Abstract
The protein-protein interaction (PPI) network offers a conceptual framework for better understanding the functional organization of the proteome. However, the intricacy of network complexity complicates comprehensive analysis. Here, we adopted a phylogenic grouping method combined with force-directed graph simulation to decompose the human PPI network in a multi-dimensional manner. This network model enabled us to associate the network topological properties with evolutionary and biological implications. First, we found that ancient proteins occupy the core of the network, whereas young proteins tend to reside on the periphery. Second, the presence of age homophily suggests a possible selection pressure may have acted on the duplication and divergence process during the PPI network evolution. Lastly, functional analysis revealed that each age group possesses high specificity of enriched biological processes and pathway engagements, which could correspond to their evolutionary roles in eukaryotic cells. More interestingly, the network landscape closely coincides with the subcellular localization of proteins. Together, these findings suggest the potential of using conceptual frameworks to mimic the true functional organization in a living cell.
Similar content being viewed by others
Introduction
Proteins are basic parts of molecular machines that usually work together to perform their biological functions in a living cell. For better understanding the underlying cellular architecture and functional organization of the proteome, the protein-protein interaction (PPI) network provides a conceptual framework that depicts a global map of protein interactions in a topological space1,2. This framework has proven useful in systematical analysis of collective dynamics3, functional inference4,5,6, module identification2,7, signaling pathway modeling8,9 and other clinical applications, such as biomarker findings, disease classification10,11 and tumor stratification12.
In a typical PPI network, proteins and their physical interactions are usually symbolized as nodes and edges, respectively, in a mathematical graph representation that describes entity relationships in the topological space. Proteins often work together to carry out their molecular functions by forming complexes or to engage in biological processes by interacting with each other in various interconnected pathways. These behaviors could be captured in the network model to detect functional modularity and protein cooperativity via in-depth topological analysis13. However, the inherent complexity of the biological network, which usually involves thousands of molecular entities and relationships, could make the systematic analysis difficult2,14. For example, due to the multi-functionality nature of proteins, a protein can play different roles and engage in a variety of biological pathway, thus creating multiple connections to various interacting partners in different biological contexts. This intricacy could limit the module detection and functional inference to relatively small local regions and also hampers in-depth investigations on global collective properties of the PPI network, such as its hierarchical structure and scale-free property, both of which are wildly conjectured on the global scale but their origins and the development processes are still unclear13,15,16.
In social science studies, a common way to decompose a social community is to classify its members into age groups, based on the general observation that people of different ages also differ in their social roles, values and positions in the community and may potentially exhibit different behaviors in response to a given event17,18,19. We propose that the same approach could be applied to the biological network analysis. Since the cellular network, just like the genome, developed through evolution16,20,21,22, the phylogenetic grouping technique could be utilized as a tool to decompose a PPI network. Phylogenetics suggests the evolutionary relationships among species and proteins. A typical approach to classify proteins by age is to search for orthologs for each protein in other sequenced genomes and subsequently, the proteins can be assigned to age categories (groups) by tracing the latest common ancestral origin of their orthologous groups across phylogeny23,24,25. In this study, we adopted the same strategy and combined it with force-directed graph simulation in the topological space, to decompose the human PPI network in a multi-dimensional manner. This approach, which we called phylogenetic decomposition (phylo-decomposition), enabled us to associate the network topological properties with evolutionary and biological implications.
Briefly, our work proceeded as follows: First, we addressed the question whether proteins at different ages would play different roles in the human PPI network. From our phylo-decomposed PPI network, we observed that the ancient proteins occupied the core of the network with high topological centrality. Next, we examined if age homophily, a typical pattern of interaction preference in social networks, also existed in the PPI network. By analyzing the temporal patterns of interaction preferences within and between age groups, we revealed the presence of age homophily in the PPI network, which in turn provided valuable clues about the evolutionary process of the network. We thus proposed a hypothesis that selection pressure may have acted on the duplication and divergence processes during the network evolution, in which proteins with higher centrality were selected to avoid perturbation by limiting the probability of a young protein connecting with old ones. Further, we found that age homogeneity prevailed over several kinds of protein communities. These results suggest that the cellular functional modules (e.g., protein complexes and biological pathways) tend to be age homogeneous. In the final part of our work, we linked the age groups to a variety of biological annotations. We found a general consistency between topological centrality and biological importance. For example, the age patterns coincided with the gene essentiality, the disease susceptibility, the evolutionary rates and the specific functional roles in typical eukaryote cells. More interestingly, the network landscape closely mimicked the subcellular localization of the cell. Together, these findings reveal the potential of using conceptual frameworks to capture the true functional organization in a living cell.
Results
Overview of the phylo-decomposition of the human PPI network
To dissect the human PPI network, we adopted a phylogenic grouping method combined with force-directed graph simulation in the topological space (Fig. 1, Supplementary Table S1). Briefly, we first estimated the approximate age group for each human protein based on protein orthology and species phylogeny. For each human protein, its orthologous group across other genomes was identified and then this protein could be classified into one of the age groups (G1–G6) based on the shared ancestral origin of its orthologous group in the phylogeny. Next, the human PPI network, compiled from several public PPI resources, could be projected onto a topological space with an additional temporal dimension that decomposed the global network into six age groups. This network model enabled us to associate the topological characteristics with evolutionary and functional implications.
Schema of the phylo-decomposition of the human PPI network.
Upper part: classification of human proteins into six age groups (represented in different colors) according to protein orthology and species phylogeny. The phylogeny tree and the estimated divergence time (Mya, millions of years ago) were based on NCBI Taxonomy and literature. Branch lengths are not proportional to time. Also note that the age group assignment only reflects the approximate conservation level of proteins, not necessarily corresponding to the estimated divergence time frames. Lower part: decomposition of the human PPI network based on the protein topological relationships and age group annotations. Force-directed layout arranged the nodes into two dimensional topological spaces, based on graph theoretic distance. Then, the global network was decomposed into six subnetworks by introducing an additional dimension based on the protein age group annotations. See main text and Methods for details.
Age-dependent core-periphery structure and network centralities
The first key question we addressed was whether proteins at different ages play different roles in the PPI network. From the phylo-decomposed human PPI network shown in Fig. 2a (see also Supplementary Fig. S1), a core-periphery structure can easily be identified through the age-group dimension, in which ancient proteins occupy the core of the network whereas young proteins tend to reside on the periphery (normalized heat maps are provided in Fig. 2b). The core is the central part of the network, typically providing the structural basis and topological essence of the entire network26,27. Indeed, topological analysis also suggested a positive correlation between protein ages and a variety of network centrality measures (Fig. 2c). The network centrality quantifies a node's relative importance within the network13. For example, the degree centrality quantifies the interaction neighbors of a node, reflecting the node's connectivity and immediate impact; the betweenness centrality of a node quantifies the relative frequency of all-paired shortest paths that rely on the given node, reflecting its potential controllability on the information flows; the closeness centrality quantifies how close in distance a given node is to other reachable nodes in the network, reflecting how fast the information can spread from the node and the stress centrality of a node measures the number of shortest paths that can pass through it, reflecting the potential information traffic load on the node. All these centrality measures can be used as proxy quantifiers of node importance inside the network with different contexts.
Phylo-decomposed human PPI network and centrality measurements.
(a) Decomposed subnetworks from G1 to G6 (shown in different colors). Node size reflects protein degree (connectivity). Edges are omitted for clarity. (b) Density heat map shows the protein localization density inside the network for each age group. (c) Network centrality measurements for proteins in each age group. Values are group means. Color intensity (red-to-green) reflects the relative magnitude of the cell values in each row (high-to-low).
Age homophily and network evolution models
Age homophily is a sociology term that describes the tendency of individuals to associate and bond with similar-aged peers. Since this tendency was found to be prevalent in various types of social networks28,29,30, we expected the PPI network may also possess this property. To this end, we computed the interaction density within and between age groups and estimated the interaction preferences using network randomization procedures (see Methods). Surprisingly, while a slightly heavier interaction density was observed for each age group toward the ancient groups (Fig. 3a), the preference pattern showed a strong tendency for each age group to interact with closely-aged groups, especially with the group itself (Fig. 3b). Furthermore, this pattern revealed a proclivity of proteins to gradually avoid interacting with aged proteins (Fig. 3b). Together, these results suggested the presence of age homophily in the human PPI network.
Interaction density and preference across age groups.
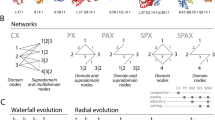
(a) The interaction density within (upper row) and between (indexed grid) age groups. Color intensity (red-to-green) reflects the relative magnitude of cell values (high-to-low). (b) The interaction preference (empirical Z scores, estimated by network randomization) within and between age groups. (c) Interaction patterns for a new hypothetical age group (Group 7) under the random duplication model (left) versus the perturbation avoidance model (right). Perturbation avoidance was implemented by setting the duplicate retention probability of a node inversely proportional to its degree centrality. (d) Composition of paralogous genes in the human genome. A total of n = 211 paralogous groups were extracted from NCBI HomoloGene.
The interaction preference profile across age groups may provide further clues to understand the evolutionary process of the human PPI network. Preferential attachment was shown to be the general mechanism used to generate a scale-free network31. In this model, a new node enters the network with a preference to establish new connections to an existing node that already has many interaction partners inside the network, eventually leading to a scale-free structure. In the evolution of the PPI network, it has been proposed that the gene duplication-and-divergence process can achieve preferential attachment implicitly because proteins with more existing interaction neighbors are more likely to gain new links from randomly duplicate genes13,20. Accordingly, the recruitment of new duplicates eventually accumulates a power-law degree distribution during network growth and thus results in a scale-free network structure. However, this canonical model does not seem to coincide with our results. As previously shown in Figure 3b, ancient proteins, though typically possessing high connectivity, were not preferentially attached by young proteins; on the contrary, proteins with similar ages preferred to interact with each other and generally avoided interacting with aged proteins. Indeed, the canonical duplication-and-divergence model was also found to be incompatible with the observations from the yeast PPI network20. Together, these results suggested a limited role of the canonical duplication-and-divergence model in explaining the evolution of PPI networks20,32.
The observed tendency for proteins to interact with similar-aged neighbors but not with aged ones suggests a possible perturbation avoidance behavior in the network growth process. Given the result that the aged proteins typically possessed high centrality in the network, their duplicates may induce greater perturbation throughout the PPI network and thus become potential threats to cellular homeostasis. For example, the dosage effect could be widely spread and thus more deleterious when the newly formed duplicate was derived from a high connectivity protein. Therefore, to prevent unfavorable perturbations, these types of high centrality duplicates may undergo higher pressure to be selectively removed or to rapidly and largely diverge. We tested the plausibility of a perturbation avoidance model by simulating the duplication process for an additional hypothetical age group (Group 7). Figure 3c illustrates the simulated interaction density patterns for the novel hypothetical age group (G7) using a random duplication model and a perturbation avoidance model. The pattern generated from the random duplication model (Fig. 3c, left) exhibits the preferential attachment mode, in which new duplicates densely connect to ancient proteins. This pattern is quite inconsistent with the true network pattern (Fig. 3a). In contrast, the pattern generated from the perturbation avoidance model (Fig. 3c, right) closely mimics the true network pattern (Fig. 3a), suggesting that perturbation avoidance could be a potential strategy adopted in the evolution of the human PPI network.
To further evaluate the plausibility of the perturbation avoidance model, we then performed a network growth simulation to examine whether the scale-free property could be acquired based on the proposed model. Briefly, a small random network was initialized to grow following a perturbation avoidance strategy until the total number of nodes reached a scale comparable to the human PPI network. The strategy applied stochastic constraints on node duplication and divergence events by setting the event probabilities inversely proportional to the node degree (see Methods for details). As expected, the final simulated network exhibits approximate scale-free property globally and locally across the temporal groups (Supplementary Fig. S2). This suggested that perturbation avoidance could be a valid strategy to grow a small random network into a sizeable scale-free network.
The perturbation avoidance model also implies the inequality of retained paralogs among different age groups. Paralogs are the homologous genes that were created by duplication events within the genome. Since the perturbation avoidance model assumes that high centrality genes are less likely to retain duplicates in the genome (most of them should have largely diverged or been rapidly removed), the ancient proteins should retain less paralogous genes than young proteins in the current genome. Figure 3d shows the age composition of paralogous genes in the human genome. In agreement with the assumption, most of the paralogous genes originated from young proteins. This tendency revealed the expected inequality of retained paralogs among age groups.
Age homogeneity prevailing over protein communities
As shown in the last section, the observed tendency for proteins to interact with similar-aged neighbors suggests that age homogeneity could prevail over protein communities. Here, we use the term “community” to refer to a group of closely-related proteins, for example, the members in a protein complex or a PPI module (physically-interacted) or in a biological pathway (functionally-related). We started with the PPI dyad, the smallest unit of a protein group that comprises only two interlinked proteins. Overall, most of the human PPI dyads were in the same or similar-aged groups (70%, empirical P < 1 × 10−5, permutation test), especially those PPI dyads inside protein complexes (82%, empirical P < 1 × 10−5, permutation test) (Fig. 4a). Noting that the presence of a PPI also implies functional similarity between the two paired proteins33, we were therefore curious whether the age homogeneity would also reflect on the degree of functional similarity. Indeed, the patterns of degree of functional similarity among the PPI dyads suggested a correlation between age homogeneity and functional similarity, in which similar-aged PPI dyads tended to have a higher degree of functional similarity (Fig. 4b).
Age homogeneity and functional similarity of PPI dyads.
(a) Composition of PPI dyads classified by the difference in age between the two paired proteins. All PPIs: all PPI dyads from the human PPI network; Complex PPIs: the PPIs within protein complexes. (b) Profile of functional similarity among PPI dyads. Cell values indicate the median. Color scheme (red-to-green) represents the magnitude of similarity degree (high-to-low). GOBP/GOMF: similarities based on the Gene Ontology namespace of the Biological Process/Molecular Function.
Age homogeneity prevailed over PPI dyads as well as larger groups of proteins. For a given group of proteins, we calculated the standard deviation (SD) of its members' ages as a proxy for estimating the degree of age homogeneity. In a PPI network, a pivot node that has relatively high connectivity can be defined as a network hub13. Each hub together with its most proximate neighbors defined a local group in the PPI network, termed a hub-spoke community. As expected, the hub-spoke communities in the human PPI network generally selected age homogeneity over randomly chosen groups, even with different degree cutoffs for hub protein definitions (Fig. 5a). Interestingly, the distribution of the age SD of the hub-spoke communities seemed to center at 1.0. We thus separated the network hubs into two classes: the school hubs and the office hubs, by setting the separate point of age SD at 1.0. The school hubs are those network hubs interacting in age-homogenous communities (age SD < 1.0), given the name because children typically form peer groups in school34,35. In contrast, the office hubs are those network hubs interacting in relatively age-heterogeneous communities (age SD ≥ 1.0), which are analogous to ordinary offices in which the colleagues are different ages. Most of the high-degree hubs were office hubs (Fig. 5b). This could be due to the fact that their community sizes were more similar to the universal population size. However, we found the communities of school hubs were slightly more functional homogeneous compared with the office hubs, regardless of the degree cutoffs (Fig. 5c). Since the interacting communities of school hubs were age homogeneous by definition, this result implied a link between age homogeneity and functional homogeneity. Indeed, the proteins engaging in the common biological pathways were found to be age homogeneous as well (Supplementary Fig. S3). Together, these results suggested that the cellular functional modules tend to be age homogeneous.
Age homogeneity in hub-spoke communities.
(a) Hub-spoke communities were homogeneous in age. Different degree threshold (top 20%, 10% and 5%; n denotes the number of hubs under that criterion) were used to test the consistency of age homogeneity (all with P < 1 × 10−16, two-tailed Mann-Whitney U test). Hubs were divided into two types: school hubs and office hubs, based on the degree of age homogeneity. (b) Composition of hub types. Each pie chart corresponds to the criterion adopted in (a). (c) Comparison of functional homogeneity between different types of hubs using a two-sample KS-test. ECDF: empirical cumulative distribution function.
Age-dependent functional landscape
The age-dependency of topological centrality suggested that age groups may play distinct roles in cellular functions. To elucidate the typical functional roles of different age groups, a series of functional analyses were conducted. First, since the aged proteins generally possessed high centralities in the PPI network, they may be more important to cell viability. Indeed, proteins with high degree centrality were more essential for yeast survival36. In agreement with that, we found that protein age could be associated with gene essentiality (Supplementary Fig. S4a) and disease susceptibility (Supplementary Fig. S4b,c): the older the proteins, the higher the essentiality and disease susceptibility. Gene essentiality could also have imposed constrains on the gene evolutionary rates. We therefore examined the human–mouse evolutionary rates for orthologous proteins. Unsurprisingly, age-dependency also reflected in the evolutionary rates (Supplementary Fig. S4d), in which aged genes were under a higher pressure of purifying selection than young proteins. A similar observation has been reported previously37. We also checked the chromosome distribution of the age groups and found that a high ratio of young genes resided on human chromosomes 19, X and Y (Supplementary Fig. S5). Consistently, genes on these chromosomes generally showed higher divergence rates (Supplementary Fig. S6). It was estimated that the differentiation of X-Y chromosomes occurred shortly after the divergence of the mammalian and avian lineages38. In addition, genes on human chromosomes 19 and Y were reported to have high divergence rates39,40. The high divergence rates of genes on human chromosome 19 were previously associated with high GC contents39; however, we found this association was not maintained on chromosome Y, on which the genes typically had low GC contents (Supplementary Fig. S6). Moreover, we found no global correlation between the divergence rates and GC contents. Unexpectedly, beyond the genes on chromosomes 19, X and Y, genes on chromosome 1 also showed high divergence rates (Supplementary Fig. S6), which failed to correspond to its uniformly distributed age group composition (Supplementary Fig. S5). Further research is needed to elucidate the evolutionary force acting on chromosome 1.
Functional analysis also revealed high specificity of enriched biological processes and pathway engagements among age groups (Fig. 6 and Supplementary Table S2). This functional specificity seemed to correspond to group-specific evolutionary roles in the eukaryotic cell. For example, the eukaryote-conserved proteins (G1) were enriched in basal cellular functions such as translation, RNA processing, oxidation reduction and protein localization; the metazoan-conserved proteins (G2) were enriched in neuron development, embryonic morphogenesis, transport and signaling cascade; the vertebrate-conserved proteins (G3) were enriched in organ development, apoptosis and signaling transduction; the mammal-conserved proteins (G5) were enriched in sensory perception, sexual reproduction and immune response and the primate-conserved proteins (G6) were enriched in defense response, transcriptional regulation and keratinization. Keratinization genes were recently found to be positively selected in primates, revealed by a comparative genome-wide sequencing of primate exomes41.
Enrichment map of biological processes across phylogenetic groups.
Nodes represent enriched biological process terms from Gene Ontology. Edges represent the associations between two enriched processes. Highly connected terms were grouped together and were annotated manually by a shared general term. Given two associated gene sets A and B, the overlap coefficient (OC) was defined as OC = |A∩B|/min(|A|, |B|).
More interestingly, the subcellular localization of proteins closely coincided with the core-periphery structure of the PPI network. As shown in Figure 7, the ancient proteins typically reside at the core of the cell, that is, the nucleus, the cytoplasm and the organelles. In contrast, young proteins are largely located at the cell periphery, namely, the cell membrane and extracellular area. Notably, besides the extracellular area, the primate-specific proteins (G6) were also slightly enriched inside the nucleus. This implies that many G6 proteins could be transcription factors or engage in transcriptional regulation, as shown in Figure 6. Together, these findings suggested the potential of the conceptual framework to mimic the true functional organization in a living cell.
Discussion
Decades of network science research have made considerable progress and numerous discoveries. One of the most striking findings is that though they differ largely in basic elements, many network systems share similar collective dynamics and structured organizations, which only emerge in the systematic view. Since the general principles behind this theme are not yet fully understood, the similarities in the collective dynamics and structured organizations could be important areas to explore in future research.
In this study, we made an attempt to utilize phylogenetic information in an analysis of the human PPI network. A temporal dimension was introduced to decompose the network structure and functional organization of the PPI network. Our results suggest a consistent pattern that the node centrality generally increases with age; in other words, aged proteins are more likely to act as pivot nodes inside the human PPI network. Notably, these patterns are generally in agreement with previous observations in the yeast PPI network, in which the degree centrality also increased with the age of the protein23. Another recent study also reported the tendency of aged proteins to have higher connectivity25.
Our results also suggest the presence of age homophily in the human PPI network. It is also of great interest to know whether the same tendency can be observed in other species. In fact, the patterns of interaction density between age groups have been studied in the yeast20,23 and human PPI networks25; however, inconsistent patterns were revealed in these studies. In the yeast PPI network, heavier interaction density was found between similar-aged groups20,23, whereas in the human PPI network, all groups possessed heavier interaction density toward ancient groups25, which was supported by our interaction density results (Fig. 3a). Nevertheless, we noticed that this inconsistency could be resolved by estimating the interaction preference instead of the density (Fig. 3b). Indeed, we found that the pattern of interaction preference coincided with the tendency found in the yeast, that is, proteins tend to interact with other similar-aged proteins. Therefore, we propose the possibility that this tendency might be a universal principle that was also applied in the evolution of PPI networks in other species.
The observed tendency for proteins to interact with similar-aged neighbors but not with aged ones suggests a possible perturbation avoidance behavior in the network growth process. Notably, this behavior was also suggested in the yeast PPI network, in which the proteins with a high degree of connectivity typically had low duplicability23. Moreover, similar observations were reported in the yeast genetic interaction network, in which the genes with many genetic interactions were found less responsive to environmental changes42. It is possible that the low responsiveness could result in avoidance of large-scale perturbation on genetically linked partners because these perturbations could ultimately trigger severe effects on fitness. Taken together, these results also suggest that perturbation avoidance could be a general principle applied in the evolution of cellular networks.
Methods
Phylogenetic decomposition of the PPI network
We adopted a similar strategy that has been applied in previous studies23,25,37 to estimate the approximate evolutionary age for each human protein. Specifically, we searched for protein orthologs from other genomes for each human protein and then the protein was assigned to an age group based on the shared ancestral origin of its orthologous group across species phylogeny. Under this strategy, human proteins could be approximately classified into six age groups (G1–G6). G1 contained those proteins well-conserved in fungi and animals (oldest group); G2 contained those proteins broadly conserved in animals but no orthologs were found in fungi; other groups were assigned using the same logic. Note that G6 was the youngest group, which included human-specific or primate-only proteins. Protein orthology was obtained from NCBI HomoloGene (release 67, Dec 2012)43 and species phylogeny was from NCBI Taxonomy and from literature44. A total of 20 species were listed in HomoloGene and 18 of them (fungi and animals) were included in this study (two plant species were excluded). The 18 included species were (alphabetical order): Anopheles gambiae, Bos taurus, Caenorhabditis elegans, Canis lupus familiaris, Danio rerio, Drosophila melanogaster, Eremothecium gossypii, Gallus gallus, Homo sapiens, Kluyveromyces lactis, Macaca mulatta, Magnaporthe oryzae, Mus musculus, Neurospora crassa, Pan troglodytes, Rattus norvegicus, Saccharomyces cerevisiae and Schizosaccharomyces pombe. We choose HomoloGene as the orthology source because a previous study reported that HomologGene showed the best performance in the phylogenetic tests over seven other orthology projects and methods45. Another reason was that HomoloGene could provide distant ortholog inferences covering most of the model organisms across fungi and animals, which was a prerequisite for this study. Nevertheless, the limited number of species available from current databases still restricted the resolution and accuracy of the age group assignment. For example, potential gene loss events in some species could have misguided the age group assignment and could not be corrected if no other species were available for that clade, though this kind of inaccuracy seldom exceeded one intergroup range. Also note that this method was based on sequence conservation, not necessarily reflecting the exact timeframe of the functional origin of a given protein in the natural history, because it is possible that some proteins have a common functional origin but have diverged extensively in sequence in each lineage. A potential area for future improvement would be on the sequence alignment algorithms. For example, a recent study showed that it is possible to achieve higher sensitivity in the case of coiled-coil protein homologs detection by adjusting domain-specific substitution matrices for sequence alignment46. The anticipated advancement in this area would improve the accuracy of homologs detection and thus benefit our methodology in the future.
The human PPI network was compiled from several public PPI resources (HPRD, DIP, IntAct, BioGRID and MINT), downloaded from the PrePPI database47. Only physical interactions were considered in this study. Note that this dataset integrated all kinds of physical PPIs derived from various experiment techniques, thus containing both direct and indirect physical interactions between proteins. Our network analysis pipeline involved two steps. First, we used Cytoscape48 to visualize and analyze the graph topology of the PPI network. Spring-embedded algorithm49 is a classic force-directed graph layout algorithm that simulates the physical dynamics of the spring force between nodes in the system. This algorithm positions the nodes (proteins) in a two-dimensional topological space using an iterative process that minimizes the free energy of the system and it generates optimized spring lengths that approximate theoretical graph distances, which makes it ideal for decomposing a network with a core-periphery structure. Next, an additional temporal dimension was introduced to the network based on the age group annotation of proteins, by which the global PPI network could be decomposed into six subnetworks. Each subnetwork corresponds to one specific age group and the PPIs could thus be classified into two classes: one as intra-group PPIs and the other as inter-group PPIs. Downstream topological and functional analysis were performed with in-house Python scripts and NetworkAnalyzer50, a Cytoscape plugin that computes specific parameters for network topology.
Interaction density and Z scores
The interaction density was defined as described before20,25. Specifically, given two age groups m and n, the interaction density Dm,n between them was calculated as


where Im,n is the number of edges between the two groups m and n; Em,n is the maximum number of edges that can possibly exist between the two groups; and Nm and Nn are the number of nodes in the group m and n, respectively. The interaction density is always a number between 0 and 1. The empirical Z scores of the interaction density were estimated from network randomization procedures 10,000 times. The procedures randomly rewired all edges in the PPI network while preserving its characteristic degree sequence. The empirical Z scores could thus be estimated from the background distributions of the interaction densities generated from the 10,000 randomized PPI networks. All homodimer interactions were removed before performing the analysis.
Network growth simulation
While the simplest form of the canonical duplication-and-divergence model assumes random duplication followed by a constant divergence rate20, the perturbation avoidance model sets the duplication and the divergence rates inversely proportional to the node degree centrality. The simulation started from a random network with n = 100 nodes and probability p = 0.1 for edge creation (namely, an Erdős-Rényi graph with parameters n = 100 and p = 0.1). Then, the initialized network started to grow following the perturbation avoidance strategy, until the total number of nodes reached n = 10,000. An additional heteromerization parameter that controls the probability of two duplicates to link can be optionally included in the model to strengthen the hierarchical structure of the network20,32, though it appeared not necessary for the emergence of the scale-free property. In addition to the Erdős-Rényi graph, we also ran the simulation on other choices of initial networks, such as Watts-Strogatz small-world graph and Barabási-Albert scaling graph. For Watts-Strogatz graph, we set initial node number = 100, each node connecting to 5 nearest neighbors and 0.1 probability of rewiring each edge. For Barabási-Albert graph, the initial node number was set to 100 nodes and the number of edges to attach from a new node to existing nodes was set to 5.
Age homogeneity and functional similarity
For PPI dyads, age homogeneity was defined by the age difference between the two paired proteins of a given PPI, as illustrated in Figure 5a. To test if most of the PPI dyads tend to be of similar age, Monte Carlo procedures were used to calculate empirical P values. The procedures randomly rewired the PPI network while maintaining a constant global degree sequence. Annotated protein complexes were obtained from the CORUM database51. Functional similarity was measured using GOSemSim52, an R package for estimating GO sematic similarities between two genes or two gene clusters. Default parameters were applied in all measurements.
For the hub-spoke communities, three stepwise degree thresholds were used to define the hub proteins, namely, the top 20%, 10% and 5% degree rank, respectively. Under each threshold, corresponding randomized communities were generated by resampling from the entire PPI network. The functional homogeneity of a protein comunity was defined as the median of the pairwise semantic similarities of a given protein group. KEGG pathways were obtained from the NCBI BioSystems database53.
Gene essentiality
Two distinct sources for the gene essentiality annotations were adopted in this study. The first source was from mouse phenotype data. Specifically, a gene knock-out in mice resulting in any lethality phenotype was defined as an essential gene and its corresponding human ortholog was thus assumed essential. This strategy to predict the essentiality of a human gene from mouse phenotype information has been widely adopted54,55,56. Mouse phenotype data were downloaded from MGI57. The second source for gene essentiality was from the COLT-Cancer database, which was based on a genome-scale pooled shRNA screening for essential genes in human cancer cell lines58.
Human–mouse evolutionary rates
The human–mouse evolutionary rates (dN, dS and dN/dS) for human protein-coding genes were obtained from Ensembl BioMart (release 72). We selected only one-to-one orthologs between human and mouse in this analysis.
Analysis and visualization of functional enrichment
Functional enrichment analysis was performed using DAVID 6.759 (GO enrichment analysis, pathway enrichment analysis, functional domain enrichment analysis). Default parameters were used in all types of analyses in DAVID. Functional landscape was visualized using Cytoscape48 and its plugin Enrichment Map60.
References
Stelzl, U. et al. A human protein-protein interaction network: a resource for annotating the proteome. Cell 122, 957–968 (2005).
Mitra, K., Carvunis, A. R., Ramesh, S. K. & Ideker, T. Integrative approaches for finding modular structure in biological networks. Nat Rev Genet 14, 719–732 (2013).
Bork, P. et al. Protein interaction networks from yeast to human. Curr Opin Struct Biol 14, 292–299 (2004).
Vazquez, A., Flammini, A., Maritan, A. & Vespignani, A. Global protein function prediction from protein-protein interaction networks. Nat Biotechnol 21, 697–700 (2003).
Nabieva, E., Jim, K., Agarwal, A., Chazelle, B. & Singh, M. Whole-proteome prediction of protein function via graph-theoretic analysis of interaction maps. Bioinformatics 21 Suppl 1, i302–310 (2005).
Sharan, R., Ulitsky, I. & Shamir, R. Network-based prediction of protein function. Mol Syst Biol 3, 88 (2007).
Chen, B., Fan, W., Liu, J. & Wu, F. X. Identifying protein complexes and functional modules--from static PPI networks to dynamic PPI networks. Brief Bioinform, (2013).
Scott, J., Ideker, T., Karp, R. M. & Sharan, R. Efficient algorithms for detecting signaling pathways in protein interaction networks. J Comput Biol 13, 133–144 (2006).
Gitter, A., Klein-Seetharaman, J., Gupta, A. & Bar-Joseph, Z. Discovering pathways by orienting edges in protein interaction networks. Nucleic Acids Res 39, e22 (2011).
Lee, E., Chuang, H. Y., Kim, J. W., Ideker, T. & Lee, D. Inferring pathway activity toward precise disease classification. PLoS Comput Biol 4, e1000217 (2008).
Ideker, T. & Sharan, R. Protein networks in disease. Genome Res 18, 644–652 (2008).
Hofree, M., Shen, J. P., Carter, H., Gross, A. & Ideker, T. Network-based stratification of tumor mutations. Nat Methods, (2013).
Barabasi, A.-L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nat Rev Genet 5, 101–113 (2004).
Albert, R. Scale-free networks in cell biology. J Cell Sci 118, 4947–4957 (2005).
Hase, T., Tanaka, H., Suzuki, Y., Nakagawa, S. & Kitano, H. Structure of protein interaction networks and their implications on drug design. PLoS Comput Biol 5, e1000550 (2009).
Jin, Y., Turaev, D., Weinmaier, T., Rattei, T. & Makse, H. A. The evolutionary dynamics of protein-protein interaction networks inferred from the reconstruction of ancient networks. PLoS One 8, e58134 (2013).
Palla, G., Barabasi, A. L. & Vicsek, T. Quantifying social group evolution. Nature 446, 664–667 (2007).
Wellman, B. & Berkowitz, S. D. Social Structures: A Network Approach. (Cambridge University Press, Cambridge, 1988).
Eisenstadt, S. N. From Generation to Generation: Age Groups and Social Structure. (Routledge & Kegan Paul, London, 1956).
Kim, W. K. & Marcotte, E. M. Age-dependent evolution of the yeast protein interaction network suggests a limited role of gene duplication and divergence. PLoS Comput Biol 4, e1000232 (2008).
Fraser, H. B., Hirsh, A. E., Steinmetz, L. M., Scharfe, C. & Feldman, M. W. Evolutionary rate in the protein interaction network. Science 296, 750–752 (2002).
Wuchty, S. Evolution and topology in the yeast protein interaction network. Genome Res 14, 1310–1314 (2004).
Prachumwat, A. & Li, W. H. Protein function, connectivity and duplicability in yeast. Mol Biol Evol 23, 30–39 (2006).
Saeed, R. & Deane, C. M. Protein-protein interactions, evolutionary rate, abundance and age. BMC Bioinformatics 7, 128 (2006).
Zhao, Y. & Mooney, S. D. Functional organization and its implication in evolution of the human protein-protein interaction network. BMC Genomics 13, 150 (2012).
Souiai, O. et al. Functional integrative levels in the human interactome recapitulate organ organization. PLoS One 6, e22051 (2011).
Csermely, P., London, A., Wu, L.-Y. & Uzzi, B. Structure and dynamics of core-periphery networks. Journal of Complex Networks 1, 93–123 (2013).
McPherson, M., Smith-Lovin, L. & Cook, J. M. Birds of a feather: homophily in social networks. Annu Rev of Sociol 27, 415–444 (2001).
Krivitsky, P. N., Handcock, M. S., Raftery, A. E. & Hoff, P. D. Representing degree distributions, clustering and homophily in social networks with latent cluster random effects models. Soc networks 31, 204–213 (2009).
Flatt, J. D., Agimi, Y. & Albert, S. M. Homophily and health behavior in social networks of older adults. Fam & community health 35, 312–321 (2012).
Barabasi, A. L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Gibson, T. A. & Goldberg, D. S. Improving evolutionary models of protein interaction networks. Bioinformatics 27, 376–382 (2011).
Pesquita, C., Faria, D., Falcao, A. O., Lord, P. & Couto, F. M. Semantic similarity in biomedical ontologies. PLoS Comput Biol 5, e1000443 (2009).
Robertson, D. & Symons, J. Do peer groups matter? peer group versus schooling effects on academic attainment. Economica 70, 31–53 (2003).
Steinberg, L. Adolescence. (McGraw-Hill, New York, 2010).
Jeong, H., Mason, S. P., Barabasi, A. L. & Oltvai, Z. N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001).
Albà, M. M. & Castresana, J. Inverse relationship between evolutionary rate and age of mammalian genes. Mol Biol Evol 22, 598–606 (2005).
Lahn, B. T. & Page, D. C. Four evolutionary strata on the human X chromosome. Science 286, 964–967 (1999).
Castresana, J. Genes on human chromosome 19 show extreme divergence from the mouse orthologs and a high GC content. Nucleic Acids Res 30, 1751–1756 (2002).
Hughes, J. F. et al. Chimpanzee and human Y chromosomes are remarkably divergent in structure and gene content. Nature 463, 536–539 (2010).
George, R. D. et al. Trans genomic capture and sequencing of primate exomes reveals new targets of positive selection. Genome Res 21, 1686–1694 (2011).
Park, S. & Lehner, B. Epigenetic epistatic interactions constrain the evolution of gene expression. Mol Syst Biol 9 (2013).
Wheeler, D. L. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 36, D13–21 (2008).
Hedges, S. B. The origin and evolution of model organisms. Nat Rev Genet 3, 838–849 (2002).
Altenhoff, A. M. & Dessimoz, C. Phylogenetic and functional assessment of orthologs inference projects and methods. PLoS Comput Biol 5, e1000262 (2009).
Kuhn, M., Hyman, A. A. & Beyer, A. Coiled-coil proteins facilitated the functional expansion of the centrosome. PLoS Comput Biol 10, e1003657 (2014).
Zhang, Q. C., Petrey, D., Garzón, J. I., Deng, L. & Honig, B. PrePPI: a structure-informed database of protein–protein interactions. Nucleic Acids Res 41, D828–D833 (2013).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P. L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432 (2011).
Kamada, T. & Kawai, S. An algorithm for drawing general undirected graphs. Inform Process Lett 31, 7–15 (1989).
Assenov, Y., Ramirez, F., Schelhorn, S. E., Lengauer, T. & Albrecht, M. Computing topological parameters of biological networks. Bioinformatics 24, 282–284 (2008).
Ruepp, A. et al. CORUM: the comprehensive resource of mammalian protein complexes. Nucleic Acids Res 36, D646–650 (2008).
Yu, G. et al. GOSemSim: an R package for measuring semantic similarity among GO terms and gene products. Bioinformatics 26, 976–978 (2010).
Geer, L. Y. et al. The NCBI BioSystems database. Nucleic Acids Res 38, D492–496 (2010).
Goh, K.-I. et al. The human disease network. P Natl Acad Sci USA 104, 8685–8690 (2007).
Liang, H. & Li, W. H. Gene essentiality, gene duplicability and protein connectivity in human and mouse. Trends Genet 23, 375–378 (2007).
Liao, B.-Y. & Zhang, J. Mouse duplicate genes are as essential as singletons. Trends Genet 23, 378–381 (2007).
Eppig, J. T. et al. The Mouse Genome Database (MGD): comprehensive resource for genetics and genomics of the laboratory mouse. Nucleic Acids Res 40, D881–886 (2012).
Koh, J. L. et al. COLT-Cancer: functional genetic screening resource for essential genes in human cancer cell lines. Nucleic Acids Res 40, D957–963 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57 (2009).
Merico, D., Isserlin, R. & Bader, G. D. Visualizing gene-set enrichment results using the Cytoscape plug-in enrichment map. Methods Mol Biol 781, 257–277 (2011).
Acknowledgements
We would like to thank Chia-Lang Hsu and Chia-Hsien Lee for valuable discussions. The image of the eukaryotic cell as the background in Figure 7 was adopted from the public domain, which was kindly provided by Mariana Ruiz Villarreal. This work was supported by the National Science Council of Taiwan (NSC 102-2628-B-002-041-MY3, 102-2627-B-002-002 and 102-2311-B-010-004) and the National Taiwan University Cutting-Edge Steering Research Project (103R7602C3).
Author information
Authors and Affiliations
Contributions
C.Y.C. designed and performed the analyses, wrote the main manuscript text and prepared Figure 1–7 & Supplementary Information. A.H. collected the materials, conducted parts of the analysis and prepared Figure 1 & Supplementary Figure S3. H.Y.H. constructed the network evolution model and performed the simulation. H.F.J. and H.C.H. conceived the study and edited the manuscript. All authors reviewed and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Chen, CY., Ho, A., Huang, HY. et al. Dissecting the Human Protein-Protein Interaction Network via Phylogenetic Decomposition. Sci Rep 4, 7153 (2014). https://doi.org/10.1038/srep07153
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07153
This article is cited by
-
How the evolution of multicellularity set the stage for cancer
British Journal of Cancer (2018)
-
Non-interacting proteins may resemble interacting proteins: prevalence and implications
Scientific Reports (2017)
-
Information processing in the CNS: a supramolecular chemistry?
Cognitive Neurodynamics (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.