Abstract
High-quality protein crystals of suitable size are an important prerequisite for applying X-ray crystallography to determine the 3-dimensional structure of proteins. However, it is often difficult to obtain protein crystals of appropriate size and quality because nucleation and growth processes can be unsuccessful. Here, we show that by adsorbing proteins onto porous polystyrene-divinylbenzene microspheres (SDB) floating on the surface of the crystallisation solution, a localised high supersaturation region at the surface of the microspheres and a low supersaturation region below the microspheres can coexist in a single solution. The crystals will easily nucleate in the region of high supersaturation, but when they grow to a certain size, they will sediment to the region of low supersaturation and continue to grow. In this way, the probability of crystallisation and crystal quality can be simultaneously increased in a single solution without changing other crystallisation parameters.
Similar content being viewed by others
Introduction
To date, X-ray crystallography remains the most widely used technique for determining the 3-dimensional structure of protein molecules1,2,3,4. The vast majority (89%) of protein structures deposited in the Protein Data Bank (PDB) have been determined using X-ray crystallography (data representing the situation on May 6, 2014, from http://www.rcsb.org/pdb/home/home.do), in which X-ray-suitable protein crystals were used as samples. Obtaining crystals suitable for X-ray diffraction is a major bottleneck in structure determination using X-ray crystallography5,6,7, which requires that not only that the proteins are crystallised successfully but also that the produced crystals are of the desired quality and internal order6,8,9. However, these goals are not easy to achieve simultaneously because the supersaturation requirements are different: to successfully obtain crystals, the protein must first form a nucleus (nucleation) in a solution of relatively high supersaturation10,11; however, to ensure that the crystal quality (i.e., the internal order of the molecule) is sufficiently high to achieve high-resolution diffraction, the crystals should be grown stably and smoothly in a solution of relatively low supersaturation12. Because the constraints regarding supersaturation are different, it is often difficult to identify crystallisation conditions that are suitable for obtaining high-quality protein crystals directly. However, ongoing investigations might help to solve this problem. For example, the nucleation and growth processes are separated in various methods13,14. A typical approach to separate the two stages is to vary the temperature15 during crystallisation: The solution can first be held at one temperature yielding high supersaturation for ease of nucleation; then, after nucleation has occurred, the temperature can changed so as to decrease the degree of supersaturation, leading to the growth of good-quality crystals. Nevertheless, this method cannot be applied to proteins for which the solubility is not affected by temperature changes; moreover, only in situ lightscattering16,17 can provide information about nucleation in crystallisation droplets, requiring the use of appropriate DLS hardware. In addition to this method, other approaches, such as seeding18,19,20 or heterogeneous nucleation12,21,22,23,24,25,26,27,28,29,30,31,32,33,34, can also be useful for identifying conditions that are suitable for the growth of high-quality protein crystals.
Seeding methods18,19,20 utilise pre-crystallised proteins or protein complexes as nucleants. Because the nucleus (i.e., the seed) already exists, the crystals can grow in the solution under low supersaturation conditions under which the nucleation process would be difficult, affording new conditions in which to grow high-quality protein crystals. Although seeding methods are in principle successful, they depend strongly on both the availability of seed crystals and simplified seeding manoeuvres. Furthermore, the need for seed crystals increases the cost and effort of crystallisation.
As an alternative, heterogeneous nucleation12,21,22,23,24,25,26,27,28,29,30,31,32,33,34 has been proposed. In particular, Chayen et al.12 have reported the best results regarding heterogeneous nucleation to date using two chemically different materials that possessed broad pore-size distributions12. Heterogeneous nucleation can improve the quality of protein crystals because protein crystals can nucleate on the surface of selected nucleants in a less supersaturated solution; thus, the crystals can grow in a solution of low supersaturation directly. The nucleants, which can lower the energy barrier for nucleation, function similarly to seed crystals. However, some problems remain to be considered. For example, attachment of the crystals to the nucleants may cause difficulties in observation and crystal harvesting or cause crystal quality degradation due to lattice mismatch between the crystals and the nucleants.
To further improve the heterogeneous nucleation method, we propose here to use a different type of heterogeneous nucleants (porous or non-porous polystyrene divinylbenzene microspheres, SDB) that can float on the upper part of the crystallisation solution due to its low density compared with that of the crystallisation solution. The nucleants can adsorb proteins when the solution concentration is high and desorb the proteins when the concentration of the surrounding solution is low. Using such adsorption and desorption properties of the material, it is possible to create a distinct and localised region of high concentration and another region of low concentration in the same solution, thereby increasing the probability of obtaining crystals and improving crystal quality within a single experiment. The attachment of crystals to the surface of the microspheres can be avoided because the crystals will sediment towards the bottom of the solution as they grow.
Results
The effect of SDB microspheres on crystallisation hits within a screening study
We analysed the effect of experiments with and without adding SDB microspheres (see Supplementary Fig. 1 and Supplementary Note 1 (1) for the method used to prepare the SDB microspheres) to crystallisation droplets (see Supplementary Fig. 2 and Supplementary Note 1 (2) for the crystallisation method). Here, positive screening results were measured and scored using the parameter “crystallisation hits” which is defined as the number of droplets that yielded detectable crystals under an 80× stereomicroscope (Nikon SMZ 1000). The conditions used in 96 crystallisation droplets for each target protein per plate were set up using IndexTM screening kits (Hampton Research, USA) according to the standard screening protocol for the sitting drop vapour diffusion method35. The crystallisation behaviour of 11 proteins was analysed for all three experiments: the control, SDB1 and SDB2 cases. All results are summarised in Table 1. The results showed that in most cases, the presence of SDB microspheres increased the number of hits remarkably. In particular, four proteins (α-chymotrypsinogen A I, ribonuclease A I, ribonuclease A III and subtilisin A VII) did not crystallise in the control experiment but showed a (non-trivial) number of hits when using SDB microspheres.
Another interesting result showed that SDB1 outperformed SDB2 in terms of the number of hits. For statistical comparison, the numbers of hits in the experiments were normalised according to the number of hits in the SDB2 case (Fig. 1). Most interestingly, the screenings in the SDB1 and the SDB2 cases achieved substantially higher number of hits (176.35% and 56.64% compared to the control experiment, respectively). The SDB1 experiments most likely provided superior results due to differences in the microsphere surfaces (e.g., in the microporous structure of the surface) (Supplementary Figs. 3 & 4).
Normalised crystallisation hits in the presence and absence of SDB microspheres.
The addition of SDB1 and SDB2 remarkably increased the number of crystallisation hits (error bars: standard error of the mean, n = 11). The number of screening hits was normalised based on the number of hits obtained using SDB2. Both SDB1 and SDB2 microspheres increased the number of hits and the effect of SDB1 was most obvious.
Improvement of crystal quality by using SDB microspheres
In this section, the potential improvement of crystal quality is validated in terms of crystal's morphology and the quality of X-ray diffraction.
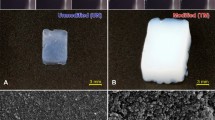
Carefully examining the crystal images, we found the differences in crystal morphology in the presence and absence of SDB microspheres. Fig. 2 shows some examples. Fig. 2a shows the morphology of typical lysozyme crystals. No difference in morphology was evident during the first few days. However, when the incubation time was extended to more than 2 months, defects (wrinkles or micro-cracks) appeared on the surface of the crystals that were grown in the absence of SDB microspheres. In contrast, the crystals grown in the presence of SDB microspheres retained the same appearance as before.
Crystal morphologies grown in the presence and absence of SDB microspheres.
(a) Lysozyme crystals grown (a1) in the presence of SDB1; (b1) in the presence of SDB2 and (c1) in the absence of SDB microspheres after 2 months. Crystallisation conditions: lysozyme: 40 mg/ml, NaCl: 40 mg/ml, buffer: 0.2 M sodium acetate buffer, pH 4.6; temperature: 20°C. (b) Catalase, concanavalin A and thaumatin crystals grown (a1-a3) in the presence of SDB1; (b1-b3) in the presence of SDB2 and (c1-c3) in the absence of SDB microspheres after 4 days at 20°C. Crystallisation conditions: (a1-c1) Catalase: 20 mg/ml, 0.2 M trimethylamine N-oxide dehydrate, 0.1 M Tris, 20% w/v polyethylene glycol monomethyl ether 2,000, pH 8.5; (a2-c2) Concanavalin A: 20 mg/ml, 0.2 M L-proline, 0.1 M HEPES, 10% w/v polyethylene glycol 3350, pH 7.5; (a3-c3) Thaumatin: 20 mg/ml, 40% tacsimate, pH 7.0. The crystals were grown at Northwestern Polytechnical University.
Fig. 2b shows examples of the crystal morphology of several other commercial proteins grown in the presence and absence of SDB microspheres during the screening experiments. In the presence of SDB microspheres (especially in the SDB1 case) fewer but larger crystals were seen with better morphology and more well-defined, faceted crystallographic surfaces. This phenomenon was frequently observed during the crystallisation experiments (for further examples from three independent laboratories, see Supplementary Figs. 5 and 6).
The morphological improvement obtained indicated that crystal quality is influenced by the presence of SDB microspheres. However, whether the intrinsic quality of crystals is improved is determined using X-ray diffraction experiments. Thus, we assessed the crystal quality of 8 proteins (concanavalin A, thaumatin, glucose isomerase, proteinase K, HSP90, TCS, ThiM36 and ML-I37) by applying X-ray diffraction analysis (the crystals were grown in three independent laboratories at Northwestern Polytechnical University, the University of Hamburg and the Shanghai Institute of Applied Physics; the quality was assessed using Beamline P14 at EMBL/Petra III/DESY/Hamburg and Beamline 17U1 at the Shanghai Synchrotron Radiation Facility, SSRF. For each group of assessments in the presence and absence of SDB, we attempted to select the crystals with the best appearance of most similar sizes. The assessment was carried out considering the diffraction resolution, Rmerge, I/σI, mosaicity and electron density map. Table 2 summarises the results of the diffraction experiments. In total we tested 8 proteins. For 5 of the 8 proteins, i.e., Concanavalin A (1.54 Å with SDB1 vs 2.10 Å without SDB1), Thaumatin (1.20 Å vs 1.49 Å), ThiM (1.97 Å vs No diffraction data), TCS (1.01 Å vs No diffraction data) and HSP90 (1.43 Å vs 2.65 Å), it can be simply seen from diffraction resolution that the SDB grown crystals showed better quality. For the remaining proteins, it is difficult to tell the quality difference directly by judging from the resolution because the diffraction resolutions with and without SDB were similar. However, we can still find the difference by carefully analyzing the other data like mosaicity, Rmerge and further, the electron density map. Thus, in the case of MLI, judging from the diffraction data we can tell that the crystal quality of SDB-grown crystals may be slightly poorer than the control because the mosaicity of the SDB-grown crystals was larger than the control. In the cases of Glucose Isomerase and Proteinase K, however, it is hard to judge, so we tried to solve the structures of these two proteins with and without SDB1 (results shown in Supplementary Fig.s 7–8). The results showed that, there are no visible differences in the electron density maps. However, the R and Rfree were marginally better in the presence of SDB1 for both proteins (for Glucose Isomerase: 19.54 (R +SDB) vs 19.39 (R -SDB) and 23.39 (Rfree +SDB) vs 22.94 (Rfree -SDB); for Proteinase K: (15.25(R +SDB) vs 16.10 (R -SDB) and 16.89 (Rfree +SDB) vs 17.02 (Rfree -SDB)), which means that, the crystal quality was slightly better in the presence of SDB1. Hence, we can say that, 5 showed improvement, 2 possibly showed a marginal improvement in crystal quality when using the SDB1 (the porous one) and only 1 protein showed slightly poorer crystal quality in the presence of the SDB1. Therefore we believe that, the crystal quality can be influenced and improved by using the SDB1 microspheres in many cases.
In summary, in many crystallisation experiments, both morphology and intrinsic quality were improved when the crystals were grown in the presence of SDB microspheres.
Concentration evolution in crystallisation solutions in the presence and absence of SDB1 microspheres
SDB microspheres are known to adsorb particular protein molecules. If these microspheres are added to a protein crystallisation solution, they will most likely cause the redistribution of protein concentrations in the solution, consequently influencing protein crystallisation.
To examine how the microspheres affect concentration in the crystallisation solution, we measured protein concentration using the bicinchoninic acid (BCA) assay. The results showed that the presence of SDB microspheres significantly reduce protein concentrations underneath the microspheres (Supplementary Fig. 9) during the crystallisation process. The BCA provided information regarding changes in concentration over a long period; however, due to the invasive nature of this assay, the concentration was measured at long intervals (to minimise the effect of measurement on crystallisation). To obtain more detailed information and further confirm the observed changes in concentration, we designed a model system to monitor how SDB1 microspheres affect the concentration distribution in a sealed crystallisation cell. In this system, a digital holographic Mach-Zehnder interferometer was built and a program was written to calculate the concentration of the solution. A model solution (lysozyme crystallisation solution) was used to measure the change in concentration in the presence and absence of SDB1 microspheres. A quartz cell was filled with the solution and the concentration distribution in the quartz cell was monitored every 5 minutes using the interferometer (for further details, see Supplementary Figs. 10–12 and Supplementary Note 1(3)).
Fig. 3a shows a time course series of reconstructed phase maps derived from the holograms in the presence (Fig. 3a, i-vi) and absence (Fig. 3b, i-vi) of SDB1 microspheres. From the phase maps, it can be clearly seen that in the presence of SDB1 microspheres, interference fringes appeared immediately under the microspheres and gradually expanded to the lower part of the cell; this observation indicated that concentration redistribution occurred in the solution after adding the microspheres. The presence of the fringes indicated that a concentration deficient region was formed immediately underneath the microspheres, indicating in turn that the microspheres did adsorb protein; expansion of the fringes to the lower part of the cell indicated protein molecules were consumed from above (both adsorption and crystallisation can consume protein molecules). The fringes then became stable, showing that the concentration gradient was maintained for a long period. At the end of the experiment, the distance between the fringes in the lower part of the cell became larger and gradually disappeared, indicating that the concentration in that region became homogeneous again. This phenomenon indicates that mass transport and/or the consumption of protein molecules from the bottom occurred. The fringe images obtained indicated that the concentration distribution was more stable and homogeneous throughout the process in the absence of microspheres (Fig. 3b, i-vi) than in the presence of SDB1 microspheres.
Concentration evolution derived from the holographic Mach-Zehnder interferometer.
(a) and (b): Time course of the reconstructed phase maps of a lysozyme crystallisation solution in the presence and absence of SDB1 microspheres as derived from the holograms. (i): Three hrs after SDB1 microspheres were added. (i)- (vi): Subsequent phase maps at time intervals of 6 hrs. Crystallisation conditions: lysozyme: 30 mg/ml, NaCl: 40 mg/ml, pH 4.60, 20°C. Solution volume: 250 μl, mass of SDB1 microspheres: 20 mg. The phase maps show clearly that adding SDB1 microspheres redistributed the concentrations in the solution. (c): Concentration evolution in the solution at various locations in the presence and absence of SDB1 microspheres. (i)- (iii): Concentration evolution at points a, b and c, respectively; (d): The rate of concentration decrease at points a, b and c, respectively. (i) and (ii): In the presence and absence of SDB1 microspheres, respectively; (e) locations of points a, b and c in the crystallisation cell. The concentration measurements verified that adsorption indeed occurred and that concentration changes during crystal growth (the second half of the experiment) were more stable in the presence of SDB1 microspheres than in the absence of SDB.
The evolution of the fringes described above provides strong evidence for the existence of an adsorption process resulting from the presence of SDB1 microspheres. To further verify this observation, we quantified the change of concentration with time. Fig. 3c shows an example of the changes in concentration in the presence and absence of SDB1 microspheres for 2,000 mins at three positions (points a, b and c in Fig. 3e) in the cell. Fig. 3d shows the derived rate of concentration decrease at the three points in the presence and absence of SDB1 microspheres.
From the results of concentration evolution presented in Fig. 3c, we can find that, in the beginning, the concentration decreased faster in the presence of SDB1 than the control at points a and b, while at point c (which is nearer to the bottom), the concentration evolution was similar to that of the control. This phenomenon strongly indicated that, upon adding SDB1 microspheres, the adsorption of proteins immediately occurred so that the protein concentration decreased faster than that without adding the microspheres. And, simultaneously, crystallisation happened soon after starting the experiments according to the immediate decrease in the concentration at all points in both cases with and without SDB1 microspheres. Apart from the above information which confirmed the observations from the fringe images, we can get more from the concentration measurement results that was not provided by the fringe images: (i) Linear concentration gradient in the cell: from the concentration data we found that a linear concentration gradient existed in the cell for the case with SDB1. In contrast, the concentration was rather homogeneous in the cell without SDB. The phenomenon indicated that adsorption of protein molecules by the microspheres occurred so that a concentration deficient region near the microspheres appeared, resulting in a concentration gradient from the upper (low concentration) to the lower part (high concentration). And the persistence of the gradient over a long time indicated that nucleation and growth of protein crystals at the surface of the microspheres occurred, so that the consumption of protein molecules could continue as shown from the concentration evolution process. (ii) Turning point at 1380 min after starting the experiments: at time 1380 min, a meeting point in concentration decreasing rate for points a, b and c was observed (Fig. 3d, i). Comparing the concentration decreasing rate at different positions, we can find that, at the time before the meeting point (i.e., from 0 min to 1380 min after starting the experiment), the consumption of protein was faster at the upper part in the cell than at the lower part, showing that there were more protein crystals at the upper part than at the lower part. This phenomenon implied that adding SDB microspheres promoted nucleation on the surface of the microspheres, so that more protein molecules were consumed at the upper part of the solution. 1380 min after starting the experiment, the consumption of protein turned to be faster at the lower part (i.e., point c) than at the upper part of the cell (i.e., point a), showing that there were more protein crystals at the lower side than at the upper side. In other words, sedimentation of protein crystals from the upper part to the lower part happened so that the number of crystals at the lower part became larger than at the upper part, as judged from the variation of protein consumption rate at points a and c.
Discussion
Heterogeneous nucleants are useful in protein crystallisation because they increase the probability of protein crystal nucleation; this occurs because the surface microstructure of the nucleants can decrease the nucleation barrier, thus generating a crystal nucleus. Furthermore, some authors have reported that crystal quality can also be improved because the use of heterogeneous nucleants can promote protein crystallisation at low levels of supersaturation, which improves the quality of protein crystals. In the literature, it has been reported that crystals often appear next to the heterogeneous nucleants because the nucleation occurs at the surface of the nucleants. This finding provides good evidence that nucleants indeed promote nucleation. Here, however, we found that the protein crystals in most cases did not have direct contact with the SDB microspheres (Fig. 4). This fact leads to uncertainty as to whether the SDB microspheres play any role in the crystallisation.
Based on the experimental results, the answer to the above question is clearly “yes”. Statistically from the experiments in this study, the addition of the SDB microspheres increased the crystallisation hits in screening study in screening study (Fig. 1 and Table 1); simultaneously, the use of SDB microspheres often improved the crystal quality (Supplementary Figs. 5 and 6). How, then, does this technique achieve these results?
In this study, the SDB microspheres are a different type of heterogeneous nucleant that exhibit special characteristics. One important characteristic of these nucleants is their ability to adsorb protein molecules in crystallisation solutions, such that regions of high and low supersaturation can coexist in a single solution. Another important characteristic of these nucleants is their low density, which ensures that they float, thus maintaining a region of high supersaturation in the upper part of the solution. Combining these two characteristics indicates an explanation, which we propose below. Fig. 5a shows a schematic hypothesis showing the adsorption of protein molecules, nucleation of the crystals, sedimentation of larger crystals and continued growth at the bottom of the solution. According to this hypothesis, the observed concentration evolution in the solution can be explained. Fig. 5b is a schematic illustration of the concentration changes with time in the presence and absence of SDB microspheres. Due to the adsorbent nature of the material that they are constructed from, SDB microspheres will adsorb protein molecules38,39 when they are added to the solution. Therefore, a localised region of high concentration will be present on the surface of the microspheres. Due to this adsorption, the concentration in the solution will be reduced. Therefore, two regions with differing concentrations will coexist in the solution: a region on the surface of the microspheres where the concentration is high and a region in the lower part of the solution where the concentration is low. In Fig. 5b, the dashed blue line shows the changes in concentration with time in the conventional crystallisation droplet in the absence of SDB microspheres. The solid lines show the changes in concentration with time when SDB microspheres are added. The red line shows the changes in concentration with time near the crystals (from the formation of the nucleus to fully grown crystals) and the black line shows the changes in concentration with time at the bottom of the droplet. In the initial stage, nucleation would occur (most likely earlier than in the absence of SDB microspheres) at the surface of the microspheres where the concentration is high. Because the density of the microspheres is smaller than that of the crystallisation solution, the microspheres float on the solution. Thus, if the crystals grow on the surface of the microspheres, the crystals would sediment to the bottom of the solution due to gravity when the crystals grow larger than several microns40,41.
Concentration evolution during a crystallisation experiment in the presence and absence of SDB microspheres.
(a) Schematic illustration of the adsorption and crystallisation processes after adding the SDB microspheres. Adsorption facilitates nucleation on the surface of the microspheres; when the crystals grow to a certain size, the crystals will sediment to the bottom of the solution and continue to grow. (b) Schematic illustration of the change in concentration with time during the crystallisation in the presence and absence of the SDB microspheres. When no microspheres are present, the concentration evolution will follow the dashed blue line; when microspheres are added to the solution, adsorption of the protein molecules will lower the concentration in the solution (the black line) and at the same time, the concentration on the surface of the microspheres will increase (the red line). After nucleation and the initial growth on the surface of the microspheres, the crystals will sediment to the bottom. Thus, the concentration near the crystals will be that at the bottom of the solution (the red line).
Hence, the concentration near the crystals would follow the curve representing the change in concentration with time at the bottom after sedimentation. Apparently, at the bottom of the solution, the change in concentration with time would be slower (implying steadier growth) than that in the absence of SDB microspheres (normal growth) because the crystals are in a region of low concentration. This conclusion is consistent with the concentrations measured using a digital holographic Mach-Zehnder interferometer (Fig. 3) and the BCA method (Supplementary Fig. 9).
In a practical vapour diffusion experiment, the change in concentration with time will not strictly follow that outlined in the hypothesis. However, we can expect that the trend will be similar because the adsorption process is certain to occur; thus, a concentration gradient will still form and the proposed phenomena will occur.
The increase in the crystallisation success rate might be attributable to the characteristics of the SDB microspheres. In the literature, we find a number of extensive studies concerning the effect of heterogeneous nucleants on protein nucleation12,21,22,23,24,25,26,27,28,29,30,31,32,33,34. Many heterogeneous nucleants (such as mineral substrates, silicon substrates, TiO2, SiO2 and polystyrene) are reported to promote the nucleation of protein crystals by lowering the energy barrier for nucleation.
In this study, the porous structure of the SDB1 microspheres was also a beneficial plus for promoting the nucleation. On the one hand, the suitable pore sizes of SDB could help in nucleation. According to Chayen et al's report12, porous nucleants with broad pore-size distributions can help in nucleation. When the pore size is in a certain range (2.5–15 nm), the pores can interact with several types of proteins and hence promote crystallisation. In the current study, the pore sizes encompassed a wide range (micropores, 9–80 nm; macropores, 5–10 μm; average pore size 59.8 nm; see Supplementary Fig. 4); thus, the SDB1 microspheres can interact with protein molecules of different sizes.
On the other hand, the area of SDB1 was determined to be approximately 107.3 m2/g (Supplementary Fig. 4). Such a large surface area can provide many nucleation sites, which might increase the probability of obtaining protein crystals.
A further important reason for the enhancement of nucleation, as mentioned above, is related to the adsorption property of the SDB microspheres. Adsorption of the protein molecules increases the supersaturation level on the surface of the microspheres, increasing the driving force for nucleation and resulting in a higher nucleation probability.
Steadier growth may be one important mechanism by which the crystal quality is increased. Because most crystals were found at the bottom of the droplet, it is clear that these crystals grew at the lower side of the droplet. This observation is important because it indicates that the crystals sedimented after nucleation and (partial) growth. It has been reported that crystals usually sediment from the solution to the bottom of the droplet when the crystal size reaches several microns40,41. Although it is currently impossible to observe the sedimentation process directly, we are certain that sedimentation occurred due to gravity because nucleation was promoted on the surface of the microspheres but the crystals were found at the bottom of the droplet at the end of the experiment.
At the bottom of the droplet, the sedimented crystals can continue to grow, but under conditions of lower supersaturation. The SDB microspheres are capable not only of adsorbing protein molecules from the crystallisation solution but can also release protein molecules back into the solution (i.e., desorption) at low protein concentrations42. Thus in a crystallisation solution, when the concentration is high, the microspheres will adsorb protein molecules and hence, reduce the concentration of the solution, enabling the crystal to grow at lower levels of supersaturation. As crystal growth continues, the protein molecules in the solution will be consumed and the concentration will be reduced. In such cases, protein molecules will be desorbed from the microspheres, such that the concentration in the solution will be maintained at a relatively stable level. In such a growth environment, steady crystal growth can be achieved. Because protein crystal quality is closely related to growth, the improvement in crystal quality observed here most likely occurred because the crystals grew for a long time in a stable, low supersaturation environment43.
The current study demonstrates a method that uses low density, porous microspheres that are capable of adsorbing protein molecules as nucleants for protein crystallisation. The method proved to be successful in the crystallisation of commercially available proteins and other target proteins in that it simultaneously increased the crystallisation success rate and the number of crystallisation hits and improved crystal quality. Other materials having similar properties (adsorption and desorption) might also be useful as nucleants. Because many materials are heavier than crystallisation solutions, porous materials will not always float on the solution. In such cases, the method described in the current study can be still applied because the materials used can still be placed in the upper part of the solution, for example, by designing the crystallisation cells in an appropriate way. Apart from the approaches mentioned, a simpler method that might find wider application is to design a hanging-drop crystallisation plate that is constructed of a material having the desired protein adsorption and desorption characteristics.
Methods
Preparation of the SDB microspheres
Two kinds types of polystyrene-divinylbenzene (SDB) microspheres were prepared: porous polystyrene-divinylbenzene microspheres (SDB1) and non-porous polystyrene-divinylbenzene microspheres (SDB2). See Supplementary Note 1(1) for more details.
Crystallisation with and without SDB microspheres
Two types of crystallisation experiments were conducted to examine the effects of the SDB microspheres on the protein crystallisation. (i) Crystallisation screening test and (ii) Crystal quality comparison study. See Supplementary Note 1(2) for more details.
Concentration distribution study during the crystallisation process with and without SDB1 microspheres
To get the information of concentration distribution in the solution during crystallisation, we studied the concentration evolution in the crystallisation solution in the presence and absence of SDB1 microspheres using interferometry. See Supplementary 1(3) for more details.
References
Kuzmanic, A., Pannu, N. S. & Zagrovic, B. X-ray refinement significantly underestimates the level of microscopic heterogeneity in biomolecular crystals. Nat. Commun. 5, 3220 (2014).
Kundrot, C. E. Which strategy for a protein crystallisation project? Cell Mol. Life Sci. 61, 525–536 (2004).
Fusco, D., Headd, J. J., DeSimone, A., Wang, J. & Charbonneau, P. Characterizing protein crystal contacts and their role in crystallization: rubredoxin as a case study. Soft Matter 10, 290–302 (2014).
Yoshikawa, H. Y. et al. Laser ablation for protein crystal nucleation and seeding. Chem. Soc. Rev. 43, 2147–2158 (2014).
Fox, B. G., Goulding, C., Malkowski, M. G., Stewart, L. & Deacon, A. Structural genomics: from genes to structures with valuable materials and many questions in between. Nat. Methods 5, 129–132 (2008).
Price, W. N. et al. Understanding the physical properties that control protein crystallization by analysis of large-scale experimental data. Nat. Biotechnol. 27, 51–57 (2009).
Redecke, L. et al. Natively Inhibited Trypanosoma brucei Cathepsin B Structure Determined by Using an X-ray Laser. Science 339, 227–230 (2013).
Newman, J. One plate, two plates, a thousand plates. How crystallisation changes with large numbers of samples. Methods 55, 73–80 (2011).
Mittl, P. R. E. & Grutter, M. G. Structural genomics: opportunities and challenges. Curr. Opin. Chem. Biol. 5, 402–408 (2001).
McPherson, A. Introduction to protein crystallization. Methods 34, 254–265 (2004).
Chayen, N. E. & Saridakis, E. Protein crystallisation: from purified protein to diffraction-quality crystal. Nat. Methods 5,147–153 (2008).
Chayen, N. E., Saridakis, E. & Sear, R. P. Experiment and theory for heterogeneous nucleation of protein crystals in a porous medium. P. Natl. Acad. Sci. USA 103, 597–601 (2006).
Gerdts, C. J. et al. Time-controlled microfluidic seeding in nL-volume droplets to separate nucleation and growth stages of protein crystallization. Angew Chem. Int. Edit. 45, 8156–8160 (2006).
Chayen, N. E. Methods for separating nucleation and growth in protein crystallization. Prog. Biophys. & Mol. Biol. 88, 329–337 (2005).
Saridakis, E., Dierks, K., Moreno, A., Dieckmann, M. W. M. & Chayen, N. E. Separating nucleation and growth in protein crystallization using dynamic light scattering. Acta Crystallogr. D 58, 1597–1600 (2002).
Dierks, K., Meyer, A., Einspahr, H. & Betzel, C. Dynamic light scattering in protein crystallisation droplets: Adaptations for analysis and optimization of crystallisation processes. Cryst. Growth Des. 8, 1628–1634 (2008).
Oberthuer, D. et al. Monitoring and Scoring Counter-Diffusion Protein Crystallisation Experiments in Capillaries by in situ Dynamic Light Scattering. PLoS One 7, E33450 (2012).
Bergfors, T. Seeds to crystals. J. Struct. Biol. 142, 66–76 (2003).
Khurshid, S., Haire, L. F. & Chayen, N. E. Automated seeding for the optimization of crystal quality. J. Appl. Crystallogr. 43, 752–756 (2010).
Stewart, P. D. S., Kolek, S. A., Briggs, R. A., Chayen, N. E. & Baldock, P. F. Random microseeding: a theoretical and practical exploration of seed stability and seeding techniques for successful protein crystallization. Cryst. Growth & Des. 11, 3432–3441 (2011).
McPherson, A. & Shlichta, P. Heterogeneous and epitaxial nucleation of protein crystals on mineral surfaces. Science 239, 385–387 (1988).
D'Arcy, A., MacSweeney, A. & Haber, A. Using natural seeding material to generate nucleation in protein crystallisation experiments. Acta Crystallogr. D 59, 1343–1346 (2003).
Edwards, A. M. et al. The mechanism of protein crystal growth from lipid layers. J. Mol. Biol. 246, 38–316 (1997).
Paxton, T. E., Sambanis, A. & Rousseau, R. W. Mineral substrates as heterogeneous nucleants in the crystallisation of proteins. J. Cryst. Growth 198, 656–660 (1999).
Cacciuto, A., Auer, S. & Frenkel, D. Onset of heterogeneous crystal nucleation in colloidal suspensions. Nature 428, 404–406 (2004).
Chayen, N. E., Saridakis, E., El-Bahar, R. & Nemirovsky, Y. Porous silicon: an effective nucleation-inducing material for protein crystallisation. J. Mol. Biol. 312, 591–595 (2001).
Derewenda, Z. S. Rational protein crystallisation by mutational surface engineering. Structure 12, 529–535 (2004).
Georgieva, D. G., Kuil, M. E., Oosterkamp, T. H., Zandbergen, H. W. & Abrahams, J. P. Heterogeneous nucleation of three-dimensional protein nanocrystals. Acta Crystallogr. D 63, 564–570 (2007).
Guo, Y. Z., Yin, D. C., Lu, Q. Q., Wang, X. K. & Liu, J. Enhancement of nucleation during hanging drop protein crystallisation using HF treatment of cover glasses. Cryst. Res. Technol. 45, 158–166 (2010).
Zhang, C. Y. et al. An investigation of the effects of self-assembled monolayers on protein crystallisation. Int. J. Mol. Sci. 14, 12329–12345 (2013).
Guo, Y. Z. et al. Surface treatment by oxidizing the plates can alter the response of protein crystallization. J. Appl. Crystallogr. 47, 228–236 (2014).
Kertis, F. et al. Heterogeneous nucleation of protein crystals using nanoporous gold nucleants. J. Mater. Chem. 22, 21928–21934 (2012).
Kallio, J. M. et al. The contribution of polystyrene nanospheres towards the crystallization of proteins. PLoS One 4, e4198 (2009).
Thakur, A. S. et al. Improved success of sparse matrix protein crystallization screening with heterogeneous nucleating agents. PLoS One 2, e1091 (2007).
Snook, C. F., Purdy, M. D. & Wiener, M. C. Use of a crystallisation robot to set up sitting-drop vapor-diffusion crystallisation and in situ crystallisation screens. J. Appl. Crystallogr. 33, 344–349 (2000).
Drebes, J., Perbandt, M., Wrenger, C. & Betzel, C. Purification, crystallization and preliminary X-ray diffraction analysis of ThiM from Staphylococcus aureus. Acta Crystallogr. F 67, 479–481 (2011).
Meyer, A. et al. The mistletoe lectin I--phloretamide structure reveals a new function of plant lectins. Biochem. Biophys. Res. Commun. 364, 195–200 (2007).
Unsworth, L. D., Sheardown, H. & Brash, J. L. Protein resistance of surfaces prepared by sorption of end-thiolated poly (ethylene glycol) to gold: effect of surface chain density. Langmuir. 21, 1036–1041 (2005).
Rabe, M., Verdes, D. & Seeger, S. Understanding protein adsorption phenomena at solid surfaces. Adv. Colloid Interfac. 162, 87–106 (2011).
Ataka, M., Katoh, E. & Wakayama, N. I. Magnetic orientation as a tool to study the initial stage of crystallisation of lysozyme. J. Cryst. Growth 173, 592–596 (1997).
Yanagiya, S. et al. Effect of a magnetic field on the orientation of hen egg-white lysozyme crystals. J. Cryst. Growth 196, 319–324 (1999).
Rabe, M., Verdes, D. & Seeger, S. Understanding protein adsorption phenomena at solid surfaces. Adv. Colloid Interfac. 162, 87–106 (2011).
Yoshizaki, I. et al. Systematic analysis of supersaturation and lysozyme crystal quality. Acta Crystallogr. D 57, 1621–1629 (2001).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31170816), the National Basic Research Program of China (973 Program) (Grant No. 2011CB710905), the National Natural Science Foundation of China (Grant Nos. 11202167, 51201137, 31200551), the Röntgen-Angström-Cluster (Project 05K12GU3), the German Federal Ministry of Education and Research (BMBF), the PhD program Foundation of the Ministry of Education of China (20116102120052) and the China Postdoctoral Science Foundation (Grant No. 2013T60890). CB was supported by the excellence cluster 'The Hamburg Centre for Ultrafast Imaging-Structure, Dynamics and Control of Matter at the Atomic Scale' of the Deutsche Forschungsgemeinschaft (DFG). We would like to thank Ji Li for providing HSP90, Zong-Xiang Xia for providing TCS, Arne Meyer for providing ML-I and Julia Drebes and Madeleine Künz for providing ThiM.
Author information
Authors and Affiliations
Contributions
D.-C.Y. and Y.-Z.G. designed research; Y.-Z.G., L.-H.S., D.O., C.B., J.-L.D., B.-L.Z., H.-L.C., Q.W. and J.-M.S. performed research, Y.-M.L., J.L., H.-H.H., J.-H.H., J.-L.Z. and Q.-Y.Z. contributed new reagents or analytic tools; Y.-Z.G., D.-C.Y., H.-H.H. and C.-Y.Z. analyzed data; Y.-Z.G., D.-C.Y., C.B. and J.-Y.S. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Guo, YZ., Sun, LH., Oberthuer, D. et al. Utilisation of adsorption and desorption for simultaneously improving protein crystallisation success rate and crystal quality. Sci Rep 4, 7308 (2014). https://doi.org/10.1038/srep07308
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07308
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.