Abstract
Drug co-prescription (or drug combination) is a therapeutic strategy widely used as it may improve efficacy and reduce side-effect (SE). Since it is impractical to screen all possible drug combinations for every indication, computational methods have been developed to predict new combinations. In this study, we describe a novel approach that utilizes clinical SEs from post-marketing surveillance and the drug label to predict 1,508 novel drug-drug combinations. It outperforms other prediction methods, achieving an AUC of 0.92 compared to an AUC of 0.69 in a previous method, on a much larger drug combination set (245 drug combinations in our dataset compared to 75 in previous work.). We further found from the feature selection that three FDA black-box warned serious SEs, namely pneumonia, haemorrhage rectum and retinal bleeding, contributed mostly to the predictions and a model only using these three SEs can achieve an average area under curve (AUC) at 0.80 and accuracy at 0.91, potentially with its simplicity being recognized as a practical rule-of-three in drug co-prescription or making fixed-dose drug combination. We also demonstrate this performance is less likely to be influenced by confounding factors such as biased disease indications or chemical structures.
Similar content being viewed by others
Introduction
The use of multiple drugs with different mechanisms or modes of action may treat the disease more effectively1,2,3. The traditional “one drug – one target – one disease” approach has been successfully used to develop drugs. However such “magic bullet” sometimes shows limited efficacy, especially for complex diseases4, which is often due to factors such as network robustness5, redundancy6, compensatory and neutralizing actions7. Polypharmacology, which focuses on multi-target drugs, has the potential to address those limitations8. High-throughput screening has been previously used to identify possible drug combinations9; however, it is impractical to screen all possible drug combinations for every indication. Therefore, computational methods10,11,12,13 have been developed to predict new drug combinations. For example, network biology was introduced to investigate drug combinations by studying the molecular networks or pathways affected by the drugs14, yet the incompleteness of molecular networks limits the practical use of such approaches for prediction of novel drug combinations.
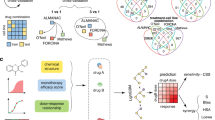
Clinical phenotypic information has not been adequately investigated for its power in predicting drug combinations. The advantages of leveraging clinical phenotypic information includes better translational power when comparing with animal models15 since it mimics a phenotypic screening of the drug effects, both therapeutic effect16,17 and toxic effect18,19,20, on humans. In this paper we leverage observed side-effects (SEs) reported in clinical findings to predict novel safe and efficacious drug co-prescriptions. The outline of this study is demonstrated in Figure 1.
The outline of this study.
Firstly, we built an integrated drug combination database by manually curating 349 fix-dose combinations approved by FDA, based on which a machine learning model has been constructed using side-effects as the feature. During the feature selection step, we found that three features contributed mostly to the model, we therefore developed a simple classification criterion called “Rule of Three”, with the aim of helping clinicians co-prescribe drugs.
Results
Construction of the drug combinations and side-effects data set
We constructed a comprehensive drug combination database (Figure 2) which contains 349 approved pairwise drug-drug co-prescriptions/combinations (DDC) from three different sources: drug combination database DCDB21, FDA approved drug combinations compiled by a recent paper13 and manual literature curation of the FDA approved or registered DDCs. The database is much larger than the DDC database in a previous publication (Figure 2). To resolve different naming issues in different data sources, DDCs were represented by their two components whose names were mapped to STITCH ID22 for comparison.
To annotate drugs with their SE features, we extracted SE information from drug labels using SIDER23 and OFFSIDES18. SIDER derives SEs from drug labels and OFFSIDES mines SEs from post-marketing surveillance system FAERS (i.e. FDA Adverse Event Reporting System). Of the 349 approved DDCs, 239 DDCs can be annotated with SEs for both components, which correspond to 245 individual drugs and 7,888 SEs. The drug frequency and SE frequency distribution are shown in Supplementary Fig. S1 and Fig. S2. As a comparison, previous work13 used 181 pairwise DDCs, out of which only 75 contains both SEs and indication annotation due to the limited data sources for DDCs, SEs and indications. Therefore the coverage of our database, available in the Supplementary Materials, is much more comprehensive.
We also constructed a negative training set consisting of unsafe drug pairs for training our DDC prediction model. We defined the unsafe co-prescriptions as those causing unexpected SEs as tracked in TWOSIDES18, a database of reported SEs only caused by the combination of marketed drugs rather than by any single drugs from FAERS. We generated all the possible pairs of the drugs that overlapped with those pairs in TWOSIDES. A resultant set of 2291 unsafe drug pairs (8% of all the possible drug combinations for the 245 drugs) were identified and used as the negative training set for training the DDC prediction model.
Evaluation of the power of predicting DDCs based on the side effects features
We used 239 marketed DDCs as positive set along with 2291 unsafe drug pairs as negative set, in total 2530 drug pairs and 245 distinct drugs. Each SE of a drug is called a feature and a drug pair can be represented as a vector of SE features with value of 0, 1 and 2 depending whether zero, one or both drugs have such SE. We applied logistic regression model with 10-fold cross validation to evaluate the performance. We measured the model performance with both AUC (area under the ROC curve) and AUPRC (area under the precision-recall curve). We repeated the cross-validation experiment 100 times with random seeds and computed the mean and the standard deviation of AUC and AUPRC over the 100 repetitions. In the experiment, logistic regression model achieved an AUC of 0.92 ± 0.01 and AUPRC of 0.86 ± 0.01 (Figure 3), outperforming existing DDC prediction model13 (AUC of 0.69). To test the impact of structural similarity on prediction results, we mimicked the method in Gottlieb's work24 by removing the drug pairs with Tanimoto similarity coefficient larger than 0.50. We re-run the logistic regression 10-fold cross-validation experiment 100 times and still achieved an AUC of 0.92 ± 0.01 (Supplementary Fig.S3) and AUPRC of 0.86 ± 0.01 (Supplementary Fig.S3), which is similar to previous results to two decimal places. Since the number of unsafe drug pairs (i.e. 2291) is larger than that of safe DDCs (i.e. 239), we randomly selected 239 unsafe drugs pairs so that the positive set and negative set were balanced and then ran the logistic regression model. The process was repeated 100 times and the reported AUC was 0.91 ± 0.01. This result shows that our model is less likely biased by the unbalanced positive set and negative set. The Supplementary Result also shows our model is less likely biased by the indication confounders.
Since the datasets are made of drug pairs, it is possible that some drugs occur in both the training and test data set. To further characterize their effect on our predictive model, we performed a hold-drug-out validation. Of the 245 drugs, we randomly chose 60 drugs for the test data set (i.e., about 25%) and 185 drugs for the training set (i.e., about 75%). From the 2530 drug pairs, we only picked the drug pairs with both drugs present in the training set to train the model. We only picked the drug pairs with both drugs present in the test data sets to test the model performance. The hold-drug-out validation experiment was carried out 100 times using random partitions and computing the mean and the standard deviation of AUC and AUPRC over these 100 repetitions. The final model achieved an AUC of 0.87 ± 0.03 (Supplementary Fig.S4) and AUPRC of 0.76 ± 0.07 (Supplementary Fig.S4).
Develop a ‘Rule of Three’ criterion with feature selections
After evaluation of the power of predicting DDCs based on the SEs features, we next aimed at constructing a simple and effective rule that can help doctors co-prescribe drugs. We choose to use the decision tree model25 to build the classifier since it is straightforward and easy to be visualized and explained. Here Figure 4 shows how AUC would change with using the top N SE features ranked by the information gain in the decision tree model. We found that the AUC increases significantly when N increases from 1 to 3 while the AUC only increases marginally when N increases from 3 to 10. Using the top three SEs as features strikes a balance between the model performance and the complexity of the model. The top three SEs are, Pneumonia, haemorrhage rectum and retinal bleeding, which happen to be the “black-box” warned adverse events featured in FDA approved different drug labels. With these three SEs features, the decision tree model (Figure 5) could achieve an AUC of 0.80 and an accuracy of 0.91. We examined the effects of different machine learning methods on the prediction performance. For the prediction performance evaluation with the three SEs as features, decision tree model gives an AUC of 0.80, Naive Bayes with an AUC of 0.84 and Logistic Regression with an AUC of 0.84. The robust performance across different machine learning methods confirms our conclusion is not biased towards a particular method.
The details of the decision tree model using the top three features to decide the candidate drug combination.
0, 1 and 2 indicates the number of drugs in the drug pair with such side effect. Pie charts indicate the percentage of correctly classified (green) and in-correctly classified (red) instances at each leaf. Safe represents the approved drug combinations while unsafe represents drug pairs from negative set.
To predict the novel drug combinations, we used all the possible pair-wise drug combinations of 239 marketed DDCs, excluding both positive and negative set. In total 27,360 drug pairs were used as prediction set. Based on the trained decision tree model with the above three SEs features, we made the prediction of the novel DDCs by only choosing pairs with predicted probability above 0.99 and co-occurred in at least 10 publications of clinical trial publications in PubMed. As a result, 1508 drug pairs were identified compared to a much higher number of 6,616 if one would apply literature co-occurrence to propose any drug combination. These 1508 drug pairs formed a well-connected network and the degree distribution is approximately a Power-law Distribution26 (Figure 6A). We further identified a condensed sub-network, highly interconnected regions in the network (Figure 6B) with Cytoscape27 and its plugin MCODE28 The connections between the hub drugs include familiar drug combinations with similar mechanism of actions like hydrocortisone and dexamethasone (immunosuppressants)29, morphine and tramadol (pain relievers)30 and could be a good starting point for further experimental validation of these novel drug combinations. Among these 1508 predicted candidate DDCs, 31 pairs contain at least one clinical trial record cording to clinicaltrial.gov as pairs, including 6 pairs in phase I, 7 in phase II, 12 in phase III and 4 in phase IV (Supplementary Fig.S5). In contrast, for the 615 drug pairs with probability less than 0.01, only 11 are supported by at least 10 publications and the network looks sparse (Figure 6C) compared to the network formed by drug pairs with predicted probability above 0.99 (p-value of 4.19 × 10−7 of Fisher's exact test). When searching the 615 drug pairs against clinicaltrial.gov, only 2 of them have clinical trial records. The different degree distributions (Supplementary Fig.S6) between network of predicted DDCs with high confidence level and predicted DDCs with low confidence show the totally different network behaviors. The predicted DDCs network with high confidence level fits the distribution of the scale-free network, similar to commonly observed biological networks31. The DDCs network with low confidence level is similar to random networks.
Network analysis of the predicted DDCs.
(a) the power-law degree distribution of the predicted drug combination network. (b) The sub-network cluster with prediction probability above 0.99 and support from at least 10 clinical type publications. (c) A network view of the 11 drug pairs with prediction probability less than 0.01 and support from at least 10 clinical type publications.
Case study
Below we selected one of the top predicted combinations as the case study.
Formoterol/Fluticasone
Formoterol, a long-acting beta-adrenoceptor agonist, exerts bronchodilatation effect and is used in the management of asthma and chronic obstructive pulmonary disease (COPD). It's already been tested and used in combination with corticosteroids, such as budesonide, to treat or prevent asthma attack and/or respiratory tract inflammation. Fluticasone, another potent glucocorticoid, has been shown to have superior or similar efficacy in improving pulmonary functions in asthma patients32,33. The predicted Formoterol/Fluticasone combination can be adopted as a new and alternative option in the management of asthma or COPD along the same combination strategy as Formoterol/Budesonide.
Discussion
In this study, we tried to address the DDC issue mainly through evaluating the safety aspect, which is critical for co-prescribing drugs or developing fix-dose combinations34,35. Several methods have been developed to predict drug-drug interactions (DDIs) based on text mining36,37, network modeling38, high-throughput screening9 and other data integrative approaches13. Our approach explored the possibility of predicting new drug pairs by representing drug combinations with their clinical SEs. It is based on the hypothesis that the drugs that can be co-prescribed usually do not have or share the serious adverse drug reactions. We tested this hypothesis in different machine learning models and identified three FDA blacklisted SEs, Pneumonia, haemorrhage rectum and retinal bleeding, as the top features contributing to the model performance. A “Rule of Three” criterion was thus developed: a drug combination with any of these three SEs has significantly high likelihood to be unsafe. We further demonstrated the robustness of such classification power based on the conclusion that the accuracy of our model is less likely to be introduced by confounding factors such as biased disease indications or chemical structures. This method provides an approach to identify novel drug combinations from clinical SEs, which should be less of a translational issue compared to animal model.
We applied this approach to identify 1,508 candidate drug combinations. Instead of testing all 27,360 combinations, a researcher looking to find novel DDCs will only test 1,508 combinations, saving an enormous amount of resources. If a researcher applies pure literature co-occurrence based filtering using “more than 10 PubMed co-occurrence” criterion, he/she still needs to test 6,616 combinations instead. On the other hand, using co-occurrence number in literature only may not be a good filter. For example, in our negative training set (unsafe drug combinations), 308 of them could have passed the “10 or more times” filter, generating unsafe predictions (false positives).
We tend to believe that our method could achieve a much better performance than a previous DDC prediction study13. To test if this improvement is only due to the better coverage of the known DDC, we re-ran our model using the dataset from their study13. The model achieved an AUC of 0.86 ± 0.01, which is much better than their best results (AUC: 0.69). However, this AUC is lower than the AUC (0.92 ± 0.01) we achieved based on the larger DDC dataset, which means the coverage of the dataset may also contribute to the model performance. We discussed the differences between our methods and previous work13 in more details in the supplemental materials (Supplementary Result 2).
To better understand the rational of using the SEs to predict DDC, we classify the SEs into two categories: efficacy-related SEs (blue) and undesired (green) as shown in Figure 7. Certain SEs contribute to the therapeutic effects of drug12 and are therefore called “efficacy-related SEs”. For example, most anti-diabetic drugs cause hypoglycemia and a decrease in blood glucose is one of the desired therapeutic effects of such drugs. An ideal drug pair is to combine drugs that can share the same SEs for the desired therapeutic effect but at the same time minimizing the number of undesirable SEs shared between them as possible. For example, if we take half dose of each drug component to make a DDC, the ideal situation would be is to reduce the potency of the undesired SEs by half while keeping the potency of the desired SEs at the current levels. In reality SEs may not combine linearly and thus this ideal situation needs to be further thoroughly tested. From the approved drug combinations, we could find many cases that come close to this ideal DDC model. For instance, the FDA approved hypertension drug Minizide is the fix-dose combination of the prazosin and polythiazide. The SEs they share, such as hypotension and impotence, are found to be associated with the therapeutic effect of the hypertension drugs12. None of the black-box warned SEs are shared and the other SEs they share are mostly like the dizziness, headache, nausea, vomiting etc., which are less likely to be associated with the serious adverse drug reaction.
An ideal model of making the DDC.
We simply halve the dose of each component to make a DDC. The potency of the efficacy-related SEs (blue) is unchanged after the two drugs are combined, but the potency of the undesired SEs (green) is decreased to the half each. Ideally, no black-box warned SEs should be shared. Such model is under the ideal assumption that the SEs are linearly addable, but the real situation may be more complicated.
We describe in this study the use of SEs data to predict new drug-drug combinations. Developing such combinations will be beneficial in three areas: (i) improving the safety profiles of drug co-prescriptions in clinic; (ii) assessing potentially hazardous drug combinations in early stage of the fix-dose combination discovery in pharmaceutical industry; and (iii) potentially reducing pill burden or bringing economics of combining the right drug pairs, e.g., one expensive drug along with a cheaper one. While our predictions were validated in-silico, they should be further tested experimentally to establish their clinical implications.
Methods
Side effect datasets
SIDER is a SE database containing information on marketed medicines and their recorded adverse drug reactions. The information is extracted from public documents and package inserts23. In this study, we downloaded the entire database from http://sideeffects.embl.de/. Besides relying on drug label as sources for drug SEs, we also checked FAERS, a database that contains information on adverse event submitted to FDA and is designed to support the FDA's post-marketing safety surveillance program for drug and therapeutic biologic products. OFFSIDES is such a SE database by mining FAERS system while controlling those confounding factors such as concomitant medications, patient demographics and patient medical histories and so on. OFFSIDES contains 1332 drugs and 10097 SEs. 438 drugs and 2322 SEs are shared between SIDER and OFFSIDE. In our final integrated SE database, drugs are represented with STITCH ID while SEs represented with MedDRA terms so that they could be integrated across databases. We tested the model performance with SEs from SIDER alone, OFFSIDES alone or OFFISDES and SIDER combined. The most predictive model was the one that included information from both OFFSIDES and SIDER(AUC:0.92), followed by OFFSIDES alone(AUC:0.77), then SIDER alone(AUC:0.69), which is consistent with previous findings18.
The TWOSIDES database identifies 59,220 pairs of drugs with 1,301 adverse events by carefully matching groups of patients in the post-marketing surveillance system FAERS. It provides a reliable and comprehensive database of SEs for drug pairs. It is thus used to identify the features enriched in approved DDCs compared to random drug pairs. In contrast, when doing the DDC prediction, we only used the SE for single drugs from drug label and OFFSIDES since it is logical to only have single drugs' SE data before such pair has come into being.
Drug combination datasets
The Drug Combination Database (DCDB) is a database collecting and organizing known examples of drug combinations. The current version contains 145 drug combinations. Zhao et al (2011)13 also lists 178 drug combinations, mainly collected from FDA orange book. We also curate 236 FDA approved or registered drugs from literature. After mapping them to STITCH ID and annotating them with SEs, we get a comprehensive list of 239 drug combinations to build the prediction model (Supplementary Table S1). We used eulerAPE (http://www.eulerdiagrams.org/eulerAPE/) to draw the area-proportional Venn diagrams for these three data sources.
Drug target, SMILES string and ATC code
DrugBank (http://www.drugbank.ca) is a unique bioinformatics and chemoinformatics resource that combines detailed drug data with comprehensive drug target information. Current version contains 6711 drugs and 4081 targets. We downloaded the full database in xml format and parsed out the drug target pairs, drug SMILES string and drug ATC pairs.
Making safe drug combination or co-prescriptions
First, we made sure what drugs can be safely put together. We hypothesize that the drugs that can be put together usually do not have overlap in some serious adverse drug reactions (ADR), but might share some SEs that contribute to the therapeutic effect16,17. Here we came up with a practical black list consisting of three SEs for clinicians to decide the safe drug pairs with high accuracy.
Machine learning models
We used logistic regression model to evaluate the power of predicting DDCs based on the SEs features. Our implementation was by Python 2.7 and the codes of logistic regression classifier are available in the Scikit-Learn package39. We considered both penalty and inverse of regularization strength (i.e., parameter C - the smaller values specify stronger regularization) parameters for the logistic regression model. The penalty can be L1 or L2 regularization and parameter C can be chosen from 0.001, 0.01, 0.1, 1, 10, 100, or 1000. In our experiment, we tuned the model parameters based on 10-fold cross validation. Finally, the logistic regression model we used in the experiments was L1-regularized logistic regression with C = 10.We used decision tree for feature selection and the development of the ‘Rule of Three’ criterion. The implementation was by J48 decision tree learner in Weka (http://www.cs.waikato.ac.nz/ml/weka/) with all the default settings.
PubMed and clinical trial validation
To validate whether the predicted drug pairs have clinical literature supports, we used the search API provided by NCBI to count the co-occurrence of the drug components for each proposed DDCs. The query term we used are ‘drug name1 AND drug name2 AND (Clinical Trial[ptyp] OR Clinical Trial, Phase I[ptyp] OR Clinical Trial, Phase II[ptyp] OR Clinical Trial, Phase III[ptyp] OR Clinical Trial, Phase IV[ptyp])’. We also checked clinicaltrial.gov to see whether predicted drug pairs are co-mentioned in the same registered clinical trials.
Structure similarity measurement
We used ChemmineR to calculate the Tanimoto similarity coefficient between drug pairs based on their SMILES string. The drug pairs with Tanimoto similarity coefficient larger than 0.5 were treated as structure similar drugs. They were removed before we re-ran the prediction model to check whether the model performance was biased by drugs' chemical similarities.
Chemical fingerprints
We used rcdk, an R interface for CDK, to calculate two different fingerprints, the 1024 hashed fingerprints from CDK and 166 MACCS keys described by MDL, for each of the drug in the drug combination.
References
Kitano, H. A robustness-based approach to systems-oriented drug design. Nat Rev Drug Discov 6, 202–210 (2007).
Zimmermann, G. R., Lehar, J. & Keith, C. T. Multi-target therapeutics: when the whole is greater than the sum of the parts. Drug Discov Today 12, 34–42 (2007).
Chou, T. C. Theoretical basis, experimental design and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 58, 621–681 (2006).
Yildirim, M. A., Goh, K. I., Cusick, M. E., Barabasi, A. L. & Vidal, M. Drug-target network. Nat. Biotechnol. 25, 1119–1126 (2007).
Smalley, K. S. et al. Multiple signaling pathways must be targeted to overcome drug resistance in cell lines derived from melanoma metastases. Mol. Cancer Ther. 5, 1136–1144 (2006).
Pilpel, Y., Sudarsanam, P. & Church, G. M. Identifying regulatory networks by combinatorial analysis of promoter elements. Nat. Genet. 29, 153–159 (2001).
Sergina, N. V. et al. Escape from HER-family tyrosine kinase inhibitor therapy by the kinase-inactive HER3. Nature 445, 437–441 (2007).
Hopkins, A. L. Drug discovery: Predicting promiscuity. Nature 462, 167–168 (2009).
Borisy, A. A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. U. S. A. 100, 7977–7982 (2003).
Wong, P. K. et al. Closed-loop control of cellular functions using combinatory drugs guided by a stochastic search algorithm. Proc. Natl. Acad. Sci. U. S. A. 105, 5105–5110 (2008).
Chou, T. C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010).
Yang, L. et al. Identifying unexpected therapeutic targets via chemical-protein interactome. PloS one 5, e9568 (2010).
Zhao, X. M. et al. Prediction of drug combinations by integrating molecular and pharmacological data. PLoS Comput. Biol. 7, e1002323 (2011).
Wang, Y. Y., Xu, K. J., Song, J. & Zhao, X. M. Exploring drug combinations in genetic interaction network. BMC Bioinformatics 13 Suppl 7S7 (2012).
Duran-Frigola, M. & Aloy, P. Recycling side-effects into clinical markers for drug repositioning. Genome Med. 4, 3 (2012).
Yang, L. & Agarwal, P. Systematic drug repositioning based on clinical side-effects. PloS one 6, e28025 (2011).
Campillos, M., Kuhn, M., Gavin, A. C., Jensen, L. J. & Bork, P. Drug target identification using side-effect similarity. Science 321, 263–266 (2008).
Tatonetti, N. P., Ye, P. P., Daneshjou, R. & Altman, R. B. Data-driven prediction of drug effects and interactions. Sci Transl Med. 4, 125ra131 (2012).
Liu, Z. et al. Translating clinical findings into knowledge in drug safety evaluation--drug induced liver injury prediction system (DILIps). PLoS Comput. Biol. 7, e1002310 (2011).
Lounkine, E. et al. Large-scale prediction and testing of drug activity on side-effect targets. Nature 486, 361–367 (2012).
Liu, Y., Hu, B., Fu, C. & Chen, X. DCDB: drug combination database. Bioinformatics 26, 587–588 (2010).
Kuhn, M. et al. STITCH 3: zooming in on protein-chemical interactions. Nucleic Acids Res. 40, D876–880 (2012).
Kuhn, M., Campillos, M., Letunic, I., Jensen, L. J. & Bork, P. A side effect resource to capture phenotypic effects of drugs. Mol. Syst. Biol. 6, 343 (2010).
Gottlieb, A., Stein, G. Y., Oron, Y., Ruppin, E. & Sharan, R. INDI: a computational framework for inferring drug interactions and their associated recommendations. Mol. Syst. Biol. 8, 592 (2012).
Quinlan, J. R. Decision Trees and Decision-Making. IEEE T Syst Man Cyb 20, 339–346 (1990).
Xu, K. J., Song, J. & Zhao, X. M. The drug cocktail network. BMC Syst Biol 6 Suppl 1S5 (2012).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P. L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432 (2011).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4, 2 (2003).
Cevc, G. & Blume, G. Hydrocortisone and dexamethasone in very deformable drug carriers have increased biological potency, prolonged effect and reduced therapeutic dosage. Biochim. Biophys. Acta 1663, 61–73 (2004).
Webb, A. R., Leong, S., Myles, P. S. & Burn, S. J. The addition of a tramadol infusion to morphine patient-controlled analgesia after abdominal surgery: a double-blinded, placebo-controlled randomized trial. Anesth. Analg. 95, 1713–1718 (2002).
Barabasi, A. L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nat Rev Genet 5, 101–113 (2004).
Derom, E., Van Schoor, J., Verhaeghe, W., Vincken, W. & Pauwels, R. Systemic effects of inhaled fluticasone propionate and budesonide in adult patients with asthma. Am. J. Respir. Crit. Care Med. 160, 157–161 (1999).
Adams, N., Lasserson, T. J., Cates, C. J. & Jones, P. W. Fluticasone versus beclomethasone or budesonide for chronic asthma in adults and children. Cochrane Database Syst. Rev. 4, CD002310 (2007).
Pirmohamed, M. Drug-drug interactions and adverse drug reactions: separating the wheat from the chaff. Wien Klin Wochenschr 122, 62–64 (2010).
Montastruc, F. et al. The importance of drug-drug interactions as a cause of adverse drug reactions: a pharmacovigilance study of serotoninergic reuptake inhibitors in France. Eur. J. Clin. Pharmacol. 68, 767–775 (2012).
Duke, J. D. et al. Literature based drug interaction prediction with clinical assessment using electronic medical records: novel myopathy associated drug interactions. PLoS Comput. Biol. 8, e1002614 (2012).
Percha, B., Garten, Y. & Altman, R. B. Discovery and explanation of drug-drug interactions via text mining. Pac. Symp. Biocomput. 410–421 (2012).
Takarabe, M., Shigemizu, D., Kotera, M., Goto, S. & Kanehisa, M. Network-based analysis and characterization of adverse drug-drug interactions. J. Chem. Inf. Model. 51, 2977–2985 (2011).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Acknowledgements
The authors would like to thank Dr. Soumitra Ghosh, Dr. Vinod Kumar and Dr. Pankaj Agarwal for critical reading of this manuscript and helpful suggestions. This work is supported by the GSK summer intern/co-op program.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: L.Y. Performed the experiments: H.H., P.Z., L.Y. Analyzed the data: H.H., L.Y., P.Z., P.S., A.Q. Contributed reagents/materials/analysis tools: L.Y., P.S. Wrote the paper: H.H., L.Y., A.Q., P.Z., P.S.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Materials
Supplementary Information
Dataset1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Huang, H., Zhang, P., Qu, X. et al. Systematic prediction of drug combinations based on clinical side-effects. Sci Rep 4, 7160 (2014). https://doi.org/10.1038/srep07160
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07160
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.