Abstract
RNA interference can induce heritable gene silencing, but it remains unexplored whether similar mechanisms play a general role in responses to cues that occur in the wild. We show that transient, mild heat stress in the nematode Caenorhabditis elegans results in changes in messenger RNA levels that last for more than one generation. The affected transcripts are enriched for genes targeted by germline siRNAs downstream of the piRNA pathway and worms defective for germline RNAi are defective for these heritable effects. Our results demonstrate that a specific siRNA pathway transmits information about variable environmental conditions between generations.
Similar content being viewed by others
Introduction
For centuries, scientists have wondered whether an organism's response to environmental change in one generation can be passed to subsequent generations. Recently, there has been much interest in the inheritance of information not encoded by differences in DNA sequence. Such effects have been seen in both plants and animals1,2,3 and may explain patterns of disease among humans4. However, these studies have often been hampered by the difficulty of controlling all environmental factors and the lack of quantitative, unbiased and reproducible assays of phenotype. Here we use the model animal C. elegans, which affords precise control of genetic and environmental factors, to investigate heritable effects triggered by ecologically relevant stimuli.
It has been known for several years that various experimental manipulations of C. elegans trigger heritable effects. These manipulations include artificially introducing exogenous RNAs5,6,7, globally disrupting chromatin8 and inserting a transgene containing non-C. elegans sequences expressing sense and antisense transcripts to simulate viral infection9. These studies may illuminate mechanisms that could enable inheritance of acquired traits. Inheritance of metastable changes is often assumed to require DNA methylation1,2,10, since this readily reversible modification is stable and can be copied between strands in a duplex. Indeed, patterns of DNA methylation are often heritable, but even in these cases the mechanism of inheritance remains obscure. As C. elegans lacks detectable DNA methylation11,12, the above examples imply the existence of DNA methylation-independent mechanisms of transgenerational “epigenetic inheritance”.
Several recent reports show that the evolutionarily conserved piRNA-silencing pathway can mediate heritable gene silencing in Drosophila and C. elegans13,14,15,16. In C. elegans, single-copy transgenes can undergo spontaneous germline silencing that is heritable for many generations14,15,16. Initiation, but not inheritance, of this silencing depends on the prg-1 gene encoding a Piwi-class argonaute protein. PRG-1 is thought to function analogously to the argonaute RDE-117,18, which binds to primary short interfering RNAs (siRNAs) generated by Dicer (DCR-1) cleavage of double-stranded RNA. The primary siRNA-RDE-1 complexes associate with target mRNA to direct production of the more abundant secondary siRNAs that are required for efficient silencing19,20. The primary short RNAs that PRG-1 binds to are not Dicer products but 21U RNAs (also known as piRNAs) that appear to be derived from endogenous short transcripts from thousands of genomic loci17,21,22. While the involvement of PRG-1 and piRNAs in transposon silencing18 has been known for some time, spontaneous silencing of endogenous non-transposon protein-coding genes by piRNAs has only recently been demonstrated23,24.
Known functions of the RNAi machinery in C. elegans include transposon silencing18,25,26,27 and resistance to viral infections28,29,30. Although circumstantial evidence suggests that endogenous RNAi conditionally alters expression of non-transposon genes in C. elegans31,32, little is known so far. Since RNAi is known to be heritable under some circumstances5,6,7, it would be of particular interest if similar regulation responds to naturally occurring environmental factors. We therefore undertook to search for heritable effects triggered by a commonly encountered environmental change.
A study reporting starvation-induced heritable changes in gene expression was published while this work was in manuscript33. Our work complements and adds to this study by showing that a different stress can induce similar inherited changes in gene expression. We also identify a specific RNAi pathway required for inheritance of environmental effects, show that extending the duration of stress increases the magnitude of the response and the number of affected generations and determine that the effect is passed via the female germline.
Results
Our approach was to culture worms at 20°C, then split the culture into 20°C control and 25°C treatment for one generation (3 days at 25°), followed by return to 20°C, transferring to fresh culture plates each generation. Animals cultured at 25°C grow faster than animals cultured at 20°C but produce fewer progeny, evidence of mild stress. We chose early-stage embryos for comparisons between environmental histories, because transcript levels for most genes stay relatively constant over time through the 4-cell stage34. While this provides high sensitivity to detect differences in transcript levels, it is unknown whether the transcripts present at this stage are important for the response to mild heat stress, but they should be reliable endogenous reporters for the phenomenon.
We found 20 genes showing notable persistence of changes in transcript abundance for at least a generation after return to 20°C (Figure 1 bottom left and top right corners and Supplementary Table S1; for source data see Supplementary Table S2 and Gene Expression Omnibus accession GSE30666). Based on permutation tests, only one gene would be expected to show such changes by chance alone (Supplementary Figure S1; see Supplementary Notes for discussions of permutation tests and outlier genes). We selected two genes, one with elevated and one with reduced transcript levels, to measure by reverse transcription quantitative PCR (RT-QPCR) in multiple independent biological replicates (Figure 1, right margin). This demonstrated the reproducibility of the observed transgenerational memory.
Inheritance of a response to environmental conditions.
Single-channel microarray analysis of 4-cell stage embryo mRNAs shows inheritance of temperature responses. Scatterplot of 15,208 genes: x-axis, generation 2 (G2); y-axis, generation 3 (G3). Values are ratios of geometric mean signal (25°C treatment/constant 20°C, six 50-embryo replicates each condition). Numbers in each quadrant count genes whose mRNA levels differ between treatment and control in both generations (>2× and ANOVA/Student's TP < 0.01, independently in each generation, larger dots; note this figure uses the statistics as a filter, not as a test). Pale dots are the lowest-signal 1/6 of genes and any genes with >4× the average inter-replicate variance. At right margin, G3 replicate RT-QPCR of additional experiments.
To investigate how long the effect persists, we measured transcript levels of the two genes over multiple 20°C generations following either one or two generations exposure to 25°C. We found that the effect can persist for at least four generations (Figure 2). Because a single C. elegans hermaphrodite worm produces hundreds of viable eggs and sperm, the dilution of material from any given developmental stage to the same stage in the next generation is expected to be at least a hundredfold. The reversibility of the heritable effects (Figure 2) makes it unlikely that they are due to mutation of genomic DNA sequences. Therefore, we surmise that the heritable signal is either self-regenerating or amplifiable from a small number of molecules; otherwise, we would have expected the effect to disappear quickly due to dilution with each passing generation.
RT-QPCR of multigenerational experiments.
Values are mRNA abundances relative to constant 20° control (dotted line) for multiple generations after one (black) or two (magenta) generations growth at 25°, geometric mean ± SEM. Two-tailed, paired T-test for difference with 20° control (shown for last two generations only), *P < 0.01 **P < 0.0001. To eliminate males and for more accurate staging, we transferred L4 hermaphrodite larvae from plate to plate (by platinum pick) instead of embryos. See Supplementary Table S5 for statistical details.
Heritable silencing of germline genes triggered by artificially introduced RNAs is similar in duration to what we see here (Figure 2), roughly two to four generations in the most detailed study so far6. To explore the possibility that an endogenous RNA interference-like phenomenon is involved in the heritable effect, we compared the mRNAs highlighted in Figure 1 with mRNAs targeted by short RNAs (sRNA) in oocytes35. We found that mRNAs showing transgenerational responses are highly enriched for genes targeted by endogenous antisense sRNA (Figure 3a and Supplementary Figure S2, P of overlap with oocyte sRNA < 10−6, by a cumulative hypergeometric distribution; see Supplementary Notes for discussion of those temperature-responsive genes that are not targeted by known sRNAs). Most of these sRNAs are 22 nucleotide residues long and start with a guanosine residue at the 5′ end and have therefore been dubbed “22G” RNAs35. Most 22G RNAs are thought to be secondary sRNAs made by RNA dependent RNA polymerases20,35,36.
Temperature-responsive early embryonic mRNAs are highly enriched for targets of endogenous 22G RNAs.
Circles labeled “G2” and “G3” represent genes whose mRNA levels in each generation differ >2× between treatment and control and ANOVA/Student's TP < 0.01 (see Figure 1). All P values are from cumulative hypergeometric distributions. Unless noted otherwise, P values represent the probabilities of the observed or greater overlap between the genes highlighted in Figure 1 and the sRNA set, relative to all genes on the array that are also represented among the annotations used by the published studies (15,19735 and 15,12036). The circle representing the set of all genes is omitted despite being required for a complete Venn diagram. The P values shown in parentheses below are for enrichment relative to the “G2” set alone, restricting analysis to only the genes identified as differing in G2. See Supplementary Notes for calculations. (a) Overlap with oocyte antisense sRNAs35 occurring at >50 (left), >100 (center) and >200 (right) reads per million. P = 6 × 10−8 (7 × 10−6), 7 × 10−10 (4 × 10−6) and 7 × 10−9 (4 × 10−5), respectively. (b) Overlap with sRNAs reported to be depleted >2× in all three of a mut-2 strain, a mut-7 strain and a strain with twelve argonaute genes disrupted35. P = 8 × 10−9 (7 × 10−5). (c) Overlap with sRNAs occurring at >10 reads per million in wild-type and depleted >20× in a mut-16 mutant36. P = 1 × 10−10 (1 × 10−5).
The production or maintenance of endogenous 22G RNAs antisense to the temperature-responsive mRNAs identified here depends on several genes required for germline RNAi, including mut-2 (also known as rde-3), mut-7, mut-16 and multiple Argonaute-encoding genes19,26,27,35,36,37,38,39,40 (Figure 3b-c and Supplementary Figure S2). MUT-2, MUT-7, MUT-16 and the RNA dependent RNA polymerase RRF-1 concentrate together in mut-16 dependent structures adjacent to germline nuclei, consistent with a proposed role for these proteins in secondary sRNA production41. As previously reported, genes targeted by 22G secondary RNAs in oocytes are composed of two complementary classes (see Figure 4a): those targeted by 22G RNAs that co-immunoprecipitate with CSR-1 and those targeted by 22G RNAs that co-immunoprecipitate with WAGO-9 (also known as HRDE-1)7,42. CSR-1 is not known to be involved in gene silencing and consistent with published results42, our microarray data shows that CSR-1 associated sRNAs target genes that tend to be highly expressed (Figure 4b). These genes do not correspond to the temperature responsive genes identified here (Figure 4c). Growing evidence indicates that CSR-1 associated sRNAs activate or license transcription of their target genes43,44,45,46. By contrast, our analysis of targets of WAGO-9, which is involved in RNAi-initiated heritable transcriptional silencing7,14, indicate that WAGO-9 binding 22G RNAs target the temperature-responsive transcripts (Figure 4d).
WAGO class 22G RNAs but not CSR-1 class 22G RNAs are associated with transcripts showing heritable temperature effects.
(a) Among transcripts targeted by 22G RNAs isolated from oocytes35, most are targeted by either WAGO-9 binding7 or CSR-1 binding 22G RNAs42, as previously reported14. (b) CSR-1 binding 22G RNAs tend to target transcripts that are highly expressed in 4-cell stage embryos. Histograms are distributions of fluorescence values on 20°C control microarray hybridizations. Dotted lines are median values for each group of genes indicated. (c) Failure of the temperature-responsive mRNAs highlighted in Figure 1 to overlap with 22G RNAs that co-immunoprecipitate with the germline argonaute CSR-142 (26% of the genes represented on our microarray). P(no overlap) = 1 × 10−3 (P = 2 × 10−4 if analysis is restricted to only transcripts targeted by oocyte 22G RNAs). (d) By contrast, overlap with 22G RNAs that co-immunoprecipitate with the germline argonaute WAGO-97. P(equal or greater overlap) = 2 × 10−5 (P = 0.03 if analysis is restricted to only transcripts targeted by oocyte 22G RNAs).
Heritable silencing can be triggered by PRG-1 and 21U RNAs14,15,16, which appear to generate WAGO-associated 22G secondary sRNA23,24. Therefore, we wondered whether the temperature-responsive transcripts are also targeted by prg-1 dependent sRNAs. Indeed, most of the temperature-responsive transcripts that are targets of oocyte 22G RNAs are also predicted 21U RNA targets where presence of 22G RNAs depends on prg-123,24 (see Supplementary Table S1).
Other primary sRNAs, specifically the 26G RNAs21,47, also direct production of 22G RNAs32,48,49. The Argonautes ALG-3 and ALG-4 act with 26G RNAs in the male germline32, while the Argonaute ERGO-1 acts with 26G RNAs found mostly in embryos and oocytes to silence genes in somatic tissues48,49,50. No known 26G RNA targets are represented among the highlighted genes in Figure 1. In conclusion, our analysis of published sRNA targets indicate that only PRG-1 dependent, WAGO bound 22G RNAs target transcripts associated with heritable effects of temperature.
Therefore, we sought to determine how well targeting by PRG-1 dependent germline sRNAs predicts heritable temperature effects. We first established a stringent statistical filter (less than one expected false positive based on permutation tests) to define a high confidence set of genes whose transcript levels vary in response to 25°C growth (G2 of Figure 1). We then defined, by consensus among published experiments, a high confidence set of transcripts that are targets of endogenous germline sRNAs. We found that among transcripts with strong evidence for an initial temperature response in G2, those that are likely targets of the PRG-1 pathway are more than ten times more likely than other transcripts to show a heritable effect upon return to 20°C (G3) (Figure 5).
Targets of PRG-1 dependent 22G RNAs are more likely than other transcripts to show effects in the following generation.
(a) Among transcripts differing in G2, likely PRG-1 targets are more likely to show a difference in G3, P = 1 × 10−8. Circle labeled “G2” represents genes whose mRNA levels differ >3× between treatment and control (one-tailed Student's T P < 0.001; this results in an estimated false positive rate for G2 of approximately 1 gene out of 15,208 based on swap tests). Circle labeled “targets of prg-1 dependent 22G RNAs” represents genes for which antisense 22G RNAs are consistently depleted in five comparisons of N2 (wild type) with prg-1 mutants23,24,72,73,74 (see Methods). (b) Same analysis as in (a), looking only at targets of oocyte 22G RNAs. Among transcripts differing in G2, likely PRG-1 targets are more likely to show a difference in G3, P = 4 × 10−3. (c) Among targets of oocyte 22G RNAs35, predicted 21U RNA targets23,24 tend to be WAGO targets7 and not CSR-1 targets42, as previously reported14.
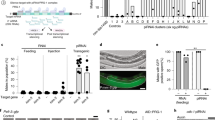
The presence of 22G RNAs antisense to the temperature-responsive transcripts raises the question of whether the 22G RNAs themselves respond to temperature as well. We used RT-QPCR to measure the abundance of one 22G RNA sequence antisense to each of B0286.1 and K10B3.5 before, during and for two generations after return to 20°C. We found that the B0286.1 22G RNA levels increase when B0286.1 transcript decreases in response to 25°C, while K10B3.5 22G RNA decreases when K10B3.5 transcript increases in response to 25°C (Figure 6a; abundances vary with generation in N2 data, ANOVA P = 2 × 10−5 for B0286.1 and 4 × 10−5 for K10B3.5). This would be expected if the changes in transcript levels are partly or wholly due to temperature-induced changes in 22G RNAs, the 22G RNAs acting as short interfering RNAs (siRNAs) to reduce transcript levels. Thus, endogenous silencing of K10B3.5 is stronger at 20°C than at 25°, while endogenous silencing of B0286.1 is stronger at 25° than at 20°.
B0286.1 and K10B3.5 transcripts are targets of temperature-responsive endogenous RNAi.
(a) Response of 22 nt RNAs (geometric mean ± SEM) to temperature. (b) Effect of inactivating germline RNAi on B0286.1 and K10B3.5 transcripts. WT, values for N2 (wild type) duplicated from Figure 2; mut-2, values for WM30 mut-2(ne298); GFP and mut-16, feeding RNAi51 of GFP::unc-22 and mut-16, respectively, from G1 to end of experiment. * & , two-way ANOVA effect & interaction, respectively P < 10−3. See Supplementary Table S6 for statistical details.
If the endogenous sRNA silences the corresponding genes, one prediction is that inactivating germline RNAi would increase transcript levels. Among germline RNAi defective mutants, the mut-2(ne298) strain WM30 is unusual in that the worms remain sufficiently fertile at 25° to allow transgenerational experiments. As expected based on sequencing of sRNAs from mut-2(ne298)40, compared to wild type (N2), WM30 has reduced levels of the selected sRNAs that target B0286.1 and K10B3.5 (Figure 6a). Comparisons between wild-type and mutator strains are complicated by likely additional unknown mutational differences. Therefore, as a complementary approach, we used RNAi to disrupt germline RNAi in wild-type worms (N2). Specifically, we fed worms E. coli food that expresses mut-16 dsRNA51, or as a negative control, food that knocks down the reporter gene GFP and the muscle gene unc-22. In most cases, we saw a substantial increase in transcript levels when germline RNAi is impaired (Figure 6b). We conclude that B0286.1 and K10B3.5 are normally silenced by germline RNAi.
A second prediction is that inactivating germline RNAi would cause the loss of the temperature effect. At least in the case of B0286.1, the absence of germline RNAi nearly eliminates the heritable temperature response (Figure 6b, 2-way ANOVA tests for interaction, P = 5 × 10−7 in the mut-2 experiment and P = 3 × 10−4 in the mut-16 experiment). We conclude that RNAi-defective worms do not respond to temperature in a wild-type manner. This result indicates that the sRNAs are not merely passive observers of transcript changes, but instead play an active role.
Recent work has suggested that siRNAs produced from artificially introduced dsRNA can be passed from parent to offspring in C. elegans9 and can persist for up to 3 or 4 generations after dsRNA exposure7, in which case siRNAs are presumably regenerated in each generation. In this light, it is interesting to note that B0286.1 22 nt RNA levels remain elevated for at least 1½ generations after heat stress is removed – through the adult stage twice at 20° after exposure to 25° (Figure 6a, P = 0.03, two-tailed Student's T-test for difference with 20° control). It is unknown whether any of this RNA is inherited, or alternately it is entirely regenerated in response to an unknown heritable molecule or mark. Nevertheless, the high levels of endogenous sRNA to specific genes in oocytes (Figure 3a) are suggestive of an inherited signal.
Finally, we note that spermatocytes contain a relatively high level of sRNAs antisense to B0286.152 (see Supplementary Table S1), leading us to wonder whether part of the heritable effect passes through the male line. To test this idea, we measured B0286.1 transcript levels in the progeny of 20°C raised hermaphrodites that were themselves the cross progeny of males and hermaphrodites raised at 20°C or 25°C (all four combinations tested; Figure 7). We found that the heritable effect passes almost entirely through the female line.
Female line transmission of the heritable effect on B0286.1 transcript levels.
We either exposed worm cultures to 25°C or kept them at 20°C and crossed male HC67 progeny with hermaphrodite N2 progeny, using the GFP transgene from HC67 to identify cross-progeny. All four possible crosses are shown. Values are B0286.1 mRNA abundances in the second generation after the crosses, geometric mean ± SEM, relative to the self-fertilized N2 20°C control from Figure 2 (dotted line). In two-way ANOVA (n = 7 cultures per cross), **P = 3 × 10−7 for the effect of the female line; P = 0.2 for the effect of the male line; and P = 0.8 for the interaction effect.
Discussion
Here we present a proof-of-principle finding that a common environmental stress triggers heritable changes in gene expression in animals. To identify genes that show a persistent change in gene expression in response to environmental change, we measured transcript levels in small populations of precisely staged animals that were transiently exposed to a mild heat stress. These data show that memory of the environmental stress persists as seen in altered transcript levels for two to three generations after return to the previous environmental condition. Extending the stress exposure to two consecutive generations doubles the magnitude of the altered transcript levels and, for at least one gene, extends by at least a generation the duration of the memory. This indicates that the information persists for multiple generations and also suggests that the inherited state is not a simple on-off switch (Supplementary Table S3).
We found that genes that show heritable changes in gene expression are normally silenced by endogenous RNAi. The affected transcripts are enriched for genes targeted by 22G germline antisense short RNAs and at least for the two transcripts chosen for further study, the abundances of 22G RNAs change in the opposite direction of their target mRNAs (Figure 6). These results imply that endogenous RNAi is important for initiating, maintaining, or expressing the heritable effects. One gene we investigated further is more efficiently silenced at 25°C than at 20°C, while the other is more efficiently silenced at 20° than at 25°C. Additionally, the effect of temperature on transcript levels is missing in RNAi defective worms. Although we analysed only two genes in detail, both categories of increasing and decreasing transcripts (Figure 1) are targeted by siRNAs (Supplementary Figure S2 and Supplementary Table S1), indicating a specific response to temperature changes rather than a global change in gene silencing efficiency. Indeed, simple numerical models we constructed show that a single mechanism deployed at different temperatures is sufficient to explain inheritance of both increased and decreased transcript (Figure 8; Supplementary Table S3)
A single-mechanism numerical model recapitulates divergent response to temperature changes.
Graphic results from a simple continuous function model (Model 2 in Supplementary Table S3). The assumptions for this numerical model are that a new temperature-dependent silencing signal is generated at the silencing temperature and that this signal is maintained at the non-silencing temperature by limited, incomplete amplification resulting in dilution of the signal with each new generation.
Consistent with a single mechanism regulating temperature responsive genes that both increase and decrease in abundance, the set of genes is strongly enriched for WAGO-associated 22G sRNA targets. It is remarkable that none of the genes are targets of CSR-1-associated sRNA. CSR-1 sRNAs predominantly target abundantly expressed maternal genes, protecting them from silencing by WAGO-dependent processes. We note that starvation induced transcripts do include CSR-1 sRNA targets33, although our study used different methods and looked at a different developmental stage and a different stress.
Analysis of our data combined with recent findings in other labs indicates that targets of PRG-1, an Argonaute protein in the Piwi family widely conserved among animal phyla, include the genes that show heritable temperature responses. Identifying the specific 21U RNAs (C. elegans piRNAs) involved is not straightforward because 21U RNAs do not need a perfect sequence match to act as primary siRNAs for secondary 22G RNA production and individual 21U species are far less abundant than their secondary 22G RNA products. In the future, it will be interesting to see whether the production or maturation of specific 21U RNAs responds to ecologically relevant stimuli.
More generally, the identification of environment-sensitive siRNA expression may serve to identify transcripts sensitized for transgenerational epigenetic reprogramming. Indeed, recent studies have identified starvation-induced and dauer-induced siRNAs33,53 with starvation being associated with transgenerational effects on targeted transcripts. Because sRNAs can target any arbitrary gene, a potentially large amount of information about environmental conditions can be encoded in sRNA profiles for transmission to future generations.
We also found that the heritable effect passes primarily through the female line. This contrasts with heritable silencing triggered by exogenous RNA in C. elegans6 and heritable epigenetic effects in mammals4,54,55 where heritable effects pass either through the male line or through both the female and male lines. However, our finding is consistent with inherited gene silencing triggered by a 21U RNA16. In an experimental system described in ref. 16, a heritably silenced locus can impose silencing on a second transgenic reporter, but only by way of the female line and not the male line. Since the two transgenes do not have to be simultaneously present for silencing to be transferred, the authors speculate that a diffusible factor in the oocyte cytoplasm is responsible for the heritable effect16.
Although many have been tempted to propose transgenerational inheritance of small RNAs, there is as yet no evidence that abundant secondary siRNAs, themselves the products of RNA directed RNA polymerase activity, can direct further amplification (see for example ref. 56) that can overcome the hundredfold or greater dilution that would occur with each generation. Thus, the precise marks or signals that mediate transgenerational inheritance remain unknown.
Variation in chromatin is a candidate for a heritable mark or signal that may act in concert with RNAi to preserve expression levels. Indeed, work on heterochromatin maintenance at centromeres in the fission yeast Schizosaccharomyces pombe and on heterochromatin-triggered gene silencing in the fruit fly Drosophila, suggests that RNAi-like mechanisms might work closely with chromatin modifications in a positive feedback loop57,58. In C. elegans effective RNAi requires heterochromatin-promoting activities and RNAi triggers the accumulation of histone H3 lysine 9 methylation at silenced loci5,51,59,60,61. Furthermore, manipulating histone modifications can trigger heritable changes in worm longevity8, although it is unknown whether this changes the abundances of siRNAs that target genes that affect aging.
Additionally, there is as yet no way to entirely rule out other signals such as transcription factors, prions, trace amounts of DNA methylation, or even hormones and nutrients deposited with the yolk as inherited temperature signals in C. elegans3. Thus, much additional work is needed to understand the mechanisms of transgenerational epigenetic inheritance.
Also of interest is whether the signal originates in somatic tissues, which then communicate with the germline62. RNAi in C. elegans is systemic; dsRNA introduced locally can spread to cause silencing in cells throughout the animals and in future generations. Thus, the involvement of an RNAi-related mechanism in the inherited response raises the possibility that gene-specific information about variable environmental conditions may be exchanged between somatic cells and the germline. In this regard we note that the effects we describe in this paper are independent of the SID-1 dsRNA channel that is responsible for systemic RNAi (see Methods), but we cannot rule out SID-1 dependent effects that may have gone undetected in this study. Additionally, there are many SID-1 independent ways that a signal could travel between somatic and germline tissues. Also pertinent to this question is the observation that RNAi is more potent in the germline than the soma63,64, raising the question as to whether the heritable effects are restricted to the germline. In one study of heritable germline silencing of a GFP-expressing transgene in C. elegans, silencing did not escape the germline into somatic tissues15, but it is unclear whether the GFP experiment is representative of all piRNA-triggered heritable gene silencing.
Temperature change is a stress that C. elegans likely encounters frequently in the wild; therefore the ability to inherit responses should provide a selective advantage. In this regard it is noteworthy that the transcriptome of 4-cell embryos varies substantially in response to temperature among different geographical isolates of C. elegans65 and seven of the transcripts highlighted in Figure 1 show an interaction between environmental conditions and natural genotypic variation65. For example, both scrm-4 and K10B3.5 transcripts increase at 25°C in strains from the UK (N2), California (CB4857) Australia (AB2) and Germany (RC301), but in a divergent strain from Hawaii (CB4856), scrm-4 transcript is constitutively high and K10B3.5 transcript is constitutively low65. However, the physiological significance of any of the heritable gene expression changes remains unknown. The list of genes (Supplementary Table S1) provides few clues in this regard. If changes in multiple transcripts each contribute a small but significant amount to fitness, it may prove to difficult to demonstrate a selective advantage for expression changes for individual genes: A recent study showed that a majority of genes in C. elegans confer a significant but small fitness advantage in a single environmental condition66. It will be interesting to see whether different kinds of environmental stimuli produce distinctly different heritable patterns of gene expression and whether the gene expression changes result in phenotypes relevant to surviving stresses.
Individuals show transcriptional responses to new environments within minutes, while populations show adaptive genetic change by natural selection over many generations. Mechanisms that enable adaptation to variable or recurring environmental conditions over intermediate timescales are poorly understood. The study of inheritance of acquired characteristics has long been plagued by controversy and irreproducible results (see for example ref. 67). Our work shows it is possible to identify specific, quantitative molecular markers of inheritance, which will facilitate replication. Studying transgenerational inheritance in C. elegans, with its wealth of molecular and genetic tools, will provide a unique perspective to this still largely unexplored territory.
Methods
Worm culture
We used standard methods to culture and handle C. elegans, except we used the nonpathogenic, sporulation-defective Bacillus subtilis strain RL1275 (spoIIAC::erm in a PY79 background, a kind gift from Richard Losick) as worm food for most experiments in order to see temperature effects independently of pathogen responses. B. subtilis is substantially less pathogenic to C. elegans than the standard C. elegans food E. coli strain OP50 at 25°C68.
The only experiment presented here that did not use RL1275 as food is the feeding RNAi experiment (right hand half of Figure 6B), where the RNase III defective E. coli strain HT115(DE3) bearing either the Ahringer library clone I-5E09 (L4440 with a mut-16 insert) or the plasmid pPD126.25 (GFP::unc-22 hairpin, lab of Andrew Fire) was used as worm food on NGM plates containing 25 μg/ml carbenicillin and 1 mM IPTG.
Before each of the experiments, we cultured the worms for at least three generations at 20°C without allowing the worms to deplete their food. Other than the 25°C treatments, done in a 25 ± 0.5° incubator, all incubations and procedures involving live worms were done in a temperature controlled room at 20 ± 0.3°C. We used HOBO miniature temperature loggers (Onset Computer Corporation, Bourne, MA, USA) to continuously monitor culture temperatures. To minimize temperature variation during handling, we used large (10 cm diameter) plates, wore insulated gloves while handling plates and used an external light source for the dissecting microscope.
For the microarray experiment, we used a platinum wire loop to transfer several hundred embryos to a fresh plate for each and every generation and used a platinum wire pick to remove any hatched worms that were carried along with the eggs. For the multigenerational follow-up experiments, we used a platinum wire pick to transfer 60 L4 larvae to a fresh plate for each generation. The purpose of transferring L4 larvae instead of embryos in the multigenerational experiment was to eliminate males from the cultures, thereby eliminating variation in the frequency of sex-specific imprinting (most C. elegans individuals are hermaphrodites, with a low rate of spontaneous males that increases with temperature).
All transfers were either to prewarmed 25° plates, or to precooled 20°C plates, as appropriate. For post-25°C steps of all experiments, we interleaved the constant 20°C (control) plates with post-25°C treatment plates in common stacks whenever possible, so that treatment and control plates experience the same microenvironment on average. In practice, the treatment plates were slightly staggered in time relative to the corresponding control plates of the same generation, because of time constraints and because development time from zygote to adult is ~17 hours faster at 25°C than at 20°C. The total time between transfers of eggs or L4 larvae to new plates — the total amount of time per generation — was 4 days for incubation at 20°C, or 3 days for incubation at 25°C.
Embryo collection
For each combination of condition and generation in the microarray experiment, we collected three replicates of the standard C. elegans laboratory strain N2 and three replicates of the strain HC445 (sid-1(qt9) in an N2 background). For all subsequent experiments (with the exception of the WM30 experiment), we used N2 alone.
We collected embryos and purified RNA as described34, except we increased the number of embryos per sample to 50. In brief, we picked young adults into water, chopped them in half, treated them 30 seconds with alkaline bleach (final concentration 80 mM KOH, 50 mM NaOCl) to destroy any RNA and protein not protected by the eggshell, stopped the bleach reaction with bovine serum albumin (final concentration 3%), collected and washed early embryos three times in nuclease-free water using a glass mouth pipette, flash-froze the embryos in liquid nitrogen in low-adhesion 600 μl microcentrifuge tubes and transferred the tubes to a −80°C freezer for storage. For the microarray experiment in Figure 1, the total time from picking of the first worm to bleaching was not allowed to exceed 15 min. For all subsequent experiments, the time was not allowed to exceed 8 min. We added 100 μl Trizol (phenol/guanidine isothiocyanate; Invitrogen Corporation, now part of Life Technologies, Carlsbad, CA, USA), with added linear polyacrylamide (5 μg) and yeast tRNA (100 ng) as carriers, to the frozen embryos for RNA extraction and isopropanol precipitation. We used the RNA pellet directly for RT (reverse transcription).
Microarrays
We used a custom microarray (Agilent Technologies, Santa Clara, CA, USA) bearing 60-mer oligonucleotide spots corresponding to 15,208 C. elegans genes. Since this is fewer than the known number of genes in C. elegans, we limited the genes to predicted protein-coding genes, prioritizing genes with 1:1 best reciprocal similarity scores to C. briggsae. We selected oligonucleotide sequences corresponding to the 3′-most predicted exon that allows design of a probe that minimizes cross-hybridization with other genes. Gene positions on the array were randomized to minimize bias due to hybridization artifacts.
For microarray hybridizations, we performed two rounds of linear amplification34,69 using the MessageAmp II kit (Ambion, now part of Life Technologies, Carlsbad, CA, USA) starting from a T7-oligo-dT RT primer. The following modifications were designed to increase the average lengths of the amplified RNA populations34: First-round cDNA construction from total RNA was carried out at 1/5 of the recommended quantities and total volume, while IVT was carried out at 1/2 of the recommended volume. 100 ng amplified RNA from the first round was used as input for the second round of amplification following the manufacturers protocol for amino-allyl modified nucleotides. We labelled 10 μg amplified RNA with Cy3 fluorescent dye using the Ambion MessageAmpII kit and used 1.65 μg labelled amplified RNA to hybridize with each array. To minimize post-amplification biases, we randomized the position and order of hybridizations. Hybridizations with obvious artefacts (a bubble, a fingerprint, or a notably skewed fluorescence profile) were redone. We scanned the arrays at 10% laser power to avoid signal saturation.
For microarray data, we used the Agilent Feature Extraction software GE1-v5_95_Feb07 to calculated fluorescence values. We log2 transformed the fluorescence values and quantile-normalized70 them among hybridizations. Normalization may introduce slight biases due to a shift in global transcript abundances at 25°C, but not enough to affect the paper's conclusions. With the exception of the 10% of genes with lowest fluorescence signal, the processed values closely fit a Gaussian distribution for each gene, allowing us to use standard parametric statistics. Supplementary Figure S1 summarizes randomization/permutation tests of the microarray data. Supplementary Figure S3 shows a test of linearity. Supplementary Figure S4 shows that the genes identified in Figure 1 have a random spatial distribution on the physical microarray, indicating that the identification of these genes is not due to common hybridization artefacts.
Initial analysis suggested the microarray data lack the statistical power to conclusively show sid-1 dependent heritable effects; therefore we set aside for the future description of such effects. A portion of the microarray data (three replicates of the first generation but none of the second) has been reported previously65.
Detailed microarray data and methods are available in Supplementary Table S2 and the Gene Expression Omnibus (GEO) GSE30666.
Reverse transcription-quantitative PCR
As reference genes for reverse transcription-quantitative PCR (RT-QPCR) we chose atf-6 and cpf-1, two well-characterized genes whose hybridization signal varies little in our microarray experiments. We performed Thermoscript RT (Invitrogen Corporation, now part of Life Technologies, Carlsbad, CA, USA) reactions using a mix of gene-specific RT primers (“RT” listed in Supplementary Table S4). A critical step to ensure proper mixing in setting up RT reactions was to mix by stirring with a pipette tip, monitored under a microscope. In order to get reliable amplification from 50-embryo samples, we needed to perform nested PCR. We amplified the cDNA in a multiplex PCR reaction containing “RT” and “outer” primers (listed in Supplementary Table S4) for 19 cycles in the presence of SYBR Green to monitor amplification. We diluted this first PCR product 1250-fold and divided it as template for separate QPCR reactions (Quantitect; Qiagen N.V., Hilden, Germany) for each gene (“QF” and “QR” primers, Supplementary Table S4).
We calculated relative mRNA levels from cycle thresholds as  . However, like with the microarray data, we calculated means and other statistics on logarithmic-scale instead of linear-scale data. Serial dilution experiments show that the RT-QPCR methods we used yield reasonably linear results (Supplementary Figure S5).
. However, like with the microarray data, we calculated means and other statistics on logarithmic-scale instead of linear-scale data. Serial dilution experiments show that the RT-QPCR methods we used yield reasonably linear results (Supplementary Figure S5).
We focused primarily on collecting samples from the second generation after return to 20° and the corresponding generation of the constant 20° control. Plates with visible contamination or excessive mineral precipitate were discarded without sampling. Occasional QPCR reactions (~2%) failed to yield fluorescence within 25 or more cycles in the second PCR and were treated as missing data.
We used custom TaqMan (Applied Biosystems, now part of Life Technologies, Carlsbad, CA, USA) RT-QPCR assays to measure abundances of short RNAs in total nucleic acid extracted from 1.5 day old adult worms. Nucleic acid was prepared by proteinase K digestion for 10 minutes at 65°, followed by two rounds of phenol-chloroform extraction, using silicone grease (vacuum grease; Dow Corning Corporation, Midland, MI, USA) to separate the aqueous phase from the organic phase and then sodium acetate-isopropanol precipitation (based on a protocol suggested by Weifeng Gu). We performed the TaqMan reverse transcriptions at 1/3 the scale of the manufacturer's protocol, but used 13.36 ng nucleic acid (as measured by absorbance at 260 nm) as template for each reaction. Estimates are 2−CT.
Short RNA sequence analysis
We compiled 22 nt sequences starting with G from datasets SRR185587-94.sra (N223), SRR185595-8.sra (prg-123), SRR513311.sra (N224), SRR513312.sra (prg-124), SRR1175718.sra (N271), SRR1175716.sra (prg-171), SRR553522.sra (N272), SRR553525.sra (F12 prg-172), SRR943469.sra (N273) and SRR943473.sra (prg-173) (retrieved through the NCBI GEO website) and aligned them for perfect antisense matches to predicted C. elegans cDNAs using bowtie 1.0.074, calculating reads per million total aligned 22G sequences (RPM). In order to be included in the set of high-confidence PRG-1 dependent targets, all of the following had to apply to the gene across the five studies: a) the average RPM in N2 > 100, b) the average RPM in N2 > 25 × the average RPM in the prg-1 strains, c) the lowest RPM in N2 > the highest RPM in the prg-1 strains and d) RPM in N2 > 25 × RPM in prg-1 in every one of the five studies. The resulting set of genes overlaps, but is not identical to, the two previously published lists of predicted 21U RNA targets23,24 (Supplementary Figure S6).
Additional information
Accession codes Microarray data and methods: Gene Expression Omnibus (GEO) GSE30666 http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc5GSE30666
References
Daxinger, L. & Whitelaw, E. Transgenerational epigenetic inheritance: more questions than answers. Genome Research 20, 1623–1628 (2010).
Daxinger, L. & Whitelaw, E. Understanding transgenerational epigenetic inheritance via the gametes in mammals. Nature Reviews Genetics 13, 153–162 (2012).
Lim, J. P. & Brunet, A. Bridging the transgenerational gap with epigenetic memory. Trends in Genetics: TIG 29, 176–186 (2013).
Kaati, G., Bygren, L. O., Pembrey, M. & Sjostrom, M. Transgenerational response to nutrition, early life circumstances and longevity. Eur J Hum Genet 15, 784–790 (2007).
Vastenhouw et al. Gene expression: long-term gene silencing by RNAi. Nature 442, 882 (2006).
Alcazar, R. M., Lin, R. & Fire, A. Z. Transmission dynamics of heritable silencing induced by double-stranded RNA in Caenorhabditis elegans. Genetics 180, 1275–1288 (2008).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012).
Greer, E. L. et al. Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans. Nature 479, 365–371 (2011).
Rechavi, O., Minevich, G. & Hobert, O. Transgenerational inheritance of an acquired small RNA-based antiviral response in C. elegans. Cell 147, 1248–1256 (2011).
Cubas, P., Vincent, C. & Coen, E. An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401, 157–161 (1999).
Simpson, V. J., Johnson, T. E. & Hammen, R. F. Caenorhabditis elegans DNA does not contain 5-methylcytosine at any time during development or aging. Nucleic Acids Research 14, 6711–6719 (1986).
Gutierrez, A. & Sommer, R. J. Evolution of dnmt-2 and mbd-2-like genes in the free-living nematodes Pristionchus pacificus, Caenorhabditis elegans and Caenorhabditis briggsae. Nucleic Acids Res 32, 6388–6396 (2004).
Grentzinger, T. et al. piRNA-mediated transgenerational inheritance of an acquired trait. Genome research 22, 1877–1888 (2012).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Ashe, A. et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Luteijn, M. J. et al. Extremely stable Piwi-induced gene silencing in Caenorhabditis elegans. The EMBO Journal 31, 3422–3430 (2012).
Batista, P. J. et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Molecular Cell 31, 67–78 (2008).
Das, P. P. et al. Piwi and piRNAs act upstream of an endogenous siRNA pathway to suppress Tc3 transposon mobility in the Caenorhabditis elegans germline. Molecular Cell 31, 79–90 (2008).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Sijen, T., Steiner, F. A., Thijssen, K. L. & Plasterk, R. H. Secondary siRNAs result from unprimed RNA synthesis and form a distinct class. Science 315, 244–247 (2007).
Ruby, J. G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Gu, W. et al. CapSeq and CIP-TAP identify Pol II start sites and reveal capped small RNAs as C. elegans piRNA precursors. Cell 151, 1488–1500 (2012).
Bagijn, M. P. et al. Function, targets and evolution of Caenorhabditis elegans piRNAs. Science 337, 574–578 (2012).
Lee, H. C. et al. C. elegans piRNAs mediate the genome-wide surveillance of germline transcripts. Cell 150, 78–87 (2012).
Tabara, H. et al. The rde-1 gene, RNA interference and transposon silencing in C. elegans. Cell 99, 123–132 (1999).
Ketting, R. F., Haverkamp, T. H., van Luenen, H. G. & Plasterk, R. H. Mut-7 of C. elegans, required for transposon silencing and RNA interference, is a homolog of Werner syndrome helicase and RNaseD. Cell 99, 133–141 (1999).
Sijen, T. & Plasterk, R. H. Transposon silencing in the Caenorhabditis elegans germ line by natural RNAi. Nature 426, 310–314 (2003).
Wilkins, C. et al. RNA interference is an antiviral defence mechanism in Caenorhabditis elegans. Nature 436, 1044–1047 (2005).
Schott, D. H., Cureton, D. K., Whelan, S. P. & Hunter, C. P. An antiviral role for the RNA interference machinery in Caenorhabditis elegans. Proc Natl Acad Sci U.S.A. 102, 18420–18424 (2005).
Felix, M. A. et al. Natural and experimental infection of Caenorhabditis nematodes by novel viruses related to nodaviruses. PLoS Biology 9, e1000586 (2011).
Welker, N. C., Habig, J. W. & Bass, B. L. Genes misregulated in C. elegans deficient in Dicer, RDE-4, or RDE-1 are enriched for innate immunity genes. RNA 13, 1090–1102 (2007).
Conine, C. C. et al. Argonautes ALG-3 and ALG-4 are required for spermatogenesis-specific 26G-RNAs and thermotolerant sperm in Caenorhabditis elegans. Proc Natl Acad Sci U.S.A. 107, 3588–3593 (2010).
Rechavi, O. et al. Starvation-induced transgenerational inheritance of small RNAs in C. elegans. Cell 158, 277–287 (2014).
Baugh, L. R., Hill, A. A., Slonim, D. K., Brown, E. L. & Hunter, C. P. Composition and dynamics of the Caenorhabditis elegans early embryonic transcriptome. Development 130, 889–900 (2003).
Gu, W. et al. Distinct argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans germline. Molecular Cell 36, 231–244 (2009).
Zhang, C. et al. mut-16 and other mutator class genes modulate 22G and 26G siRNA pathways in Caenorhabditis elegans. Proc Natl Acad Sci U.S.A. 108, 1201–1208 (2011).
Grishok, A., Tabara, H. & Mello, C. C. Genetic requirements for inheritance of RNAi in C. elegans. Science 287, 2494–2497 (2000).
Dernburg, A. F., Zalevsky, J., Colaiacovo, M. P. & Villeneuve, A. M. Transgene-mediated cosuppression in the C. elegans germ line. Genes & Development 14, 1578–1583 (2000).
Tijsterman, M., Ketting, R. F., Okihara, K. L., Sijen, T. & Plasterk, R. H. RNA helicase MUT-14-dependent gene silencing triggered in C. elegans by short antisense RNAs. Science 295, 694–697 (2002).
Phillips, C. M. et al. MUT-14 and SMUT-1 DEAD box RNA helicases have overlapping roles in germline RNAi and endogenous siRNA formation. Current Biology: CB 24, 839–844 (2014).
Phillips, C. M., Montgomery, T. A., Breen, P. C. & Ruvkun, G. MUT-16 promotes formation of perinuclear mutator foci required for RNA silencing in the C. elegans germline. Genes & Development 26, 1433–1444 (2012).
Claycomb, J. M. et al. The Argonaute CSR-1 and its 22G-RNA cofactors are required for holocentric chromosome segregation. Cell 139, 123–134 (2009).
Seth, M. et al. The C. elegans CSR-1 argonaute pathway counteracts epigenetic silencing to promote germline gene expression. Developmental Cell 27, 656–663 (2013).
Wedeles, C. J., Wu, M. Z. & Claycomb, J. M. Protection of germline gene expression by the C. elegans Argonaute CSR-1. Developmental Cell 27, 664–671 (2013).
Conine, C. C. et al. Argonautes promote male fertility and provide a paternal memory of germline gene expression in C. elegans. Cell 155, 1532–1544 (2013).
Cecere, G., Hoersch, S., O'Keeffe, S., Sachidanandam, R. & Grishok, A. Global effects of the CSR-1 RNA interference pathway on the transcriptional landscape. Nat Struct Mol Biol 21, 358–365 (2014).
Han, T. et al. 26G endo-siRNAs regulate spermatogenic and zygotic gene expression in Caenorhabditis elegans. Proc Natl Acad Sci U.S.A. 106, 18674–18679 (2009).
Gent, J. I. et al. Distinct phases of siRNA synthesis in an endogenous RNAi pathway in C. elegans soma. Molecular Cell 37, 679–689 (2010).
Vasale, J. J. et al. Sequential rounds of RNA-dependent RNA transcription drive endogenous small-RNA biogenesis in the ERGO-1/Argonaute pathway. Proc Natl Acad Sci U.S.A. 107, 3582–3587 (2010).
Fischer, S. E. et al. The ERI-6/7 helicase acts at the first stage of an siRNA amplification pathway that targets recent gene duplications. PLoS Genetics 7, e1002369 (2011).
Kim, J. K. et al. Functional genomic analysis of RNA interference in C. elegans. Science 308, 1164–1167 (2005).
Gent, J. I. et al. A Caenorhabditis elegans RNA-directed RNA polymerase in sperm development and endogenous RNA interference. Genetics 183, 1297–1314 (2009).
Hall, S. E., Chirn, G. W., Lau, N. C. & Sengupta, P. RNAi pathways contribute to developmental history-dependent phenotypic plasticity in C. elegans. RNA 19, 306–319 (2013).
Rassoulzadegan, M., Grandjean, V., Gounon, P., Vincent, S., Gillot, I. & Cuzin, F. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006).
Anway, M. D., Cupp, A. S., Uzumcu, M. & Skinner, M. K. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 308, 1466–1469 (2005).
Pak, J., Maniar, J. M., Mello, C. C. & Fire, A. Protection from feed-forward amplification in an amplified RNAi mechanism. Cell 151, 885–899 (2012).
Todeschini, A. L., Teysset, L., Delmarre, V. & Ronsseray, S. The epigenetic trans-silencing effect in Drosophila involves maternally-transmitted small RNAs whose production depends on the piRNA pathway and HP1. PloS One 5, e11032 (2010).
Moazed, D. Mechanisms for the inheritance of chromatin states. Cell 146, 510–518 (2011).
Robert, V. J., Sijen, T., van Wolfswinkel, J. & Plasterk, R. H. Chromatin and RNAi factors protect the C. elegans germline against repetitive sequences. Genes & Development 19, 782–787 (2005).
Burton, N. O., Burkhart, K. B. & Kennedy, S. Nuclear RNAi maintains heritable gene silencing in Caenorhabditis elegans. Proc Natl Acad Sci U.S.A. 108, 19683–19688 (2011).
Gu, S. G. et al. Amplification of siRNA in Caenorhabditis elegans generates a transgenerational sequence-targeted histone H3 lysine 9 methylation footprint. Nature Genetics 44, 157–164 (2012).
Weismann, A. Prüfung der Hypothese einer Vererbung funktioneller Abänderungen. (Gustav Fischer Jena, 1902).
Wang, D. et al. Somatic misexpression of germline P granules and enhanced RNA interference in retinoblastoma pathway mutants. Nature 436, 593–597 (2005).
Wu, X., Shi, Z., Cui, M., Han, M. & Ruvkun, G. Repression of germline RNAi pathways in somatic cells by retinoblastoma pathway chromatin complexes. PLoS Genetics 8, e1002542 (2012).
Grishkevich, V. et al. A genomic bias for genotype-environment interactions in C. elegans. Molecular Systems Biology 8, 587 (2012).
Ramani, A. K. et al. The majority of animal genes are required for wild-type fitness. Cell 148, 792–802 (2012).
Vargas, A. O. Did Paul Kammerer discover epigenetic inheritance? A modern look at the controversial midwife toad experiments. J Exp Zoo B Mol Dev Evol 312, 667–678 (2009).
Garsin, D. A. et al. Long-lived C. elegansdaf-2 mutants are resistant to bacterial pathogens. Science 300, 1921 (2003).
Yanai, I. & Hunter, C. P. Comparison of diverse developmental transcriptomes reveals that coexpression of gene neighbors is not evolutionarily conserved. Genome Research 19, 2214–2220 (2009).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
de Albuquerque, B. F. et al. PID-1 is a novel factor that operates during 21U-RNA biogenesis in Caenorhabditis elegans. Genes & Development 28, 683–688 (2014).
Sarkies, M. et al. Reduced insulin/IGF-1 signaling restores germ cell immortality to Caenorhabditis elegans Piwi mutants. Cell Reports 7, 762–773 (2014).
Weick, E. M. et al. PRDE-1 is a nuclear factor essential for the biogenesis of Ruby motif-dependent piRNAs in C. elegans. Genes & Development 28, 783–796 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biology 10, R25 (2009).
Acknowledgements
We thank Merck for seed funding and National Institutes of Health GM089795 grant for support and many colleagues for comments and suggestions. We also thank the Harvard Faculty of Arts and Sciences Center for Systems Biology and multiple laboratories at Harvard's Department of Molecular and Cellular Biology, for the use of equipment and reagents. We thank the Caenorhabditis Genetics Center for strains and T. Chin-i and Y. Kohara for the use of unpublished in situ hybridization images.
Author information
Authors and Affiliations
Contributions
D.S., I.Y. and C.H. designed the experiments. I.Y. designed the custom microarray and D.S. and I.Y. performed the microarray experiments from culturing worms to scanning hybridized arrays. I.Y. extracted and assembled fluorescence values from array images. D.S. analysed the data, developed RT QPCR assays and performed the follow-up experiments. D.S. and C.H. wrote the paper. All authors discussed the data and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Table S1
Supplementary Information
Table S2
Supplementary Information
Table S3
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Schott, D., Yanai, I. & Hunter, C. Natural RNA interference directs a heritable response to the environment. Sci Rep 4, 7387 (2014). https://doi.org/10.1038/srep07387
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07387
This article is cited by
-
Molecular mechanisms of transgenerational epigenetic inheritance
Nature Reviews Genetics (2022)
-
poly(UG)-tailed RNAs in genome protection and epigenetic inheritance
Nature (2020)
-
Tissue-specific effects of temperature on proteasome function
Cell Stress and Chaperones (2020)
-
Intergenerational and transgenerational epigenetic inheritance in animals
Nature Cell Biology (2019)
-
The piRNA pathway responds to environmental signals to establish intergenerational adaptation to stress
BMC Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.