Abstract
The centromere-specific histone H3 variant, CENP-A, is overexpressed in particular aggressive cancer cells, where it can be mislocalized ectopically in the form of heterotypic nucleosomes containing H3.3. In the present study, we report the crystal structure of the heterotypic CENP-A/H3.3 particle and reveal its “hybrid structure”, in which the physical characteristics of CENP-A and H3.3 are conserved independently within the same particle. The CENP-A/H3.3 nucleosome forms an unexpectedly stable structure as compared to the CENP-A nucleosome and allows the binding of the essential centromeric protein, CENP-C, which is ectopically mislocalized in the chromosomes of CENP-A overexpressing cells.
Similar content being viewed by others
Introduction
In eukaryotes, chromatin compacts genomic DNA with the nucleosome, as the basic repeating unit. Histones H2A, H2B, H3 and H4 are the protein components of the nucleosome. The histone octamer, containing two each of histones H2A, H2B, H3 and H4, left-handedly wraps about 150 base pairs of DNA around itself in the nucleosome1,2. In addition to the canonical types of histones, non-allelic isoforms of histones H2A, H2B and H3 have been identified as histone variants in many species3 and are suggested to have specific functions in the regulation and maintenance of genomic DNA in chromatin4,5,6.
In humans, eight types of histone H3 genes, encoding H3.1, H3.2, H3.3, H3T (also named H3.4), H3.5, H3.X, H3.Y and CENP-A (also named CenH3), have been identified3,7. H3.1, H3.2 and H3.3 are constantly produced in somatic cells. H3.1 and H3.2 are the canonical types of histone H3 and are expressed in the S-phase of the cell cycle8. On the other hand, H3.3 is continuously produced throughout the cell cycle. During the transcription and DNA repair processes, H3.3 functions as a replacement histone that is incorporated into chromatin regions by the HIRA histone chaperone, after the depletion of the canonical H3.18,9,10. In addition, H3.3 is specifically localized in the telomeric and pericentromeric regions of chromosomes, by a complex containing the histone chaperone DAXX and the nucleosome remodeler ATRX proteins11,12. Therefore, H3.3 may have dual functions as a replacement histone and an architectural component of distinct chromosome domains.
Among the histone H3 variants, CENP-A is the most distant isoform and is conserved from yeasts to humans. CENP-A specifically localizes to centromeres and epigenetically specifies the sites for kinetochore assembly13,14,15,16,17,18. CENP-A expression is tightly controlled in normal cells and its chromosome localization is strongly restricted within centromeric regions. However, CENP-A is reportedly overexpressed in particular aggressive cancer cells4,19,20,21,22. In Drosophila cells, overexpressed CENP-A (CID) is mislocalized into noncentromeric euchromatic regions and leads mitotic delays, anaphase bridges and chromosome fragmentation23. Similarly, in human cells, CENP-A overexpression can lead to its ectopic localization to chromosome regions with active histone turnover, as shown in cancer cell lines19,24. At these ectopic loci, CENP-A forms heterotypic nucleosomes, containing one each of the histone H3 variants, CENP-A and histone H3.3, within a single nucleosome24. These heterotypic nucleosomes occlude CTCF binding and their presence may increase DNA damage tolerance in cancer cells24. However, the structural features of the heterotypic CENP-A/H3.3 nucleosome are unknown.
In the present study, we report the crystal structure of the human CENP-A/H3.3 nucleosome at 2.67 Å resolution. The CENP-A/H3.3 nucleosome forms a “hybrid structure”, in which the physical characteristics of CENP-A and H3.3 are independently conserved within the same nucleosome. Unexpectedly, we found that the heterotypic CENP-A/H3.3 nucleosome is more stable than the CENP-A nucleosome and its stability is very similar to that of the H3.3 nucleosome. In addition, the CENP-A/H3.3 nucleosome retains the ability to bind to the essential centromeric protein, CENP-C, whose ectopic localization may also be harmful for the proper regulation of cell division and chromosome integrity.
Results and Discussion
Preparation of the heterotypic CENP-A/H3.3 nucleosome
To decipher its structural features, we reconstituted the heterotypic CENP-A/H3.3 nucleosome using purified histones. For this purpose, we established a method to prepare this heterotypic nucleosome (Fig. 1a), based on the method for the heterotypic nucleosome reconstitution with yeast H2A and FLAG-tagged HtZ25. Human histones H2A, H2B, H3.3, H4 and CENP-A were prepared as described previously26,27 and histone H3.3 was also prepared as a His6-SUMO tagged protein expressed in bacteria (Supplementary Fig. 1a). The nucleosomes were then reconstituted on a 146 base-pair palindromic DNA1 by the salt-dialysis method (Fig. 1b, lane 1). In this procedure, three types of nucleosomes were simultaneously formed: two homotypic nucleosomes with either two CENP-As or two His6-SUMO-H3.3s and a heterotypic nucleosome containing one CENP-A and one His6-SUMO-H3.3. In non denaturing polyacrylamide gel electrophoresis (native PAGE), the heterotypic CENP-A/His6-SUMO-H3.3 nucleosome migrated faster than the homotypic His6-SUMO-H3.3/His6-SUMO-H3.3 nucleosome, but slower than the homotypic CENP-A/CENP-A nucleosome, which allowed us to isolate the CENP-A/His6-SUMO-H3.3 nucleosome by preparative native PAGE (Fig. 1b). After the purification, the His6-SUMO portion of the His6-SUMO-H3.3 fusion protein was proteolytically removed by PreScission protease treatment (Supplementary Fig. 1b, lanes 1–2). We further purified the heterotypic CENP-A/H3.3 nucleosome by a second round of preparative native PAGE (Fig. 1c and Supplementary Fig. 1b). The purified heterotypic CENP-A/H3.3 nucleosome contained all five histones, H2A, H2B, H3.3, H4 and CENP-A (Fig. 1d).
Preparation of the CENP-A/H3.3 nucleosome.
(a) Schematic representation of the CENP-A/His6-SUMO-H3.3 nucleosome preparation and the CENP-A/H3.3 nucleosome preparation with the His6-SUMO removal step. The CENP-A and His6-SUMO-H3.3 molecules are colored red and blue, respectively. (b) The reconstituted nucleosomes (lane 1) were separated by native PAGE with the Prep Cell apparatus. Purified nucleosome fractions were analyzed by 6% native PAGE with ethidium bromide staining. The fractions shown in lanes 5–8 were collected. (c) The purified H3.3, CENP-A and CENP-A/H3.3 nucleosomes were analyzed by 6% PAGE with ethidium bromide staining. (d) The protein compositions of the CENP-A/His6-SUMO-H3.3 nucleosome (lane 2) and the CENP-A/H3.3 nucleosome (lane 3) were analyzed by 18% SDS-PAGE with Coomassie Brilliant Blue staining.
The structural characteristics of CENP-A and H3.3 are conserved independently within the heterotypic CENP-A/H3.3 nucleosome
We then determined the crystal structure of the heterotypic CENP-A/H3.3 nucleosome at 2.67 Å resolution (Fig. 2a and Supplementary Table 1). In the crystal structure, the CENP-A and H3.3 molecules were clearly distinguishable. For example, the side chain moiety of the CENP-A-specific His104 residue, corresponding to the H3.3 Gly102 residue, was clearly observed in the CENP-A molecule in the heterotypic nucleosome (Fig. 2b and Supplementary Fig. 2b and c). On the other hand, the H3.3-specific Gln68 residue, corresponding to the CENP-A Ser68 residue, was distinctly visible in the H3.3 molecule (Fig. 2b and Supplementary Fig. 2b and c). In the homotypic CENP-A nucleosome, the CENP-A αN helices were shorter than the canonical H3.1 αN helices in H3.1 homotypic nucleosomes27. This CENP-A-specific αN structure is perfectly conserved in the CENP-A/H3.3 nucleosome, in which the CENP-A αN helix is one helical turn shorter than the H3.3 αN helix (Fig. 2b and Supplementary Fig. 2a).
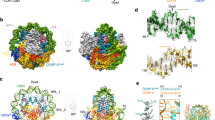
Crystal structure of the human CENP-A/H3.3 nucleosome.
(a) The CENP-A/H3.3 nucleosome structure is presented. CENP-A and H3.3 molecules are colored red and blue, respectively. The 2mFo - DFc maps of the two DNA end regions of the CENP-A/H3.3 were calculated and contoured at the 2.0σ level. (b) Close-up views of the CENP-A αN helix, the H3.3 αN helix, the CENP-A Ser68 residue, the H3.3 Gln68 residue, the CENP-A His104 residue and the H3.3 Gly102 residue. Electron density maps are presented at the 1.5σ level.
The structural asymmetry of the CENP-A and H3.3 molecules induces the asymmetric wrapping of the DNA in the CENP-A/H3.3 nucleosome (Fig. 2a). The different electron densities observed at the two ends suggested that the DNA end on the CENP-A side is more flexible than that on the H3.3 side in the CENP-A/H3.3 nucleosome. This was confirmed by differential micrococcal nuclease (MNase) digestion at the ends (Fig. 3a and Supplementary Fig. 3a) and by a treatment with Exonuclease III (ExoIII), which digests one strand of duplex DNA from the 3′ end and degraded only one end of the DNA in the CENP-A/H3.3 nucleosome (Fig. 3b and Supplementary Fig. 3b). In contrast, with the homotypic CENP-A nucleosome (Fig. 3a and Supplementary Fig. 3a), MNase attacked about 20 base pairs of DNA (Fig. 3a and Supplementary Fig. 3a) and ExoIII degraded about 10 bases from both DNA ends (Fig. 3b and Supplementary Fig. 3b).
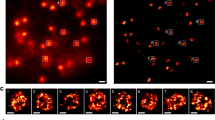
The DNA end close to CENP-A is asymmetrically flexible in the CENP-A/H3.3 nucleosome.
(a) MNase assay. The H3.3, CENP-A and CENP-A/H3.3 nucleosomes were treated with MNase (0, 0.3, 0.5 and 0.7 units) and the resulting DNA fragments were analyzed by native PAGE. The gel image shown is a representative of four independent experiments, in which similar results were obtained. (b) ExoIII assay. The H3.3, CENP-A, or CENP-A/H3.3 nucleosomes were incubated with or without 2.0 units of ExoIII for 2.5, 5 and 7.5 minutes at 37°C and the resultant DNA fragments were extracted and analyzed by 14% denaturing PAGE with 7 M urea. The gel image shown is a representative of three independent experiments, in which similar results were obtained.
Therefore, in the CENP-A/H3.3 nucleosome, the DNA end close to CENP-A could be more flexible than the DNA end close to H3.3. These results are consistent with a previous in vivo MNase analysis, which demonstrated that the average DNA length tightly wrapped within the CENP-A/H3.3 nucleosomes is shorter than that of the canonical H3 nucleosome, but longer than that of the homotypic CENP-A nucleosome24.
The heterotypic CENP-A/H3.3 nucleosome is more stable than the homotypic CENP-A nucleosome
To assess the biological significance of the CENP-A/H3.3 nucleosome, we examined its thermal stability. For this, we utilized a fluorescent dye, SYPRO Orange, which binds to denatured proteins by hydrophobic interactions28. In the assay, the fluorescence signal from SYPRO Orange was monitored as a function of increasing temperature. An increase in the fluorescence signal indicates that the histones have dislodged from the nucleosome and become denatured29,30 (Fig. 4a).
Stability of the CENP-A/H3.3 nucleosome.
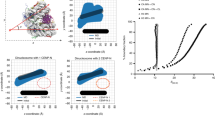
(a) Schematic representation of the thermal stability assay. (b) Thermal stability curve of the H3.3 nucleosome. The fluorescence intensity was plotted against each temperature (from 55°C to 90°C). The derivative values of the H3.3 stability curve presented in the upper panel are plotted against the temperatures (bottom panel). Means ± s.d. (n = 3) are shown. (c) A thermal stability curve of the CENP-A nucleosome. The fluorescence intensity was plotted against each temperature (from 55°C to 90°C). The derivative values of the CENP-A stability curve presented in the upper panel are plotted against the temperatures (bottom panel). Means ± s.d. (n = 3) are shown. (d) Tetrasomes, reconstituted with the H3.3-H4 tetramer or the CENP-A-H4 tetramer and DNA, were analyzed by 6% native PAGE. Lane 1 indicates the H3.3 tetrasome before incubation. Lanes 2, 3, 4, 5 and 6 indicate the H3.3 tetrasomes after 25°C, 35°C, 45°C, 55°C and 65°C incubations, respectively. Lane 7 indicates the CENP-A tetrasome before incubation. Lanes 8, 9, 10, 11 and 12 indicate the CENP-A tetrasomes after 25°C, 35°C, 45°C, 55°C and 65°C incubations, respectively. DNA was visualized by ethidium bromide staining. The gel image is a representative of seven independent experiments with similar results. (e) A thermal stability curve of the CENP-A/H3.3 nucleosome. The fluorescence intensity was plotted against each temperature (from 55°C to 90°C). The derivative values of the CENP-A/H3.3 stability curve presented in the upper panel are plotted against the temperatures (bottom panel). Means ± s.d. (n = 3) are shown. (f) The H3.3, CENP-A and CENP-A/H3.3 nucleosomes were incubated for 1 min at each temperature from 25°C and the samples at 65°C, 72°C, 79°C and 86°C were analyzed by 6% native PAGE. DNA was visualized by ethidium bromide staining. The gel image is a representative of three independent experiments with similar results.
For the H3.3 nucleosome, a biphasic histone dissociation curve (Fig. 4b, upper panel) was observed, with first and second Tm values of 70–71°C and 83–84°C, respectively (Fig. 4b, lower panel). The first and second thermal dissociation curves reflect the stepwise dissociation of the H2A-H2B dimer and the H3.3-H4 tetramer, respectively30. The CENP-A nucleosome exhibited a monophasic dissociation curve (Fig. 4c, upper panel) with a Tm value of about 71–72°C (Fig. 4c, lower panel). This suggested that the CENP-A-H4 tetramer may be unstably incorporated into the nucleosome, as compared to the H3.3-H4 tetramer and may simultaneously dissociate with the H2A-H2B dimer.
To test the unstable association of the CENP-A-H4 tetramer with DNA, we reconstituted tetrasomes, containing the CENP-A-H4 tetramer or the H3.3-H4 tetramer with a 146 base-pair DNA fragment (CENP-A tetrasome and H3.3 tetrasome, respectively), in the absence of the H2A-H2B dimer. After the salt dialysis reconstitution, the tetrasomes were fractionated by native PAGE. The bands corresponding to the H3.3 tetrasomes were smeared, probably by the unusual histone-DNA binding and/or the multiple positions of the H3.3-H4 tetramer in the tetrasomes (Fig. 4d, lanes 1–3). The H3.3 tetrasomes ran as a single band, when the sample was treated at 45–65°C for 2 hr (Fig. 4d, lanes 4–6), suggesting the formation of the stably positioned H3.3 tetrasome. In contrast, the CENP-A tetrasomes had a tendency to form positioned tetrasomes at lower temperatures (Fig. 4d, lanes 7–10), but the positioned CENP-A tetrasomes were substantially disrupted by the 55–65°C treatment (Fig. 4d, lanes 11–12). These results confirmed that the association of the CENP-A-H4 tetramer with DNA was less stable than that of the H3.3-H4 tetramer. Therefore, the homotypic CENP-A nucleosome ectopically mislocalized in the chromosome arms could easily be removed in cells, although it should be stably maintained with additional proteins in functional centromeres.
Surprisingly, although the DNA end is asymmetrically detached in the CENP-A/H3.3 nucleosome, we found that the CENP-A/H3.3 nucleosome was very stable, in contrast to the CENP-A nucleosome (Fig. 4e). The CENP-A/H3.3 nucleosome generated a biphasic histone dissociation curve (Fig. 4e, upper panel) and the first and second Tm values (70–72°C and 82–84°C) were very similar to those of the H3.3 nucleosome (Fig. 4e, lower panel). Consistently, a gel retardation assay revealed that the CENP-A nucleosome was disrupted at lower temperatures (72 and 79°C), as compared to the H3.3 and CENP-A/H3.3 nucleosomes (Fig. 4f).
The CENP-C fragment binds to the heterotypic CENP-A/H3.3 nucleosome
We next tested the binding of CENP-C to the CENP-A/H3.3 nucleosome. The CENP-C(426–537) fragment (a CENP-A binding region)31,32,33,34 efficiently bound to the CENP-A/H3.3 nucleosome (Fig. 5a and b). As anticipated, only one CENP-C bound to the CENP-A/H3.3 nucleosome (Fig. 5b, lanes 3–9), while two CENP-C peptides bound to the homotypic CENP-A nucleosome (Fig. 5b, lanes 10–16) and no CENP-C binding was detected with the homotypic H3.3 nucleosome (Fig. 5b, lanes 1 and 2).
CENP-C binding to the CENP-A/H3.3 nucleosome.
(a) Schematic representations of CENP-C binding to the H3.3 nucleosome, the CENP-A nucleosome and the CENP-A/H3.3 nucleosome. (b) The binding of CENP-C to the CENP-A/H3.3 nucleosome. CENP-C(426–537) peptide binding to the H3.3 nucleosome (0.2 μM), the CENP-A nucleosome (0.2 μM) and the CENP-A/H3.3 nucleosome (0.2 μM) was evaluated by a gel mobility shift assay. The CENP-C(426–537) peptide concentrations are 0 μM (lanes 1, 3 and 10), 0.2 μM (lanes 4 and 11), 0.4 μM (lanes 5 and 12), 0.6 μM (lanes 6 and 13), 0.8 μM (lanes 2, 7 and 14), 1.0 μM (lanes 8 and 15) and 1.2 μM (lanes 9 and 16). The gel image is a representative of three independent experiments with similar results.
Conclusion
We conclude that the structural characteristics of CENP-A and H3.3 are conserved in the CENP-A/H3.3 nucleosome, which forms a surprisingly stable structure. This stable existence of the CENP-A/H3.3 nucleosome may cause ectopic kinetochore assembly, which could lead to neocentromere formation and chromosome instability in cancer cells19,20,21,22,23,24. The unique structural and physical properties of the CENP-A/H3.3 nucleosome provide important insights toward understanding the chromosome instability observed in cancer progression, thus offering a basis for potential drug development.
Methods
Purification of recombinant human histones
The DNA fragment encoding human histone H3.3 was inserted between the NdeI and BamHI sites of the pET21a-NHSP2 vector35, in which the His6-tag sequence, the Saccharomyces cerevisiae SUMO homolog, Smt3 and a PreScission protease cleavage sequence are located just upstream of the H3.3 coding sequence. Then, the N-terminally His6-SUMO tagged H3.3 was expressed in Escherichia coli BL21(DE3) cells in the presence of isopropyl-β-D-thiogalactopyranoside (final concentration 1 mM). His6-SUMO tagged H3.3 was recovered from inclusion bodies with 50 mM Tris-HCl (pH 8.0) buffer, containing 7 M guanidine hydrochloride, 500 mM NaCl, 1 mM phenylmethylsulfonyl fluoride and 5% glycerol and was purified by nickel-nitrilotriacetic acid (Ni-NTA) agarose chromatography (Qiagen) under denaturing conditions with 6 M urea. His6-SUMO tagged H3.3 was then purified by Mono S column chromatography (GE Healthcare) under denaturing conditions with 6 M urea. The purified His6-SUMO tagged H3.3 was dialyzed against water containing 2 mM 2-mercaptoethanol, freeze-dried and stored at 4°C.
Human histones H2A, H2B, H3.3, CENP-A and H4 were expressed and purified as described previously26,27,36.
Preparation of the histone octamer containing H2A, H2B, His6-SUMO-H3.3, CENP-A and H4
Purified histones H2A, H2B, His6-SUMO-H3.3, CENP-A and H4 were mixed in a 1:1:0.6:0.4:1 ratio and the histone octamer was reconstituted in 20 mM Tris-HCl (pH 7.5) buffer, containing 7 M guanidine-HCl and 20 mM 2-mercaptoethanol. To curtail the formation of the histone octamer containing two CENP-A molecules, we reduced the amount of CENP-A relative to His6-SUMO-H3.3 in the histone octamer reconstitution mixture. The guanidine-HCl was removed by dialysis against 10 mM Tris-HCl (pH 7.5) buffer, containing 1 mM EDTA, 5 mM 2-mercaptoethanol and 2 M NaCl (500 ml) for 4 hr and this dialysis step was repeated four times. The resulting mixtures of histone octamers containing two CENP-A, two His6-SUMO-H3.3, or one each of CENP-A and His6-SUMO-H3.3 were further purified by Superdex 200 gel filtration chromatography.
Preparation of the heterotypic nucleosome containing CENP-A and H3.3
The nucleosome reconstitution was performed with the histone octamers prepared above in the presence of the 146 base-pair palindromic α-satellite DNA derivative1, by the salt dialysis method. Three nucleosomes were thus formed: 1) the homotypic His6-SUMO-H3.3 nucleosome, 2) the homotypic CENP-A nucleosome and 3) the heterotypic CENP-A/His6-SUMO-H3.3 nucleosome. These three nucleosomes were separated by native PAGE and the heterotypic CENP-A/His6-SUMO-H3.3 nucleosome was purified by preparative native PAGE (Fig. 1b). The His6-SUMO portion of H3.3 was then proteolytically removed by PreScission protease (GE Healthcare) (Supplementary Fig. 1b, lanes 1–2) and the heterotypic CENP-A/H3.3 nucleosome was further purified by preparative native PAGE (Supplementary Fig. 1b, lanes 3–12). The purified heterotypic CENP-A/H3.3 nucleosome and its histone composition were analyzed by native PAGE (Fig. 1c, lane 4) and SDS-PAGE (Fig. 1d, lane 3).
Crystallization and structure determination
The purified CENP-A/H3.3 nucleosome was dialyzed against 20 mM potassium cacodylate (pH 6.0) buffer, containing 1 mM EDTA and 1 μl of the 3 mg/ml CENP-A/H3.3 nucleosome solution (concentration of DNA) was mixed with 1 μl of 20 mM potassium cacodylate (pH 6.0) buffer, containing 50 mM KCl and 90 mM MnCl2. The sample was equilibrated against a reservoir solution of 20 mM potassium cacodylate (pH 6.0), 40 mM KCl and 65 mM MnCl2 for a month. The resulting CENP-A-H3.3 nucleosome crystals were cryoprotected with a 30% polyethylene glycol 400 solution, containing 20 mM potassium cacodylate (pH 6.0), 40 mM KCl, 50 mM MnCl2 and 5% trehalose and were flash-cooled in a stream of N2 gas (100 K). The diffraction data were collected at the beamline BL17A (wavelength: 0.97319 Å) at the Photon Factory (Tsukuba, Japan) and were processed using the HKL2000 and CCP4 programs37,38. The CENP-A/H3.3 nucleosome structure was determined by the molecular replacement method, with the human H3.1 nucleosome39 (PDB ID: 2CV5) as the search model, using the PHASER program40. Crystallographic refinement was performed using the PHENIX program41. The model rebuilding was performed using the Coot program42. All structural graphics were displayed using the PyMOL program (http://pymol.org). The atomic coordinates of the CENP-A/H3.3 nucleosome have been deposited in the Protein Data Bank, with the ID code 3WTP. In the refined model, 97.5% of the residues are in the favored regions of the Ramachandran plot, with 0.1% in the disallowed regions.
MNase and exonuclease III treatment assays
The H3.3, CENP-A, or CENP-A/H3.3 nucleosomes (0.4 μM) were treated with MNase (Takara) or ExoIII (Takara). For the MNase assay, the nucleosome samples were incubated with MNase (0, 0.3, 0.5 and 0.7 units) for 5 minutes at 25°C in 10 μl of 44 mM Tris-HCl (pH 8.0) buffer, containing 15 mM NaCl, 2.5 mM CaCl2 and 1.9 mM dithiothreitol. After the MNase treatment, the reactions were stopped by the addition of 55 μl of 0.5 mg/ml proteinase K (Roche) solution, containing 20 mM Tris-HCl (pH 8.0), 20 mM EDTA and 0.25% SDS. The resultant DNA fragments were extracted by phenol/chloroform/isoamyl alcohol and were precipitated by ethanol. The purified DNA fragments were subjected to 10% native PAGE in 0.5 × TBE buffer (11.1 V/cm for 1 hr). For the ExoIII assay, the nucleosome samples were incubated with or without 2.0 units of ExoIII for 2.5, 5 and 7.5 minutes at 37°C in 10 μl of 63 mM Tris-HCl (pH 8.0) buffer, containing 5 mM MgCl2, 5 mM KCl and 2.45 mM dithiothreitol. After the ExoIII treatment, the reactions were stopped by the addition of 55 μl of 0.5 mg/ml proteinase K (Roche) solution, containing 20 mM Tris-HCl (pH 8.0), 20 mM EDTA and 0.25% SDS. The resultant DNA fragments were extracted by phenol/chloroform/isoamyl alcohol and were precipitated by ethanol. The purified DNA fragments were subjected to 14% native PAGE in 0.5 × TBE buffer containing 7 M urea (11.1 V/cm for 3.5 hr).
Thermal stability assay of nucleosomes
The stabilities of the H3.3, CENP-A and CENP-A/H3.3 nucleosomes were evaluated by a thermal stability assay in the presence of SYPRO Orange, by the previously described method29,30. The thermal stability assay was performed in 10 μl of 20 mM Tris–HCl (pH 7.5) buffer, containing 1 mM dithiothreitol and 100 mM NaCl. The StepOnePlus™ Real-Time PCR unit (Applied Biosystems) was used to detect the fluorescence signals with a temperature gradient from 26°C to 95°C, in steps of 1°C/min. Raw fluorescence data were adjusted to normalized % values as (F(T) − F26°C)/(F95°C − F26°C), where F(T), F26°C and F95°C indicate each fluorescence at a particular temperature, the fluorescence at 26°C and the fluorescence at 95°C, respectively.
Purification of the human CENP-C fragment
The DNA fragment encoding human CENP-C(426–537) was ligated between the NdeI and BamHI sites of the pET15b (Novagen) expression vector. The N-terminally His6-tagged CENP-C(426–537) fragment was expressed in Escherichia coli BL21(DE3) cells in the presence of isopropyl-β-D-thiogalactopyranoside (final concentration 1 mM). The cells producing the His6-tagged CENP-C(426–537) fragment were cultured overnight at 30°C. The cells were harvested and disrupted by sonication in 50 mM Tris–HCl (pH 8.0) buffer, containing 5% glycerol, 0.5 M NaCl, 1 mM phenylmethylsulfonyl fluoride and 10 mM imidazole. His6-tagged CENP-C(426–537) was purified by Ni-NTA agarose chromatography (Qiagen), eluted with 50 mM Tris–HCl (pH 8.0) buffer, containing 5% glycerol, 0.5 M NaCl, 1 mM phenylmethylsulfonyl fluoride and 500 mM imidazole. The His6-tag portion was removed by thrombin proteinase (GE Healthcare) during the dialysis step against 20 mM Tris-HCl (pH 7.5) buffer, containing 5% glycerol and 2 mM 2-mercaptoethanol, at 4°C overnight. CENP-C(426–537) was further purified by Mono S column chromatography (GE Healthcare) and was dialyzed against 50 mM Tris-HCl (pH 7.5) buffer, containing 100 mM NaCl, 5% glycerol and 2 mM 2-mercaptoethanol. The purified CENP-C(426–537) was concentrated to 500 μM and stored at −80°C.
CENP-C binding assay
CENP-C(426–537) was mixed with the H3.3, CENP-A, or CENP-A/H3.3 nucleosomes (final concentration 0.2 μM) and incubated at 37°C for 30 minutes in 10 μl of 35 mM Tris-HCl (pH 7.5) buffer, containing 50 mM NaCl, 2.5% glycerol, 1 mM 2-mercaptoethanol and 0.5 mM dithiothreitol. The final CENP-C(426–537) concentrations were 0, 0.2, 0.4, 0.6, 0.8, 1.0 and 1.2 μM. The samples were then subjected to 6% native PAGE in 0.2 × TBE buffer (8.3 V/cm for 1 hr).
Additional Information
Accession codes: The atomic coordinates of the CENP-A/H3.3 nucleosome have been deposited in the Protein Data Bank, with the ID code 3WTP.
References
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Tan, S. & Davey, C. A. Nucleosome structural studies. Curr. Opin. Struct. Biol. 21, 128–136 (2011).
Talbert, P. B. et al. A unified phylogeny-based nomenclature for histone variants. Epigenetics Chromatin 5, 7 (2012).
Vardabasso, C. et al. Histone variants: emerging players in cancer biology. Cell. Mol. Life Sci. 71, 379–404 (2013).
Maze, I., Noh, K. M., Soshnev, A. A. & Allis, C. D. Every amino acid matters: essential contributions of histone variants to mammalian development and disease. Nat. Rev. Genet. 15, 259–271(2014).
Volle, C. & Dalal, Y. Histone variants: the tricksters of the chromatin world. Curr. Opin. Genet. Dev. 25C, 8–14 (2014).
Kurumizaka, H., Horikoshi, N., Tachiwana, H. & Kagawa, W. Current progress on structural studies of nucleosomes containing histone H3 variants. Curr. Opin. Struct. Biol. 23, 109–115 (2013).
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004).
Ray-Gallet, D. et al. HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell 9, 1091–1100 (2002).
Adam, S., Polo, S. E. & Almouzni, G. Transcription recovery after DNA damage requires chromatin priming by the H3.3 histone chaperone HIRA. Cell 155, 94–106 (2013).
Goldberg, A. D. et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140, 678–691 (2010).
Drané, P., Ouararhni, K., Depaux, A., Shuaib, M. & Hamiche, A. The death-associated protein DAXX is a novel histone chaperone involved in the replication-independent deposition of H3.3. Genes. Dev. 24, 1253–1265 (2010).
Palmer, D. K., O'Day, K., Wener, M. H., Andrews, B. S. & Margolis, R. L. A 17-kD centromere protein (CENP-A) copurifies with nucleosome core particles and with histones. J. Cell Biol. 104, 805–815 (1987).
Black, B. E. & Cleveland, D. W. Epigenetic centromere propagation and the nature of CENP-A nucleosomes. Cell 144, 471–479 (2011).
De Rop, V., Padeganeh, A. & Maddox, P. S. CENP-A: the key player behind centromere identity, propagation and kinetochore assembly. Chromosoma 121, 527–538 (2012).
Quénet, D. & Dalal, Y. The CENP-A nucleosome: a dynamic structure and role at the centromere. Chromosome Res. 20, 465–479 (2012).
Stellfox, M. E., Bailey, A. O. & Foltz, D. R. Putting CENP-A in its place. Cell. Mol. Life Sci. 70, 387–406 (2013).
Catania, S. & Allshire, R. C. Anarchic centromeres: deciphering order from apparent chaos. Curr. Opin. Cell Biol. 26, 41–50 (2014).
Tomonaga, T. et al. Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res. 63, 3511–3516 (2003).
Biermann, K. et al. Gene expression profiling identifies new biological markers of neoplastic germ cells. Anticancer Res. 27, 3091–3100 (2007).
Wu, Q. et al. Expression and prognostic significance of centromere protein A in human lung adenocarcinoma. Lung Cancer 77, 407–414 (2007).
McGovern, S. L., Qi, Y., Pusztai, L., Symmans, W. F. & Buchholz, T. A. Centromere protein-A, an essential centromere protein, is a prognostic marker for relapse in estrogen receptor-positive breast cancer. Breast Cancer Res. 14, R72 (2012).
Heun, P. et al. Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev. Cell. 10, 303–315 (2006).
Lacoste, N. et al. Mislocalization of the centromeric histone variant CenH3/CENP-A in human cells depends on the chaperone DAXX. Mol. Cell 53, 631–644 (2014).
Luk, E. et al. Stepwise histone replacement by SWR1 requires dual activation with histone H2A.Z and canonical nucleosome. Cell 143, 725–736 (2010).
Tanaka, Y. et al. Expression and purification of recombinant human histones. Methods 33, 3–11 (2004).
Tachiwana, H. et al. Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 476, 232–235 (2011).
Lo, M. C. et al. Evaluation of fluorescence-based thermal shift assays for hit identification in drug discovery. Anal Biochem. 332, 153–159 (2004).
Iwasaki, W. et al. Contribution of histone N-terminal tails to the structure and stability of nucleosomes. FEBS Open Bio. 3, 363–369 (2013).
Taguchi, H., Horikoshi, N., Arimura, Y. & Kurumizaka, H. A Method for Evaluating Nucleosome Stability with a Protein-Binding Fluorescent Dye. Methods, in press.
Saitoh, H. et al. CENP-C, an autoantigen in scleroderma, is a component of the human inner kinetochore plate. Cell 70, 115–125 (1992).
Yang, C. H., Tomkiel, J., Saitoh, H., Johnson, D. H. & Earnshaw, W. C. Identification of overlapping DNA-binding and centromere-targeting domains in the human kinetochore protein CENP-C. Mol. Cell. Biol. 16, 3576–3586 (1996).
Carroll, C. W., Milks, K. J. & Straight, A. F. Dual recognition of CENP-A nucleosomes is required for centromere assembly. J. Cell Biol. 189, 1143–1155 (2010).
Kato, H. et al. A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 340, 1110–1113 (2013).
Ichikawa, Y. et al. Purification and characterization of the fission yeast telomere clustering factors, Bqt1 and Bqt2. Protein Expr. Purif. 88, 207–213 (2013).
Tachiwana, H. et al. Structures of human nucleosomes containing major histone H3 variants. Acta Crystallogr. D Biol. Crystallogr. 67, 578–583 (2011).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project Number 4, The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Tsunaka, Y., Kajimura, N., Tate, S. & Morikawa, K. Alteration of the nucleosomal DNA path in the crystal structure of a human nucleosome core particle. Nucleic Acids Res. 33, 3424–3434 (2005).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
We thank the beamline scientists at the BL17A station of the Photon Factory and the BL41XU station of SPring-8 for their assistance with data collection. We thank Sebastian Muller (Institut Curie) for critical reading. This work was supported in part by MEXT KAKENHI Grant Number 25116002 [to H.K.] and the Platform for Drug Discovery, Informatics and Structural Life Science from MEXT, Japan [to H.K.]. H.K. was also supported by the Waseda Research Institute for Science and Engineering and by the Uehara Foundation and the Naito Foundation. Y.A. was supported by Research Fellowships of the Japan Society for the Promotion of Science for Young Scientists [25-3931].
Author information
Authors and Affiliations
Contributions
Y.A. and K.S. reconstituted the CENP-A/H3.3 nucleosome and performed structural and biochemical experiments. N.H. and Y.A. established the reconstitution method for the heterotypic nucleosome. R.F. purified the histones and CENP-A. H.T. established thermal stability assay and K.S. and H.T. tested the thermal stability of the nucleosomes. Y.A., N.H. and W.K. collected X-ray diffraction data and performed the structural analysis of the CENP-A/H3.3 nucleosome. T.F. and G.A. provided unpublished results for the CENP-A/H3.3 nucleosome formation in cells and contributed to the initiation of this project. H.K. conceived, designed and supervised all of the work and H.K., G.A., T.F. and Y.A. wrote the paper. All of the authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary figures
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Arimura, Y., Shirayama, K., Horikoshi, N. et al. Crystal structure and stable property of the cancer-associated heterotypic nucleosome containing CENP-A and H3.3. Sci Rep 4, 7115 (2014). https://doi.org/10.1038/srep07115
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07115
This article is cited by
-
In vitro co-expression chromatin assembly and remodeling platform for plant histone variants
Scientific Reports (2024)
-
CENP-A and CENP-B collaborate to create an open centromeric chromatin state
Nature Communications (2023)
-
Centromere protein N promotes lung adenocarcinoma progression by activating PI3K/AKT signaling pathway
Genes & Genomics (2022)
-
Histone variant H2A.B-H2B dimers are spontaneously exchanged with canonical H2A-H2B in the nucleosome
Communications Biology (2021)
-
Internal modifications in the CENP-A nucleosome modulate centromeric dynamics
Epigenetics & Chromatin (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.