Abstract
The modification of proteins with polyubiquitin chains alters their stability, localization and activity, thus regulating various aspects of cellular functions in eukaryotic cells. The ER quality control protein E3 gp78 catalyzes Lys48-linked polyubiquitin-chain- assembly on the Ube2g2 active site and is capable of transferring preassembled ubiquitin chains to its substrates. However, the underlying mechanism of polyubiquitin- chain-assembly remains elusive. Here, we demonstrate that the active site-linked ubiquitin chain is extended from the distal end by the cooperative actions of the G2BR and CUE domains of gp78. The G2BR domain is involved in ubiquitin chain synthesis by binding to the donor Ube2g2~Ub and promoting ubiquitin transfer from the E2 in cis. The CUE domain shows preferential binding to the ubiquitin chain compared to monoubiquitin and helps to position the distal ubiquitin in the correct orientation to attack the Ube2g2~Ub thioester bond. Our studies reveal that two interactions, one between the donor Ube2g2~Ub and the gp78 G2BR domain and another between the Ube2g2-linked ubiquitin chain and the gp78 CUE domain, cooperatively drive polyubiquitin-chain-assembly on the Ube2g2 active site.
Similar content being viewed by others
Introduction
The modification of proteins with ubiquitin (Ub) regulates various aspects of the cellular functions of proteins by changing their activity, stability and localization1,2,3. This reaction requires the sequential actions of the following three types of enzymes: (1) an activating enzyme (E1) that forms a thioester linkage between its catalytic cysteine and the carboxyl group of Gly76 in ubiquitin to activate it; (2) a conjugating enzyme (E2) that receives ubiquitin from the E1; and (3) a ubiquitin ligase (E3) that transfers ubiquitin molecules from the E2 to the substrate4,5. An unusual feature of ubiquitination is its ability to form a chain consisting of multiple ubiquitin molecules linked by isopeptide bonds6.
Ubiquitin is a 76-amino acid polypeptide that is evolutionary conserved throughout the eukaryotic kingdom7. There are seven lysine residues in ubiquitin and any of them can be used to assemble polyubiquitin chains8. The C-terminus of ubiquitin can be linked to the N-terminal amino group of another ubiquitin molecule to form linear ubiquitin chains9. Among the eight known ubiquitin linkages, the functions of the Lys48- and the Lys63-linked polyubiquitin chains are best characterized. In general, Lys63-linked polyubiquitination performs non-proteolytic functions such as protein trafficking, DNA repair and signal transduction, whereas Lys48-linked ubiquitin chains target protein substrates to the 26S proteasome for degradation10,11,12,13. It is generally believed that a chain containing a minimum of four ubiquitin molecules linked by Lys48 is required for substrate targeting to the proteasome13.
Recently, several models have been proposed to explain the mechanism underlying linkage-specific polyubiquitination6,14,15,16,17,18,19,20,21. The extension of polyubiquitin chains was previously thought to occur via the sequential addition model, in which the most distal ubiquitin on the substrate provides the attacking lysine for additional rounds of the ubiquitination cycle6. In sharp contrast to the sequential addition model, a unique seesaw mechanism has also been proposed, which predicts that the ubiquitin chain will be extended by the incorporation of ubiquitin at the proximal end of a chain linked to the catalytic cysteine of an E2. In the seesaw model, ubiquitin chains are transferred back and forth from the active site of one E2 to a neighboring E2. Ubiquitin chains were indeed detected in conjugation with the catalytic cysteine of E2 Ube2g2, as well as its yeast homolog Ubc7p, a key E2 involved in the degradation of misfolded proteins in the endoplasmic reticulum22,23,24,25. Intriguingly, polyubiquitin chains preassembled on the active site of Ube2g2 can be subsequently transferred to a substrate24. Similarly, polyubiquitin chains linked to the active site of Ubc7p can be transferred to a nearby lysine residue in Ubc7p for its own degradation25. In this reaction, the oligomerization of gp78 and the high affinity of gp78 to its cognate E2 Ube2g2 bring multiple Ube2g2~ubiquitin complexes into close proximity, thereby enhancing the chain assembly activity of Ube2g226,27. In addition, dimerization of Ube2g2 could engage two charged ubiquitins to specifically synthesize Lys48-linked ubiquitin chains28. These results establish that polyubiquitination can be achieved by ligating a preassembled ubiquitin chain to the substrate from a cysteine in the E2 active site. However, it remains unclear whether the ubiquitin chains on the Ube2g2 active site are built by the seesaw mechanism or via the sequential addition model.
In this study, we further dissected the mechanism of ubiquitin chain assembly by the Ube2g2-gp78 enzyme pair. Our results show that the Ube2g2-linked polyubiquitin chain can only be elongated from the distal end but not from the proximal end. Moreover, we find that in addition to the RING domain29, two other gp78 domains, the G2BR and CUE domains, also play important roles in ubiquitin chain formation. The G2BR domain directly binds to the donor Ube2g2 to promote the discharge of ubiquitin from the E2. The CUE domain helps to engage an acceptor Ube2g2 molecule by binding to the ubiquitin chain on the acceptor Ube2g2. Together, our results reveal that two domains of the ER-associated protein degradation (ERAD)-related E3 gp78 cooperatively elongate polyubiquitin chains on the catalytic cysteine of Ube2g2 from the distal end by the sequential addition model.
Results
Ubiquitin chains on the Ube2g2 active site are elongated from the distal end
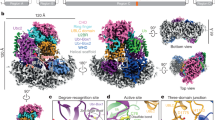
Previously, an interesting “seesaw” model was proposed to explain how to build longer polyubiquitin chains, which is distinct with the canonical sequential addition model6. In the sequential addition model, ubiquitin molecules are added one at a time, first to the substrate and then to the distal end of a growing ubiquitin chain (Fig. 1a)6. In contrast, in the seesaw model, the latest ubiquitin is always added to the proximal end of the ubiquitin chain rather than to the distal end. Additionally, the seesaw model predicts that an E2 should be able to transfer a ubiquitin chain from its active site to another E2 in a homodimeric E2 complex. First, we tested whether a short ubiquitin oligomer conjugated to the active site of Ube2g2 could be transferred to a neighboring Ube2g2~Ub complex. To determine this, we used a single-round ubiquitin transfer assay modified from a previous report (Fig. 1b)24. Ubiquitin oligomers containing a FLAG-tagged Ub K48R (F∧UbK48R) mutant at the distal end were synthesized in vitro (see experimental procedures) (Fig. 1c) and charged onto Ube2g2 (Fig. 1d). Because the distal F∧UbK48R mutant prevented chain extension from this end, these E2-ubiquitin oligomer complexes could serve only as potential donors in the transfer reaction. Upon incubating these complexes with Ube2g2 precharged with an untagged wildtype ubiquitin (acceptor), the transfer of these ubiquitin oligomers to an untagged ubiquitin acceptor should then lead to the extension of the chains by one ubiquitin molecule, which could be detected by immunoblotting with the FLAG antibody. Our results showed that a fraction of the F∧UbK48R monomer was transferred to the Ube2g2~Ub acceptor after incubation, as was previously demonstrated (Fig. 1e, lanes 1–3)24. However, neither di- nor tri-ubiquitin oligomers could be transferred to the acceptor under the same conditions (Fig. 1e, lanes 4–9). These data indicate that ubiquitin oligomers conjugated to the active site of Ube2g2 cannot be transferred to another Ube2g2~Ub thioester complex, which is inconsistent with the “seesaw” model.
Ubiquitin can only be added to the distal end of ubiquitin oligomers conjugated to the Ube2g2 active site.
(a) Schematic representation of the proposed sequential and seesaw models for ubiquitin chain extension on the active site of an E2 enzyme. (b) The experimental scheme of (e). (c) Ubiquitin oligomers carrying F∧UbK48R on their distal ends were synthesized and analyzed by immunoblotting. (d) Conjugation of the synthesized ubiquitin oligomers to the Ube2g2 active site were reduced by the addition of the reducing reagent dithiothreitol (DTT)24. (e) As indicated in (b), Ube2g2 precharged with the indicated ubiquitin oligomers was mixed with Ube2g2 precharged with untagged wildtype Ub in the presence of GST-gp78c. The transfer of ubiquitin oligomers carrying F∧UbK48R to the Ube2g2~Ub complex was analyzed by immunoblotting. The red arrowhead shows Ube2g2 carrying a hybrid di-ubiquitin on its active site as a result of the above mentioned transfer reaction between Ube2g2~F∧UbK48R and Ube2g2~Ub. *, indicates non-specific bands. (f) The experimental scheme of (h). (g) The charging of ubiquitin oligomers to Ube2g2. Charging reactions were performed with E1, Ube2g2 and ubiquitin oligomers carrying 1-4 ubiquitin moieties (Ub1-4) and analyzed under non-reducing condition by immunoblotting. (h) As indicated in the experimental scheme in (f), Ube2g2 precharged with the indicated ubiquitin oligomers (Ub1-4) was mixed with Ube2g2 precharged with F∧UbK48R in the presence of GST-gp78c. The transfer of F∧UbK48R to the ubiquitin oligomers linked to the Ube2g2 active site was analyzed by immunoblotting. *, indicates Ube2g2 conjugated with two F∧UbK48Rs that was generated in the precharging reaction by transferring F∧UbK48R to a lysine residue in Ube2g224. Western blots (e) and (h) in this figure are representative blots from 3 replicates.
To demonstrate that ubiquitin chains on Ube2g2 are extended by the sequential addition model, we performed an experiment reciprocal to that shown in Fig. 1f. We used Ube2g2 precharged with chemically synthesized ubiquitin oligomers comprising one to four untagged wildtype ubiquitin moieties as the acceptor (Fig. 1g) and Ube2g2 carrying the F∧UbK48R mutant on its active site as the donor. In this experiment, the ligation of F∧UbK48R to the acceptor ubiquitin oligomer would further extend the chains by one ubiquitin moiety. Indeed, F∧UbK48R was efficiently transferred to the distal end of these ubiquitin oligomers, regardless of the chain length (Fig. 1h). Collectively, these data suggested that ubiquitin chains linked to the Ube2g2 active site are extended from the distal end, which supports the sequential addition model.
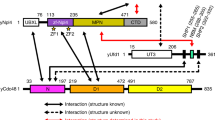
The gp78 G2BR domain is required for Ube2g2-gp78-mediated ubiquitination
We previously showed that Ube2g2 assembles di-ubiquitin in the absence of any E3s but that di-ubiquitin cannot be further extended to form longer ubiquitin chains28. These results suggest that an E3 is required to elongate polyubiquitin chains on the catalytic cysteine of Ube2g2. To further understand the role of E3 in this reaction, we dissected the function of gp78, the cognate E3 ligase of Ube2g2, using in vitro ubiquitination assays. gp78 contains several transmembrane (TM) segments in its N-terminal region and a p97-binding motif in the C-terminus, but these domains are not required for gp78-mediated ubiquitination reaction (data not shown). In addition, it has the following three other domains: (1) RING, (2) CUE and (3) G2BR. The RING domain is the catalytic domain essential for ubiquitination, the G2BR domain is a motif that binds Ube2g2 with high affinity and the CUE domain can bind to ubiquitin or ubiquitin-like domains. In this study, we applied a GST-tagged gp78 carboxy-terminal cytosolic domain (GST-gp78c), which contains the three domains mentioned above24.
The interaction of G2BR with Ube2g2 appears to be essential for building ubiquitin chains at the active site of Ube2g2 because mutating two Ube2g2 residues (L163 and L165), which are involved in G2BR binding, abolishes chain formation on Ube2g227. To further confirm that the interaction between Ube2g2 and G2BR is essential for polyubiquitin-chain-assembly, we created a GST-gp78c A593R/F597R mutant (referred to as G2BRA593R/F597R), the introduction of these two mutations did not affect its secondary structure and its binding to polyubiquitin chains (Figure S1a and S1b). However, according to the reported Ube2g2-G2BR structures, both of these two residues are involved in binding to Ube2g226,27,30. Indeed, the interaction of this mutant with Ube2g2 was significantly reduced compared with that of wildtype GST-gp78c (Fig. 2a). In the absence of a functional G2BR domain, the E3 activity of gp78 was almost completely abolished, as the mutant failed to promote ubiquitin discharging from Ube2g2 (Fig. 2b) in the single-round ubiquitin turnover assay (Fig. 2c). In addition, the G2BRA593R/F597R mutant failed to assemble polyubiquitin chains on the catalytic cysteine of the Ube2g2 (Fig. 2d) and to ubiquitinate the gp78 substrate Herp in vitro (Fig. 2e). These results indicated that the interaction between Ube2g2 and the G2BR domain of gp78 is essential for ubiquitin chain formation on the Ube2g2 active site and on Ube2g2 substrates.
The G2BR domain is required for Ube2g2-gp78 mediated polyubiquitination.
(a) The gp78 G2BRA593R/F597R mutant abolishes the interaction between gp78c and Ube2g2. Ube2g2 was pulled-down by the indicated gp78c variants with GST beads. (b) Defectivity of the G2BRA593R/F597R mutant in discharging F∧UbK48R from Ube2g2. WT ubiquitin was mixed with Ube2g2 precharged with F∧UbK48R in the presence of the indicated gp78c variants. The transfer of F∧UbK48R to ubiquitin was analyzed by immunoblotting with an anti-FLAG antibody. (c) Defectivity of the G2BRA593R/F597R mutant in the ubiquitin single-round turnover assay. Ube2g2 precharged with WT ubiquitin was mixed with Ube2g2 precharged with F∧UbK48R in the presence of the indicated gp78c variants. The transfer of F∧UbK48R to ubiquitin linked to the Ube2g2 active site was analyzed by immunoblotting with an anti-FLAG antibody. (d) Effects of the WT gp78c and the G2BRA593R/F597R mutant on polyubiquitin-chain-assembly on the Ube2g2 active site. The assembly of the Ube2g2-linked ubiquitin chains was determined at 37°C in the presence of E1, Ube2g2, FLAG-Ub and the indicated gp78 variants. Samples were taken at the indicated time points and polyubiquitin chains were detected by immunoblotting with an anti-FLAG antibody. (e) Effects of the WT gp78c and the G2BRA593R/F597R mutant on Herpc polyubiquitination. Herpc was incubated at 37°C in the presence of E1, Ube2g2, ATP, FLAG-Ub and the indicated gp78 variants. Samples were taken at the indicated time points and polyubiquitinated Herpc was detected by immunoblotting with an anti-RGS-His antibody. *, indicates Herpc dimerization. All of the western blots in this figure are representative blots from 2 replicates.
gp78 bound Ube2g2~Ub donor and relative free Ube2ge~Ub acceptor are required for ubiquitin chain formation
We previously showed that gp78 oligomerization is essential for forming ubiquitin chains on the Ube2g2 active site27. Conceptually, multiple G2BR domains in a gp78 oligomer can simultaneously recruit several Ube2g2s, which can serve as either donor or acceptor E2s (Model I). Ubiquitin transfer between the E2 molecules within a single gp78 oligomer may drive chain formation on the active site of an Ube2g2 molecule within the same complex. However, it is possible that ubiquitin is transferred between Ube2g2 molecules bound to two distinct gp78 oligomers to form active site-linked ubiquitin chains (Model II). Furthermore, it is also possible that only the donor (Model III) or the acceptor Ube2g2 (Model IV) molecule needs to form a high affinity interaction with gp78 (Fig. 3a).
Acceptor Ube2g2~Ub does not need to bind to the gp78 G2BR domain.
(a) Four possible models of E2 and E3 interaction in the asymmetric polyubiquitin chain assembly. Model I, Donor E2 and Acceptor E2 bind with the same gp78 oligomers; Model II, Donor E2 and Acceptor E2 bind with the neighboring gp78 oligomers separately; Model III, Donor E2 binds with gp78 oligomers, but Acceptor E2 does not; Model IV, Acceptor E2 binds with gp78 oligomers, but Donor E2 does not. (b) Ube2g2 L163A/L165A was defective in its donor activity, but not its acceptor function. (c) The acceptor Ube2g2~Ub did not need E3 in the single-round turnover assay. The transfer of F∧UbK48R to Ube2g2~Ub was analyzed by immunoblotting. G2BR M is a G2BRA593R/F597R mutant. Numbers on the gels show the relative amount of Ube2g2~Di-Ub, calculated from intensities of the bands (the intensity of the Ube2g2~Di-Ub band in lane 2 was set to 1.00). (d) Ubiquitin conjugated to Ube2g2 bound to gp78 can be efficiently transferred to the acceptor. (e) Once bound to the GST-gp78c G2BR domain, Ube2g2~Ub cannot accept ubiquitin from the donor. (f) The E2:E3 ratio of 2:1 gave the highest di-ubiquitin chain formation efficiency in the single-round turnover assay. Ube2g2 precharged with F∧UbK48R was mixed with Ube2g2 precharged with untagged wildtype Ub in the presence of different amounts of GST-gp78c. The transfer of F∧UbK48R to Ube2g2~Ub was analyzed by immunoblotting. (g) Quantification results of (f). (h) Extra inactive WT Ube2g2 (but not inactive Ube2g2 L163A/L165A) inhibited di-ubiquitin chain assembly due to its competitive binding with GST-gp78c. After pretreatment with NEM and EDTA at RT for 15 min, WT or L163A/L165A Ube2g2 was mixed with Ube2g2 precharged with F∧UbK48R and GST-gp78c on ice at different ratios compared with the donor. The donor was mixed with Ube2g2 precharged with untagged wildtype Ub. The transfer of F∧UbK48R to Ube2g2~Ub was analyzed by immunoblotting with an anti-FLAG (Ub) antibody. Western blots (b) to (f) and (h) in this figure are representative blots from 2 replicates.
To distinguish between these possibilities, we performed a single-round ubiquitin turnover assay using a Ube2g2 mutant (L163A/L165A) that cannot interact with the G2BR domain of gp78. By arbitrarily placing the mutant at either the donor or the acceptor position, we showed that the E2 mutant could not function as a ubiquitin donor but could still serve as an acceptor of ubiquitin from gp78-bound Ube2g2 molecules (Fig. 3b). We concluded that the donor Ube2g2~Ub needs to bind to the gp78 G2BR domain, whereas the acceptor E2 does not need to form a high affinity interaction with the gp78 G2BR domain, thus excluding the third possible model.
We next wished to test whether the acceptor Ube2g2 needed to physically contact the gp78 G2BR domain. The G2BRA593R/F597R mutant, which cannot bind to Ube2g2, was applied to the following modified single-round turnover assay: (1) Ube2g2 was charged with F∧UbK48R and then (2) incubated with an equal molar amount of GST-gp78c to form a complex of gp78c with the E2~ubiquitin (K48R) thioester. The reaction would not yield di-ubiquitin or longer ubiquitin chains because gp78-catalyzed ubiquitination is strictly dependent on the ubiquitin Lys48 residue. The resulting complex of E2~ubiquitin (K48R) thioester with gp78c (incubated in an ice bath for 30 min) then served as the donor by mixing with Ube2g2~Ub (WT) alone or in the presence of either wildtype gp78c or the G2BRA593R/F597R gp78c mutant. A short incubation was carried out to mitigate the confounding effect of gp78c dissociation from the donor E2~ubiquitin complex. Under our experimental conditions, dissociation of the gp78 oligomer did not occur, as demonstrated by the lack of E3 mixtures identified using two differently tagged gp78cs (Figure S2). Surprisingly, the first two reactions yielded similar amounts of di-ubiquitin on the Ube2g2 active site, but the third group showed a decreased amount of di-ubiquitin (Fig. 3c). Together, these results suggested that the acceptor Ube2g2~Ub may not need to bind to E3 at all, thus excluding the second possible model.
To further test whether Ube2g2~F∧UbK48R, which binds to gp78, could still be efficiently transferred to the acceptor, Ube2g2~F∧UbK48R was incubated in an ice bath for 30 min and pulled down by GST-gp78c with glutathione beads and then incubated with the acceptor Ube2g2~Ub complex. Active site-linked di-ubiquitin was efficiently formed, similarly to that in solution (Fig. 3d). In a reciprocal experiment, when the acceptor Ube2g2~Ub was immobilized on beads with the GST-gp78c RING mutant, its ability to accept ubiquitin from the donor Ube2g2 was completely lost (Fig. 3e). The GST-gp78c RING mutant lost its catalytic activity but could still interact with Ube2g2 (Figure S3) and polyubiquitin chains (Figure S1b) without changing its secondary structure (Figure S1a). These results suggested that Ube2g2~Ub bound to the gp78 G2BR domain cannot be used as an acceptor at all, thus excluding both the first and the second possible models.
Consistent with the fourth model, we found that a molar E2:E3 ratio of 2:1 gave the highest di-ubiquitin chain assembly efficiency and that excess gp78c actually inhibited rather than promoted active site associated polyubiquitin-chain-assembly (Fig. 3f–g). Presumably, under these conditions, all of the E2s are in the gp78-bound state and no functional Ube2g2 acceptors are available. In addition, according to the fourth model, if there were too much Ube2g2 to compete with the acceptor Ube2g2~Ub, the reaction would not occur, either. This prediction has been further demonstrated by the following experiments: with the addition of increasing amount of NEM/EDTA-pretreated WT Ube2g2, which was inactive in accepting ubiquitin from E1 but still could functionally interact with the gp78 G2BR domain, to the single round turnover assay, di-ubiquitin chain formation was inhibited significantly. While the NEM/EDTA-pretreated Ube2g2 L163A/L165A mutant, which lost its catalytic activity and could not bind to the gp78 G2BR domain, failed to do so (Figure S4 and Fig. 3h). Together, all of these results suggested that the reaction likely occurs between a gp78-bound Ube2g2~Ub donor and a relative free Ube2g2~Ub acceptor.
The gp78 G2BR domain promotes the E3 activity of gp78 in cis
Because the donor Ube2g2 needs to interact with both the RING and the G2BR domains of gp78 for proper function, we asked whether the donor Ube2g2~Ub binds the RING and G2BR domains in the same molecule (in cis,Fig. 4a) or in two neighboring gp78 molecules in a gp78 oligomer (in trans,Fig. 4b). If intermolecular cooperation between the G2BR and RING domains promotes gp78-mediated ubiquitination, combining a RING and a G2BR mutant into a single gp78 oligomer complex should rescue E3 activity at least in part. We then created a G2BR deletion mutant (G2BR Del), a RING mutant (RING M) and a RING/G2BR double mutant (RING M/G2BR Del). We co-expressed the RING M and G2BR Del mutants in bacteria and purified a gp78c complex containing both mutants (Fig. 4c). These hybrid E3 complexes showed similar characteristics to the WT E3 oligomers, such as its distribution in Superose 6 column (data not shown). Mutating either the RING or the G2BR domain resulted in a complete loss of gp78 activity, as demonstrated by the single-round turn over assay (Fig. 4d, lanes 5–8 and Figure S5 Lanes 5–8). The ‘hybrid complex’ containing both mutant proteins still could not assemble di-ubiquitin chains on the Ube2g2 active site (Fig. 4d, lanes 9–12). In contrast, the RING M/G2BR Del double mutant co-purified with wildtype gp78 impaired di-ubiquitin chain formation on the Ube2g2 active site by 50% compared with wildtype gp78 (Fig. 4d, lanes 13–16 & 1–4). These results suggested that the G2BR and RING domains need to function within the same gp78 molecule; in other words, the G2BR domain works with the RING domain in cis. To further distinguish whether model a or b was used in Ube2g2/gp78-mediated polyubiquitin chain assembly, we changed to a relatively simple system to dissect whether the RING domain and the G2BR domain within the same gp78 molecule (in cis, Fig. 4e) or in two neighboring gp78 molecules (in transFig. 4f) mediated the discharge of the donor ubiquitin to the accepter. F∧UbK48R was charged onto Ube2g2, quenched by adding NEM/EDTA and then added to the above mentioned E3 or combined E3 hybrid protein together with an excess amount of wildtype ubiquitin to allow the charged F∧UbK48R to be transferred to free ubiquitin to form free di-ubiquitin chains. Again, the co-purified RING plus G2BR mutant proteins failed to discharge F∧UbK48R and transfer it to free ubiquitin; however, in the presence of wild-type gp78, this reaction occurred (Fig. 4g). These results suggested that the RING and G2BR domains in the same gp78 molecule mediated donor ubiquitin discharging (Fig. 4e), thus supporting model a in Fig. 4.
The RING and G2BR domains from the same protein catalyze donor ubiquitin transfer to the acceptor Ube2g2~Ub and ubiquitin.
(a) The G2BR domain works with the RING domain in cis. (b) The G2BR domain works with the RING domain in trans. (c) gp78c variants and copurified gp78c complexes were analyzed by SDS-PAGE and stained with Coomassie. RING M is the RINGC341A/C375A mutant and G2BR Del is the gp78c truncation (1-510aar). (d) Co-purified GST-gp78c RING M + G2BR Del protein failed to assemble di-ubiquitin chains on the Ube2g2 active site. The amounts of the E3 complex in the four group experiments are equal. Numbers on the gels show the relative amounts of Ube2g2~Di-Ub, as calculated from intensities of the bands (the intensity of the Ube2g2~Di-Ub band in lane 2 was set to 1.00). (e)–(f) Similar to (a) and (b), but extra ubiquitin was used to accept F∧UbK48R from Ube2g2. (g) Copurified GST-gp78c RING M + G2BR Del protein failed to transfer F∧UbK48R from Ube2g2 to free wildtype ubiquitin. The amounts of E3 complex in the four group experiments are equal. Numbers on the gels show the relative amounts of the Di-Ub, as calculated from intensities of the bands (the intensity of the Di-Ub band in lane 2 was set to 1.00). Western blots (d) and (g) in this figure are representative blots from 2 replicates.
The gp78 CUE domain determines the processivity of polyubiquitin-chain-assembly
In addition to the G2BR domain, gp78 contains a coupling of ubiquitin conjugation to ER degradation (CUE) domain. Although the CUE domain is essential for the biological function of gp7831,32, its mechanistic role(s) remain unknown. Recently, the structure of the gp78 CUE domain in complex with ubiquitin was determined and the results hint that the CUE domain might be involved in polyubiquitin chain elongation33.
To test the role of the CUE domain in polyubiquitin-chain-assembly, we created three CUE domain mutants (M467A/F468A, D479A/L480A and I493A/L494A) by site-directed mutagenesis. These residues were selected because they are highly conserved among various CUE domains and they either stabilize the folding of the gp78 CUE domain or form important contacts with ubiquitin33. Interestingly, the ubiquitin ligase activity of these mutants was similar to that of wildtype gp78, as demonstrated by their abilities to discharge F∧UbK48R from the cysteine of the Ube2g2 active site to free Ub or by their single-round ubiquitin turnover assay results (Figure S6). However, the processivity of polyubiquitin-chain-assembly on either the Ube2g2 active catalytic cysteine of Ube2g2 or the model substrate Herpc was significantly affected by the mutations in the CUE domain (Fig. 5a–b). To further study the effects of the CUE domain mutants on the processivity of polyubiquitin-chain-assembly, we then applied these CUE domain mutants to the single-round ubiquitin turnover assay. Similarly to the experiment in Fig. 1f, Ube2g2 precharged with chemically synthesized ubiquitin oligomers comprising one to four untagged wildtype ubiquitin moieties were used as acceptors and Ube2g2 carrying a F∧UbK48R monomer on its active site was used as the donor. After adding E3 to the reaction buffer, the wildtype GST-gp78c could efficiently transfer the donor ubiquitin to the distal end of the mono- to tetra-ubiquitin chains, which had been precharged on Ube2g2. Although the CUE mutants still could efficiently transfer F∧UbK48R from the donor Ube2g2 to the acceptor Ube2g2~Ub(n) (n = 1 or 2) (Fig. 5c, lanes 2–3; 7–8 and data not shown), the transfer activity reduced with the elongation of the chain (Fig. 5c, lanes 4–5; 9–10 and data not shown). To confirm the processivity defect of the CUE mutants in longer polyubiquitin-chain-assembly, chemically synthesized ubiquitin oligomers (Ub2-7) were successfully charged onto Ube2g2 as acceptors (Fig. 5d–e). They were used to compare the polyubiquitin-chain-assembly activity of WT GST-gp78c and the CUED479A/L480A mutant by the single-round ubiquitin turnover assay. Consistent with previous results, although the donor ubiquitin could be efficiently transferred to Ube2g2~Ub2, the elongation of Ube2g2~Ub(n) (n ≥ 3) catalyzed by the CUED479A/L480A mutant was significantly reduced compared with the wildtype E3 (Fig. 5f–g). Together, these results established that the gp78 CUE domain participates in polyubiquitin-chain-assembly by improving the processivity of ubiquitination.
The gp78 CUE domain determines the processivity of polyubiquitin-chain-assembly.
(a) Effect of the gp78 CUED479A/L480A mutant on the assemblage of polyubiquitin chains on the E2 active site. In vitro ubiquitination was performed using the indicated E3s incubated at 37°C. Samples were taken at the indicated time points and the polyubiquitin chains were detected by immunoblotting with an anti-FLAG antibody under non-reducing conditions. (b) Effect of the gp78 CUED479A/L480A mutant on Herpc polyubiquitination. Herpc was incubated at 37°C in the presence of E1, Ube2g2, ATP, FLAG-Ub and the indicated gp78 variants. Samples were taken at the indicated time points and the polyubiquitinated Herpc was detected by immunoblotting with an anti-RGS-His antibody. *, indicates Herpc dimerization. (c) As indicated in Figure 1f, Ube2g2 precharged with the indicated ubiquitin oligomers (Ub1-4) was mixed with Ube2g2 precharged with F∧UbK48R in the presence of WT and CUED479A/L480A GST-gp78c. The transfer of F∧UbK48R to ubiquitin oligomers linked to the Ube2g2 active site was analyzed by immunoblotting. *, indicates Ube2g2 conjugated with two F∧UbK48Rs that was generated in the precharging reaction. (d) The experimental scheme of (f). (e) The charging of ubiquitin oligomers to Ube2g2. Charging reactions were performed with E1, Ube2g2, ATP and ubiquitin oligomers carrying 2-7 ubiquitin moieties (Ub2-7). The charging efficiency was analyzed by immunoblotting with an anti-Ube2g2 antibody under non-reducing condition. (f) The transfer of F∧UbK48R to ubiquitin oligomers (2-7) linked to the Ube2g2 active site was analyzed by immunoblotting. *, indicates Ube2g2 conjugated with two F∧UbK48Rs that was generated in the precharging reaction. (g) Quantification of the experiment shown in (f). Error bars, standard error (n = 2). (h) Effect of the gp78 CUE mutants on the interaction with polyubiquitin chains. Polyubiquitin chains were mixed with the GST-gp78c variants and incubated in an ice bath for 60 min. The mixture was pulled-down with GST beads at 4°C and then washed 3 times with IP buffer. Samples were detected by immunoblotting with anti-GST (gp78) and anti-FLAG (Ub) antibodies. Western blots (a) to (c), (f) and (h) in this figure are representative blots from 2 replicates.
We then asked why those CUE mutants are defective in elongating the ubiquitin chain but do not affect chain initiation. One possibility is that the CUE domain prefers to bind longer ubiquitin chains but binds to mono-ubiquitin with a lower affinity. To test this possibility, we synthesized polyubiquitin chains on the active site of Ube2g2 using gp78 in vitro. Once the reaction was finished, DTT was added to reduce the ubiquitin chains from Ube2g2. The samples were then heated to precipitate E1, E2 and E3. Following centrifugation, the free ubiquitin chains were used to test their interaction with either WT GST-gp78c or the CUE mutants. In a GST pull-down experiment, longer ubiquitin chains were preferentially co-precipitated with wild-type GST-gp78c (Fig. 5h, compare lanes 2 and 7), but this trend was completely abrogated in all three CUE mutants (Fig. 5h, compare lanes 7 and 8–10). These results suggested that the gp78 CUE domain prefers to bind longer ubiquitin chains and that this binding might help to position the last ubiquitin in the right orientation for attacking the thioester-bond at the donor Ube2g2 active site.
Discussion
Our previous study demonstrates that polyubiquitination can be achieved by ligating a ubiquitin chain preassembled on an Ube2g2 active site cysteine to a substrate24 and that Ube2g2 dimerization allows the E2 to effectively engage two ubiquitins to specifically synthesize Lys48-linked short ubiquitin chains28. In this study, we further dissected the functional roles of two gp78 domains in the polyubiquitin-chain-assembly on the E2 active site.
Because the E2 or E3 active site cysteine-linked ubiquitin chain is the key intermediate in the seesaw model, the recent discovery that Ube2g2 and Ubc7 allow ubiquitin chains to be preassembled on their active site cysteines before transferring these chains to substrates22,23,24,25 sheds light on this amazing model. Unexpectedly, we found that even a di-ubiquitin molecule conjugated to the Ube2g2 active site cannot be transferred to another Ube2g2~ubiquitin complex to form tri-ubiquitin-linked Ube2g2 (Fig. 1d) and that Ube2g2-linked ubiquitin oligomers containing one to four ubiquitin moieties can all serve as acceptors to efficiently receive additional ubiquitin molecules from a donor E2~ubiquitin complex (Fig. 1g). Together, these data indicate that an Ube2g2 active site-linked ubiquitin chain is likely built by the sequential incorporation of ubiquitin molecules onto the distal end of the chain. This conclusion is in line with recent computer modeling studies suggesting that, in theory, ubiquitin can be added to the distal ends of ubiquitin chains of varying lengths, given the geometric flexibility of Lys48-linked ubiquitin chains15,18,19,34.
In contrast to most other E3 ligases, which do not have strong interactions with their corresponding E2, gp78 has an E2 binding domain, G2BR, which binds Ube2g2 with very high affinity26,27,30. Although we and others have shown that this domain is essential for polyubiquitin-chain-assembly on either the E2 or the substrate26,27, the detailed role of the G2BR domain in polyubiquitin-chain-assembly on the Ube2g2 active site remains elusive. Here, we found that only the donor Ube2g2~Ub needs to bind to the gp78 G2BR domain and that the acceptor Ube2g2~Ub moiety does not need to firmly bind to gp78 at all. Although it has been reported that even G2BR peptides can promote RING domain activity26, we found that G2BR promotes gp78 RING assembly of the polyubiquitin chain on the Ube2g2 active site in cis. The difference between our results and those published by Das et al. might be explained by the following two possibilities: (1) the readout that they used was the autoubiquitination of gp78 and the mechanism underlying this autoubiquitination may be completely different from the polyubiquitin-chain-assembly on the Ube2g2 active site or (2) too many G2BR peptides were used in their experiment, suggesting a crowded molecular environment in which G2BR may also work in trans. In physiological conditions, it is unlikely to have so much G2BR in the reaction. Thus, we favor a model where the RING domain with the G2BR domain in the same gp78 molecule catalyzes the discharging of the donor ubiquitin. The dimerization of Ube2g2~Ub and/or a neighboring RING domain together with the donor Ube2g2~Ub mediates the acceptor Ube2g2~Ub to orientate itself into the correct position to form di-ubiquitin chains on the Ube2g2 active site (Fig. 6).
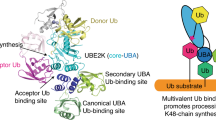
A model for Ube2g2-gp78-mediated polyubiquitin-chain-assembly on the Ube2g2 active site.
The gp78 dimer shown in this figure comprises two neighboring gp78 molecules in a gp78 oligomer. (1), The initiation of ubiquitin chain synthesis. The dimerization of Ube2g2~Ub and/or a neighboring RING domain with the donor Ube2g2~Ub mediates the acceptor Ube2g2~Ub to orientate itself into the correct position to form the di-ubiquitin chain on the active site of Ube2g2. (2), Elongation of the ubiquitin chain. The gp78 CUE domain might recruit and correctly position the distal end of the ubiquitin moiety of the growing polyubiquitin chain for an attack on the thioester-linked ubiquitin at the active site of Ube2g2 to promote polyubiquitin chain elongation.
The CUE domain is a type of ubiquitin binding domain that comprises three helices31,33. Similar structures exist in mammalian E2-25K and yeast Ubc1. The UBA of Ubc1 has been shown to affect both the length and the ubiquitin-ubiquitin linkage specificity of the chain35,36. Thus, it has been speculated that noncovalent binding of a second ubiquitin molecule to the E2 might dictate certain features of ubiquitin-chain assembly. However, the functional role(s) of the CUE domain in an E3 such as gp78 in the polyubiquitin-chain-assembly on the Ube2g2 active site remain enigmatic. In the current study, we show that the gp78 CUE domain participates in the processivity of polyubiquitin-chain-assembly by preferentially binding to longer ubiquitin chains. Thus, similar to the UBA in Ubc1, the gp78 CUE domain might recruit and correctly position the distal end of the ubiquitin moiety of the growing polyubiquitin chain for attacking the thioester-linked ubiquitin on the Ube2g2 active site to promote polyubiquitin chain elongation (Fig. 6).
It has been shown previously that the gp78 CUE domain is involved in the polyubiquitination of protein fused to ubiquitin in vitro and the pathogenic CFTRΔ508 after initial ubiquitination by another ERAD E3 RMA1/RNF5 in vivo26,37. Thus, once one E3 initiates substrate ubiquitination, substrates bearing one or more ubiquitin moieties may be recruited to the gp78 CUE domain for polyubiquitination. As for Ube2g2-gp78-mediated polyubiquitination on the active site of Ube2g2, the initiation and elongation of the polyubiquitin chain on the E2 are catalyzed by a single E3: gp78. According to our results, together with others38, we propose a polyubiquitin-chain-assembly model in which, upon capture by G2BR, Ube2g2~Ub induces allosteric effects and increases its affinity for the RING domain in cis. The interaction between the RING domain and Ube2g2~Ub promotes the formation of di-ubiquitin chains on the acceptor Ube2g2 and, concurrently, impairs Ube2g2-G2BR at multiple contact sites, thus promoting the rapid release of Ube2g2 from gp78. Upon elongation of the ubiquitin chain, Ube2g2~Ub(n) (where n≥3) is recruited by the CUE domain of a neighboring gp78 molecule, which is in a oligomerized state and the distal ubiquitin of the chain is well positioned in the correct orientation for an attack on the thioester-linked ubiquitin at the Ube2g2 active site, which has been loaded onto the previous G2BR domain with the help of the RING domain. Then, an additional ubiquitin is added to the distal end of the polyubiquitin chain and with each cycle of ubiquitination, a new ubiquitin molecule is added to the end of the growing polyubiquitin chain. Thus, a dynamic interaction between the donor Ube2g2~Ub and gp78 G2BR domain and a very stable interaction between the acceptor Ube2g2~Ub(n) (where n≥3) and the gp78 CUE domain cooperatively drive the processivity of polyubiquitin-chain-assembly on the Ube2g2 active site (Fig. 6). This model of polyubiquitin-chain-assembly is consistent with the notion that the initiation and elongation of polyubiquitin chain formation are governed by different mechanisms in the Cdc34-SCF complex-mediated polyubiquitination system14,39. Basically, the gp78 G2BR domain mainly promotes the initiation of ubiquitination and the CUE domain is in charge of the processivity of ubiquitination in an E4-like manner.
The combination of the initiation and elongation of the polyubiquitin chain into a single E3 ligase has obvious advantages compared with those other examples in which one E2-E3 pair monoubiquitinates the target protein and another pair subsequently assembles the polyubiquitin chains16,20,40,41. The switch between different E2-E3 pairs might provide new layers of regulation of the modification, but this inevitably slows down the processivity of the ubiquitination of the substrate. In yeast, Cue1p contains a CUE domain and a U7BR domain and they are very similar to the gp78 CUE and G2BR domains, respectively, in both structure and function30,31. Thus, gp78 may have emerged as a fusion of the Hrd1 and Cue1 genes during evolution. The fusion of these two domains into a single ERAD E3 ligase greatly enhances the efficiency of the polyubiquitin-chain-assembly by this E3 ligase.
Methods
Antibodies and proteins
The His and GST antibodies were purchased from Abmart (Beijing, China). The GAPDH antibody was purchased from Sanjian (Tianjin, China). The FLAG and RGS-His antibodies were purchased from Abmart and Qiagen, respectively. The E1, FLAG-Ub, WT Ub, Lys48-linked Ub2, Ub3, Ub4 and Ub2-7 proteins were purchased from Boston Biochem (Cambridge, MA, USA).
Plasmids, strains and cell lines
The pET28-Ube2g2, pGEX-gp78c, pET28-flag-UbK48R and pET3-Ub plasmids were described previously24. The GST-gp78c fragment was amplified and cloned into the pET28a plasmid. gp78 was amplified and cloned into the pcDNA3.1 plasmid. pET28-Ube2g2 and pGEX-gp78c single/double/multiple point(s) mutants were generated by site-directed mutagenesis. The pGEX-gp78c G2BR deletion mutant, pET28-GST-gp78c RING mutant, pET28-GST-gp78c RING mutant and G2BR deletion double mutant were described previously27. All of the plasmids were amplified in E. Coli Top10 cells and the recombinant proteins were expressed in E. Coli BL 21 or Rosetta 2 cells.
Protein purification
HF∧UbK48R, his-Ub, Ube2g2 and gp78c were purified as previously described42. The purified E2 and E3 variants were further fractionated by size exclusion chromatography on Superdex75 and Superose 6 columns, respectively, in a buffer containing 50 mM Tris-HCl, pH 8.0, 150 mM potassium chloride, 5% glycerol and 2 mM magnesium chloride. For copurification of the hybrid E3 complex, His- and GST-tagged gp78c mutants were coexpressed in E. Coli. Rosetta 2 cells and copurified via a two-step (His and GST) affinity pull down. The purified hybrid E3 complexes were further fractionated by size exclusion chromatography on a Superose 6 column.
In vitro ubiquitination assay
The ubiquitin charging experiment was previously described24. E1 (60 nM) and UBE2G2 (E2, 300 nM) were incubated with FLAG-tagged Ub (10 µM) at 37°C in the reaction buffer (25 mM Tris-HCl, pH 7.4, 2 mM magnesium/ATP). Samples taken at the indicated time points were mixed with Laemmli buffer in the absence or presence of a reducing reagent. The polyubiquitination assay was conducted using the same conditions as described above with the addition of purified E3 gp78c (300 nM) and the substrate Herpc (300 nM).
Free polyubiquitin chain preparation
Polyubiquitin chains were synthesized using the in vitro ubiquitination assay. Upon completion of the reaction, 5 mM DL-dithiothreitol (DTT) was added to reduce the polyubiquitin chains from Ube2g2. The samples were then heated at 85°C for 15 min to precipitate E1, E2 and E3. Following centrifugation, the free polyubiquitin chains were used to test their interaction with either WT GST-gp78c or CUE mutants.
Synthesis of ubiquitin oligomers
Ubiquitin oligomers carrying F∧UbK48R at the distal end were synthesized in vitro using a protocol adapted from a previously published one43. Briefly, Ub (D77) and F∧UbK48R (8 mg/ml each) were incubated with 20 µM E2-25K in a buffer containing 25 mM Tris-HCl, pH 7.4, 2 mM magnesium, 0.1 mM DTT and an ATP regenerating system. The reaction was initiated by the addition of 0.1 µM mammalian E1 and incubated at 37°C for 3 hrs. At the end of the incubation, the reaction was treated with 1 mM DTT and 1 mM EDTA at 25°C for 20 min. Yuh1 was then added to the reaction to a final concentration of 20 µg/ml and the reaction was further incubated at 37°C for 1 hr. Acetic acid (200 µl/reaction, 2 N) was added to the reaction to adjust the pH to ~4. The acidified reaction mixture was applied to a Resource S column equilibrated with buffer A (50 mM ammonium acetate, pH 4.5, 1 mM EDTA and 5 mM DTT). The column was washed with 3 column volumes of buffer A and the bound proteins were eluted with 10 column volumes of buffer A containing a linear sodium chloride gradient (0–1.0 M). The fractions were collected and examined by immunoblotting with the FLAG antibody. The reaction yielded mostly di-ubiquitins with F∧UbK48R at the distal end. Because a small fraction of Ub (D77) appeared to have the Asp77 residue removed by carboxypeptidase44, a small amount of tri-ubiquitin with F∧UbK48R at the distal end was also obtained. The fractions enriched with di-Ub and tri-Ub, respectively, were used in the single-round ubiquitin turnover experiments.
Single-round ubiquitin turnover and discharging assays
The single-round ubiquitin turnover and discharging assays were described previously. To detect the single-round turnover of Ub by gp78 or its variants, Ube2g2 (300 nM) was incubated with E1 (60 nM), together with either F∧UbK48R (as a donor) or WT Ub (K48-Ub2, K48-Ub3, K48-Ub4 or K48-Ub2-7, as an acceptor), at 37°C for 15 min in reaction buffer (25 mM Tris-HCl, pH 7.4, 2 mM magnesium/ATP and 0.1 mM DTT). The reaction was treated with 50 mM EDTA and 10 mM NEM at room temperature for 15 min. Equal volumes of the donor and acceptor were mixed into the reaction and catalyzed by GST-gp78c or its variants at 37°C for 15 min. The reaction was stopped by the addition of Laemmli buffer and analyzed by immunoblotting with anti-FLAG antibodies. The discharging assay was performed using the same conditions as described above, except that free wild-type ubiquitin was used as an acceptor.
Protein interaction assay (GST pull down)
The GST-gp78c variants were incubated with GST beads at 4°C for 2 hrs (GST protein was used as a negative control). The GST beads were centrifuged and washed with IP buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl and 0.1% Triton) 3 times. Then, Ube2g2 was added to the beads and they were incubated at 4°C for 2 hrs. The beads were centrifuged and washed 3 times with IP buffer and then SDS loading buffer was added. The samples were heated at 65°C for 5 min. After centrifuging the beads, the supernatants were loaded onto SDS-PAGE and detected by immunoblotting.
Circular dichroism spectroscopy (CD)
To determine the secondary structure of gp78 variants, the CD spectra of the proteins were taken at 0.2 mg/ml in 50 mM Tris, using a Chirascan Plus CD spectrometer (Applied Photophysics, Surrey, UK) and a 1 mm quartz sample cell equilibrated at 20°C. Scans between 200 and 260 nm were performed at a scan rate of 50 nm per min.
Polyubiquitin chain pull-down by GST-gp78c variants
The polyubiquitin chains were mixed with the GST-gp78c variants and incubated in an ice bath for 60 min. The mixture was incubated with GST beads at 4°C for 2 hrs and then washed 3 times with IP buffer. The samples were detected by immunoblotting.
References
Mukhopadhyay, D. & Riezman, H. Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science 315, 201–5 (2007).
Weissman, A. M. Themes and variations on ubiquitylation. Nat Rev Mol Cell Biol 2, 169–78 (2001).
Welchman, R. L., Gordon, C. & Mayer, R. J. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat Rev Mol Cell Biol 6, 599–609 (2005).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol 22, 159–80 (2006).
Pickart, C. M. & Eddins, M. J. Ubiquitin: structures, functions, mechanisms. Biochim Biophys Acta 1695, 55–72 (2004).
Hochstrasser, M. Lingering mysteries of ubiquitin-chain assembly. Cell 124, 27–34 (2006).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu Rev Biochem 67, 425–79 (1998).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat Biotechnol 21, 921–6 (2003).
Kulathu, Y. & Komander, D. Atypical ubiquitylation - the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat Rev Mol Cell Biol 13, 508–23 (2012).
Chau, V. et al. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243, 1576–83 (1989).
Haglund, K. & Dikic, I. Ubiquitylation and cell signaling. EMBO J 24, 3353–9 (2005).
Hicke, L. & Dunn, R. Regulation of membrane protein transport by ubiquitin and ubiquitin-binding proteins. Annu Rev Cell Dev Biol 19, 141–72 (2003).
Thrower, J. S., Hoffman, L., Rechsteiner, M. & Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. EMBO J 19, 94–102 (2000).
Deffenbaugh, A. E. et al. Release of ubiquitin-charged Cdc34-S - Ub from the RING domain is essential for ubiquitination of the SCF(Cdc4)-bound substrate Sic1. Cell 114, 611–22 (2003).
Kleiger, G., Saha, A., Lewis, S., Kuhlman, B. & Deshaies, R. J. Rapid E2-E3 assembly and disassembly enable processive ubiquitylation of cullin-RING ubiquitin ligase substrates. Cell 139, 957–68 (2009).
Koegl, M. et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96, 635–44 (1999).
Petroski, M. D. & Deshaies, R. J. Function and regulation of cullin-RING ubiquitin ligases. Nat Rev Mol Cell Biol 6, 9–20 (2005).
Petroski, M. D., Kleiger, G. & Deshaies, R. J. Evaluation of a diffusion-driven mechanism for substrate ubiquitination by the SCF-Cdc34 ubiquitin ligase complex. Mol Cell 24, 523–34 (2006).
Pierce, N. W., Kleiger, G., Shan, S. O. & Deshaies, R. J. Detection of sequential polyubiquitylation on a millisecond timescale. Nature 462, 615–9 (2009).
Rodrigo-Brenni, M. C. & Morgan, D. O. Sequential E2s drive polyubiquitin chain assembly on APC targets. Cell 130, 127–39 (2007).
Wickliffe, K. E., Lorenz, S., Wemmer, D. E., Kuriyan, J. & Rape, M. The mechanism of linkage-specific ubiquitin chain elongation by a single-subunit E2. Cell 144, 769–81 (2011).
Bazirgan, O. A. & Hampton, R. Y. Cue1p is an activator of Ubc7p E2 activity in vitro and in vivo. J Biol Chem 283, 12797–810 (2008).
Cao, J. et al. Ufd1 is a cofactor of gp78 and plays a key role in cholesterol metabolism by regulating the stability of HMG-CoA reductase. Cell Metab 6, 115–28 (2007).
Li, W., Tu, D., Brunger, A. T. & Ye, Y. A ubiquitin ligase transfers preformed polyubiquitin chains from a conjugating enzyme to a substrate. Nature 446, 333–7 (2007).
Ravid, T. & Hochstrasser, M. Autoregulation of an E2 enzyme by ubiquitin-chain assembly on its catalytic residue. Nat Cell Biol 9, 422–7 (2007).
Das, R. et al. Allosteric activation of E2-RING finger-mediated ubiquitylation by a structurally defined specific E2-binding region of gp78. Mol Cell 34, 674–85 (2009).
Li, W. et al. Mechanistic insights into active site-associated polyubiquitination by the ubiquitin-conjugating enzyme Ube2g2. Proc Natl Acad Sci U S A 106, 3722–7 (2009).
Liu, W. et al. Dimeric Ube2g2 simultaneously engages donor and acceptor ubiquitins to form Lys48-linked ubiquitin chains. EMBO J 33, 46–61 (2014).
Chen, B. et al. The activity of a human endoplasmic reticulum-associated degradation E3, gp78, requires its Cue domain, RING finger and an E2-binding site. Proc Natl Acad Sci U S A 103, 341–6 (2006).
Metzger, M. B. et al. A structurally unique E2-binding domain activates ubiquitination by the ERAD E2, Ubc7p, through multiple mechanisms. Mol Cell 50, 516–27 (2013).
Bagola, K. et al. Ubiquitin binding by a CUE domain regulates ubiquitin chain formation by ERAD E3 ligases. Mol Cell 50, 528–39 (2013).
Tsai, Y. C. et al. The ubiquitin ligase gp78 promotes sarcoma metastasis by targeting KAI1 for degradation. Nat Med 13, 1504–9 (2007).
Liu, S. et al. Promiscuous interactions of gp78 E3 ligase CUE domain with polyubiquitin chains. Structure 20, 2138–50 (2012).
Ryabov, Y. & Fushman, D. Interdomain mobility in di-ubiquitin revealed by NMR. Proteins 63, 787–96 (2006).
Merkley, N., Barber, K. R. & Shaw, G. S. Ubiquitin manipulation by an E2 conjugating enzyme using a novel covalent intermediate. J Biol Chem 280, 31732–8 (2005).
Pichler, A. et al. SUMO modification of the ubiquitin-conjugating enzyme E2-25K. Nat Struct Mol Biol 12, 264–9 (2005).
Morito, D. et al. Gp78 cooperates with RMA1 in endoplasmic reticulum-associated degradation of CFTRDeltaF508. Mol Biol Cell 19, 1328–36 (2008).
Das, R. et al. Allosteric regulation of E2:E3 interactions promote a processive ubiquitination machine. EMBO J 32, 2504–16 (2013).
Varelas, X., Stuart, D., Ellison, M. J. & Ptak, C. The Cdc34/SCF ubiquitination complex mediates Saccharomyces cerevisiae cell wall integrity. Genetics 174, 1825–39 (2006).
Christensen, D. E., Brzovic, P. S. & Klevit, R. E. E2-BRCA1 RING interactions dictate synthesis of mono- or specific polyubiquitin chain linkages. Nat Struct Mol Biol 14, 941–8 (2007).
Ulrich, H. D. & Jentsch, S. Two RING finger proteins mediate cooperation between ubiquitin-conjugating enzymes in DNA repair. EMBO J 19, 3388–97 (2000).
Ye, Y., Meyer, H. H. & Rapoport, T. A. Function of the p97-Ufd1-Npl4 complex in retrotranslocation from the ER to the cytosol: dual recognition of nonubiquitinated polypeptide segments and polyubiquitin chains. J Cell Biol 162, 71–84 (2003).
Raasi, S. & Pickart, C. M. Ubiquitin chain synthesis. Methods Mol Biol 301, 47–55 (2005).
Pickart, C. M. & Raasi, S. Controlled synthesis of polyubiquitin chains. Methods Enzymol 399, 21–36 (2005).
Acknowledgements
We would like to thank Yihong Ye for critically reading and editing this manuscript. This work was supported by the National Natural Science Foundation of China (Grant No. 30970603), the Major Basic Research Program (Grant No. 2012CB944404) and the Knowledge Innovation Program (Grant No. KSCX2-YW-N-071) and the One Hundred Talents Program of the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
L.W.X. planned and performed most of the experiments and helped to write the manuscript. S.Y.L. performed and analyzed some of the biochemical experiments. L.W. designed the study and wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemental Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Liu, W., Shang, Y. & Li, W. gp78 elongates of polyubiquitin chains from the distal end through the cooperation of its G2BR and CUE domains. Sci Rep 4, 7138 (2014). https://doi.org/10.1038/srep07138
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07138
This article is cited by
-
Neuroleukin/Autocrine Motility Factor Receptor Pathway Promotes Proliferation of Articular Chondrocytes through Activation of AKT and Smad2/3
Scientific Reports (2015)
-
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.