Abstract
For self-assembly of the bacterial flagellum, a specific protein export apparatus utilizes ATP and proton motive force (PMF) as the energy source to transport component proteins to the distal growing end. The export apparatus consists of a transmembrane PMF-driven export gate and a cytoplasmic ATPase complex composed of FliH, FliI and FliJ. The FliI6FliJ complex is structurally similar to the α3β3γ complex of FOF1-ATPase. FliJ allows the gate to efficiently utilize PMF to drive flagellar protein export but it remains unknown how. Here, we report the role of ATP hydrolysis by the FliI6FliJ complex. The export apparatus processively transported flagellar proteins to grow flagella even with extremely infrequent or no ATP hydrolysis by FliI mutation (E211D and E211Q, respectively). This indicates that the rate of ATP hydrolysis is not at all coupled with the export rate. Deletion of FliI residues 401 to 410 resulted in no flagellar formation although this FliI deletion mutant retained 40% of the ATPase activity, suggesting uncoupling between ATP hydrolysis and activation of the gate. We propose that infrequent ATP hydrolysis by the FliI6FliJ ring is sufficient for gate activation, allowing processive translocation of export substrates for efficient flagellar assembly.
Similar content being viewed by others
Introduction
AAA+ family ATPases, which are involved in various cellular activities such as DNA replication, proteolysis and membrane fusion, usually form ring-shaped oligomers with a narrow central channel. AAA+ ATPases couple ATP binding and hydrolysis to the translocation of their substrates to the central channel. Coordination and cooperativity among subunits in the ring-shaped ATPases are critical for their biological activities1,2.
The bacterial flagellum is a rotary nanomachine powered by PMF across the cytoplasmic membrane. It is composed of about 30 different proteins with their copy numbers ranging from a few to tens of thousands. The flagellum is divided into at least three parts: the basal body, the hook and the filament. Flagellar assembly begins with the basal body, followed by the hook and finally the filament. The flagellar export apparatus of Salmonella enterica is a type III secretion system and consists of a membrane-embedded export gate made of FlhA, FlhB, FliO, FliP, FliQ and FliR and a cytoplasmic ATPase complex consisting of FliH, FliI and FliJ and transports flagellar proteins from the cytoplasm to the distal end of the growing flagellar structure for self-assembly. The flagellar export apparatus is evolutionally related to the following two nanomachines: the injectisome of pathogenic bacteria, which directly inject virulence factors into their host cells; and the F- and V-type ATPases3,4.
The flagellar export apparatus utilizes both ATP and PMF as the energy sources for protein export5,6. FliI is the ATPase of the export apparatus7 and forms a homo-hexamer with a narrow central pore8,9. The FliI6 ring has been structurally identified at the flagellar base by electron cryotomography10. FliJ binds to the center of the FliI6 ring to form the FliI6FliJ ring, which looks very similar to F1-ATPase where the α/β and γ subunits correspond to FliI and FliJ, respectively11,12. FliH binds to FliI13 and anchors the FliI6FliJ ring complex to the export gate through interactions of FliH with a C ring protein FliN and FlhA14,15. The export gate is intrinsically a proton-protein antiporter that uses the two components of PMF, Δψ and ΔpH, for different steps of the export process16. An interaction between FliJ and FlhA turns the export gate into a highly efficient Δψ-driven export apparatus16. Although FliH, FliI and FliJ are dispensable for protein export, they make the export gate highly more efficient than their absence by which most of Salmonella cells cannot form flagella at all5,6. However, the roles of PMF and the ATPase are still under strong debate because the actual mechanistic role of the ATPase has remained unclear.
In vivo fluorescent imaging of FliI-YFP by fluorescence microscopy with single molecule precision has shown that not only the FliI6 ring but also several FliH2FliI complexes are associated with the flagellar basal body (FBB) through interactions of FliH with FliN and FlhA and that about 90% of the FliI-YFP spots show turnover between the FBB-localized and free-diffusing ones after photobleaching17. Neither the number of FliI-YFP associated with the FBB nor FliI-YFP turnover rate are affected by catalytic mutations in FliI, indicating that ATP hydrolysis by FliI does not drive the assembly-disassembly cycle of FliI during flagellar assembly17. In this study, to clarify the actual mechanistic role of ATP hydrolysis in flagellar protein export, we characterized the SalmonellafliI(E211Q) and fliI(E211D) catalytic mutants and an in-frame fliI deletion mutant with a reduced ATPase activity. We show that the export gate processively transports flagellar proteins during flagellar assembly even with extremely infrequent ATP hydrolysis. We also show that deletion of residues 401 to 410 of FliI considerably affects the energy coupling between ATP hydrolysis and activation of the gate by the FliI6FliJ ring.
Results
Effect of the E211Q and E211D mutations on flagellar protein export
The carboxyl group of Glu-211 in FliI, which corresponds to Glu-190 in the thermophilic Bacillus F1-ATPase, polarizes a water molecule for the nucleophilic attack to the γ-phosphate of ATP (Fig. 1)11,18,19. To investigate whether ATP hydrolysis by FliI limits the rate of flagellar protein export, we replaced Glu-211 with Asp and measured the ATPase activity. We used FliI(E211Q) as a negative control because the E211Q substitution results in the complete loss of its ATPase activity19. The ATPase activity of FliI(E211D) was measured quantitatively by subtracting the data from FliI(E211Q) as a background and it was about 100 times lower than that of wild-type FliI (Fig. 2a). The E211Q mutation in FliI considerably reduced both motility (Fig. 2b) and flagellar protein export (Fig. 2c, lane 7). In contrast, more than 80% of the fliI(E211D) mutant cells were motile and their swimming speed was about 50% of the wild-type level (Fig. 2b and Fig. S1). Consistently, most flagellar proteins were detected in the culture supernatant of this fliI(E211D) mutant (Fig. 2c, lane 8). Interestingly, the levels of FlgD (1st row), FliK (3rd row) and FliD (6th row) secreted by the fliI(E211D) mutant were comparable to the wild-type levels while the secretion levels of FlgE (2nd row), FlgK (4th row) and FlgL (5th row) were four, two and three-fold lower than wild-type levels, respectively. The E211Q and E211D mutations did not affect the cellular levels of FliI at all (Fig. 2c, 7th row). Therefore, these results indicate that the rate of ATP hydrolysis by FliI does not determine the rate of flagellar protein export.
Structure-base sequence alignment of FliI and F1-β subunit from the thermophilic Bacillus PS3 (1SKYB).
The regions of secondary structural elements are shown below each sequence: magenta line, α-helix; blue line, β-structure. The secondary structural elements are labeled with initials of three domains and numbers. A highly conserved Glu reside is highlighted by red. In-frame deletion of residues 401–410 in FliI (indicated as Δ40) is indicated by a red box.
Characterization of the fliI(E211Q) and fliI(E211D) mutants.
(a) Effect of the E211Q and E211D mutations on the FliI ATPase activity. Measurements were carried out on purified His-FliI (indicated as WT, black filled circle), His-FliI(E211Q) (indicated as E211Q, red filled triangle) or His-FliI(E211D) (indicated as E211D, blue filled square) using malachite green assay. (b) Swarming motility of a ΔfliI mutant transformed with pTrc99A (V), pMM1702 (WT), pKK211 (E211Q) or pMM1702(E211D) (E211D) in soft agar plates. The plate was incubated at 30°C for 6 hours. (c) Effect of the E211Q and E211D mutations on flagellar protein export. Whole cell proteins (Cell) and culture supernatant fractions (Sup) were prepared from the above transformants, subjected to SDS-PAGE and analyzed by immunoblotting with polyclonal anti-FlgD (1st row), anti-FlgE (2nd row), anti-FliK (3rd row), anti-FlgK (4th row), anti-FlgL (5th row), anti-FliD (6th row) or anti-FliI (7th row) antibody. The positions of molecular mass markers are indicated on the left.
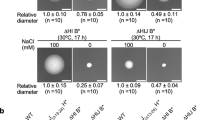
We then quantitatively analyzed the number of the flagellar filaments produced by these two fliI mutants (Fig. 3a). The number of the filaments produced by wild-type cells ranged from 1 to 8 with an average of 4.4 ± 1.6 (Fig. 3b). About 83% of the fliI(E211Q) cells had no filaments while the remaining population had one or two filaments with length only about 25% of the wild-type (Fig. 3b and c). In contrast, more than 90% of the fliI(E211D) cells produced filaments of which number ranged from 1 to 7 with an average of 2.3 ± 1.5 (Fig. 3b). The average filament length was about half of the wild-type (Fig. 3c). These results suggest that the export gate can processively export flagellar proteins during flagellar assembly even with extremely infrequent ATP hydrolysis.
Effect of FliI mutations on the number and length of the flagellar filaments.
(a) Fluorescent images of the ΔfliI mutant harboring pTrc99A (V), pMM1702 (WT), pMM1702(E211D) (E211D), pKK211 (E211Q) or pMMI40 (Δ40). Flagellar filaments were labeled with Alexa Fluor 594. The fluorescence images of the filaments labeled with Alexa Fluor 594 (red) were merged with the bright field images of the cell bodies. (b) Distribution of the number of the flagellar filaments in the ΔfliI mutant carrying pTrc99A (V, gray), pMM1702 (WT, red), pMM1702(E211D) (E211D, green), pKK211 (E211Q, orange) or pMMI40 (Δ40, light blue). More than 100 cells for each transformants were counted. (c) Measurements of the length of the flagellar filaments produced by each transformants. Filament length is the average of more than 100 cells for the wild-type, fliI(E211D) and fliI(E211Q) cells and 20 cells for the fliIΔ40 cell and vertical lines are standard deviations.
To test whether FliI(E211Q) and FliI(E211D) are incorporated into the export apparatus, we analyzed their dominant negative effect on motility of wild-type cells. Because FliI with a catalytic mutation requires FliH for its negative dominance13, we used a ΔfliH-fliI flhB(P28T) bypass mutant that can form some flagella even in the absence of FliH and FliI5 as a control. FliI(E211Q) and FliI(E211D) both inhibited the motility of wild-type cells (Fig. S2a) but not that of the ΔfliH-fliI flhB(P28T) bypass mutant (Fig. S2b), suggesting their assembly into the export apparatus in the presence of FliH.
Effect of an in-frame deletion of FliI residues 401 to 410 on flagellar protein export
The export gate requires an interaction of FliJ with FlhA to drive protein export in a Δψ-dependent manner16, raising the possibility that ATP hydrolysis by FliI activates the export gate through the FliJ-FlhA interaction. Because FliJ binds to the C1 α-helix of the C-terminal domain of FliI12, we constructed a series of FliI variants with sequential 10-amino-acid deletions within the C-terminal domain from residue 381 to 456 (FliIC). All the deletion variants were too insoluble to measure the ATPase activity, except for FliIΔ40 lacking residues 401 to 410 (Fig. 1). The ATPase activity of FliIΔ40 was about 40% of wild-type FliI (Fig. 4a). To test whether this reduction in the ATPase activity resulted from a decrease in the probability of FliI ring formation, we analyzed the capability of FliIΔ40 to form the ring in the presence of Mg2+-ADP-AlF4, which stabilizes the FliI6 ring structure19. FliIΔ40 formed a homo-hexamer at the wild-type level (Fig. 4b). These results indicate that residues 401–410 of FliI are required for the full ATPase activity but not for FliI6 ring formation.
Characterization of the fliIΔ40 mutant.
(a) Relative ATPase activity of His-FliIΔ40. The ATPase activity of His-FliIΔ40 (indicated as Δ40) is normalized to that of His-FliI (indicated as WT). These data are the average of four independent experiments and vertical lines are the standard deviations. (b) Electron micrographs of negatively stained samples of purified His-FliI and His-FliIΔ40 preincubated with 5 mM MgCl2, 5 mM ATP (labeled Mg-ATP) or 5 mM MgCl2, 5 mM ADP, 5 mM AlCl3, 5 mM NaF and 100 μg/ml acidic phospholipids. (c) Swarming motility of the ΔfliI mutant transformed with pTrc99A (V), pMM1702 (WT), or pMMI40 (Δ40) in soft agar plates. (d) Immunoblotting, using polyclonal anti-FlgD antibody, of whole cell proteins and culture supernatant fractions prepared from the same transformants.
We then measured the protein export activity of the fliIΔ40 mutant. The fliIΔ40 mutant displayed a very weak motility phenotype only after prolonged incubation (Fig. 4c). Consistently, very low levels of FlgD were detected in the culture supernatant (Fig. 4d, lane 6). The majority of fliIΔ40 cells had no flagellar filament and only a few % of cells had one short filament (Fig. 3). FliIΔ40 exerted a dominant negative effect on motility of wild-type cells (Fig. S3a) but not on that of the fliH-fliI flhB(P28T) bypass mutant (Fig. S3b). Therefore, FliIΔ40 is not functional in driving efficient protein export even though the (FliIΔ40)6 ring is assembled into the export apparatus.
Since FliI binds to the C-terminal cytoplasmic domains of FlhA (FlhAC) and FlhB (FlhBC), FliJ and chaperone substrate complexes, the FliI6 ring is proposed to act as a static substrate loader to facilitate the initial entry of export substrate into the export gate17. To test whether the deletion of residues 401–410 in FliI affects these interactions, we carried out pull-down assays by GST affinity chromatography. FliI co-purified with GST-FlhAC, GST-FlhBC, GST-FliJ and GST-FliT94/FliD complex but not with GST alone, in agreement with previous reports14,20,21 (Fig. 5, left panels). The amounts of FliIΔ40 co-purified with GST-FlhAC, GST-FlhBC, GST-FliJ and GST-FliT94/FliD complex were essentially the same as those of FliI (Fig. 5, right panels). These results indicate that the impaired export activity of the fliIΔ40 mutant does not result from the impaired interactions of FliI with them. Therefore, we suggest that the (FliIΔ40)6 ring is simply wasting the energy of ATP hydrolysis even with its proper incorporation into the export apparatus through interactions with FliJ and FlhA.
Effect of in-frame deletion of residues 401–410 in FliI on the interactions of FliI with FlhAC, FlhBC, FliJ and the FliT94/FliD complex.
(a) Interactions of GST-FlhAC with FliI (left panel) and FliIΔ40 (right panel). Purified His-FliI or His-FliIΔ40 was mixed with GST-FlhAC and dialyzed overnight against PBS. These mixtures (L) were subjected to GST affinity chromatography. Flow through fraction (F.T.), wash fractions (W) and elution fractions (E) were analyzed by CBB staining (1st row) and immunoblotting with polyclonal anti-FliI antibody (2nd row). (b) Interactions of GST-FlhBC with FliI (left panel) and FliIΔ40 (right panel). GST-FlhBC is cleaved between Asp-269 and Pro-270 in an autocatalytic way41 but FlhBCC remains to associate with GST-FlhBCN even after cleavage. (c) Interactions of GST-FliJ with FliI (left panel) and FliIΔ40 (right panel). (d) Interactions of the GST-FliT94/FliD complex with FliI (left panel) and FliIΔ40 (right panel). The positions of molecular mass markers are shown on the left.
Discussion
ATPases converts the chemical energy released by ATP hydrolysis into mechanical works required for their biological activities. In general the rate of ATP hydrolysis is coupled to their biological activity. SecA ATPase utilizes free energy derived from ATP hydrolysis to drive preprotein translocation through the SecYEG protein channel. Each catalytic cycle of ATP binding and hydrolysis is coupled to conformational changes in SecA that drive the stepwise translocation of preproteins across the membrane in a way similar to AAA+ ATPases such as ClpX and FtsH22. InvC, a FliI homolog of the Salmonella injectisome, forms a homo-hexamer, as has been visualized by electron cryomicroscopy at the base of the Salmonella injectisome23 and is thought to induce the release of chaperone from the chaperone-substrate complex and to unfold the substrate for efficient protein export in a way similar to the AAA+ ATPases24. In contrast, flagellar assembly occurs even in the absence of FliI, indicating that FliI is not essential5,6. Furthermore, FliI does not induce the release of flagellar chaperone from the chaperone-substrate complex in an ATP-dependent manner21. These observations suggest that the FliI6 ring does not act as an unfoldase to facilitate the substrate entry into the export gate. In this study, we showed that more than 90% of the fliI(E211D) mutant cells have a couple of flagella with length nearly half of the wild-type (Fig. 3) despite the 100-fold lower ATPase activity than that of wild-type FliI (Fig. 2a). Interestingly, the levels of FlgD, FliK and FliD secreted by the fliI(E211D) mutant were at the wild-type levels (Fig. 2c, lane 8). Furthermore, the fliI(E211Q) mutant, in which the mutation does not affect ATP binding although resulting in the complete loss of the ATPase activity19, exported flagellar proteins to some degree (Fig. 2c, lane 7) and formed flagella albeit at very low probability (Fig. 3). Therefore, we suggest that the rate of ATP hydrolysis by FliI is not at all coupled with the protein export activity and that ATP consumption could be relatively small despite that 20,000–30,000 flagellin molecules have to be exported for the assembly of each flagellum.
Although deletion of FliH and FliI results in almost complete loss of flagellar formation, most of ΔfliH-fliI flhB(P28T) bypass mutant cells are capable of forming a couple of flagella even in the absence of FliH and FliI. This indicates that the flhB(P28T) bypass mutation increases the probability of entry of flagellar proteins into the export gate. Because an increased gate-opening probability for higher efficiency of protein entry could be deleterious to the cells due to leakage of small solutes, the interaction of export substrate with the gate would presumably induce the gate opening, with or without the FliI6 ring5. Deletion of residues 401–410 in FliI (FliIΔ40) resulted in a considerably weak motility phenotype despite that the ATPase activity of FliIΔ40 was about 40% of the wild-type level (Fig. 4). Pull-down assays by GST affinity chromatography revealed that FliIΔ40 retained the ability to bind to FlhAC, FlhBC, FliJ and the FliT-FliD chaperone-substrate complex at the wild-type levels (Fig. 5). These results indicate that the Δ40 deletion causes uncoupling between ATP hydrolysis by the FliI6 ring and flagellar protein export. Therefore, we suggest that this deletion affects an activation step of the export gate by the FliI6 ring rather than a gate-opening step. The export gate itself is an inefficient proton-protein antiporter and since a specific interaction of FlhA with FliJ promoted by FliH and FliI switches the export gate to a highly efficient Δψ-driven export engine16, we propose that the FliI6FliJ ring complex acts as an ATP-driven ignition key to turn on the export engine to be a Δψ-dependent, rapid and efficient protein transporter (Fig. 6).
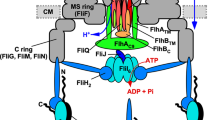
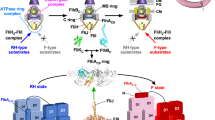
An ignition key mechanism for flagellar protein export.
ATP hydrolysis by the FliI6 ring induces the FliJ rotation within the ring to cause a conformational change of the export gate to activate it, allowing rapid and efficient entry of the substrates into the gate. The export gate then efficiently utilizes the membrane potential component (Δψ) of proton motive force across the cytoplasmic membrane to unfold and transport the substrates into the central channel of the growing flagellar structure.
How does the FliI6FliJ ring complex act as the ignition key for the Δψ-driven export engine? The FliI6FliJ ring complex is structurally very similar to the α3β3γ complex of F1-ATPase11,12. F1-ATPase is an ATP-driven rotary motor, driving the γ subunit rotation within the α3β3 ring. Three catalytic β-subunits in the α3β3 ring undergo highly cooperative and sequential conformational changes in their C-terminal domains of the β-subunits coupled with ATP hydrolysis25. It has been shown that ATP binds to the P-loop of FliI to induce a conformational change in FliIC similarly to that observed in the β-subunits11. FliJ binds to the C-terminal region of the C1 α-helix of FliIC, which corresponds to the region of the β subunit responsible for the interaction with the γ subunit in F1-ATPase12. These similarities suggest that FliJ may also rotate within the FliI6 ring. In fact FliJ actually shows a rotor like function within the central pore of the A3B3 complex of the Thermus thermophilus V-type ATPase26. Because a highly conserved surface of FliJ is responsible for the interaction with FlhA27, we propose that FliJ is a rotary ignition key driven by the FliI6 ring to activate the export gate through interaction of the conserved surface of FliJ with FlhA. The ATP hydrolysis by FliI in the ring is likely to induce cooperative conformational changes in the FliIC domains to rotate the FliJ ignition key.
This model can explain many aspects of our present results by analogy to F-ATPase. The E190D mutation in the β subunit considerably reduces the rotation rate of the γ subunit within the α3β3 ring18. Therefore, markedly reduced efficiency of flagellar assembly by the fliI(E211D) mutant would be a consequence of a significant reduction in the rotation frequency of FliJ. The elementary step size of γ rotation is 120°, which consists of 80° and 40° substeps driven by ATP binding–ADP release and ATP hydrolysis–Pi release, respectively25. So, just an ATP binding to the FliI(E211Q)6 ring could rotate FliJ by 80° and this may be sufficient to trigger the processive protein export to form a flagellum, although the frequency of this event would be very rare because no ATP hydrolysis occurs to make the FliI(E211Q)6 ring ready for the next rotation event. Since the β − γ contact at the C-terminal domain of the β subunit is responsible for torque generation25, it is possible that the Δ40 deletion impairs torque generation at the FliI-FliJ interface, resulting in a limited FliJ function as a rotary ignition key and therefore a very weak motility phenotype of the mutant cell. Thus, our data provides a clear insight into the mechanistic implication of the structural similarity between the FliI6FliJ ring and F1-ATPase and presents a very unique use of a rotary ATPase in a biological mechanism.
How does the export apparatus processively transport flagellar proteins even with extremely infrequent ATP hydrolysis? The cytoplasmic domain of FlhA (FlhAC) forms a nonameric ring structure in the export apparatus28,29 and provides binding-sites for chaperone-substrate complexes30,31,32. FliI also forms the FliH2FliI complex together with a FliH2 homo-dimer in the cytoplasm13. Not only the FliI6 ring but also several FliH2FliI complex associate with the FBB through interactions of FliH with FlhA and FliN17. The chaperone-substrate complexes bind to the FliH2FliI complex through co-operative interactions among FliI, chaperone and export substrate20,21,30,33. The FliH2FliI complex shows rapid turnovers between the FBB and the cytoplasmic pool in an ATP-independent manner17, suggesting that the FliH2FliI complex acts as a dynamic carrier to deliver the chaperone-substrate complexes from the cytoplasm to the FlhAC9 ring. Because FlhAC is directly involved in the translocation of the export substrate34, we propose that the FlhAC9 ring may play an important role in processsive protein transport along with the FliI6 ring that acts as a substrate loader upon the activation of the export gate by ATP hydrolysis by the FliI6FliJ ring complex.
Methods
Bacterial strains, plasmids and media
Salmonella strains and plasmids used in this study are listed in Table 1. L-broth (LB) and soft tryptone agar plates were prepared as described35,36.
DNA manipulations
DNA manipulations were carried out as described beforeref. 37. Site-directed mutagenesis was performed as described previously37. DNA sequencing was performed using BigDye ver 3.1 (Applied Biosystems) and the reaction mixtures were analyzed by a 3130 Genetic Analyzer (Applied Biosystems).
Protein expression and purification
His-FliI, His-FliI(E211Q), His-FliI(E211D) and His-FliIΔ40 were over-expressed in SJW1368 carrying pMM1702, pKK211, pMM1702(E211D) and pMMI40, respectively and purified by Ni affinity chromatography using a Ni-NTA agarose column (QIAGEN) as described36. GST-FlhAC, GST-FlhBC, FliJ and GST-FliT94 were overproduced in SJW1368 transformed with pMMHA1001, pMMHB1001, pMMJ1001 and pMMT002, respectively and purified by GST affinity chromatography using a glutathione Sepharose 4B column (GE Healthcare) as described beforeref. 21. FliD was overproduced in BL21 (DE3) pLysS harboring pKOT134 and purified as described previously38.
Measurements of FliI ATPase activity
The malachite green ATPase assay was carried out as describe previously7. At least four independent measurements were carried out.
Motility assays
For motility assay in soft ager, fresh colonies were inoculated on soft tryptone agar plates and incubated at 30°C. For measurements of free-swimming speeds of motile cells, overnight culture of Salmonella cells was inoculated into fresh LB and incubated at 30°C with shaking for 4 hours. The cells were washed twice with motility buffer (10 mM potassium phosphate pH 7.0, 0.1 mM EDTA, 10 mM L-sodium lactate) and resuspended in the motility buffer. Salmonella cells were observed under a phase contrast microscope (CH40, Olympus) at room temperature and their motile behavior was recoded on hard disk drive. The swimming speed of individual cells was analyzed as described before39.
Secretion assays
Salmonella cells were grown with shaking in 5 ml of LB at 30°C until the cell density had reached an OD600 of ca. 1.4–1.6. Cultures were centrifuged to obtain cell pellets and culture supernatants. Cell pellets were resuspended in the SDS-loading buffer, normalized to a cell density to give a constant amount of cells. Proteins in the culture supernatants were precipitated by 10% trichloroacetic acid, suspended in the Tris/SDS loading buffer and heated at 95°C for 5 min. After SDS-PAGE, immunoblotting with polyclonal anti-FlgD, anti-FlgE, anti-FliK, anti-FlgK, anti-FlgL and anti-FliD antibodies were carried out as described beforeref. 35. Detection was performed with an ECL plus immunoblotting detection kit (GE Healthcare). At least three independent experiments were carried out.
Observation of flagellar filaments with a fluorescent dye
Salmonella cells were attached to a cover slip (Matsunami glass, Japan) and unattached cells were washed away with motility buffer. A 1 μl aliquot of polyclonal anti-FliC serum was mixed with 50 μl of motility buffer and then 50 μl of the mixture was applied to the cells attached to the cover slip. After washing with the motility buffer, 1 μl of anti-rabbit IgG conjugated with Alexa Fluor® 594 (Invitrogen) were mixed with 50 μl of motility medium and then the mixture was applied. After washing with the motility buffer, the cells were observed as described before40. Fluorescence images were analyzed using ImageJ software version 1.48 (National Institutes of Health). To measure the filament length, each filament was traced by ImageJ freehand line tool and smoothed using the spline fit function.
in vitro reconstitution of the FliI6 ring and electron microscopy
Purified His-FliI or His-FliIΔ40 were incubated in 35 mM Tris–HCl (pH 8.0), 113 mM NaCl, 1 mM DTT, 5 mM ADP, 5 mM AlCl3, 15 mM NaF, 5 mM MgCl2 and 100 μg/ml acidic phospholipids at 37°C for a few minutes. Samples were applied to carbon-coated copper grids and negatively stained with 2% (w/v) uranyl acetate. Micrographs were recorded at a magnification of ×35,000 with a JEM-1011 transmission electron microscope (JEOL) operated at 100 kV.
Pull-down assays using GST affinity chromatography
Pull-down assays using GST affinity chromatography were performed as described previously32. Flow through fraction, wash fractions and eluted fractions were analyzed by SDS-PAGE with Coomassie Brilliant blue (CBB) staining and immunoblotting with polyclonal anti-FliI antibody. At least three independent experiments were carried out.
References
Ogura, T. & Wilkinson, A. J. AAA+ superfamily ATPases: common structure–diverse function. Genes Cells 6, 575–597 (2001).
Iino, R. & Noji, H. Intersubunit coordination and cooperativity in ring-shaped NTPases. Curr. Opin. Struct. Biol. 23, 229–234 (2013).
Minamino, T. Imada, K. & Namba, K. Mechanisms of type III protein export for bacterial flagellar assembly. Mol. BioSyst. 4, 1105–1115 (2008).
Minamino, T. Protein export through the bacterial flagellar type III export pathway. Biochim. Biophys. Acta. 1843, 1642–1648 (2014).
Minamino, T. & Namba, K. Distinct roles of the FliI ATPase and proton motive force in bacterial flagellar protein export. Nature 451, 485–488 (2008).
Paul, K., Erhardt, M., Hirano, T., Blair, D. F. & Hughes, K. T. Energy source of the flagellar type III secretion. Nature 451, 489–492 (2008).
Fan, F. & Macnab, R. M. Enzymatic characterization of FliI: an ATPase involved in flagellar assembly in Salmonella typhimurium. J. Biol. Chem. 271, 31981–31988 (1996).
Claret, L., Susannah, C. R., Higgins, M. & Huges, C. Oligomerisation and activation of the FliI ATPase central to the bacterial flagellum assembly. Mol. Microbiol. 48, 1349–1355 (2003).
Minamino, T. et al. Oligomerization of the bacterial flagellar ATPase FliI is controlled by its extreme N-terminal region. J. Mol. Biol. 360, 510–519 (2006).
Chen, S. et al. Structural diversity of bacterial flagellar motors. EMBO J. 30, 2972–2981 (2011).
Imada, K., Minamino, T., Tahara, A. & Namba, K. Structural similarity between the flagellar type III ATPase FliI and F1-ATPase subunits. Proc. Natl. Acad. Sci. USA 104, 485–490 (2007).
Ibuki, T. et al. Common architecture between the flagellar protein export apparatus and F- and V-ATPases. Nat. Struct. Mol. Biol. 18, 277–282 (2011).
Minamino, T. & Macnab, R. M. FliH, a soluble component of the type III flagellar export apparatus of Salmonella, forms a complex with FliI and inhibits its ATPase activity. Mol. Microbiol. 37, 1494–1503 (2000).
Minamino, T. et al. Roles of the extreme N-terminal region of FliH for efficient localization of the FliH-FliI complex to the bacterial flagellar type III export apparatus. Mol. Microbiol. 74, 1471–1483 (2009).
Hara, N., Morimoto, Y. V., Kawamoto, A., Namba, K. & Minamino, T. Interaction of the extreme N-terminal region of FliH with FlhA is required for efficient bacterial flagellar protein export. J. Bacteriol. 194, 5353–5360 (2012).
Minamino, T., Morimoto, Y. V., Hara, N. & Namba, K. An energy transduction mechanism used in bacterial type III protein export. Nat. Commun. 2, 475 (2011).
Bai, F. et al. Assembly dynamics and the roles of FliI ATPase of the bacterial flagellar export apparatus. Sci. Rep. 4, 6528 (2014).
Shimabukuro, K. et al. Catalysis and rotation of F1 motor: cleavage of ATP at the catalytic site occurs in 1 ms before 40 degree substep rotation. Proc. Natl. Acad. Sci. USA 100, 14731–14736 (2003).
Kazetani, K., Minamino, T., Miyata, T., Kato, T. & Namba, K. ATP-induced FliI hexamerization facilitates bacterial flagellar protein export. Biochem. Biophys. Res. Commun. 388, 323–327 (2009).
Imada, K., Minamino, T., Kinoshita, M., Furukawa, Y. & Namba, K. Structural insight into the regulatory mechanisms of interactions of the flagellar type III chaperone FliT with its binding partners. Proc. Natl. Acad. Sci. USA 107, 8812–8817 (2010).
Minamino, T., Kinoshita, M., Imada, K. & Namba, K. Interaction between FliI ATPase and a flagellar chaperone FliT during bacterial flagellar export. Mol. Microbiol. 83, 168–178 (2012).
Park, E. & Rapoport, T. M. Mechanisms of Sec61/SecY-mediated protein translocation across membranes. Ann. Rev. Biophys. 41, 21–40 (2012).
Kawamoto, A. et al. Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci. Rep. 3, 3369 (2013).
Akeda, Y. & Galán, J. E. Chaperone release and unfolding of substrates in type III secretion. Nature 473, 911–915 (2005).
Watanabe, R. & Noji, H. Catalysis-enhancement via rotary fluctuation of F1-ATPase. FEBS lett. 587, 1030–1035 (2013).
Kishikawa, J. et al. Common evolutionary origin for the rotor domain of rotary ATPases and flagellar protein export apparatus. PLoS One 8, e64695 (2013).
Ibuki, T. et al. Interaction between FliJ and FlhA, components of the bacterial flagellar type III export apparatus. J. Bacteriol. 195, 466–473 (2013).
Abrusci, P. et al. Architecture of the major component of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 20, 99–104 (2013).
Morimoto, Y. V. et al. Assembly and stoichiometry of FliF and FlhA in Salmonella flagellar basal body. Mol. Microbiol. 91, 1214–1226 (2014).
Bange, G. et al. FlhA provides the adaptor for coordinated delivery of late flagella building blocks to the type III secretion system. Proc. Natl. Acad. Sci. USA 107, 11295–11300 (2010).
Minamino, T. et al. Interaction of a bacterial flagellar chaperone FlgN with FlhA is required for efficient export of its cognate substrates. Mol. Microbiol. 83, 775–788 (2012).
Kinoshita, M., Hara, N., Imada, K., Namba, K. & Minamino, T. Interactions of bacterial flagellar chaperone-substrate complexes with FlhA contribute to co-ordinating assembly of the flagellar filament. Mol. Microbiol. 90, 1249–1261 (2013).
Thomas, J., Stafford, G. P. & Hughes, C. Docking of cytosolic chaperone-substrate complexes at the membrane ATPase during flagellar type III protein export. Proc. Natl. Acad. Sci. USA 101, 3945–3950 (2004).
Minamino, T. et al. Role of the C-terminal cytoplasmic domain of FlhA in bacterial flagellar type III protein export. J. Bacteriol. 192, 1929–1936 (2010).
Minamino, T. & Macnab, R. M. Components of the Salmonella flagellar export apparatus and classification of export substrates. J. Bacteriol. 181, 1388–1394 (1999).
Minamino, T. & Macnab, R. M. Interactions among components of the Salmonella flagellar export apparatus and its substrates. Mol. Microbiol. 35, 1052–1064 (2000).
Saijo-Hamano, Y., Minamino, T., Macnab, R. M. & Namba, K. Structural and functional analysis of the C-terminal cytoplasmic domain of FlhA, an integral membrane component of the type III flagellar protein export apparatus in Salmonella. J. Mol. Biol. 343, 457–466 (2004).
Imada, K., Vonderviszt, F., Furukawa, Y., Oosawa, K. & Namba, K. Assembly characteristics of flagellar cap protein HAP2 of Salmonella: decamer and pentamer in the pH-sensitive equilibrium. J. Mol. Biol. 277, 883–891 (1998).
Minamino, T., Imae, Y., Oosawa, F., Kobayashi, Y. & Oosawa, K. Effect of intracellular pH on the rotational speed of bacterial flagellar motors. J. Bacteriol. 185, 1190–1194 (2003).
Morimoto, Y. V., Nakamura, S., Kami-ike, N., Namba, K. & Minamino, T. Charged residues in the cytoplasmic loop of MotA are required for stator assembly into the bacterial flagellar motor. Mol. Microbiol. 78, 1117–1129 (2010).
Minamino, T. & Macnab, R. M. Domain structure of Salmonella FlhB, a flagellar export component responsible for substrate specificity switching. J. Bacteriol. 182, 4906–4919 (2000).
Yamaguchi, S., Fujita, H., Sugata, K., Taira, T. & Iino, T. Genetic analysis of H2, the structural gene for phase-2 flagellin in Salmonella. J. Gen. Microbiol. 130, 255–265 (1984).
Ohnishi, K., Ohto, Y., Aizawa, S., Macnab, R. M. & Iino, T. FlgD is a scaffolding protein needed for flagellar hook assembly in Salmonella typhimurium. J. Bacteriol. 176, 2272–2281 (1994).
Ikeda, T., Oosawa, K. & Hotani, H. Self-assembly of the filament capping protein, FliD, of bacterial flagella into an annular structure. J. Mol. Biol. 259, 679–686 (1996).
Acknowledgements
We acknowledge Ken-ichi Kazetani, Tatsuya Ibuki and Yasuyo Abe for technical help and Makoto Miyata and Masahiro Ueda for continuous support and encouragement. This work was supported in part by JSPS KAKENHI Grant Numbers 21227006 and 25000013 (to K.N.) and 26293097 (to T.M.) and MEXT KAKENHI Grant Numbers 23115008, 24117004 and 25121718 (to T.M.) and 26115720 (to Y.V.M). An International Project Grant awarded jointly by the Daiwa Foundation and Royal Society (UK) funded T.M. and P.D.A during this study.
Author information
Authors and Affiliations
Contributions
T.M. and K.N. conceived and designed research; T.M., Y.V.M., M.K. and P.D.A. preformed research; T.M., Y.V.M., M.K. and P.D.A. analysed the data; and T.M. and K.N. wrote the paper based on discussion with other authors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Minamino, T., Morimoto, Y., Kinoshita, M. et al. The bacterial flagellar protein export apparatus processively transports flagellar proteins even with extremely infrequent ATP hydrolysis. Sci Rep 4, 7579 (2014). https://doi.org/10.1038/srep07579
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07579
This article is cited by
-
The FlgN chaperone activates the Na+-driven engine of the Salmonella flagellar protein export apparatus
Communications Biology (2021)
-
Control of membrane barrier during bacterial type-III protein secretion
Nature Communications (2021)
-
A positive charge region of Salmonella FliI is required for ATPase formation and efficient flagellar protein export
Communications Biology (2021)
-
The FlhA linker mediates flagellar protein export switching during flagellar assembly
Communications Biology (2021)
-
FliI6-FliJ molecular motor assists with unfolding in the type III secretion export apparatus
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.