Abstract
We present a microfluidic approach that allows simultaneous interrogation of RBC properties in physiological flow conditions at a single cell level. With this method, we studied healthy hemoglobin A (HbA) and homozygous sickle hemoglobin (HbS) containing RBCs using whole blood samples from twelve subjects. We report that HbS-containing RBCs are heterogeneous in terms of adhesion and deformability in flow.
Similar content being viewed by others
Introduction
Red Blood Cell (RBC) deformability, adhesion, hemolysis and alterations in flow are root pathophysiologic underpinnings of Sickle Cell Disease (SCD)1,2. Many facets of sickle RBCs have been investigated, including hemoglobin polymerization3,4,5, cellular deformability6,7,8,9, adhesion8,10,11,12,13,14, hemodynamic changes15,16 and clinical heterogeneity17,18. Specifically, sickle RBC adhesion (stickiness) and deformability have been identified as critical biophysical factors involved in vaso-occlusion14,19 and have been shown to correlate with disease severity20,21,22. These factors have been studied in isolation due to significant technological barriers faced in directly analyzing blood samples from diverse clinical phenotypes. Technical barriers included complicated custom designed systems, requirement for trained personnel and extensive sample manipulation. These challenges have posed significant limitations to the cogent integration of complex biophysical phenomena into our understanding of a heterogeneous multi-faceted clinical condition, such as SCD. Even though SCD is rooted in a single gene mutation, it is a clinically heterogeneous disease17,18. The aim of this study is to analyze sickle RBCs in terms of adhesion and deformability as a potential source of clinical heterogeneity at the cellular level. Here, we present a microfluidic approach that allows interrogation of RBC properties of single cells at physiological flow velocities. With this method, we studied healthy hemoglobin A (HbA) and homozygous sickle hemoglobin S (HbS) containing RBCs in physiological flow using blood samples from twelve subjects. We report for the first time that HbS-containing sickle RBCs are heterogeneous, displaying subpopulations within, in terms of adhesion and deformability in physiological flow conditions.
SCD is the first recognized molecular disease, identified more than sixty years ago23 and is caused by a mutation of the beta globin gene in hemoglobin. Replacement of a hydrophilic amino acid with a hydrophobic amino acid in the 6th position of the β-globin chain leads to polymerization of intracellular HbS and to the formation of stiff hemoglobin polymer structures within the cell1. This mutation afflicts millions of people worldwide and is associated with considerable morbidity and mortality24. The pathophysiology of SCD is a consequence of abnormal hemoglobin polymerization and its deleterious effects on RBC membrane, shape, density, adhesion and deformability1,11,14,19,21,25,26. Deoxygenated HbS alone contributes to the pathophysiologically important hemoglobin polymer1. A healthy RBC has a characteristic biconcave shape that allows cells to easily deform and pass through minuscule vessels and capillaries in the body27,28,29. However, sickled RBCs undergo a radical morphological transformation that leads to reduced deformability, increased stiffness and abnormal adhesion causing a blockage of blood vessels known as vaso-occlusion1,21,26. The consequences of this blood vessel blockade include painful crises, wide-spread organ damage and early mortality24,30.
Recent advances in micro and nano fabrication technologies have yielded microfluidic platforms that can rapidly probe single cell behavior under precisely controlled biological, biophysical and flow conditions, mimicking physiological systems at baseline and with disease31,32,33,34,35. Therefore, microfluidic tools have been adapted to the study of blood cells36,37,38, blood flow39,40,41 and blood-to-endothelium interactions42. Sickle RBC adherence to blood vessel walls has been shown to take place in post-capillary venules14,19,29. Fibronectin (FN) plays a role in HbS RBC adhesion to the endothelial wall12,43,44. FN is an adhesive glycoprotein that circulates in plasma and is present in the endothelial cell membrane44,45.
As a model of physiological flow, we designed the microfluidic channels with immobilized FN to mimic the size scale (50 µm) and bulk flow velocities (0.24-40 mm/sec) of post-capillary venules19,29,46. Studies using similar flow conditions, based on extrapolations from physiological states in SCD, have been reported in the literature10,12,16. However, no study analyzes HbS-containing RBC adhesion and deformability using whole blood at the micro-vasculature scale of ≤ 60 µm, as we do here. We show heterogeneity in HbS-containing RBC adhesion and deformability measured at a single cell level in SCD blood samples examined in microfluidic channels mimicking microvasculature.
Results
The microfluidic system was composed of a FN functionalized glass surface, Poly(methyl methacrylate) plastic top (encompassing micromachined inlets and outlets) and a 50 µm thick double sided adhesive film in the middle that defines the outlines and the thickness of the microchannels (Figure 1a). Multiple microchannels enable increased throughput and parallel processing of bloods samples in the microfluidic system on demand. The height of the microchannels were designed to be 50 µm, mimicking the post-capillary venule size and the width of the microchannels were designed to be 4 mm to provide a sufficiently large sampling area. The microfluidic system was placed on an Olympus IX83 inverted motorized microscope stage for high resolution live single cell image recording and analysis (Figure 1a).
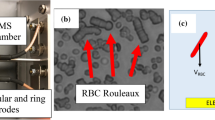
Overview of microfluidic system for evaluation of red blood cell (RBC) adhesion and deformability in physiological flow conditions.
(a) Microfluidic chip contains 50 µm high channels and can interrogate unprocessed whole blood. A microchip was placed on an automated microscope stage for high resolution image recording and analysis of single RBCs. (b) Depiction of flowing and adhered RBCs on Fibronectin functionalized surface in the presence of 3D laminar flow velocity profile in the microfluidic chip. (c) Heterogeneity in adhered sickle RBC morphology was observed in microfluidic channels. RBCs from the same blood sample with different levels of sickling effect is shown: (i) mildly affected RBC, (ii) moderately affected RBC and (iii) highly affected RBC (Scale bar represents 5 µm length).
In this study, FN was immobilized on microfluidic channel surfaces to mimic in part endothelial wall characteristics and whole blood was passed over it. Microfluidic design that we utilized allowed precise control of flow velocities similar to physiological conditions in microvasculature29, resulting in laminar flow conditions with straight and parallel streamlines on the surface (Figure 1b). This flow profile in microfluidic channels allowed interaction of flowing RBCs with FN immobilized surface and enabled attachment of those RBCs with increased adhesive properties (Figure 1b). We observed adhesion of a morphologically heterogeneous RBC population in blood samples from subjects with HbS (Figure 1c). On the other hand, as expected, we did not observe morphologically heterogeneous RBC populations from subjects with HbA (not shown). Adhered RBCs included mildly sickled (Figure 1c-i), moderately sickled (Figure 1c-ii) and highly sickled (Figure 1c-iii) cell morphologies within the same field of view from a single HbS-containing blood sample. Of note, nuclear staining performed in microchannels showed no significant white cell adhesion to FN immobilized surfaces in these experiments.
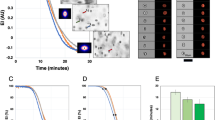
We analyzed the aspect ratio and deformability of single HbA- or HbS-containing RBCs in three conditions: (1) no flow, (2) flow and (3) at detachment instant (Figure 2a) (Table 1). Cell aspect ratio change (with respect to no flow condition) was used as a measure of deformability, in which a greater change in cell aspect ratio translates to more deformability and hence, less stiffness. Flow velocities were increased in a step-wise manner. Cell deformability for each RBC was assessed at a maximal flow velocity just prior to detachment in a time-lapse experiment. This approach allowed us to estimate maximum deformation of an adhered cell (Figure 2b&c). We observed two sub-groups of HbS-containing RBCs in terms of deformability: HbS deformable and HbS non-deformable (Figure 2c). Representative videos showing the behavior of each cell type in flow are provided in Supporting Information (Supplementary Videos 1a, b, & c).
Aspect ratio and deformability of healthy (HbA-containing) and sickle RBCs (HbS-containing deformable and non-deformable) are presented under no flow, flow and at detachment instant.
(a) Healthy and sickle RBCs at no flow, flow and at detachment conditions are shown. Flow velocities that result in detachment of RBCs in different experimental groups are noted, as bulk and as in the vicinity of the cells (in parentheses), below each column. White dashed lines denote the initial positions of RBCs at no flow condition. Flow direction is denoted with arrowhead. Scale bar represents 5 µm length. Aspect Ratio (AR) of cells in each frame are provided at the lower left of the images. Representative videos showing the behavior of each cell type in flow are provided in Supporting Information. (b) Cell AR of different RBC groups were measured at no flow, flow and at detachment. While HbA cells present a continuous decrease in AR from no flow to detachment, HbS non-deformable cells conserved their initial AR all the way through detachment. (c) Deformability (% cell AR change with respect to no flow condition) of healthy and sickle RBCs. Deformability of HbA and HbS deformable RBCs increased significantly from flow to detachment, whereas HbS non-deformable RBCs stayed the same. Data shown is based on 14 individual RBCs (3 HbA, 6 HbS deformable and 5 HbS non-deformable RBCs) from 12 different subjects (3 normal and 9 with SCD). The horizontal lines between individual groups represent statistically significant difference based on one way ANOVA test with Fisher's post-hoc test for multiple comparisons (n = 3–6, p<0.05). Error bars represent the standard error of the mean.
HbA-containing RBCs showed significantly greater cell aspect ratios (circularity) than HbS-containing RBCs at no-flow (p<0.05, one way ANOVA with Fisher's post-hoc test, Figure 2b) (Table 1). The aspect ratio of HbA RBCs significantly decreased (p<0.05) in the presence of flow, implying higher deformability. HbS-containing deformable RBCs presented a significant decrease (p<0.05) in aspect ratio only at the detachment instant, compared to no-flow conditions. Indeed, the aspect ratio of HbS non-deformable RBCs did not change in any flow condition (p>0.05). In addition, HbS non-deformable RBCs did not exhibit the characteristic biconcave morphology (Figure 2a).
The HbS deformable RBCs displayed a significantly greater cell aspect ratio at no-flow (Figure 2b) and significantly greater deformability at the detachment instant (Figure 2c), compared with HbS non-deformable RBCs (p<0.05, one way ANOVA with Fisher's post-hoc test) (Table 1). The deformability of HbA-containing RBCs was significantly greater than HbS-containing RBCs in all flow conditions (p<0.05). The deformability of both HbA and HbS-containing deformable RBCs was significantly different when measured during flow and when measured at the detachment instant (p<0.05). However, under these same conditions, HbS non-deformable RBCs did not display any significant difference in deformability during this interval (p>0.05). While adhered on a surface that mimics features of the normal blood stream in SCD, HbS-containing RBCs were heterogeneous in aspect ratio and in deformability.
We validated the accuracy of the predicted flow velocities in comparison with measured local flow velocities through particle image tracking around the adhered cells and using the non-adhered flowing cells as free flowing particles with which to measure the local flow velocities (Figure 3a). We observed that measured local flow velocities displayed significant correlation (Pearson correlation coefficient of 0.94, p<0.001, n = 9, R2 = 0.88) with the predicted flow velocities (Figure 3b). Hence, we utilized the mean theoretical flow velocities (Figure 3c) in determining the shear stress and drag force levels for detachment of HbA and HbS-containing RBCs (Figure 3d and 3e).
HbS-containing non-deformable RBCs detach at relatively higher flow velocity, shear stress and drag force compared with HbA and HbS-containing deformable RBCs.
(a) Sequential images of RBCs during flow were recorded using an inverted microscope in phase contrast mode and a charge coupled device (CCD) camera. We observed both adhered and free flowing RBCs in image analysis. (b) Shown is correlation between the locally measured flow velocity and calculated mean flow velocity within the microchannels (Pearson correlation coefficient of 0.94, p<0.001). HbS non-deformable RBCs detached at relatively higher flow velocity, shear stress and drag force compared to HbA or HbS deformable RBCs. (c) Relative positions of adhered RBCs in microchannels with respect to channel width (x- axis) and flow velocities (dotted lines), as bulk and as in the vicinity of the cells (in parentheses), at detachment instant. HbS non-deformable RBCs were significantly different than HbA and HbS deformable RBCs in terms of: (c) flow velocity, (d) shear stress and (e) drag force at detachment. There is no difference between HbA and HbS deformable RBCs in terms of shear stress and drag force levels (p>0.05). Data shown is based on 14 individual RBCs (3 HbA, 6 HbS deformable and 5 HbS non-deformable RBCs) from 12 different subjects (3 normal and 9 with SCD). The horizontal lines between individual groups represent statistically significant difference based on one way ANOVA parametric test with Fisher's post-hoc test for multiple comparisons (n = 3–6, p<0.05). Error bars represent the standard error of the mean.
We then determined the relative positions of adhered RBCs and flow velocities on the adhered RBCs (Figure 3c), shear stresses (Figure 3d) and drag forces (Figure 3e) acting on the cells at the detachment instant. Adhered RBCs that we analyzed were positioned in the mid-region of the microchannels where the flow velocity was uniform across the channel (Figure 3c). We observed that up to 4.3 times greater flow velocity (Figure 3c), 4.3 times greater shear stress (Figure 3d) and 4.1 times greater drag force (Figure 3e) was required to detach non-deformable HbS-containing RBCs compared to HbS-containing deformable RBCs (p<0.05, one way ANOVA with Fisher's post-hoc test). In contrast, HbA-containing RBCs and deformable HbS-containing RBCs did not differ in terms of flow velocity, shear stress and drag force at detachment (p>0.05). These results confirmed that HbS-containing RBCs are heterogeneous in terms of their adhesion strength to FN.
Next, we studied the motion of adhered RBCs at the flow initiation instant using consecutive high resolution microscopic images taken over 0.28 seconds, in order to analyze sites of adhesion in HbA (Figure 4a–d), HbS deformable (Figure 4e–h) and HbS non-deformable (Figure 4i–l) RBCs. We observed that HbA and HbS deformable cells displayed rotational motion in response to fluid flow direction, indicating a single adhesion pivot point (Figure 4d and h). To confirm these results, we utilized counter-direction flow on an HbS-containing deformable RBC (Supplementary Figure 1). We observed clockwise and counter-clockwise rotation of the cell around a pivot point in response to change in fluid flow direction, which further indicated a single adhesion site. On the other hand, HbS non-deformable cells did not display a rotational motion in response to fluid flow direction, implying multiple adhesion sites (Figure 4l). Importantly, these observations suggest that the higher adhesion strength of HbS non-deformable cells may be due to a greater number of adhesion sites. Representative videos showing the motion of each cell type in response to flow are provided in Supporting Information (Supplementary Videos 2a, 2b, & 2c, 3).
Determination of cell adhesion sites based on analysis of projected cell outlines at flow initiation for HbA- and HbS-containing RBCs.
Outlines of individual RBCs in three consecutive frames taken over 0.28 seconds were projected to reflect the motion of the cells in response to initiation of fluid flow, for: (a–d) HbA-containing RBCs, (e–h) HbS-containing deformable RBCs and (i–l) HbS-containing non-deformable RBCs (see representative videos, supporting information). HbA (a–d) and HbS deformable (e–h) cells displayed cell adhesion at a single site, which acts as a pivot (d, h), resulting in rotational motion (denoted with a white arrow in d and h) in response to fluid flow (horizontal yellow arrows). HbS non-deformable cells displayed no rotational motion and showed multiple sites of adhesion, as determined in projected cell outline analysis (l). Scale bars represent 5 µm length.
Discussion
Here, we present a microfluidic approach that allows interrogation of RBC properties of single cells at physiological flow velocities. Dramatic changes in morphologic, physical and hemodynamic properties of RBCs are caused by HbS polymerization1. Earlier studies have shown that RBCs in SCD patients are heterogeneous in density, morphology and function15,47,48,49,50. Results presented in this study show that HbS-containing RBCs are heterogeneous in: deformability, adhesion strength and number of adhesion sites. We defined heterogeneity in sickle RBCs as heterogeneity in terms of adhesion and deformability characteristics, albeit with relative homogeneity within each subset. In addition, deformable HbS-containing RBCs were less adherent, while non-deformable cells were more adherent, to FN. Our findings suggest a significant difference in deformability of normal and sickled RBCs which adhere through a single adhesion site. Number of adhesion sites may affect the way a RBC deforms in the presence of flow. Greater number of adhesion sites observed in a subpopulation of sickled RBCs may play a role in deformability of adhered cells in microvasculature. In a recent study, decreased deformability and RBC aggregation, measured using ektacytometry and laser backscatter of Percoll-separated sickle RBCs, were shown to correlate with hemolysis51. RBC adhesion and deformability are critical components of vaso-occlusion and hemolysis in subjects with SCD.
Most of the earlier studies investigating HbS RBC adhesion utilized open systems with static conditions or parallel plate flow chambers and employed blood preprocessing, separation of blood cell populations and washing of erythrocytes10,11,19. HbS polymerization critically depends on oxygen concentration of blood1,40. These earlier reports attempted to maintain physiological conditions by using environmental controls and additional manipulation. Furthermore, various reports indicated that plasma proteins play a role in abnormal adhesion of sickle RBCs to the endothelium8,52. Since these studies often exposed the blood cells to ambient conditions and to artificial buffers during preprocessing and washing, physiological oxygen and plasma protein levels may have been compromised. Therefore, disparate results are present in the literature in terms of adhesion characteristics of sickle RBCs on endothelium8,14. Heterogeneity of sickle RBC adhesion in relation to cellular deformability has been suggested8,14,21, but not shown in a physiological flow condition at a single cell level using unprocessed whole blood. The microfluidic system presented in this study allows fully closed, single step, preprocessing-free and direct analysis of blood samples after collection. This method minimizes the artifactual changes in oxygenation or plasma proteins in blood samples while they pass through the microchannels, that extensive pre-processing (e.g., washing, centrifugation and dilution) utilized in prior analyses may have conferred.
There are a number of processes, in addition to intracellular polymerization, that may have an impact on cellular adhesion. In this study, we emphasize the adhesive qualities of the cell membrane that are the sum of various intracellular processes, including hemoglobin content and composition (i.e., proportion of fetal hemoglobin)53,54,55, hemoglobin polymerization1,56, as well as the consequences of cytokine exposure14,15 and beta adrenergic stimulation57,58. Our results reflect the accumulated membrane damage over the life of a sickle RBC due to these factors.
It is known that adhesive interactions of RBCs are increased by the exposure of adhesive proteins on cell membranes due to RBC damage resulting from hemoglobin polymerization1,56. Our results showed that HbS non-deformable RBCs exhibit increased adhesion sites compared to HbS deformable RBCs. Polymerization grade is a potential factor that could directly affect this increase in adhesive properties of RBCs. Lower cell aspect ratio of the HbS non-deformable RBCs compared to HbS deformable RBCs suggests differences in Hb polymerization levels. Spectroscopic characterization methods, including Raman microspectroscopy59 and UV-vis absorption spectroscopy60, have been used in the analysis of these important intracellular phenomena. Any of these processes could be investigated and spectroscopic analyses could be fruitfully coupled to microfluidic approach in future studies.
The microfluidic chip that we introduced here has the potential to be utilized in a single use and disposable manner due its simple and cost effective design, which still allows detailed single cell level analysis of important biophysical phenomena relevant to sickle cell disease. The results presented in this manuscript establish the groundwork of a new approach and method that would benefit sickle cell disease patients with a new clinically adoptable microfluidic technology that would provide a functional blood test in a range of resource-limited settings.
In the future, the study of cellular heterogeneity within subjects in larger clinically diverse SCD populations may provide important insights into complex disease phenotypes in SCD. The microfluidic technology presented here has the potential to be widely available and routinely used in research laboratories, as well as in clinical settings. Further, these sensitive studies may allow us to better understand additional (more clinically subtle) SCD phenotypes, such as compound heterozygous HbSC or sickle cell trait (HbAS). Furthermore, FN has been used as a mimic for activated endothelium in SCD research12, although this method partially recapitulates endogenous vascular surfaces. Finally, abnormal RBC adhesion to microvascular surfaces has been implicated in other multi-system diseases, such as β-thalassemia, diabetes mellitus, hereditary spherocytosis, polycythemia vera and malaria51,61,62. Future studies focused on modifications to the microchannel design via functionalization by other endothelium associated proteins or endothelial cells may improve the fidelity of the system in replicating disease physiology.
In summary, we present a microscale approach with which to investigate important biophysical phenomena, in SCD and potentially in other disorders, namely the adhesion (stickiness) and deformation characteristics of single RBCs in microvasculature. We report heterogeneous HbS-containing RBC adhesion and deformability, which may have significant implications for understanding vaso-occlusion and clinical phenotypes in SCD.
Methods
Microfluidic Chip Fabrication
Poly(methyl methacrylate) (PMMA) top parts were prepared by cutting an inlet and outlet (0.61 mm in diameter and 26 m apart) using a VersaLASER system (Universal Laser Systems Inc., Scottsdale, AZ). Double sided adhesive (DSA) film (iTapestore, Scotch Plains, NJ) was cut to fit the size of the PMMA part and 28 × 4 mm channels. DSA was then attached to the PMMA top part to include an inlet and outlet between the outline of the DSA film. Gold Seal glass slide (adhesion coating: APTES, 3-Aminopropyl Triethoxysilane, Electron Microscopy Sciences (Hatfield, PA)) was then assembled with the PMMA–DSA structure to form a microfluidic channel. The microfluidic chips were designed to enable disposable use, which was possible through a cost-efficient strategy to employ widely available materials and fabrication methods (i.e., less than $5 material and fabrication cost per microfluidic chip). Disposable use of the microfluidic chips prevents potential cross-contamination between blood samples.
Surface Chemistry
N-g-Maleimidobutyryloxy succinimide ester (GMBS) stock solution was prepared by dissolving 25 mg of GMBS in 0.25 mL dimethyl sulfoxide (DMSO) and stock solution was diluted with ethanol to obtain 0.28% v/v GMBS working solution. FN was diluted with phosphate buffered saline (PBS, pH = 7.4, 1X) to create a (1:10) FN working solution. Bovine serum albumin (BSA) solution was prepared by dissolving 3 mg of lyophilized BSA in 1 mL PBS. The channels were washed with 30 µL of PBS and ethanol after assembly. Next, 20 µL of cross-linker agent GMBS working solution was injected into the channels twice and incubated for 15 min. at room temperature. Following GMBS incubation, channels are washed twice with 30 µL of ethanol and PBS. Next, 20 µL of Fibronectin (FN) solution (0.1 mg/mL) was injected into the channels and incubated for 1.5 h at room temperature. The surface was then passivated by injecting 30 µL of BSA solution and overnight incubation at 4°C. Before processing blood samples, channels were rinsed with PBS.
Blood Processing
Discarded de-identified patient blood samples were obtained from University Hospital's Hematology and Oncology Division under institutional review board (IRB) approval. Blood samples were collected into EDTA-containing purple cap Vacutainer tubes. Before using in experiments, blood samples were aliquoted into sealed microtubes and sealed syringes to minimize exposure to ambient air. Blood flow through the channels and following flow cytometry staining buffer (FCSB, 1X) flow steps were applied using New Era NE-300 syringe pump system (Farmingdale, NY). Blood samples were kept sealed in 1 mL vertically aligned disposable syringes before and during injection into microchannels. Next, blood was introduced into microchannels at 2.38 mm/s until the channel was filled with blood and then 15 µL of blood sample was injected at a flow rate of 0.24 mm/s. Next, the syringe was changed and 120 µL of FCSB at a flow rate of 0.83 mm/s was introduced into the channel to remove the non-adhered cells. Adhered RBCs in channels were visualized using an inverted microscope (Olympus IX83) and microscopy camera (EXi Blue EXI-BLU-R-F-M-14-C). During real time microscope imaging and high resolution video recording, controlled fluid flow with stepwise increments were applied until RBC detachment was observed from the microchannel surface. We performed all the experiments within 24 hours of blood collection from subjects at the clinic. A typical experiment takes less than an hour to complete, which includes the following main steps: (1) setting up the microfluidic system for blood processing takes approximately 5 minutes, (2) processing of blood samples in microchannels takes approximately 15 minutes, (3) washing out non-adherent cells from microchannels takes approximately 20 minutes and (4) live video recording of single adherent cells takes approximately 10 minutes.
Microfluidic channel visualization and image processing
An Olympus IX83 inverted motorized microscope with Olympus Cell Sense live-cell imaging and analysis software was used to obtain real-time microscopic recordings in this study. Olympus (20×/0.45 ph2 and 40×/0.75 ph3) long working distance objective lenses were utilized for phase contrast imaging of single RBCs in microchannels (Figure 1a). Videos were recorded at 7 frames per second and converted to single frame images for further processing and analysis. Cell dimensions were analyzed by using Adobe Photoshop software (San Jose, CA). Use of natural particle tracers, such as RBCs, is widely adapted in the literature for flow velocity measurements63,64.
Biophysical probing of individual cells in microchannels necessitates accurate control, measurement and estimation of flow velocities in close vicinity to the adhered cells. These measured and estimated values allowed us to make theoretical calculations of flow in microfluidic channels. When free flowing particles, such as cells, are imaged at relatively long camera exposure times, they appear as straight lines due to motion blur, which is also known as streaking65. The length of these streaks is proportional to the flowing particle velocity (Figure 3a). Local flow velocities (vp) for flowing cells were determined by dividing the streaking line length (Lp) minus average cell size (dp) to camera exposure duration (tp) using Equation (1).

Then, we analyzed the correlation between the measured local flow velocity and the predicted mean flow velocity (vm) determined by the volumetric flow rate (Q) and the dimensions of the microchannels (width: wc and height: hc) using Equation (2).

We analyzed the adhered RBCs in terms of biophysical properties in flow in the recorded images and we utilized the free flowing cells (appearing as a blurry line at 1.5 ms camera exposure time) for determining local flow velocities (Figure 3a). In addition, cellular adhesion, cellular deformation in flow and cellular detachment were analyzed in the recorded sequential images taken at 7 frames per second.
Data Analysis
Flow velocity on the adhered RBCs were calculated using equations of pressure-driven flow in a rectangular channel (Equation (3) and (4)).


where x,y and z are the principal axes, h and w are the channel height and width, η is the fluid viscosity (Table 2),  is the pressure change along the x axis and Q is the volumetric flow rate.
is the pressure change along the x axis and Q is the volumetric flow rate.
Drag force applied on the adhered RBCs was calculated using drag force equation66 (Equation (5)).

where Fd is the drag force, ρ is the fluid density (Table 2), Cd is the drag coefficient and A is the reference area. Cd was calculated as  , where Re is the Reynolds number, according to the circular disk parallel to flow assumption of cell at low Reynolds number flow66. A, reference area, is the projection of the RBC in the plane that is perpendicular to the fluid flow direction and calculated by using the typical RBC thickness and the measured RBC width at detachment (Table 2). A was calculated by using the typical RBC thickness and the measured RBC width at detachment (Table 2). Shear stress (τ) on the adhesion surface was calculated using Equation (6):
, where Re is the Reynolds number, according to the circular disk parallel to flow assumption of cell at low Reynolds number flow66. A, reference area, is the projection of the RBC in the plane that is perpendicular to the fluid flow direction and calculated by using the typical RBC thickness and the measured RBC width at detachment (Table 2). A was calculated by using the typical RBC thickness and the measured RBC width at detachment (Table 2). Shear stress (τ) on the adhesion surface was calculated using Equation (6):

Statistical analysis
Data obtained in this study were reported as mean ± standard error of the mean. Cell aspect ratio, aspect ratio change (deformability), flow rate, shear stress and drag forces were statistically assessed (Minitab 16 software, Minitab Inc., State College, PA) using Analysis of Variance (ANOVA) with Fisher's post hoc test for multiple comparisons (n = 3–6 blood samples per group). Statistical significance was set at 95% confidence level for all tests (p< 0.05). Error bars in figures represent the standard error of the mean.
References
Barabino, G. A., Platt, M. O. & Kaul, D. K. Sickle cell biomechanics. Annu Rev Biomed Eng 12, 345–367 (2010).
Alexy, T. et al. Rheologic behavior of sickle and normal red blood cell mixtures in sickle plasma: implications for transfusion therapy. Transfusion 46, 912–918 (2006).
Hebbel, R. P. Beyond hemoglobin polymerization: the red blood cell membrane and sickle disease pathophysiology. Blood 77, 214–237 (1991).
Ferrone, F. A. Polymerization and sickle cell disease: a molecular view. Microcirculation 11, 115–128 (2004).
Noguchi, C. T. & Schechter, A. N. Sickle hemoglobin polymerization in solution and in cells. Annu Rev Biophys Biophys Chem 14, 239–263 (1985).
Nash, G. B., Johnson, C. S. & Meiselman, H. J. Mechanical properties of oxygenated red blood cells in sickle cell (HbSS) disease. Blood 63, 73–82 (1984).
Brandao, M. M. et al. Optical tweezers for measuring red blood cell elasticity: application to the study of drug response in sickle cell disease. Eur J Haematol 70, 207–211 (2003).
Mohandas, N. & Evans, E. Sickle erythrocyte adherence to vascular endothelium - morphologic correlates and the requirement for divalent-cations and collagen-binding plasma-proteins. J Clin Inves 76, 1605–1612 (1985).
Byun, H. et al. Optical measurement of biomechanical properties of individual erythrocytes from a sickle cell patient. Acta Biomaterialia 8, 4130–4138 (2012).
Montes, R. A., Eckman, J. R., Hsu, L. L. & Wick, T. M. Sickle erythrocyte adherence to endothelium at low shear: role of shear stress in propagation of vaso-occlusion. Am J Hematol 70, 216–227 (2002).
Hillery, C. A., Du, M. C., Montgomery, R. R. & Scott, J. P. Increased adhesion of erythrocytes to components of the extracellular matrix: isolation and characterization of a red blood cell lipid that binds thrombospondin and laminin. Blood 87, 4879–4886 (1996).
Kasschau, M. R., Barabino, G. A., Bridges, K. R. & Golan, D. E. Adhesion of sickle neutrophils and erythrocytes to fibronectin. Blood 87, 771–780 (1996).
Bartolucci, P. et al. Erythrocyte density in sickle cell syndromes is associated with specific clinical manifestations and hemolysis. Blood 120, 3136–3141 (2012).
Kaul, D. K., Finnegan, E. & Barabino, G. A. Sickle Red Cell-Endothelium Interactions. Microcirculation 16, 97–111 (2009).
Kaul, D. K., Fabry, M. E., Windisch, P., Baez, S. & Nagel, R. L. Erythrocytes in sickle cell anemia are heterogeneous in their rheological and hemodynamic characteristics. J Clin Invest 72, 22–31 (1983).
Lei, H. & Karniadakis, G. E. Probing vasoocclusion phenomena in sickle cell anemia via mesoscopic simulations. Proc Natl Acad Sci U S A 110, 11326–11330 (2013).
Bunn, H. F. Pathogenesis and treatment of sickle cell disease. N Engl J Med 337, 762–769 (1997).
Ballas, S. K. & Mohandas, N. Sickle red cell microrheology and sickle blood rheology. Microcirculation 11, 209–225 (2004).
Kaul, D. K., Fabry, M. E. & Nagel, R. L. Microvascular sites and characteristics of sickle cell adhesion to vascular endothelium in shear flow conditions: pathophysiological implications. Proc Natl Acad Sci U S A 86, 3356–3360 (1989).
Hebbel, R. P. Adhesive interactions of sickle erythrocytes with endothelium. J Clin Inves 100, S83–S86 (1997).
Hebbel, R. P., Boogaerts, M. A., Eaton, J. W. & Steinberg, M. H. Erythrocyte adherence to endothelium in sickle-cell anemia. A possible determinant of disease severity. N Engl J Med 302, 992–995 (1980).
Ballas, S. K. & Smith, E. D. Red-blood-cell changes during the evolution of the sickle-cell painful crisis. Blood 79, 2154–2163 (1992).
Pauling, L., Itano, H. A., Singer, S. J. & Wells, I. C. Sickle cell anemia a molecular disease. Science 110, 543–548 (1949).
Platt, O. S. et al. Mortality in sickle-cell disease - life expectancy and risk-factors for early death. N Engl J Med 330, 1639–1644 (1994).
Stuart, M. J. & Nagel, R. L. Sickle-cell disease. Lancet 364, 1343–1360 (2004).
Hoover, R., Rubin, R., Wise, G. & Warren, R. Adhesion of normal and sickle erythrocytes to endothelial monolayer cultures. Blood 54, 872–876 (1979).
An, X. & Mohandas, N. Disorders of red cell membrane. Br J Haematol 141, 367–375 (2008).
Mohandas, N. & Gallagher, P. G. Red cell membrane: past, present and future. Blood 112, 3939–3948 (2008).
Lipowsky, H. H. Microvascular rheology and hemodynamics. Microcirculation 12, 5–15 (2005).
Hofrichter, J., Ross, P. D. & Eaton, W. A. Kinetics and mechanism of deoxyhemoglobin S gelation: a new approach to understanding sickle cell disease. Proc Natl Acad Sci U S A 71, 4864–4868 (1974).
Tasoglu, S. et al. Exhaustion of racing sperm in nature-mimicking microfluidic channels during sorting. Small 9, 3374–3384 (2013).
Rizvi, I. et al. Flow induces epithelial-mesenchymal transition, cellular heterogeneity and biomarker modulation in 3D ovarian cancer nodules. Proc Natl Acad Sci U S A 110, E1974–1983 (2013).
Tasoglu, S., Gurkan, U. A., Wang, S. & Demirci, U. Manipulating biological agents and cells in micro-scale volumes for applications in medicine. Chem Soc Rev 42, 5788–5808 (2013).
Gurkan, U. A. et al. Miniaturized lensless imaging systems for cell and microorganism visualization in point-of-care testing. Biotechnol J 6, 138–149 (2011).
Zhang, X. et al. Lensless imaging for simultaneous microfluidic sperm monitoring and sorting. Lab Chip 11, 2535–2540 (2011).
Moon, S. et al. Enumeration of CD4+ T-cells using a portable microchip count platform in Tanzanian HIV-infected patients. PLoS One 6, e21409 (2011).
Gurkan, U. A. et al. Controlled viable release of selectively captured label-free cells in microchannels. Lab Chip 11, 3979–3989 (2011).
Gurkan, U. A. et al. Smart interface materials integrated with microfluidics for on-demand local capture and release of cells. Adv Healthc Mater 1, 661–668 (2012).
Cohen, S. I. & Mahadevan, L. Hydrodynamics of hemostasis in sickle-cell disease. Phys Rev Lett 110, 138104 (2013).
Higgins, J. M., Eddington, D. T., Bhatia, S. N. & Mahadevan, L. Sickle cell vasoocclusion and rescue in a microfluidic device. Proc Natl Acad Sci U S A 104, 20496–20500 (2007).
Wood, D. K., Soriano, A., Mahadevan, L., Higgins, J. M. & Bhatia, S. N. A biophysical indicator of vaso-occlusive risk in sickle cell disease. Sci Transl Med 4, 123ra126 (2012).
Tsai, M. et al. In vitro modeling of the microvascular occlusion and thrombosis that occur in hematologic diseases using microfluidic technology. J Clin Invest 122, 408–418 (2012).
Kumar, A., Eckmam, J. R., Swerlick, R. A. & Wick, T. M. Phorbol ester stimulation increases sickle erythrocyte adherence to endothelium: a novel pathway involving alpha 4 beta 1 integrin receptors on sickle reticulocytes and fibronectin. Blood 88, 4348–4358 (1996).
Wick, T. M. & Eckman, J. R. Molecular basis of sickle cell-endothelial cell interactions. Curr Opin Hematol 3, 118–124 (1996).
Mosher, D. F. Physiology of fibronectin. Annu Rev Med 35, 561–575 (1984).
Turitto, V. T. Blood viscosity, mass transport and thrombogenesis. Prog Hemost Thromb 6, 139–177 (1982).
Fabry, M. E. & Nagel, R. L. The effect of deoxygenation on red cell density: significance for the pathophysiology of sickle cell anemia. Blood 60, 1370–1377 (1982).
Kaul, D. K., Nagel, R. L. & Baez, S. Pressure effects on the flow behavior of sickle (HbSS) red cells in isolated (ex-vivo) microvascular system. Microvasc Res 26, 170–181 (1983).
Lipowsky, H. H., Sheikh, N. U. & Katz, D. M. Intravital microscopy of capillary hemodynamics in sickle cell disease. J Clin Invest 80, 117–127 (1987).
Kaul, D. K., Fabry, M. E. & Nagel, R. L. Vaso-occlusion by sickle cells: evidence for selective trapping of dense red cells. Blood 68, 1162–1166 (1986).
Connes, P. et al. Haemolysis and abnormal haemorheology in sickle cell anaemia. Br J Haematol 165, 564–572 (2014).
Yang, Y., Koo, S., Lin, C. S. & Neu, B. Specific Binding of Red Blood Cells to Endothelial Cells Is Regulated by Nonadsorbing Macromolecules. J Biol Chem 285, 40489–40495 (2010).
Fathallah, H. & Atweh, G. F. Induction of fetal hemoglobin in the treatment of sickle cell disease. Hematology Am Soc Hematol Educ Program, 58–62 (2006).
Saunthararajah, Y. et al. Effects of 5-aza-2'-deoxycytidine on fetal hemoglobin levels, red cell adhesion and hematopoietic differentiation in patients with sickle cell disease. Blood 102, 3865–3870 (2003).
Setty, B. N., Kulkarni, S., Dampier, C. D. & Stuart, M. J. Fetal hemoglobin in sickle cell anemia: relationship to erythrocyte adhesion markers and adhesion. Blood 97, 2568–2573 (2001).
Steinberg, M. H. Management of sickle cell disease. N Engl J Med 340, 1021–1030 (1999).
Zennadi, R. et al. Erythrocyte plasma membrane-bound ERK1/2 activation promotes ICAM-4-mediated sickle red cell adhesion to endothelium. Blood 119, 1217–1227 (2012).
Hines, P. C. et al. Novel epinephrine and cyclic AMP-mediated activation of BCAM/Lu-dependent sickle (SS) RBC adhesion. Blood 101, 3281–3287 (2003).
Wood, B. R., Hammer, L., Davis, L. & McNaughton, D. Raman microspectroscopy and imaging provides insights into heme aggregation and denaturation within human erythrocytes. J Biomed Opt 10, 14005 (2005).
Wojdyla, M., Raj, S. & Petrov, D. Absorption spectroscopy of single red blood cells in the presence of mechanical deformations induced by optical traps. J Biomed Opt 17, 97006–97001 (2012).
Colin, Y., Le Van Kim, C. & El Nemer, W. Red cell adhesion in human diseases. Curr Opin Hematol 21, 186–192 (2014).
Cooke, B. M., Morrisjones, S., Greenwood, B. M. & Nash, G. B. Adhesion of parasitized red-blood-cells to cultured endothelial-cells - a flow-based study of isolates from gambian children with falciparum-malaria. Parasitology 107, 359–368 (1993).
Hove, J. R. et al. Intracardiac fluid forces are an essential epigenetic factor for embryonic cardiogenesis. Nature 421, 172–177 (2003).
Lee, J. Y., Ji, H. S. & Lee, S. J. Micro-PIV measurements of blood flow in extraembryonic blood vessels of chicken embryos. Physiol Meas 28, 1149–1162 (2007).
Goda, K. et al. High-throughput single-microparticle imaging flow analyzer. Proc Natl Acad Sci U S A 109, 11630–11635 (2012).
Munson, B. R., Rothmayer, A. P., Okiishi, T. H. & W, H. W. Fundamentals of Fluid Mechanics. 7 edn, 580 (John Wiley & Sons Canada, 2012).
Jay, A. W. L. Geometry of Human Erythrocyte .1. Effect of Albumin on Cell Geometry. Biophys J 15, 205–222 (1975).
Oh, J. et al. Microfluidic fabrication of cell adhesive chitosan microtubes. Biomed Microdevices 15, 465–472 (2013).
Acknowledgements
This work was supported by Innovations in Clinical Research Award, Grant # 2013126 from the Doris Duke Charitable Foundation. Authors acknowledge Ismail Sayin for his help in experimental protocol development. We acknowledge with gratitude the contributions of patients and clinicians at Seidman Cancer Center (University Hospitals, Cleveland).
Author information
Authors and Affiliations
Contributions
Y.A., J.A.L. and U.A.G. developed the idea, Y.A., J.A.L. and U.A.G. designed the experiments, Y.A. performed the experiments, Y.A. and U.A.G. analyzed the results, Y.A. and U.A.G. prepared the figures and the Supporting Information, Y.A., J.A.L. and U.A.G. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Video S1a
Supplementary Information
Video S1b
Supplementary Information
Video S1c
Supplementary Information
Video S2a
Supplementary Information
Video S2b
Supplementary Information
Video S2c
Supplementary Information
Video S3
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Alapan, Y., Little, J. & Gurkan, U. Heterogeneous Red Blood Cell Adhesion and Deformability in Sickle Cell Disease. Sci Rep 4, 7173 (2014). https://doi.org/10.1038/srep07173
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07173
This article is cited by
-
Partitioning of dense RBC suspensions in single microfluidic bifurcations: role of cell deformability and bifurcation angle
Scientific Reports (2024)
-
Red blood cell distribution width is a risk factor for multiple organ dysfunction syndrome in elderly patients with infection: a case control study
Aging Clinical and Experimental Research (2023)
-
Simultaneous Determination of Human Erythrocyte Deformability and Adhesion Energy: A Novel Approach Using a Microfluidic Chamber and the “Glass Effect”
Cell Biochemistry and Biophysics (2021)
-
Tethering, evagination, and vesiculation via cell–cell interactions in microvascular flow
Biomechanics and Modeling in Mechanobiology (2021)
-
Characterization of red blood cell microcirculatory parameters using a bioimpedance microfluidic device
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.