optimization process. We have aimed to highlight the high degree of variability in resulting SMLM images and the differences between normal fluorescence images and SMLM images. Accordingly, we have stressed the importance of protocol development and benchmarking, as well as furthering our understanding of the mechanisms of fixation.
Methods
Cell Culture
COS-7 cells were cultured routinely in Dulbecco's modified eagle's medium (DMEM) containing no phenol red and supplemented with 10% foetal bovine serum (FBS) and penicillin (100 units/ml) and streptomycin (100 μg/ml) at 37°C and 5% CO2. Cells were regularly passaged to keep confluence below 70% using TryPLE Express (Life Technologies). For all experiments cells were seeded at very low confluences (<20%) directly into 8 well Lab Tek II chambered coverglass (No. 1.5 borosilicate, Nunc). Prior to seeding, these chambers were cleaned by sonication for 15 minutes in 1 M KOH. Each chamber held 500 μl culture medium and was returned to the incubator for 36–48 hours prior to fixation.
Antibodies and Variations to Protocols
For the purposes of the figures and discussion presented in this paper the fully optimized protocols were varied somewhat to induce artifacts (Full protocols for each experiment and Figure available in SI). Stated ranges in concentration indicate that little difference was observed. The concentration and time of the fixation steps themselves were also varied to find the ideal balance (data not shown). Both the PFA and GA fixations were also done with an optional pre-extraction step using 37°C 0.3% Triton X100 in PBS or CB for 30 seconds. For all images of MTs mouse anti-α/β-tubulin (Sigma) and Alexa Fluor 647 rabbit-anti-mouse (Life Sciences) were used. For images of MC rabbit-anti-Tom20 (Santa Cruz) and Alexa Fluor 647 cow-anti-rabbit (Life Sciences) were used. For actin staining Alexa Fluor 647-phalloidin (Life Sciences) was used at a 1:500 dilution of 6.6 μM stock solution.
dSTORM Measurement and Analysis
Samples were kept at 4°C in the dark and were imaged within a week of fixation and staining. Switching buffer was made up from stock solutions using 2X PBS and milli-Q water and pH adjusted immediately before measurements. The stock solutions were 20% w/v glucose in milli-Q water, 1 M mercaptoethylamine (MEA) in milli-Q water and an enzyme cocktail containing catalase and glucose oxidase in PBS. Switching buffer was made up to a concentration of 10% glucose, 120 mM MEA, 400 μg/mL glucose oxidase and 35 μg/mLcatalase in isotonic PBS and 1 M KOH added to adjust the pH to a starting point of 8. The buffer was measured to shift to a pH of 6 during the experiment before significant distortions to the photophysics of the Alexa Fluor 647 dyes was observed.
All measurements were conducted on a home-built widefield microscope as detailed previously37. Briefly, a 150 mW 638 nm (Oxxius) diode laser is expanded and then focused using two lenses (focal lengths 25 and 100 mm) onto the back focal aperture of a 1.49 NA 100X TIRF (total internal reflection fluorescence) objective (Köhler illumination) via a dichroic filter. A quasi-TIRF illumination pattern is achieved by translating the excitation beam so that it is entering the objective off-centre, but parallel to the optical axis. Fluorescence is collected back through the objective and dichroic filter and imaged directly onto an electron-multiplying charge-coupled device (EM-CCD) camera (Andor Ixon Ultra).
All images were obtained using the Andor Solis software with the microscope in Q-TIRF mode using a power density of approximately 3–6 kW cm−2. Between ten and twenty thousand frames were collected at a frame rate of 80–100 Hz with the EM-Gain held at 200 (For an example of single frames showing single molecule emissions see SI Figure 1b–d and f–h). TIFF stacks were directly imported into the rapidSTORM software suite and used to generate super resolution coordinate files that were rendered into SMLM images using 10 × 10 nm pixels. The input pixel size was set at 105 nm and the PSF FWHH diameter found to be 350 nm by using the rapidSTORM ‘Estimate PSF form’. Single molecule emissions were identified only when more than 600 photons were detected within a single frame and localized to a single spot. The rapidSTORM software identified single molecule emissions as sufficiently intense local maxima in the noise-reduced (using an average mask) image and localized the centroid using a 350 nm 2D Gaussian function45. SI Figure 1b–d and f–h show example frames with well-separated single molecule emissions. Localization precision was estimated using the equation derived by Thompson46 and MT widths quoted are full width at half maximum of an intensity cross-section averaged over 400 nm. For cluster analysis of the distribution of Tom20 on the MC membrane, the images shown in Figure 6 were smoothed and analysed using “Analyse Particles” in the FIJI software suite47.
References
Hess, S. T., Girirajan, T. P. K. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Heilemann, M. et al. Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew. Chem. Int. Edit. 47, 6172–6176 (2008).
Rust, M. J., Bates, M. & Zhuang, X. W. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Jones, S. A., Shim, S.-H., He, J. & Zhuang, X. Fast, three-dimensional super-resolution imaging of live cells. Nat. Methods 8, 499–U496 (2011).
Xu, K., Babcock, H. P. & Zhuang, X. W. Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton. Nat. Methods 9, 185–188 (2012).
Klein, T., Proppert, S. & Sauer, M. Eight years of single-molecule localization microscopy. Histochem. Cell Biol. 141, 561–575 (2014).
Gould, T. J., Verkhusha, V. V. & Hess, S. T. Imaging biological structures with fluorescence photoactivation localization microscopy. Nat. Protoc. 4, 291–308 (2009).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Huang, B., Jones, S. A., Brandenburg, B. & Zhuang, X. W. Whole-cell 3D STORM reveals interactions between cellular structures with nanometer-scale resolution. Nat. Methods 5, 1047–1052 (2008).
Klein, T. et al. Live-cell dSTORM with SNAP-tag fusion proteins. Nat. Methods 8, 7–9 (2011).
Wombacher, R. et al. Live-cell super-resolution imaging with trimethoprim conjugates. Nat. Methods 7, 717–U772 (2010).
Kamiyama, D. & Huang, B. Development in the STORM. Dev. Cell 23, 1103–1110 (2012).
Bennett, B. T., Bewersdorf, J. & Knight, K. L. Immunofluorescence imaging of DNA damage response proteins: Optimizing protocols for super-resolution microscopy. Methods 48, 63–71 (2009).
Loeschberger, A. et al. Super-resolution imaging visualizes the eightfold symmetry of gp210 proteins around the nuclear pore complex and resolves the central channel with nanometer resolution. J. Cell Sci. 125, 570–575 (2012).
Szymborska, A. et al. Nuclear pore scaffold structure analyzed by super-resolution microscopy and particle averaging. Science 341, 655–658 (2013).
Dempsey, G. T., Vaughan, J. C., Chen, K. H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Schnell, U., Dijk, F., Sjollema, K. A. & Giepmans, B. N. G. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Small, J. V., Rottner, K., Hahne, P. & Anderson, K. I. Visualising the actin cytoskeleton. Microsc. Res. Techniq. 47, 3–17 (1999).
Xu, K., Zhong, G. S. & Zhuang, X. W. Actin, spectrin and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Small, J. V., Isenberg, G. & Celis, J. E. Polarity of actin at the leading edge of cultured cells. Nature 272, 638–639 (1978).
Bates, M., Huang, B., Dempsey, G. T. & Zhuang, X. W. Multicolor super-resolution imaging with photo-switchable fluorescent probes. Science 317, 1749–1753 (2007).
Wurm, C. A. et al. Nanoscale distribution of mitochondrial import receptor Tom20 is adjusted to cellular conditions and exhibits an inner-cellular gradient. Proc. Natl. Acad. Sci. U. S. A. 108, 13546–13551 (2011).
Doughty, M. J., Bergmanson, J. P. G. & Blocker, Y. Shrinkage and distortion of the rabbit corneal endothelial cell mosaic caused by a high osmolality glutaraldehyde-formaldehyde fixative compared to glutaraldehyde. Tissue Cell 29, 533–547 (1997).
Vielkind, U. & Swierenga, S. H. A simlpe fixation procedure for immunofluorescent detection of different cytoskeletal components within the same cell. Histochemistry 91, 81–88 (1989).
Stadler, C., Skogs, M., Brismar, H., Uhlen, M. & Lundberg, E. A single fixation protocol for proteome-wide immunofluorescence localization studies. J. Proteomics 73, 1067–1078 (2010).
Hoetelmans, R. W. M. et al. Effects of acetone, methanol, or paraformaldehyde on cellular structure, visualized by reflection contrast microscopy and transmission and scanning electron microscopy. Appl. Immunohisto. M. M. 9, 346–351 (2001).
Kim, H. R., Gallant, C., Leavis, P. C., Gunst, S. J. & Morgan, K. G. Cytoskeletal remodeling in differentiated vascular smooth muscle is actin isoform dependent and stimulus dependent. Am. J. Physiol.-Cell Ph. 295, C768–C778 (2008).
Schroede, T. E. Actin in dividing cells – contractive ring filaments bind heavy meromyosin. Proc. Natl. Acad. Sci. U. S. A. 70, 1688–1692 (1973).
Kumar, S. et al. Viscoelastic retraction of single living stress fibers and its impact on cell shape, cytoskeletal organization and extracellular matrix mechanics. Biophys. J. 90, 3762–3773 (2006).
Cooke, P. Filamentous cytoskeleton in vertebrate smooth-muscle fibers. J. Cell Bio. 68, 539–556 (1976).
Srinivasan, M., Sedmak, D. & Jewell, S. Effect of fixatives and tissue processing on the content and integrity of nucleic acids. Am. J. Pathol. 161, 1961–1971 (2002).
Noguchi, M., Furuya, S., Takeuchi, T. & Hirohashi, S. Modified formalin and methanol fixation methods for molecular biological and morphological analyses. Pathol. Int. 47, 685–691 (1997).
Fox, C. H., Johnson, F. B., Whiting, J. & Roller, P. P. Formaldehyde fixation. J. Histochem. Cytochem. 33, 845–853 (1985).
Bussolati, G., Annaratone, L., Medico, E., D'Armento, G. & Sapino, A. Formalin fixation at low temperature better preserves nucleic acid integrity. Plos One 6, 10.1371/journal.pone.0021043 (2011).
Zakout, Y. M. A., Salih, M. M. & Ahmed, H. G. The effect of fixatives and temperature on the quality of glycogen demonstration. Biotech. Histochem. 85, 93–98 (2010).
Whelan, D. R., Holm, T., Sauer, M. & Bell, T. D. M. Focus on super-resolution imaging with direct stochastic optical reconstruction microscopy (dSTORM) Aust. J. Chem. 67, 179–183 (2014).
Lippincott-Schwartz, J. & Manley, S. Putting super-resolution fluorescence microscopy to work. Nat. Methods 6, 21–23 (2009).
Mortensen, K. I., Churchman, L. S., Spudich, J. A. & Flyvbjerg, H. Optimized localization analysis for single-molecule tracking and super-resolution microscopy. Nat. Methods 7, 377–381 (2010).
Wolter, S. et al. rapidSTORM: accurate, fast open-source software for localization microscopy. Nat. Methods 9, 1040–1041 (2012).
Holm, T. et al. A blueprint for cost-efficient localization microscopy. Chemphyschem 15, 651–654 (2013).
Dedecker, P., Duwe, S., Neely, R. K. & Zhang, J. Localizer: fast, accurate, open-source and modular software package for superresolution microscopy. J. Biomed. Opt. 17 (2012).
Henriques, R. et al. QuickPALM: 3D real-time photoactivation nanoscopy image processing in ImageJ. Nat. Methods 7, 339–340 (2010).
Kechkar, A., Nair, D., Heilemann, M., Choquet, D. & Sibarita, J.-B. Real-time analysis and visualization for single-molecule based super-resolution microscopy. Plos One 8 10.1371/journal.pone.0062918 (2013).
Wolter, S., Schüttpelz, M., Tscherepanow, M., van der Linde, S., Heilemann, M. & Sauer, M. Real-time computation of subdiffraction-reolution fluorescence images. J. Microsc-Oxford 237, 12–22 (2009).
Thompson, R. E., Larson, D. R. & Webb, W. W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
TDMB is grateful for a Monash University Faculty of Science Dean's Strategic Funding Initiative. TDMB and DRW thank the German Laboratory Exchange Program (DAAD). DRW acknowledges an Australian Synchrotron Scholarship.
Author information
Authors and Affiliations
Contributions
D.R.W and T.D.M.B conceived, designed and performed the experiments, analyzed the data and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Whelan, D., Bell, T. Image artifacts in Single Molecule Localization Microscopy: why optimization of sample preparation protocols matters. Sci Rep 5, 7924 (2015). https://doi.org/10.1038/srep07924
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07924
This article is cited by
-
Genome-wide ATAC-see screening identifies TFDP1 as a modulator of global chromatin accessibility
Nature Genetics (2024)
-
SEMORE: SEgmentation and MORphological fingErprinting by machine learning automates super-resolution data analysis
Nature Communications (2024)
-
Rapid in-solution preparation of somatic and meiotic plant cell nuclei for high-quality 3D immunoFISH and immunoFISH-GISH
Plant Methods (2023)
-
4polar-STORM polarized super-resolution imaging of actin filament organization in cells
Nature Communications (2022)
-
ArtSeg—Artifact segmentation and removal in brightfield cell microscopy images without manual pixel-level annotations
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.