Abstract
Real-time amplification and quantification of specific nucleic acid sequences plays a major role in medical and biotechnological applications. In the case of infectious diseases, such as HIV, quantification of the pathogen-load in patient specimens is critical to assess disease progression and effectiveness of drug therapy. Typically, nucleic acid quantification requires expensive instruments, such as real-time PCR machines, which are not appropriate for on-site use and for low-resource settings. This paper describes a simple, low-cost, reaction-diffusion based method for end-point quantification of target nucleic acids undergoing enzymatic amplification. The number of target molecules is inferred from the position of the reaction-diffusion front, analogous to reading temperature in a mercury thermometer. The method was tested for HIV viral load monitoring and performed on par with conventional benchtop methods. The proposed method is suitable for nucleic acid quantification at point of care, compatible with multiplexing and high-throughput processing and can function instrument-free.
Similar content being viewed by others
Introduction
Real-time amplification and quantification of specific nucleic acid sequences has revolutionized genetic research, medical diagnostics and environmental monitoring1,2,3. In the case of infectious diseases, quantification of the pathogen-load in patient specimens is critical to assess disease progression, effectiveness of drug therapy and emergence of drug-resistance. HIV-1 is an important example4,5. Currently, nucleic acid quantification requires sophisticated and expensive instruments6,7, such as real-time PCR machines, for continuous monitoring of fluorescence emission from intercalating dye. Instrument-free, endpoint quantitative methods are highly desirable for nucleic acids-based, molecular diagnostics at the point of care, at home and in resource-poor settings. Although enzymatic amplification products can be detected without an instrument with lateral flow strips8,9,10, this method suffers from low accuracy and low sensitivity.
Over the past decade, microfluidic technology has enabled significant progress towards developing low-cost, point-of-care (POC) devices for molecular diagnostics11,12,13,14. However, current microfluidic-based, nucleic acid quantification still depends on conventional, continuous real-time monitoring of fluorescence emission intensity with benchtop optical instruments15,16,17. This requirement is a significant obstacle to enabling simple, affordable, on-site diagnostics based on nucleic acid amplification test (NAAT).
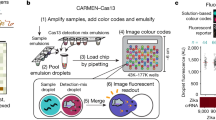
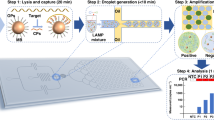
Herein, we propose a new paradigm for quantifying the number of target nucleic acid molecules in a sample. We develop a simple, low-cost, reaction-diffusion device, dubbed the “nuclemeter”, affording endpoint, quantitative detection of target nucleic acids based on the position of a reaction front. The nuclemeter is comprised of a sample chamber and a reaction-diffusion conduit, containing all the reagents needed for enzymatic amplification, as well as intercalating dye reporter (Fig. 1a–c). A sample laden with target nucleic acids is introduced into the sample chamber and the amplification reaction is triggered thermally. As time progresses, amplicons diffuse into the reaction-diffusion conduit, where they continue to react and amplify. After a certain time threshold, the conduit consists of two distinct regions (Fig. 1b): the bright, left segment (0 < x < XF), where the amplification reaction has already generated a sufficient number of amplicons to emit detectable fluorescence emission and the dark, right section (x > XF) into which amplicons have not yet diffused. As time proceeds, the reaction front (XF), separating between the bright and dark regions, propagates to the right with a constant velocity (v0). We hypothesize that the position of the reaction front indicates target analyte concentration. Many nuclemeters can be housed on a single chip and imaged simultaneously for concurrent monitoring of multiple amplification processes, calibration standards and controls. Fig. 1c features a plastic chip with four nuclemeters, many more can be housed on a single substrate. Additionally, many reaction-diffusion conduits (not shown) can branch from a single sample chamber.
The nuclemeter.
(a) A schematic depiction of the cross-section of the nuclemeter, consisting of a sample chamber and a reaction-diffusion conduit. (b) An illustration of the nuclemeter's operation. Initially, only the sample chamber contains the nucleic acid template (top). The template amplifies and diffuses into the conduit, where it continues to amplify at the appropriate amplification temperature (middle). XF indicates the position of the reaction front that propagates with a constant velocity (v0) (bottom). (c) A photograph of a plastic chip housing four nuclemeters.
Results
The nuclemeter and its portable processor
To prove the concept, we fabricated a 46 mm long × 36 mm wide Polymethyl methacrylate (PMMA) chip housing four nuclemeters (Fig. 1c). Each nuclemeter consists of a 2 mm diameter × 2.80 mm deep sample chamber (~9 μL) connected to a 330 μm wide × 330 μm deep × 12 mm long reaction-diffusion conduit (Fig. 1a and Supplementary Fig. 1). When desired, each nuclemeter can be customized to process a different target or to serve as a control. A detailed description of the nuclemeter chip's fabrication process is provided in the Methods section.
We used a custom-made, portable, processor (Fig. 2a) for nucleic acids isothermal amplification and detection. Our processor can be powered either with four AA batteries or grid power. A flexible, polyimide-based, thin film heater (inset of Fig. 2a) maintained the nuclemeter at the temperature needed for enzymatic amplification of nucleic acids, typically 62.5–65°C for the RT-LAMP process that we used in our experiments. A portable, USB-based, fluorescence microscope monitored the fluorescence emission from the various reaction-diffusion conduits and a Matlab™ program determined the emission intensity as a function of position along the reaction-diffusion conduit.
The custom-made, portable, processor for nucleic acid isothermal amplification and detection.
(a) A photograph of the processor. Inset: a flexible, polyimide-based, thin film heater. (b) A thermograph of the nuclemeter chip's surface taken with an infrared camera T360. The four reaction-diffusion reactors are located within the dashed square. (c) A mask made with black 3M Scotch electrical tape to block background emission. A ruler was fixed on the mask to assist in determining the position of the reaction front (XF).
To evaluate temperature uniformity of the nuclemeters, a thermograph of the nuclemeter chip's surface was taken with an infrared camera. The four nuclemeters located within the dashed square showed excellent temperature uniformity, within ±0.5% (Fig. 2b). To block any background emission, a mask was made with black 3M Scotch electrical tape. The mask is equipped with a ruler to assist in reading the position of the reaction front (XF) by eye (Fig. 2c).
HIV viral load test
To demonstrate the performance of the nuclemeter, we used reverse transcription, loop-mediated isothermal amplification (RT-LAMP)20,21 to quantify HIV viral load. Samples containing 0, 102, 103 and 104 HIV-1 RNA molecules were inserted into the four sample chambers (Fig. 1b) and incubated at 62.5°C using our custom-made, portable, processor (Fig. 2a). 0.04% (w/v) hydroxypropyl-methyl-cellulose (HPMC) was added to the RT-LAMP reaction mixture to slow amplicons' diffusion and obtain a well-defined reaction front (Supplementary Note 1). Upstream of the front, the amplification process had reached its conclusion due to depletion of reaction components and the fluorescence emission intensity was nearly independent of target type and concentration. The emission from the reaction-diffusion conduits was monitored with the USB fluorescent microscope (Fig. 3 and Video 1). At any given time, the greater the number of target molecules, the larger XF. Thus, with appropriate calibration, the number of initial target molecules can be inferred from XF. Although XF increases as time increases at any target concentration, the differences between XF values associated with different concentrations are time-independent. In Fig. 3, we monitored fluoresce emission for nearly an hour. However, the information needed for viral load determination is available within less than 30 minutes.
Fluorescence emission imaging from the nuclemeters used for HIV viral load testing.
The images are at 8, 24, 32, 40, 48 and 56 min after the start of incubation. The sample chambers connected to reaction-diffusion conduits 1, 2, 3 and 4 contained 104, 103, 102 and 0 (negative control) HIV-1 RNA templates.
The reproducibility of the nuclemeter was evaluated by introducing identical target concentrations (102 copies of HIV-1 RNA) into all four sample chambers (Fig. 4a). All four conduits exhibited nearly identical length emission columns XF (±4.5%) at any given time.
The reproducibility and sensitivity of the nuclemeter.
(a) Four nuclemeters each containing identical target concentrations (102 copies HIV-1 RNA) to illustrate reproducibility. (b) Evaluation of the limits of detection of the nuclemeter. Sample chambers connected to reaction-diffusion conduits 1, 2, 3 and 4 contain, respectively, 50, 50, 5 and 5 copies of HIV RNA target.
Furthermore, we tested the limit of detection of the nuclemeter by reducing the number of target molecules. We consistently detected as few as 50 RNA copies and were unable to detect 5 RNA copies (Fig. 4b). This is comparable to the performance of the benchtop, “tubed-based”, RT-LAMP method (Supplementary Fig. 4). Thus, our limit of detection is smaller than 50 copies per sample.
Experimental data analysis
Fig. 5 analyzes the experimental data and compares it with the predictions of a simple theoretical model (to be described later). We take the emission intensity to be proportional to the amplicons' concentration c(x,t), assumed uniform in each cross-section of the conduit. Fig. 5a depicts  as a function of position x at various times t. The lines and symbols correspond, respectively, to predictions and experimental data. We define the location of the reaction front XF(t) as the position at which
as a function of position x at various times t. The lines and symbols correspond, respectively, to predictions and experimental data. We define the location of the reaction front XF(t) as the position at which  . When x < XF,
. When x < XF,  and the amplification reaction is nearly complete (the bright regions with fluorescent emission in Fig. 3). When x > XF,
and the amplification reaction is nearly complete (the bright regions with fluorescent emission in Fig. 3). When x > XF,  and no amplification has yet occurred (the dark regions in Fig. 3).
and no amplification has yet occurred (the dark regions in Fig. 3).
Experimental data and theoretical predictions of nuclemeter's performance.
(a) Normalized emission intensity  as a function of position along the reaction-diffusion conduit at various times. The solid lines and symbols correspond, respectively, to predictions and experimental data. The number of target molecules is 103 copies. (b) Normalized emission intensity
as a function of position along the reaction-diffusion conduit at various times. The solid lines and symbols correspond, respectively, to predictions and experimental data. The number of target molecules is 103 copies. (b) Normalized emission intensity  as a function of time at positions x = 1.2, 1.8 and 2.4 mm along the length of the conduit. The solid lines and symbols correspond, respectively, to the predictions and experimental data. The number of target molecules is 103 copies. (c) The experimental rate of the reaction
as a function of time at positions x = 1.2, 1.8 and 2.4 mm along the length of the conduit. The solid lines and symbols correspond, respectively, to the predictions and experimental data. The number of target molecules is 103 copies. (c) The experimental rate of the reaction  as a function of position (x) at various times. (d) The measured width of the reaction-rate peak at midheight Λexp as a function of time. (e) The measured position of the reaction front XF, exp as a function of time for various template concentrations (error bars = s.d.; n = 3; R2 = 0.998). (f) The intercept (t0, exp) of the line in Fig. 5e and the threshold time Ct of real time, benchtop RT-LAMP curves as functions of the number of templates (error bars = s.d.; n = 3; R2 = 0.99). (g) XF, exp-XF,exp(3) as a function of the template number at various times t (error bars = s.d.; R2 = 0.99, n = 15). (h) The predicted position of the reaction front (XF, th) as a function of time for various numbers of templates. (i) The predicted intercept (t0, th) of the asymptotes in Fig. 5h as a function of template number.
as a function of position (x) at various times. (d) The measured width of the reaction-rate peak at midheight Λexp as a function of time. (e) The measured position of the reaction front XF, exp as a function of time for various template concentrations (error bars = s.d.; n = 3; R2 = 0.998). (f) The intercept (t0, exp) of the line in Fig. 5e and the threshold time Ct of real time, benchtop RT-LAMP curves as functions of the number of templates (error bars = s.d.; n = 3; R2 = 0.99). (g) XF, exp-XF,exp(3) as a function of the template number at various times t (error bars = s.d.; R2 = 0.99, n = 15). (h) The predicted position of the reaction front (XF, th) as a function of time for various numbers of templates. (i) The predicted intercept (t0, th) of the asymptotes in Fig. 5h as a function of template number.
Fig. 5b depicts  as a function of time at various positions x. An observer located at a position x will not see a signal until after a certain time delay. The greater the magnitude of x, the larger the delay is. Fig. 5c depicts the experimentally-determined rate of the reaction
as a function of time at various positions x. An observer located at a position x will not see a signal until after a certain time delay. The greater the magnitude of x, the larger the delay is. Fig. 5c depicts the experimentally-determined rate of the reaction  as a function of position (x) at various times. The rate of the reaction resembles a propagating peak that travels at a fixed velocity v0. The peak's width at midheight (Λ) did not vary with time (Fig. 5d), i.e., the reaction front is non-dispersive. In other words, the precision with which we can determine the reaction front's position does not deteriorate with time.
as a function of position (x) at various times. The rate of the reaction resembles a propagating peak that travels at a fixed velocity v0. The peak's width at midheight (Λ) did not vary with time (Fig. 5d), i.e., the reaction front is non-dispersive. In other words, the precision with which we can determine the reaction front's position does not deteriorate with time.
Next, to analyze the propagation speed of the reaction front, we depict the position of the reaction front XF(t) as a function of time when the number of target molecules is 102, 103 and 104 (Fig. 5e, n = 3). For sufficiently large times, t > t1 > t0, the experimental data correlates well with straight lines (R2 = 0.998).

where t (s) is the observation time, t0 (s) is the intercept with the horizontal axis and t1 (s) is the delay time until a visible signal is observed anywhere in the conduit. All the lines in Fig. 5e have nearly the same slope, indicating that the front propagates at a nearly constant speed of  μm/s (n = 9) independent of target concentration. In contrast, t0 decreases as the target concentration increases (Fig. 5f), playing a similar role to the threshold time (Ct) in a standard real-time, quantitative amplification. For comparison, Ct is also shown in Fig. 5f (and in Supplementary Fig. 5). We find
μm/s (n = 9) independent of target concentration. In contrast, t0 decreases as the target concentration increases (Fig. 5f), playing a similar role to the threshold time (Ct) in a standard real-time, quantitative amplification. For comparison, Ct is also shown in Fig. 5f (and in Supplementary Fig. 5). We find

where A and B are constants and c0 is the number of target molecules in the sample (R2 = 0.996).
Although the position of the front XF is time-dependent, the distances between the positions of any two fronts associated with different numbers of target molecules are not (Supplementary Fig. 6). Thus, time-dependence can be eliminated by subtracting the position of the reaction front of a calibration lane (XF(c)) containing a known target concentration from that of the test lane. To demonstrate this, we denote variables associated with conduits 1, 2 and 3 (Fig. 3) with superscripts 1, 2 and 3. Fig. 5g depicts XF(i)-XF(3) as a function of the number of target molecules c0(i). Witness that all the data collapses to a single straight line (n = 15, R2 = 0.99), eliminating any explicit dependence on the time at which XF was measured.

In the above, the nuclemeter 3 serves as the calibration nuclemeter and we replaced superscript (3) with c. In other words, in the presence of one or more calibration nuclemeters, one can rely on the differences among reaction front positions to determine target concentration, independent of measurement time. Although not essential, it is expected that a practical device would include at least one calibration nuclemeter. The calibration nuclemeters can, of course, double up as positive controls.
With the aid of equation (2), we can rewrite equation (3) to express explicitly the dependence of ΔXF on target analyte concentration.

Theoretical model
To gain further insights into the operation of the nuclemeter, we propose a simple reaction-diffusion mathematical model to simulate our experiment. We approximate the amplicon production during enzymatic amplification with the production rate  , where the reaction rate constant k ~ 0.008 s−1 was determined empirically by fitting theoretical predictions based on the above production rate with real time RT-LAMP amplification curves (Supplementary Note 2). We estimated cmax ~ 1.1 × 10−10 mol/m3. We model the reaction diffusion process in the nuclemeter with the dimensionless equation22
, where the reaction rate constant k ~ 0.008 s−1 was determined empirically by fitting theoretical predictions based on the above production rate with real time RT-LAMP amplification curves (Supplementary Note 2). We estimated cmax ~ 1.1 × 10−10 mol/m3. We model the reaction diffusion process in the nuclemeter with the dimensionless equation22

In the above, we scaled distance with  and time with k−1. The diffusion coefficient D ~ 10−10 m2/s was estimated by monitoring the diffusion of labeled primers in the conduit in the absence of amplification reaction (Supplementary Note 3). The boundary and interfacial conditions are:
and time with k−1. The diffusion coefficient D ~ 10−10 m2/s was estimated by monitoring the diffusion of labeled primers in the conduit in the absence of amplification reaction (Supplementary Note 3). The boundary and interfacial conditions are:  ;
;  (at the interface between the well and the conduit); and
(at the interface between the well and the conduit); and  . The initial conditions are
. The initial conditions are  when
when  (sample chamber) and
(sample chamber) and  when
when  (reaction-diffusion conduit).
(reaction-diffusion conduit).
The predictions of equation (5) (solid lines in Figs. 5a and b) closely resemble the experimental data. Fig. 5h depicts the position of the predicted reaction front as a function of time for different initial concentrations  . Although at short times, the front velocity varies as a function of
. Although at short times, the front velocity varies as a function of  , soon enough all the curves asymptote to straight lines with a slope independent of time and the initial target concentration. The dimensionless predicted reaction front velocity is 2. The dimensional predicted reaction front velocity
, soon enough all the curves asymptote to straight lines with a slope independent of time and the initial target concentration. The dimensionless predicted reaction front velocity is 2. The dimensional predicted reaction front velocity  is very close to the experimentally measured one. Moreover, consistent with experiments, the theory predicts a constant reaction front velocity independent of target concentration. When
is very close to the experimentally measured one. Moreover, consistent with experiments, the theory predicts a constant reaction front velocity independent of target concentration. When  , the front location can be estimated with equation (1), where
, the front location can be estimated with equation (1), where  depends on the initial concentration through equation (2). Fig. 5i depicts the predicted t0 as a function of the number of target molecules using an estimated value of cmax. Fig. 5i is in qualitative agreement with the experimental data (blue) of Fig. 5f.
depends on the initial concentration through equation (2). Fig. 5i depicts the predicted t0 as a function of the number of target molecules using an estimated value of cmax. Fig. 5i is in qualitative agreement with the experimental data (blue) of Fig. 5f.
Discussion
We have described a new paradigm for simple, endpoint quantification of target nucleic acids undergoing enzymatic amplification. Our method is based on inferring the number of target nucleic acid molecules from the position of the amplification reaction front. The position of the front can be read at a prescribed time or preferably, in the presence of a calibration column, at any time. Since the reaction front is non-dispersive, the quality of the data is insensitive to the time when it is read. In contrast to traditional quantitative enzymatic amplification methods that require continuous monitoring of fluorescent emission intensity as a function of time, the nuclemeter requires one to observe the signal only at a single instant in time. Here we carried our experiments using the RT-LAMP process, but the nuclemeter can operate with any other amplification scheme. We demonstrated the utility of our method to quantify HIV RNA, achieving performance on par with benchtop equipment.
Although in our experiments we used a custom made, portable processor to control the amplification reaction temperature and to monitor fluorescent emission, the nuclemeter can operate without any instrumentation. The heating to maintain the amplification temperature can be provided by an exothermic reaction and the temperature controlled with a phase change material, as we have previously described18, eliminating the need for electrical power, which may not be reliably available in resource poor settings and in the field and a potentially costly thermal control. The excitation for the fluorescence can be provided with light emitting diodes (LED) and the position of the reaction front can be read by eye without any optical reader. Alternatively, one can use a filtered flash light of a cell phone and a cell phone camera19,23 to record, analyze and transmit the test results. Thus, the nuclemeter enables one to quantify the number of target molecules, nearly as simply as one would infer the temperature from the length of a mercury column in a “mercury in glass” thermometer.
We described here the basic operating principle of the nuclemeter module. The nuclemeter can be readily combined with a module for nucleic acid isolation, concentration and purification25 and with a self-heating module18 to facilitate inexpensive non-instrumented, quantitative molecular detection technology for low resource settings, on-site and home applications. Many other extensions of the nuclemeter concept are possible. Numerous nuclemeters, containing different sets of primers, can be housed on a single substrate to concurrently detect and quantify different targets. Alternatively, mixtures or primers can be placed in a single nuclemeter to amplify multiple targets. The intercalating dye that we used in our work can be replaced with molecular beacons, each type of beacon specific to a different target and emitting in a distinct region of the spectrum, enabling concurrent monitoring of multiple targets propagating in a single conduit. Multiple reaction-diffusion conduits with different functionalizations can be connected to a single sample chamber. Additionally, one can apply a linear temperature gradient along the reaction-diffusion conduits to determine the melting temperature24. And these are just a few examples of numerous possibilities.
Methods
Nuclemeter chip fabrication
The 46 mm long × 36 mm wide × 3.0 mm thick, Poly(methyl methacrylate) (PMMA, Acrylic glass) body of the chip was milled with a precision, computer-controlled (CNC, HAAS Automation Inc., Oxnard, CA) milling machine (Fig. 1c). An inlet port and an exit port were connected to the sample chamber and a third port was connected to the distal end of the microconduit (Fig. 1a). After milling, the chip body was sonicated in 100% ethanol for 15 minutes, rinsed with water and air-dried at room temperature. Then, to eliminate any RNase and DNase that could degrade nucleic acids or interfere with enzymatic reactions, the chip body was dipped in Decon™ ELIMINase™ decontaminant (Thermo Fisher Scientific Inc., Walthman, MA) for 2 minutes, rinsed twice with sterile molecular biology-grade water (Thermo Fisher Scientific Inc.) and air-dried at room temperature.
The chip body was, respectively, ceiled and floored with PMMA film (top) and PCR Sealers™ tape (bottom) (Supplementary Fig. 2). Both were cut with a CO2 laser (Universal Laser Systems). The top PMMA film was solvent-bonded to the chip body with acetonitrile (Sigma-Aldrich) at room temperature. The bonded chip was heated overnight (Isotemp Vacuum Oven Model 280A, Fisher Scientific Inc., Pittsburgh, PA) at 55°C to remove any residual solvent. Finally, the PCR Sealers™ tape was used to seal the bottom of the chip.
HIV RNA purification from plasma samples
Viral RNA was extracted from HIV-1 standards (AcroMetrix® HIV-1 High Control, Benicia, CA) with QIAamp Viral RNA Mini Kit (Qiagen, Valencia, CA) according to the manufacturer's protocol. Briefly, 140 μL of virus suspension was lysed with 560 μL virus lysis buffer containing carrier RNA. A 560 μL ethanol was added to the lysate and the mixture was centrifuged in a spin column (630 μL aliquots) at 10,000 rpm for 2 minutes. Prior to eluting the HIV viral RNA, wash buffers were loaded into the spin column and centrifuged at 14,000 rpm for 5 minutes. The RNA was eluted with 60 μL of elution buffer. Negative controls were prepared from a de-identified HIV-negative plasma sample (provided by the Penn Center for AIDS Research CFAR with the Institutional Review Board approval (protocol: 814752)) using the same extraction procedures as described above.
RT-LAMP reagents
The RT-LAMP primers were designed by Curtis et al.26 at the Center for Disease Control and Prevention (CDC) and were synthesized by Sigma-Aldrich. The real time benchtop RT-LAMP experiments were carried out with 15 μL reaction volumes. The reaction mixture consisted of 0.2 μM of F3 and B3, each; 0.8 μM Loop F and Loop B, each; and 1.6 μM of FIP and BIP, each, 1.25 U AMV reverse transcriptase (Life Technologies, Carlsbad, CA); 0.53 × EvaGreen dye (Biotium, Hayward, CA); 0.04% (w/v) hydroxypropyl-methyl-cellulose (HPMC); and 9 μL Isothermal Master Mix (ISO-001nd, OptiGene, Horsham, UK).
The HPMC was dissolved in Isothermal Master Mix, centrifuged at 10,000 rpm for 2 minutes and filtered through a Corning Costar® Spin-X® centrifuge tube equipped with cellulose acetate membrane filters with a pore size of 0.45 μm to remove any traces of insoluble HPMC.
A ten-fold dilution series of HIV viral RNA extracted from a HIV-1 standard panel and a negative control without template prepared from a HIV-negative plasma sample were tested in parallel. The real time, “tubed-based” RT-LAMP was carried out in a Peltier Thermal Cycler PTC-200 (Bio-Rad DNA Engine, Hercules, CA). Reactions were carried out at 62.5°C for 60 minutes with real-time fluorescence monitoring. Real-time RT-LAMP results were analyzed and the threshold time Ct (the time needed for the emission intensity to exceed a predetermined value) was obtained.
Device operation
5 μL of RT-LAMP master mixture, comprised of all the reagents necessary for the RT-LAMP and 0.04% HPMC (excluding the HIV RNA template), was inserted into each reaction-diffusion microconduit through inlet port 1 (Fig. 1a). Then, inlet ports 1 of all four nuclemeters were sealed with PCR Sealers™ tape. Next, 15 μl of RT-LAMP master mixture and HIV RNA template of various concentrations were injected into the sample chambers through the inlet ports 2 (Fig. 1a). Subsequently, both the inlet ports 2 and outlet ports were sealed with PCR Sealers™ tape to minimize evaporation during the amplification process. The nuclemeter chip was placed on a custom, portable heater and incubated at 62.5°C for about 60 minutes to enable isothermal amplification.
Portable processor for RT-LAMP
The custom made, portable processor (Fig. 2 and Supplementary Fig. 3) for the nuclemeter consisted of a chip holder equipped with a flexible, polyimide-based, thin film heater (Model HK5572R7.5L23A, Minco Products, Inc., Minneapolis, MN) (inset in Fig. 2a), an electronic circuit board and a thermocouple positioned at the interface between the thin film heater and the nuclemeter chip. When the nuclemeter chip, filled with LAMP master mixture, was inserted into the processor, the reaction chambers and diffusion conduits were in thermal contact with the thin film heater.
To calibrate the device, we constructed a calibration chip with a type-K thermocouple (Omega Engr., each wire 75 mm in diameter and a junction diameter of 170 μm) in the reaction-diffusion conduit. The sample chambers and reaction-diffusion conduits were filled with water. The thermocouple reading was monitored with a HH506RA multilogger thermometer (Omega Engr., Stamford, CT, USA). In addition, an infrared image of the microfluidic chip heated by our processor was taken with an infrared thermography camera T360 (FLIR Systems, Wilsonville, USA) to evaluate temperature uniformity (Fig. 2b).
Endpoint, fluorescence image for quantitative detection
The fluorescence excitation and emission imaging were carried out with a handheld, USB-based, fluorescence microscope (AM4113T-GFBW Dino-Lite Premier, AnMo Electronics, Taipei, Taiwan) (Fig. 2 and Supplementary Fig. 3). The USB-based, fluorescence microscope has built-in, filtered blue LEDs for excitation, a 510 nm emission filter and a CCD camera for fluorescence imaging. The microscope was interfaced with a computer through a USB interface. Images were acquired with a DinoCapture 2.0 software program. The images were processed with MatLab to remove background noise and uneven illumination effects. A normalized and averaged fluorescence intensity signal for each lane was extracted from each processed image (Supplementary Note 4). The locations of the reaction fronts of different samples were directly read out by eye with the fluorescence ruler (Fig. 2c).
References
Higuchi, R., Fockler, C., Dollinger, G. & Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Nat. Biotechnol. 11, 1026–1030 (1993).
Heid, C. A., Stevens, J., Livak, K. J. & Williams, P. M. Real time quantitative PCR. Genome Res. 6, 986–994 (1996).
Toumazou, C. et al. Simultaneous dna amplification and detection using a ph-sensing semiconductor system. Nat Methods. 10, 641–646 (2013).
Rouet, F. et al. Transfer and evaluation of an automated, low-cost real-time reverse transcription-PCR test for diagnosis and monitoring of human immunodeficiency virus type 1 infection in a West African resource-limited setting. J. Clin. Microbiol. 43, 2709–2717 (2005).
Granich, R. M., Gilks, C. F., Dye, C., De Cock, K. M. & Williams, B. G. Universal voluntary HIV testing with immediate antiretroviral therapy as a strategy for elimination of HIV transmission: a mathematical model. Lancet 373, 48–57 (2009).
Christensen, D. R. et al. Detection of biological threat agents by real-time PCR: comparison of assay performance on the R.A.P.I.D., the LightCycler and the Smart Cycler platforms. Clin Chem. 52, 141–145 (2006).
Bell, A. S. & Ranford-Cartwright, L. C. Real-time quantitative PCR in parasitology. Trends Parasitol 18, 337–342 (2002).
Jaroenram, W., Kiatpathomchai, W. & Flegel, T. W. Rapid and sensitive detection of white spot syndrome virus by loop-mediated isothermal amplification combined with a lateral flow dipstick. Mol Cell Probes. 23, 65–70 (2009).
Rohrman, B. A., Leautaud, V., Molyneux, E. & Richards-Kortum, R. R. A Lateral Flow Assay for Quantitative Detection of Amplified HIV-1 RNA. PLoS ONE 7, e45611 (2012).
Roskos, K. et al. Simple system for isothermal DNA amplification coupled to lateral flow detection. PLoS One. 8, e69355 (2013).
Yager, P. et al. Microfluidic diagnostic technologies for global public health. Nature 442, 412–418 (2006).
Hart, R. W. et al. Point-of-care oral-based diagnostics. Oral Dis. 17, 745–52 (2011).
Song, Y. et al. Point-of-care technologies for molecular diagnostics using a drop of blood. Trends Biotechnol. 32, 132–139 (2014).
Niemz, A., Ferguson, T. M. & Boyle, D. S. Point-of-care nucleic acid testing for infectious diseases. Trends Biotechnol. 29, 240–250 (2011).
Zhang, C. & Xing, D. Miniaturized PCR chips for nucleic acid amplification and analysis: latest advances and future trends. Nucleic Acids Res. 35, 4223–4237 (2007).
Lee, J. G. et al. Microchip-based one step DNA extraction and real-time PCR in one chamber for rapid pathogen identification. Lab Chip 6, 886–895 (2006).
Dimov, I. K. et al. Integrated microfluidic tmRNA purification and real-time NASBA device for molecular diagnostics. Lab Chip 8, 2071–2078 (2008).
Liu, C., Mauk, M. G., Hart, R., Qiu, X. & Bau, H. H. A self-heating cartridge for molecular diagnostics. Lab Chip 11, 2686–2692 (2011).
Selck, D. A., Karymov, M. A., Sun, B. & Ismagilov, R. F. Increased robustness of single-molecule counting with microfluidics, digital isothermal amplification and a mobile phone versus real-time kinetic measurements. Analytical Chemistry 85, 11129–11136 (2013).
Notomi, T. et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, E63 (2000).
Tomita, N., Mori, Y., Kanda, H. & Notomi, T. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat. Protoc. 3, 877–882 (2008).
Fisher, R. A. The wave of advance of advantageous genes. Ann. Eugenics 7, 353–369 (1937).
Liu, C. et al. A low-cost microfluidic chip for rapid genotyping of malaria-transmitting mosquitoes. PLoS ONE 7, e42222 (2012).
Mao, H., Holden, M. A., You, M. & Cremer, P. S. Reusable platforms for high-throughput on-chip temperature gradient assays. Anal Chem. 74, 5071–5075 (2002).
Liu, C. et al. An isothermal amplification reactor with an integrated isolation membrane for point-of-care detection of infectious diseases. Analyst 136, 2069–2076 (2011).
Curtis, K. A., Rudolph, D. L. & Owen, S. M. Rapid detection of HIV-1 by reverse-transcription, loop-mediated isothermal amplification (RT-LAMP). J. Virol. Meth. 151, 264–270 (2008).
Acknowledgements
C.L. was supported by NIH/NIAID K25AI099160; MMS was supported by the Nanotechnology Institute of the Pennsylvania Department of Community and Economic Development, Ben Franklin Technology Development Authority; P.H.E., F.D.B., R.G. and H.H.B. were funded, in part, by NIH/NIAID 1R41AI104418-01A1. C.L., R.G. and H.H.B. were also funded, in part, by Penn Center for AIDS Research (CFAR) Pilot Grant (No. AI045008). Curtis (Centers for Disease Control and Prevention, CDC) provided Hex-labeled oligonucleotides. W. Zhao, a UPenn undergraduate student, machined the chips and constructed the portable processor.
Author information
Authors and Affiliations
Contributions
C.L. and H.H.B. conceived the study. C.L. performed experiments. M.M.S. and H.H.B. carried out the theoretical calculations. C.L., H.H.B., M.M.S., M.G.M., P.H.E., F.D.B. and R.G. discussed the results and analyzed the data. C.L., M.G.M. and H.H.B. wrote the manuscript. M.M.S., P.H.E., F.D.B. and R.G. edited the manuscript. All authors approved the final version.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Video 1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Liu, C., Sadik, M., Mauk, M. et al. Nuclemeter: A Reaction-Diffusion Based Method for Quantifying Nucleic Acids Undergoing Enzymatic Amplification. Sci Rep 4, 7335 (2014). https://doi.org/10.1038/srep07335
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07335
This article is cited by
-
CFD simulations for paper-based DNA amplification reaction (LAMP) of Mycobacterium tuberculosis—point-of-care diagnostic perspective
Medical & Biological Engineering & Computing (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.