Abstract
Formalin-fixed paraffin-embedded (FFPE) tissues are important resources for molecular medical research. However, long-chain RNA analysis is restricted in FFPE tissues due to high levels of degradation. To explore the possibility of long RNA quantification in FFPE tissues, we selected 14 target RNAs (8 mRNAs and 6 long noncoding RNAs) from literatures and designed short (~60 bp) and long (~200 bp) amplicons for each of them. Colorectal carcinomas with adjacent normal tissues were subjected to quantitative reverse-transcription PCR (quantitative RT-PCR) in 3 cohorts, including 18 snap-frozen and 83 FFPE tissues. We found that short amplicons were amplified more efficiently than long amplicons both in snap-frozen (P = 0.0006) and FFPE (P = 0.0152) tissues. Nonetheless, comparison of colorectal carcinomas with their adjacent normal tissues demonstrated that the consistency of fold-change trends in a single short amplicon between snap-frozen and FFPE tissues was only 36%. Therefore, we innovatively performed quantitative RT-PCR with 3 non-overlapping short amplicons for 14 target RNAs in FFPE tissues. All target RNAs showed a concordance of 100% of fold-change trends in at least two short amplicons, which offers sufficient information for accurate quantification of target RNAs. Our findings demonstrated the possibility of long-chain RNA analysis with 3 non-overlapping short amplicons in standardized-preserved FFPE tissues.
Similar content being viewed by others
Introduction
Large numbers of surgically resected tissues and biopsy specimens are preserved as formalin-fixed and paraffin-embedded (FFPE) blocks. Therefore, FFPE tissues have been and still are the most abundant supply of clinical samples, which are often accompanied with sufficient follow-up data. Due to well-preserved morphology, FFPE is the standard process to handle post-mortem tissues which are very important sources of rare disease1. In addition, FFPE tissues also play an irreplaceable role in the fast growing translation medical research through bridging laboratory work and clinical applications. Full exploration of FFPE tissues is essential for medical research.
Unfortunately, the nucleic acids present extensive degradation in FFPE tissues and random fragmentation differs in normal and tumour tissues, making it more problematic to measure long RNA molecules. The average fragment length of DNA in FFPE tissues is often less than 300 bp, while the degradation of RNA is even worse due to the inherent instability as a single-stranded molecule2. In contrast, the degraded RNA cannot be restored like DNA using the other DNA strand as a template1. Due to these reasons, researchers prefer frozen tissues to FFPE tissues in long RNA exploration. Nonetheless, frozen tissues may not easily replace FFPE tissues because: (i) RNA degradation is unavoidable in archived snap-frozen tissues even though frozen clinical samples are considered to be well-preserved sources of nucleic acids3; (ii) a lack of sufficient clinical outcomes (e.g. prognosis and response to treatment) exists in the use of frozen tissues. Taken together, there is a pressing need to explore the possibility of RNA analysis in FFPE tissue.
Recently, the identification of abundant long non-coding RNAs (lncRNAs) (> 200 bp in length) has catalysed their roles as drivers of tumour suppressive and oncogenic functions4,5,6,7,8,9,10,11. LncRNAs are a novel class of mRNA-like transcripts which represent the leading edge of cancer research. Their identity, function and dysregulation in various cancer types are beginning to be discovered. Increased research on these RNAs may lead to novel clinical applications in cancer biogenesis and prognosis. Unfortunately, the inherent instability of lncRNA makes the current studies mainly limited in frozen tissues12.
Short RNA molecule fragments are stable and detectable in quantitative reverse-transcription PCR (quantitative RT-PCR)13,14,15,16,17,18. Meanwhile, previous studies have proved that amplicon size can affect the quantification of mRNA and smaller size can increase the yield of amplifiable mRNA18,19,20,21. However, the quantification reliability of such short amplicons in long-chain RNA molecules has not been systematically evaluated in FFPE tissues.
In this study, we investigated the possibility of long RNA quantification in FFPE tissues based on the broken molecular fragments. Inspired by previous studies, we selected 14 target RNAs (8 mRNAs and 6 lncRNAs) from literatures and designed short and long amplicons for each of them to evaluate the efficiency and reliability of quantifications in frozen and FFPE tissues. To address the problem of random fragmentation at different extents in tumour and normal tissues, we explored the expression profiles of long RNA quantification with 3 non-overlapping short amplicons in FFPE tissues from 65 patients with non-metastasis colorectal cancer (CRC).
Results
Evaluation of amplification efficiency
Amplification efficiency is an important indicator that shows whether an amplicon works or not. To evaluate the quantification efficiency of short amplicons, parallel quantitative RT-PCR was initially performed with short-A and long amplicons on the mixed frozen and FFPE RNA samples to generate standard curves. The amplification efficiency and coefficient of determination (R2) of short and long amplicons are presented in Table 1. The results in Fig. 1 demonstrated that short amplicons were more efficient than long amplicons both in frozen (P = 0.0006) and FFPE tissues (P = 0.0152).
Box-and-whisker plots of amplification efficiency and coefficient of determination (R2) obtained from short and long amplicons both in frozen and FFPE tissues.
(a): Box-and-whisker plot of amplification efficiency. (b): Box-and-whisker plot of coefficient of determination (R2). Frozen-short A: quantification with short-A amplicons in frozen tissues. Frozen-long: quantification with long amplicons in frozen tissues. FFPE-short A: quantification with short-A amplicons in FFPE tissues. FFPE-long: quantification with long amplicons in FFPE tissues.
In frozen tissues, the short amplicon of the endogenous control β-actin had full amplification efficiency (100%) with R2 = 0.999, while 11 (79%) of 14 target RNAs achieved optimal efficiencies between 90% and 110% with R2 > 0.960. In contrast, the amplification efficiency of long amplicons differed remarkably amongst different target RNAs and only SMAD4 reached the optimal efficiency. Additionally, the R2 of long amplicons was 0.885 ± 0.1 (Mean ± SD), which was significantly lower than that of short amplicons (0.990 ± 0.0, P < 0.0001).
When it came to FFPE tissues, 11 (73%) of 15 short amplicons successfully achieved the optimal efficiencies between 90% and 110% and the R2 of short amplicons was 0.957 ± 0.1 (Mean ± SD). In contrast, the amplification efficiency of most long amplicons was overblown and even β-actin failed to achieve an optimal efficiency. The R2 of long amplicons was 0.577 ± 0.3 (Mean ± SD), which was significantly lower than that of short amplicons (0.957 ± 0.1, P < 0.0001).
Evaluation of quantification correlation
To determine the quantification consistency with short amplicons, the expression profiles of the 14 target RNAs with short-A amplicons were examined in frozen tissues and compared with those of the paired long amplicons. Figure 2A intuitively shows the mean Ct values of the studied RNAs. The results demonstrated that the mean Ct values of short amplicons were significantly lower than that of long amplicons (P < 0.0001). Generally, short amplicons had 1.8 cycles less than that of long amplicons in quantitative RT-PCR, suggesting that short amplicons were more sensitive for the quantification in frozen tissues. Moreover, the short-A amplicons were evaluated in FFPE tissues from the exact CRC cases of aforementioned frozen tissues. The results presented in Fig. 2B indicated that the Ct values of short amplicons did not show significantly difference between frozen and FFPE tissues (P = 0.1455), suggesting that short amplicons quantified effectively in FFPE tissues.
Comparisons of Ct values obtained from short and long amplicons in frozen and FFPE tissues.
(a): Comparison of Ct values obtained from short and long amplicons in frozen tissues. (b): Comparison of Ct values obtained from short amplicons in frozen and FFPE tissues. Frozen-short A: quantification with short-A amplicons in frozen tissues. FFPE-short A: quantification with short-A amplicons in FFPE tissues. Error bars indicate the SD of the Ct values.
To test the validity of short amplicons in detecting the different expression patterns, the expression profiles of the target RNAs were generated with paired short and long amplicons in frozen and FFPE tissues. Table 2 shows the relative expression levels of target RNAs in CRC tissues compared with their adjacent normal tissues. The quantification consistency between short and long amplicons was 64% in frozen tissues. Worse than this, the quantification consistency of short amplicons between frozen and FFPE tissues was only 36%. The results suggested that the quantification with a single short amplicon was unreliable.
Evaluation of quantification reliability
Considering that the aforementioned poor consistency might be caused by the random RNA fragmentation at different extent in tumour and normal tissues, primer sets for two additional short amplicons (short-B and C) were designed for each of the studied RNAs. Thereafter, quantitative RT-PCR with 3 non-overlapping short amplicons (short-A, B and C) was performed in 65 colorectal tumour-normal pairs of FFPE tissues, including 32 Dukes' A and 33 Dukes' B.
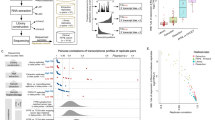
As Fig. 3 shows, a close relativity existed amongst 3 short amplicons. The correlation between the mean Ct values and individual short-A, B and C amplicon was 0.9386, 0.8437 and 0.9581, respectively. The correlation between the mean Ct values and short-B amplicon was much lower than that of the other two short amplicons, which was also reflected by the separate data points in Fig. 3A. In the comparison of colorectal carcinomas with their adjacent normal tissues, the correlation between the mean fold change values and individual short-A, B and C amplicon was 0.9608, 0.9064 and 0.8874, respectively (Figure 3B). As it can be seen from Table 3, 14 target RNAs showed a concordance of 57% on the fold change trends (regulation or no change) with 3 short amplicons, while the quantification consistency obtained from at least two short amplicons was 100%. The results demonstrated the possibility of long RNA analysis in FFPE tissues using 3 non-overlapping short amplicons.
Correlation of Ct values and fold changes obtained from 3 non-overlapping short amplicons in FFPE tissues.
(a): Correlation of Ct values obtained from 3 non-overlapping short amplicons in 65 FFPE colorectal surgical tissues. (b): Correlation of fold changes (tumour/normal) obtained from 3 non-overlapping short amplicons in 65 FFPE colorectal surgical tissues. Mean Ct values of 3 short amplicons for each target RNA were calculated by individual sample.
Discussion
FFPE tissues are important resources for biomedical research and play an irreplaceable role in translational medicine research. However, there exists extensive degradation of nucleic acids especially for long-chain RNA molecules, such as mRNA and lncRNA12. To explore the possibility of long RNA analysis in FFPE tissues, we evaluated the quantification efficiency and reliability of short amplicons. Our major discovery in this study is that we can obtain reliable quantification data of long-chain RNA from FFPE tissues by performing quantitative RT-PCR with 3 non-overlapping short amplicons, which makes it possible to study mRNA and lncRNA in FFPE tissues.
The effect of amplicon size on quantification has been revealed previously18,19,20,21, however, in this study, we evaluated the effect in a more rigorous way. As we mentioned in primer design, the short amplicon and corresponding long amplicon shared a forward or a reverse primer sequence, therefore, the short amplicon was always a segment within the long amplicon. Our results indicated that short amplicons achieved optimal efficiency, however, most long amplicons had poor amplification efficiency both in frozen and FFPE tissues. In general, 90%–110% is considered to be the optimal efficiency. Lower efficiency of the amplicon suggests that the corresponding primer set barely work, while higher efficiency outside this range suggests possible PCR inhibition22. It shall be noted that most PCR procedures assume that the amplification efficiency for the amplicon of interest is constant and nearly 100%. Furthermore, the commonly used comparative Ct method even assumes equal efficiency in reference and target amplicons. However, such a comparative Ct method is very sensitive to variation in PCR efficiency23. The parameter R2 denotes the repeatability of efficiency, not only amongst triplicate samples of same diluted gradient, but also amongst serial dilution samples. The significantly higher R2 of short amplicon is a further proof of stable efficiency, which implies that the good amplification efficiency is maintained throughout different RNAs.
Short amplicons are more efficient than long amplicons in quantitative RT-PCR. The Ct value is the PCR cycle at which a statistically significant increase in target amplicon fluorescence can be detected above background. Our results indicated that short amplicons generally required 1.8 cycles less than that of long amplicons in frozen tissues. The smaller size has offered the possibility of higher sensitivity for short amplicons since shorter sequence will be less affected by the fragmentations. It is generally believed that long-chain RNA is often better preserved and less frequently broken in frozen tissues compared to FFPE tissues. Our results demonstrated that the Ct values obtained from the two types of tissues with short amplicons did not show significant differences. This implies that with short amplicons, FFPE and frozen tissues are promising to be used interchangeably in lncRNA and mRNA analysis by quantitative RT-PCR.
Random fragmentation of RNA to different extent in matched normal and tumour tissues compromised the quantification reliability of short amplicons. In our previous study14, we used RNA integrity number (RIN) to measure the quality of snap-frozen RNA samples and it turned out to be 7.3 ± 1.1 (Mean ± SD) in CRC tissues and 3.3 ± 1.6 (Mean ± SD) in adjacent normal tissues. The significant difference of RNA quality between normal and tumour samples also occurs in FFPE tissues. This could be due to the fact that RNase activity in CRC tissues is lower than that in normal tissues24. The fragmentation occurred in amplicon sequence or in primer sequence will cause the failure of amplification and further lead to the poor quantification consistency. It is impossible to get the degradation ratio between tumour and normal tissues and therefore, varying degrees of degradation might not be systematically corrected3.
Using 3 non-overlapping short amplicons for one target RNA, we can reliably determine the expression profiles of lncRNA and mRNA in FFPE tissues by quantitative RT-PCR. Degradation of different extent in tumour and normal tissues is a troublesome problem in quantification. As such, using one amplicon, no matter long or short, cannot reliably validate the expression of a target RNA even in frozen tissues. Three short amplicons can address this problem and ultimately determine the expression profiles of target RNAs. Nevertheless, slight differences existed between our results and the published data for some of the RNA expression profiles. In the final analysis, there are 3 main reasons for the differences. First, for the previous reports, the quality of their studied RNA samples and the details of primer sets in quantitative RT-PCR are not available. As shown in this study, RNA integrity and primer design are two critical points to guarantee the quantification reliability. Second, some studies used cultured human CRC cells. The change of microenvironment in vitro might affect the RNA expression profile. Last but not the least, different stages of CRC would present different RNA expression profiles. Dukes' A and B were selected in our study for screening lncRNAs and mRNAs related to tumorigenesis. In late stages of CRC, the differentially expressed RNAs may be more related to tumour metastasis. On the other hand, necrosis in late stage tumour tissues is more severe and brings about extensive RNA degradation.
A previous study on plasma-based lncRNA demonstrated significant difference in expression levels amongst different segments of MALAT113. Likewise, short-B amplicon we designed for MALAT1 seemed to be at a much lower expression level than short-A and C amplicons in CRC. The mechanism for this remained unclear. Further studies are needed to confirm the characteristics of short-B amplicon in MALAT1.
There still exists a limitation for the quantification of mRNA using short amplicons. One widely applied guideline for mRNA amplification is that the amplicon should span one or more introns to avoid possible interference from genomic DNA. However, since the primer set we designed only amplify a product of 50–70 bp, it is impossible to span introns unless the intron is short enough. For this point, short amplicons are not as good as long amplicons. Considering this, it is important to ensure there is no genomic DNA contamination in the RNA.
In conclusion, we have investigated a short-amplicon approach in quantitative RT-PCR to quantify lncRNA and mRNA in FFPE tissues. Our study shows that 3 non-overlapping short amplicons for one target RNA provide the possibility of long-chain RNA analysis in standardized- preserved FFPE tissues.
Methods
Clinical specimens and study design
The study was approved by the Institutional Review Board of Shanghai Medical College in Fudan University, with written informed consent obtained from all patients. All experiments in this study were performed in accordance with the approved guidelines and regulations. Colorectal carcinomas with adjacent normal tissues, including 18 snap-frozen and 83 FFPE surgical specimens, were collected from Zhongshan and Huashan Hospitals in Fudan University between 2007 and 2013. For the CRC sample selection, routine histological classification was used according to the WHO classification of tumours25. All cases were diagnosed by two pathologists and independently reviewed by an expert CRC pathologist. Patients who had received preoperative radiotherapy or chemotherapy were excluded. Supplementary Table S1 summarizes the clinical characteristics of the patients and tumours in the study.
The study consisted of 3 different phases. All samples were allocated to the 3 phases without overlapping between any two phases (Figure 4). In phase 1, the amplification efficiency of short and long amplicons was evaluated in 6 independent frozen and FFPE surgical colorectal tissues from 12 patients. To eliminate the possible interference from individual differences on the final results, the total RNA samples from the frozen and FFPE tissues were mixed, respectively. The standard curves on the 14 target RNAs and an endogenous control were generated in the mixed frozen and FFPE RNA samples using quantitative RT-PCR with paired short and long amplicons.
In phase 2, the quantification correlation between short and long amplicons was determined in paired frozen and FFPE surgical colorectal tissues from 12 patients. Manual macrodissection was performed on both frozen and FFPE tissues to ensure the presence of > 75% cancer cells or normal epithelial cells. The expression profiles of the 14 target RNAs in the comparison of colorectal carcinomas with their adjacent normal tissues were assessed using quantitative RT-PCR with paired short and long amplicons.
In phase 3, the quantification reliability of 3 non-overlapping short amplicons was investigated on the 14 target RNAs in FFPE tissues from 65 non-metastasis CRC patients. The manual macrodissection was also performed as mentioned in the phase 2 before the RNA extraction. The expression profiles of the 14 target RNAs in the comparison of colorectal carcinomas with their adjacent normal tissues were determined using quantitative RT-PCR with 3 non-overlapping short amplicons.
Macrodissection
Macrodissection was essentially as described in our previous studies15,16. H&E-stained sections of each of the snap-frozen and FFPE tissue blocks were prepared to check the proportion of tumour material. If a tumour had more than 75% neoplastic cells, it was deemed suitable for analysis without further purification of tumour cells. If, however, histology showed the tumour having < 75% cancer cells, it was selected and marked as tumour for manual macrodissection. Similarly, if histology showed the normal tissue having < 75% epithelial cells, it was also selected and marked epithelial cells for manual macrodissection.
RNA isolation
Total RNA of the frozen tissue sections was extracted using the RNeasy Plus Mini Kit according to the instructions of the manufacturer (QIAGEN, Hilden, Germany). For the frozen tissues, genomic DNA was eliminated by the gDNA Eliminator spin column. Total RNA of the FFPE tissue sections was isolated using the RecoverAll Total Nucleic Acid Isolation Kit according to the instructions of the manufacturer (Ambion, Austin, Texas, USA). For the FFPE tissues, DNase digestion to the nucleic acid samples was included before the final purification of RNA. The concentration of RNA sample was quantified by NanoDrop 2000 Spectrophotometers (Thermo Fisher Scientific, Waltham, MA, USA). The quality of RNA sample was determined by OD 260/280 ratio. A RNA sample was discarded for further analysis if OD 260/280 ratio was less than 1.8.
Target RNA selection and primer design
Fourteen tumour-related long chain RNAs were selected as target molecules from literatures6,7,8,9,10,11,15,26,27,28,29,30,31,32,33,34, including 8 mRNAs (CDK6, JUN, MAPK3, PDGFRA, PDGFRB, PIK3R3, SMAD3, SMAD4) and 6 lncRNAs (DD3, H19, MALAT1, MEG3, p15AS, UCA1). β-actin mRNA was used as an endogenous control in quantitative RT-PCR. Supplementary Table S2a and S2b present the primer sequences and transcript variants of the target RNAs, respectively.
Two groups of primer sets were designed to amplify RNA short (~60 bp, short-A) and long (~200 bp) segments within the target sequences. For each of the target RNAs, the paired primer sets for the short and long amplicons shared either a forward or a reverse primer sequence. Such primer design ensured that the resulting short amplicon was always a segment of the long amplicon. Furthermore, additional primer sets were design to amplify other two short-amplicons (short-B and short-C) for each of the target RNAs. All 3 short amplicons were non-overlapping, arbitrarily chosen segments within the RNA sequence.
All the primer sets were designed in line with the general principles and all splice variants were targeted for the amplification. The primer specificity was screened by performing BLAST using the Human genomic plus transcript database (Human G+T) and further verified through melting curve obtained in quantitative RT-PCR. Supplementary Figure S3 shows the melting curves on all the investigated primers. The primers were synthetized by Sangon Biotech (Shanghai, China) and the purification was performed by high affinity purification.
Quantitative RT-PCR
Following the ‘Minimal Information for Publication of Quantitative Real-Time PCR Experiments’ (MIQE) guidelines, quantitative RT-PCR of the 14 target RNAs and β-actin gene was performed on 7900HT Fast Real-Time PCR System (Applied Biosystems) using High Capacity cDNA Reverse Transcription Kit (Invitrogen, Foster City, California, USA) and Power SYBR Green PCR Master Mix (Applied Biosystems, Warrington, UK) according to the instructions of manufacturers.
For the reverse transcription, 500 ng of total RNA sample was reverse transcripted into cDNA by the random primers in a total volume of 50 μl. For real-time PCR, 6 μl of the cDNA solution was amplified with 16 μl 2X SYBR Green mix and 2 μl target-specific primers (5 μM/L) in a final volume of 32 μl. All assays were carried out in triplicate. The Ct values were determined using 40 cycles.
Statistical analysis
The amplification efficiency and coefficient of determination (R2) of an amplicon were generated from standard curve. Amplification efficiency was calculated using the equation: E = 10-1/k-1 in which E is the amplification efficiency and k is the slope of standard curve. Mann-Whitney unpaired test was used to determine the significant difference of amplification efficiency between short and long amplicons.
Target RNA expression level was normalized to β-actin gene. The relative RNA expression between tumour and normal samples was calculated using the equation: ΔCt = Ct (target) -Ct (β-actin); ΔΔCt = ΔCt (tumour) -ΔCt (normal); Fold change was calculated using the equation: Folds = 2-ΔΔCt. Wilcoxon test for paired samples was used to determine the significant difference of target RNA expression level between colorectal carcinomas and their adjacent normal tissues. The target RNA was considered as being up-regulated if the fold change exceeded 2.0 or as being down-regulated if the fold change was below 0.5. The fold change between 0.5 and 2.0 was defined as unchanged. MedCalc software (version 10.4.7.0; MedCalc, Mariakerke, Belgium) was used to perform the statistical analysis. All P values were two-tailed, with P value < 0.05 considered significantly different.
References
Bonin, S., Petrera, F., Niccolini, B. & Stanta, G. PCR analysis in archival postmortem tissues. Mol Pathol 56, 184–186 (2003).
Carninci, P. Molecular biology,The long and short of RNAs. Nature 457, 974–975 (2009).
Ibberson, D., Benes, V., Muckenthaler, M. U. & Castoldi, M. RNA degradation compromises the reliability of microRNA expression profiling. BMC Biotechnol 9, 102 (2009).
Prensner, J. R. & Chinnaiyan, A. M. The emergence of lncRNAs in cancer biology. Cancer Discov 1, 391–407 (2011).
Brunner, A. L. et al. Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a tdiverse panel of archived human cancers. Genome Biol 13, R75 (2012).
Zhang, X. et al. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. J Clin Endocrinol Metab 88, 5119–5126 (2003).
Pasmant, E., Sabbagh, A., Vidaud, M. & Bièche, I. ANRIL, a long, noncoding RNA, is an unexpected major hotspot in GWAS. FASEB J 25, 444–448 (2011).
Bussemakers, M. J. et al. DD3: a new prostate-specific gene, highly overexpressed in prostate cancer. Cancer Res 59, 5975–5979 (1999).
Lin, R., Maeda, S., Liu, C., Karin, M. & Edgington, T. A large noncoding RNA is a marker for murine hepatocellular carcinomas and a spectrum of human carcinomas. Oncogene 26, 851–858 (2006).
Ji, P. et al. MALAT-1, a novel noncoding RNA and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003).
Wang, X. S. et al. Rapid identification of UCA1 as a very sensitive and specific unique marker for human bladder carcinoma. Clin Cancer Res 12, 4851–4858 (2006).
Clark, M. B. et al. Genome-wide analysis of long noncoding RNA stability. Genome Res 22, 885–898 (2012).
Ren, S. et al. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer 49, 2949–2959 (2013).
Wang, S. et al. Improvement of tissue preparation for laser capture microdissection: application for cell type-specific miRNA expression profiling in colorectal tumors. BMC Genomics 11, 163 (2010).
Wang, S. et al. A microRNA panel to discriminate carcinomas from high-grade intraepithelial neoplasms in colonoscopy biopsy tissue. Gut 62, 280–289 (2013).
Huang, W. et al. Two microRNA panels to discriminate three subtypes of lung carcinoma in bronchial brushing specimens. Am J Respir Crit Care Med 186, 1160–1167 (2012).
Abrahamsen, H. N., Steiniche, T., Nexo, E., Hamilton-Dutoit, S. J. & Sorensen, B. S. Towards quantitative mRNA analysis in paraffin-embedded tissues using real-time reverse transcriptase-polymerase chain reaction: a methodological study on lymph nodes from melanoma patients. J Mol Diagn 5, 34–41 (2003).
Antonov, J. et al. Reliable gene expression measurements from degraded RNA by quantitative real-time PCR depend on short amplicons and a proper normalization. Lab Invest 85, 1040–1050 (2005).
Specht, K. et al. Quantitative gene expression analysis in microdissected archival formalin-fixed and paraffin-embedded tumor tissue. Am J Pathol 158, 419–429 (2001).
Godfrey, T. E. et al. Quantitative mRNA expression analysis from formalin-fixed, paraffin-embedded tissues using 5' nuclease quantitative reverse transcription-polymerase chain reaction. J Mol Diagn 2, 84–91 (2000).
Lehmann, U. & Kreipe, H. Real-time PCR analysis of DNA and RNA extracted from formalin-fixed and paraffin-embedded biopsies. Methods 25, 409–418 (2001).
Suslov, O. & Steindler, D. A. PCR inhibition by reverse transcriptase leads to an overestimation of amplification efficiency. Nucleic Acids Res 33, e181 (2005).
Ramakers, C., Ruijter, J. M., Deprez, R. H. L. & Moorman, A. F. M. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339, 62–66 (2003).
Roth, J. S. Ribonuclease activity and cancer: A review. Cancer res 23, 657–666 (1963).
Hamilton, S. R. & Aaltonen, L. A. (eds.): Pathology and Genetics of Tumours of the Digestive System. (IARC Press, Lyon, 2003).
Tsang, W. P., Wong, T. W., Cheung, A. H. & Kwok, T. T. Induction of drug resistance and transformation in human cancer cells by the noncoding RNA CUDR. RNA 13, 890–898 (2007).
Tsang, W. P. et al. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis 31, 350–358 (2010).
Zhu, X. et al. miR-137 inhibits the proliferation of lung cancer cells by targeting Cdc42 and Cdk6. FEBS Lett 587, 73–81 (2013).
Xu, L. et al. MicroRNA-7–regulated TLR9 signaling–enhanced growth and metastatic potential of human lung cancer cells by altering the phosphoinositide-3-kinase, regulatory subunit 3/Akt pathway. Mol Biol Cell 24, 42–55 (2013).
Verhaak, R. G. et al. Integrated Genomic Analysis Identifies Clinically Relevant Subtypes of Glioblastoma Characterized by Abnormalities in PDGFRA, IDH1, EGFR and NF1. Cancer cell 17, 98–110 (2010).
Miyaki, M. et al. Higher frequency of Smad4 gene mutation in human colorectal cancer with distant metastasis. Oncogene 18, 3098–3103 (1999).
Kuwai, T. et al. Targeting the EGFR, VEGFR and PDGFR on colon cancer cells and stromal cells is required for therapy. Clin Exp Metastasis 25, 477–489 (2008).
Baba, Y. et al. Prognostic significance of AMP-activated protein kinase expression and modifying effect of MAPK3/1 in colorectal cancer. Br J Cancer 103, 1025–1033 (2010).
Arai, T. et al. Genomic structure of the human Smad3 gene and its infrequent alterations in colorectal cancers. Cancer Lett 122, 157–163 (1998).
Author information
Authors and Affiliations
Contributions
H.K. designed, carried out the experiments, analysed the data and prepared the draft of manuscript. M.Z., F.C. and S.W. participated the experiments. X.G. and S.L. provided the samples for experiments. Y.W. and H.Z. conceived the idea, supervised all research and revised the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary data
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kong, H., Zhu, M., Cui, F. et al. Quantitative assessment of short amplicons in FFPE-derived long-chain RNA. Sci Rep 4, 7246 (2014). https://doi.org/10.1038/srep07246
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07246
This article is cited by
-
Selection of endogenous control genes for normalising gene expression data derived from formalin-fixed paraffin-embedded tumour tissue
Scientific Reports (2020)
-
Quantitative analysis of noncoding RNA from paired fresh and formalin-fixed paraffin-embedded brain tissues
International Journal of Legal Medicine (2020)
-
Systematic evaluation of RNA quality, microarray data reliability and pathway analysis in fresh, fresh frozen and formalin-fixed paraffin-embedded tissue samples
Scientific Reports (2018)
-
Straightforward and sensitive RT-qPCR based gene expression analysis of FFPE samples
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.