Abstract
5-methylcytosine (5-mC) is a well-characterized epigenetic regulator in mammals. Recent studies showed that Ten-eleven translocation (Tet) proteins can catalyze the stepwise oxidation of 5-mC to produce 5-hydroxymethylcytosine (5-HmC), 5-formylcytosine (5-FoC) and 5-carboxylcytosine (5-CaC). The exciting discovery of these novel cytosine modifications has stimulated substantial research interests about their roles in epigenetic regulation. Here we systematically examined the effects of the oxidized 5-mC derivatives on the efficiency and fidelity of DNA transcription using a recently developed competitive transcription and adduct bypass assay. Our results showed that, when located on the transcribed strand, 5-FoC and 5-CaC exhibited marginal mutagenic and modest inhibitory effects on DNA transcription mediated by single-subunit T7 RNA polymerase or multi-subunit human RNA polymerase II in vitro and in human cells. 5-HmC displayed relatively milder blocking effects on transcription and no mutant transcript could be detectable for 5-HmC in vitro or in cells. The lack of considerable mutagenic effects of the oxidized 5-mC derivatives on transcription was in agreement with their functions in epigenetic regulation. The modest blocking effects on transcription suggested that 5-FoC and 5-CaC may function in transcriptional regulation. These findings provided new evidence for the potential functional interplay between cytosine methylation status and transcription.
Similar content being viewed by others
Introduction
Methylation at the C5 position of cytosine residues represents a major epigenetic mark that has important regulatory functions in gene expression and developmental processes1,2. It was recently discovered that Ten-eleven translocation (Tet) family dioxygenases can catalyze the sequential oxidation of 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine (5-HmC), 5-formylcytosine (5-FoC) and 5-carboxylcytosine (5-CaC) (Figure 1)3,4,5,6,7,8,9. The resulting 5-FoC and 5-CaC in DNA can be recognized and excised by thymine DNA glycosylase (TDG) and ultimately converted back to unmodified cytosine8,10,11. Thus, 5-HmC, 5-FoC and 5-CaC constitute important intermediates of a new pathway of active cytosine demethylation in mammals12.
These Tet-mediated oxidation products of 5-mC can be readily detected in various mammalian tissues and cells. In this regard, it was reported that 5-HmC is present in mouse brain genomic DNA at a level of 5.6 modifications per 104 nucleosides and occurs at substantial frequencies in other types of mammalian tissues and cells13,14,15,16. Although much less abundant than 5-HmC, the levels of 5-FoC and 5-CaC in mammalian genomic DNA are comparable to those of the common types of DNA damage induced by endogenous and environmental sources3,5,7,8,13,17,18,19,20. Emerging evidence indicates that 5-HmC, 5-FoC and 5-CaC may carry distinct regulatory functions in addition to being demethylation intermediates1,12,21,22,23. For instance, 5-HmC was reported to have important roles in epigenetic regulation and reprogramming of somatic cells to pluripotent cells24,25,26,27,28. In addition, a proteome-wide analysis revealed that 5-HmC, 5-FoC and 5-CaC can be recognized by some cellular proteins, which may confer specific biological functions29. Moreover, a recent study showed that the presence of 5-FoC and 5-CaC on a template DNA strand are able to reduce the rate and substrate specificity of transcription mediated by yeast or rat RNA polymerase II in vitro19. It is therefore of interest to investigate how these oxidized 5-mC derivatives modulate the process of transcription in human cells.
Results
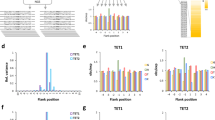
We employed a recently developed competitive transcription and adduct bypass (CTAB) assay30 to investigate how the oxidation products of 5-mC affect the efficiency and fidelity of DNA transcription in vitro and in human cells. To this end, we constructed nonreplicative double-stranded plasmids containing a single site-specifically incorporated 5-HmC, 5-FoC, or 5-CaC (Figure 2a), as well as the corresponding unmodified control and competitor plasmids. These cytosine derivatives were located on the transcribed strand 58 and 39 nucleotides downstream of the cytomegalovirus (CMV) and T7 promoters, respectively (Figure 2b). We premixed the cytosine derivative-bearing or control plasmid with a competitor construct and used them as templates for the transcription assays. The run-off transcripts of interest were subjected to reverse transcription PCR (RT-PCR), followed by polyacrylamide gel electrophoresis (PAGE) and liquid chromatography-tandem mass spectrometry (LC-MS/MS) analyses of the restriction digestion mixture of RT-PCR products (Figure 2b).
Experimental outline.
(a) Schematic diagrams showing the procedures for the construction of the plasmids harboring a site-specifically incorporated 5-HmC, 5-FoC, or 5-CaC. (b) CTAB assay for assessing the impact of the oxidized 5-mC derivatives on DNA transcription. “X” indicates 5-HmC, 5-FoC or 5-CaC, which was located on the transcribed strand of TurboGFP gene downstream of the CMV and T7 promoters. The +1 transcription start sites are indicated by arrowheads. Oxidized 5-mC derivative-bearing or unmodified control plasmids were mixed individually with the competitor genome as DNA templates for in vitro or in vivo transcription. Although truncated RNA may be produced when transcription arrests at or near a lesion site, only run-off RNA is shown and used for RT-PCR. Among the RT-PCR products, only the wild-type sequence arising from the cytosine derivative-containing vector is shown. The arrows indicate the cleavage sites of Nt.BstNBI, NcoI and SfaNI. The last two enzymes were used to digest the RT-PCR products for subsequent PAGE and LC-MS/MS analyses.
We firstly used the CTAB assay to quantitatively assess the effects of the oxidation products of 5-mC on transcription catalyzed by purified T7 RNA polymerase (T7 RNAP) or human RNA polymerase II (hRNAPII) in HeLa cell nuclear extract. In this regard, we chose to use T7 RNAP, a structural homologue of single-subunit eukaryotic mitochondrial RNA polymerases31, as a model to examine the impact of the 5-mC oxidation products on transcription of the mitochondrial genome.
PAGE analysis showed that, when located on the template DNA strand, a single 5-HmC did not substantially compromise the efficiency of transcription by T7 RNAP, whereas 5-CaC and 5-FoC impeded modestly the T7 RNAP-mediated transcription elongation, with relative bypass efficiencies (RBE) being ~50% and 66%, respectively (Figure 3a–c). Similarly, 5-FoC and 5-CaC, but not 5-HmC, inhibited considerably the transcription by hRNAPII (Figure 3a–c). We also employed LC-MS/MS analysis to detect the potential mutant products induced by these cytosine derivatives during transcription in vitro. We found that transcriptional bypass of 5-HmC with T7 RNAP or hRNAPII did not generate any detectable mutant transcripts; however, 5-FoC and 5-CaC were able to induce at least one type of mutant transcript (G→A), which contains an adenosine misincorporation opposite the 5-FoC and 5-CaC and occurs at frequencies of ~1–1.7% (Figure 4a,b and Supplementary Figure S1–4).
The effects of cytosine derivatives on transcriptional efficiency.
(a) Sample processing for PAGE analysis (p* indicates 32P-labeled phosphate group). Only the wild-type sequence arising from the cytosine derivative-containing vector is shown. (b) Representative gel images showing the restriction fragments of interest. ‘13mer-C’ represents the standard ODN d(CATGGCGCGCTAT). ‘16mer-Comp’ represents the standard ODN d(CATGGCGATATGCTAT), which corresponds to the restriction fragment arising from the competitor vector. (c) The RBE values of cytosine derivatives in in vitro transcription systems using T7 RNAP and HeLa nuclear extract (hRNAPII). (d) The RBE values of cytosine derivatives in 293T cells treated with siRNA targeting TDG (siTDG) or nontargeting control siRNA (siControl). (e) The RBE values of cytosine derivatives in XPA-deficient (XP12RO) and XPA-complemented (GM15876A) cells. The data represent the mean and standard error of results from three independent experiments.
The effects of 5-FoC and 5-CaC on transcriptional fidelity.
(a) The sequences of wild-type and mutant (G→A) transcripts are indicated above the double-stranded DNA construct. The underlined base (i.e., A) indicates an adenosine misincorporation opposite a 5-FoC or 5-CaC. (b). The mutation (G→A) frequencies of 5-FoC and 5-CaC in in vitro transcription systems using T7 RNAP and hRNAPII. (c) The mutation (G→A) frequencies of 5-FoC and 5-CaC in 293T cells treated with siRNA targeting the TDG gene (siTDG).
We further asked how 5-HmC, 5-FoC and 5-CaC affect DNA transcription in human cells. To this end, we premixed either cytosine derivative-bearing or control plasmids with the competitor vector and co-transfected the mixed DNA substrates into human cells. After a 24-h incubation, we extracted the RNA products from the cells and prepared the restriction digestion mixture of RT-PCR products for PAGE and LC-MS/MS analyses as described above.
Our results showed that the bypass efficiency for 5-HmC is comparable to that of unmodified control, whereas 5-FoC and 5-CaC constitute modest blocks to DNA transcription machinery in human 293T cells, with the RBE values being ~69% and 55%, respectively (Figure 3d and Supplementary Figure S5). Given that 5-FoC and 5-CaC are considered as substrates for TDG-mediated base excision repair (BER) pathway8,10,11, we used siRNAs to efficiently knockdown the expression of TDG in 293T cells (Supplementary Figure S6) and tested its effect on the transcription bypass of 5-FoC and 5-CaC. We found that, compared with the control siRNA treatment, depletion of TDG did not cause significant changes in the transcriptional bypass efficiencies of these modified cytosines (Figure 3d and Supplementary Figure S5). With the LC-MS/MS method, we also determined the mutagenic properties of these cytosine derivatives in TDG siRNA-treated human cells. Similar to what we observed from in vitro transcription reaction, we found that 5-HmC did not induce any detectable transcription errors, whereas transcriptional bypass of 5-FoC and 5-CaC could induce G→A mutations at frequencies of ~0.7–0.9% in the tested human cells (Figure 4c).
We next examined the potential role of nucleotide excision repair (NER) in transcriptional alternations induced by the oxidized 5-mC derivatives in human cells. Xeroderma pigmentosum group A (XPA) is a core and essential component of NER machinery32, we therefore compared the RBE values for 5-HmC, 5-FoC or 5-CaC in XPA-deficient (XP12RO) and XPA-complemented (GM15876A) cells. The results from PAGE analysis revealed no significant difference in transcriptional bypass efficiencies between XPA-deficient and XPA-complemented cells for 5-HmC, 5-FoC or 5-CaC (Figure 3e and Supplementary Figure S7). This result suggests that NER pathway may not be involved in the removal of these cytosine modifications from the template strand of an actively transcribed gene in human cells.
Discussion
The Tet-mediated oxidation products of 5-mC, including 5-HmC, 5-FoC and 5-CaC, have attracted substantial research interests regarding their potential biological functions1,12. In this study, we systematically investigated the effects of these cytosine derivatives on the process of DNA transcription in vitro and in human cells.
It has been recently shown that 5-FoC and 5-CaC, but not 5-HmC, are able to reduce the rate of DNA transcription mediated by yeast or rat RNA polymerase II in vitro19. 5-FoC and 5-CaC were also found to decrease the rate of DNA replication in mammalian cells33,34. In agreement with these studies, our results revealed that, when located on the transcribed strand, 5-FoC and 5-CaC, but not 5-HmC, constituted modest blocks to transcription mediated by single-subunit T7 RNA polymerase (T7 RNAP) or multi-subunit human RNA polymerase II in vitro. Moreover, we demonstrated the modest inhibitory effects of 5-FoC and 5-CaC on DNA transcription in human cells. Therefore, our findings provided new evidence to support the notion that, apart from their roles as intermediates of active cytosine demethylation, 5-FoC and 5-CaC may have important functions in other biological processes including regulation of gene expression in mammalian cells. On the other hand, it was reported that mitochondrial DNA is rich in 5-HmC and Tet proteins can be detected in mammalian mitochondrial extracts, suggesting the potential formation of Tet-mediated 5-mC oxidation products in mitochondria35,36. Our findings that 5-FoC and 5-CaC had substantial impact on T7 RNAP-mediated transcription are of interest, due to the high degree of homology between T7 RNAP and eukaryotic mitochondrial RNA polymerases31. Although future studies are needed, it is reasonable to deduce that, aside from regulating nuclear gene expression, 5-FoC and 5-CaC might also be involved in transcriptional control in mitochondria.
We also found that 5-FoC and 5-CaC could weakly induce G→A mutations opposite the modified cytosine sites in vitro and in human cells. It was suggested that the imino tautomer of 5-FoC or 5-CaC can be stablized by intramolecular hydrogen bonding between the 5-carbonyl oxygen and the imino hydrogen37,38 and the imino tautomer is expected to form a base pair favorably with an incoming adenine nucleotide during transcription. Thus, the preferential formation of the imino tautomer of 5-FoC and 5-CaC over 5-HmC and unmodified C may account partly for the G→A mutation induced by the former modified cytosine derivatives. In addition, a previous study showed that transcriptional bypass of abasic sites preferentially induces the incorporation of adenine nucleotide opposite the lesion site39. Therefore, RNAP slippage over BER intermediates of 5-FoC and 5-CaC such as abasic sites may also contribute, in part, to the observed transcriptional mutations8,10,11.
Our results are consistent with previously published data indicating that 5-FoC and 5-CaC can affect the fidelity of transcription mediated by yeast or rat RNA polymerase II in vitro19. Similarly, it was reported that these cytosine derivatives exhibit no or marginal mutagenic potential during DNA replication in vitro and in vivo33,34,40,41,42. In this vein, accumulating evidence suggests that, apart from errors in DNA replication, transient transcriptional mutagenesis can trigger stable phenotypic changes and may also serve as important inducers of cancer and other human diseases43,44,45,46,47,48,49. Thus, the weak mutagenic effects of 5-FoC and 5-CaC during transcription and replication are in keeping with the roles of 5-FoC and 5-CaC as epigenetic marks or as intermediates for active cytosine demethylation in mammals.
It has been suggested that 5-FoC and 5-CaC can be recognized and repaired by TDG-mediated BER pathway to regenerate unmodified cytosine8,10,11. However, we found that siRNA knockdown of TDG did not lead to significant changes in the effects of 5-FoC and 5-CaC on transcription in human cells. This result may be attributed in part to the incomplete knockdown of TDG and the removal of 5-FoC and 5-CaC by residual TDG in human cells. Alternatively, the knockdown of TDG may be compensated by the presence of other cellular proteins that can remove these cytosine derivatives50,51,52.
In conclusion, we showed here that 5-FoC and 5-CaC, two recently identified oxidative products of 5-mC, exhibited marginal mutagenic and modest inhibitory effects on transcription mediated by single-subunit T7 RNA polymerase or multi-subunit human RNA polymerase II in vitro and in human cells. This result indicates that these two oxidized 5-mC derivatives may play a role on transcriptional regulation. In addition, the presence of 5-HmC on template DNA does not perturb appreciably the efficiency or fidelity of transcription in vitro or in cells. Recent LC-MS/MS quantification studies revealed the presence of markedly higher levels of 5-HmC than 5-FoC and 5-CaC in genomic DNA from mammalian cells and tissues3,5,7,8,13,18,19. The lack of adverse effects of 5-HmC on transcription and replication, as discussed above, may enable mammalian cells to maintain a relatively high level of 5-HmC in their genome during evolution. In addition, the weak mutagenic effects of 5-FoC and 5-CaC on transcription and replication may constitute another justification for the mammalian cells to evolve with a BER-based machinery for the efficient removal of these modified nucleobases from their genome. Together, our findings provide important insights into the potential functional interplay between active cytosine demethylation and transcription.
Methods
Materials
Unmodified oligodeoxyribonucleotides (ODNs) were purchased from Integrated DNA Technologies. [γ-32P]ATP was obtained from Perkin-Elmer. All enzymes and chemicals unless otherwise specified were obtained from New England BioLabs and Sigma-Aldrich, respectively. ON-TARGETplus SMARTpool siRNA against human TDG (L-003780) and Non-Targeting control siRNA (D-001210) were from Thermo Scientific Dharmacon. The 293T human embryonic kidney epithelial cells were purchased from ATCC. XPA-deficient (XP12RO) and XPA-complemented (GM15876A) human fibroblast cell lines were kindly provided by Prof. Karlene A Cimprich53. Cells were cultured in Dulbecco's Modified Eagle's medium supplemented with 10% fetal bovine serum (Invitrogen), 100 U/mL penicillin and 100 µg/mL streptomycin (ATCC) and incubated at 37°C in 5% CO2 atmosphere.
Transcription template preparation
To construct the unmodified control vector, a 50-mer ODN with the sequence of 5′-CTAGCGGATGCATCGACTCCCACATAGCGCGCCATGGAT-GACTCGCTGCG-3′ was annealed with its complementary strand and ligated to a restriction fragment from the NheI-EcoRI-treated pTGFP-T7-Hha10 plasmids30. Using the same method, a competitor vector was constructed to carry three more nucleotides than the control plasmid between the two restriction sites used for CTAB assay (Figure 2). We next nicked the unmodified control plasmid with Nt.BstNBI and produced a gapped vector by removing the Nt.BstNBI-generated 25-mer single-stranded ODN, followed by filling the gap with a 13-mer unmodified ODN (5′-GTGGGAGTCGATG-3′) and a 12-mer modified cytosine-containing ODN (5′-ATGGCGXGCTAT-3′, X = 5-HmC, 5-FoC or 5-CaC)33,54. We incubated the ligation mixture with ethidium bromide and purified the resulting supercoiled cytosine derivative-bearing plasmids by agarose gel electrophoresis, as described elsewhere55,56,57. Purified control or cytosine derivative-bearing plasmids were mixed with the competitor genome at 3:1 ratio and used as DNA templates for all transcription assays in this study.
In Vitro Transcription assay
Multiple rounds of transcription reactions catalyzed by purified T7 RNA polymerase (T7 RNAP) or human RNA polymerase II (hRNAPII) in HeLa nuclear extract (Promega) were performed as described previously30,58. Briefly, T7 RNAP-mediated transcription reaction was carried out at 37°C for 1 h in a 20-μL mixture containing 50 ng of NotI-linearized DNA templates, 20 U of T7 RNAP, 10 U of RNase inhibitor and 0.5 mM each of ATP, CTP, GTP and UTP. The hRNAPII-mediated transcription reaction was performed at 30°C for 1 h in a 25-μL mixture containing 50 ng of NotI-linearized DNA templates, 8 U of HeLa nuclear extract, 10 U of RNase inhibitor and 0.4 mM each of the four ribonucleotides.
In Vivo transcription assay
XPA-complemented (GM15876A) and XPA-deficient (XP12RO) cell lines were grown to about 70% confluence in 24-well plates and co-transfected with 50 ng mixed DNA templates and 400 ng carrier plasmid (self-ligated pGEM-T, Promega) with Lipofectamine 2000 (Invitrogen), according to the manufacturer's protocol. For siRNA experiments, 293T cells were grown to 40–60% confluence in 24-well plates and transfected with 25 pmol siRNAs directed against TDG or non-targeting control siRNAs. After a 48-h incubation, 50 ng DNA templates, 400 ng carrier DNA and another aliquot of siRNAs were co-transfected into the cells with Lipofectamine 2000 (Invitrogen). For all in vivo transcription assays, the cells were harvested for RNA extraction 24 h after transfection with the mixed DNA templates.
RNA extraction and RT-PCR
We used the Total RNA Kit I (Omega) to purify the RNA products arising from in vitro and in vivo transcription and we subjected the RNA products to two rounds of DNase I treatment using the DNA-free kit (Ambion) to eliminate DNA contamination. We then produced cDNA by using a gene-specific primer (5′-TCGGTGTTGCTGTGAT-3′) and M-MLV reverse transcriptase (Promega). RT-PCR amplification was then performed by using Phusion high-fidelity DNA polymerase and a pair of primers spanning the lesion site as described in a recent study30. Real-time quantitative RT-PCR for evaluating the efficiency of siRNA knockdown was conducted by using the iQ SYBR Green Supermix kit (Bio-Rad) and gene-specific primers for TDG or the control gene GAPDH as described elsewhere33.
PAGE analysis
PAGE analysis of transcription products was performed as described elsewhere30,58. Briefly, a portion of the above RT-PCR fragments was incubated in a 10-μL mixture containing 1× NEB buffer 3, 1 U shrimp alkaline phosphatase and 5 U NcoI at 37°C for 1 h and then at 70°C for 20 min. The resulting dephosphorylated DNA was incubated in a 15-μL solution containing 1× NEB buffer 3, 10 U T4 polynucleotide kinase and ATP (50 pmol cold, premixed with 1.66 pmol [γ-32P]ATP). The mixture was incubated at 37°C for 30 min and then at 70°C for 20 min, after which 2 U SfaNI was added and incubated at 37°C for 2 h. The resulting 32P-labeled restriction fragments were resolved by using 30% polyacrylamide gel (acrylamide:bis-acrylamide = 19:1) with 5 M urea and quantified by phosphorimager analysis. The bypass efficiency was calculated using the following formula, bypass efficiency = (cytosine derivative signal/competitor signal)/(unmodified control signal/competitor signal)30,58,59.
LC-MS/MS analysis
To identify the transcription products using LC-MS/MS, RT-PCR products were treated with 50 U NcoI, 50 U SfaNI and 20 U shrimp alkaline phosphatase in 250 μL NEB buffer 4 at 37°C for 4 h, followed by heating at 80°C for 20 min. The resulting solution was extracted with phenol/chloroform/isoamyl alcohol (25:24:1, v/v) and the aqueous portion was dried with Speed-vac, desalted with HPLC and dissolved in water. The resultant ODN mixture was subjected to LC-MS/MS analysis following previously described conditions30,55,58. Briefly, a 0.50 × 150 mm Zorbax SB-C18 column (Agilent Technologies) was used. The flow rate was 8.0 μL/min and a 5-min linear gradient of 5–20% methanol followed by a 25 min of 20–60% methanol in 400 mM 1,1,1,3,3,3-hexafluoro-2-propanol buffer (pH was adjusted to 7.0 by the addition of triethylamine) was employed for the separation. The LTQ linear ion trap mass spectrometer (Thermo Fisher Scientific) was set up for monitoring the fragmentation of the [M-3H]3− ions of 13-mer ODNs, i.e., d(CCACATAGCMCGC), where “M” designates A, T, C, or G.
References
Shen, L., Song, C. X., He, C. & Zhang, Y. Mechanism and function of oxidative reversal of DNA and RNA methylation. Annu. Rev. Biochem. 83, 585–614 (2014).
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–32 (2007).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–5 (2009).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–30 (2009).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–3 (2011).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–33 (2010).
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. Int. Ed. Engl. 50, 7008–12 (2011).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–7 (2011).
Wu, H. & Zhang, Y. Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev. 25, 2436–52 (2011).
Maiti, A. & Drohat, A. C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–8 (2011).
Zhang, L. et al. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat. Chem. Biol. 8, 328–30 (2012).
Song, C. X. & He, C. Potential functional roles of DNA demethylation intermediates. Trends Biochem. Sci. 38, 480–4 (2013).
Liu, S. et al. Quantitative assessment of Tet-induced oxidation products of 5-methylcytosine in cellular and tissue DNA. Nucleic Acids Res. 41, 6421–9 (2013).
Jin, S. G. et al. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer Res. 71, 7360–5 (2011).
Kudo, Y. et al. Loss of 5-hydroxymethylcytosine is accompanied with malignant cellular transformation. Cancer Sci. 103, 670–6 (2012).
Chen, M. L. et al. Quantification of 5-methylcytosine and 5-hydroxymethylcytosine in genomic DNA from hepatocellular carcinoma tissues by capillary hydrophilic-interaction liquid chromatography/quadrupole TOF mass spectrometry. Clin. Chem. 59, 824–32 (2013).
Lord, C. J. & Ashworth, A. The DNA damage response and cancer therapy. Nature 481, 287–94 (2012).
Raiber, E. A. et al. Genome-wide distribution of 5-formylcytosine in embryonic stem cells is associated with transcription and depends on thymine DNA glycosylase. Genome Biol. 13, R69 (2012).
Kellinger, M. W. et al. 5-formylcytosine and 5-carboxylcytosine reduce the rate and substrate specificity of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 19, 831–3 (2012).
Cadet, J., Douki, T., Gasparutto, D. & Ravanat, J. L. Oxidative damage to DNA: formation, measurement and biochemical features. Mutat. Res. 531, 5–23 (2003).
Shen, L. et al. Genome-wide analysis reveals TET- and TDG-dependent 5-methylcytosine oxidation dynamics. Cell 153, 692–706 (2013).
Song, C. X. et al. Genome-wide profiling of 5-formylcytosine reveals its roles in epigenetic priming. Cell 153, 678–91 (2013).
Koh, K. P. & Rao, A. DNA methylation and methylcytosine oxidation in cell fate decisions. Curr. Opin. Cell Biol. 25, 152–61 (2013).
Branco, M. R., Ficz, G. & Reik, W. Uncovering the role of 5-hydroxymethylcytosine in the epigenome. Nat. Rev. Genet. 13, 7–13 (2012).
Kriukiene, E., Liutkeviciute, Z. & Klimasauskas, S. 5-Hydroxymethylcytosine--the elusive epigenetic mark in mammalian DNA. Chem. Soc. Rev. 41, 6916–30 (2012).
Gao, Y. et al. Replacement of Oct4 by Tet1 during iPSC induction reveals an important role of DNA methylation and hydroxymethylation in reprogramming. Cell Stem Cell 12, 453–69 (2013).
Doege, C. A. et al. Early-stage epigenetic modification during somatic cell reprogramming by Parp1 and Tet2. Nature 488, 652–5 (2012).
Costa, Y. et al. NANOG-dependent function of TET1 and TET2 in establishment of pluripotency. Nature 495, 370–4 (2013).
Spruijt, C. G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–59 (2013).
You, C. et al. A quantitative assay for assessing the effects of DNA lesions on transcription. Nat. Chem. Biol. 8, 817–22 (2012).
Masters, B. S., Stohl, L. L. & Clayton, D. A. Yeast mitochondrial RNA polymerase is homologous to those encoded by bacteriophages T3 and T7. Cell 51, 89–99 (1987).
Hanawalt, P. C. & Spivak, G. Transcription-coupled DNA repair: two decades of progress and surprises. Nat. Rev. Mol. Cell Biol. 9, 958–70 (2008).
Ji, D., You, C. & Wang, Y. Effects of Tet-induced Oxidation Products of 5-Methylcytosine on DNA Replication in Mammalian Cells. Chem. Res. Toxicol. 27, 1304–9 (2014).
Kamiya, H. et al. Mutagenicity of 5-formylcytosine, an oxidation product of 5-methylcytosine, in DNA in mammalian cells. J. Biochem. 132, 551–5 (2002).
Shock, L. S., Thakkar, P. V., Peterson, E. J., Moran, R. G. & Taylor, S. M. DNA methyltransferase 1, cytosine methylation and cytosine hydroxymethylation in mammalian mitochondria. Proc. Natl. Acad. Sci. U S A 108, 3630–5 (2011).
Dzitoyeva, S., Chen, H. & Manev, H. Effect of aging on 5-hydroxymethylcytosine in brain mitochondria. Neurobiol. Aging 33, 2881–91 (2012).
Shibutani, T. et al. Guanine- 5-carboxylcytosine base pairs mimic mismatches during DNA replication. Sci. Rep. 4, 5220 (2014).
Hashimoto, H., Hong, S., Bhagwat, A. S., Zhang, X. & Cheng, X. Excision of 5-hydroxymethyluracil and 5-carboxylcytosine by the thymine DNA glycosylase domain: its structural basis and implications for active DNA demethylation. Nucleic Acids Res. 40, 10203–14 (2012).
Clauson, C. L., Oestreich, K. J., Austin, J. W. & Doetsch, P. W. Abasic sites and strand breaks in DNA cause transcriptional mutagenesis in Escherichia coli. Proc. Natl. Acad. Sci. U S A 107, 3657–62 (2010).
Xing, X. W. et al. Mutagenic and cytotoxic properties of oxidation products of 5-methylcytosine revealed by next-generation sequencing. PLoS One 8, e72993 (2013).
Karino, N., Ueno, Y. & Matsuda, A. Synthesis and properties of oligonucleotides containing 5-formyl-2′-deoxycytidine: in vitro DNA polymerase reactions on DNA templates containing 5-formyl-2′-deoxycytidine. Nucleic Acids Res. 29, 2456–63 (2001).
Munzel, M. et al. Improved synthesis and mutagenicity of oligonucleotides containing 5-hydroxymethylcytosine, 5-formylcytosine and 5-carboxylcytosine. Chemistry 17, 13782–8 (2011).
Gordon, A. J. et al. Transcriptional infidelity promotes heritable phenotypic change in a bistable gene network. PLoS Biol. 7, e44 (2009).
Gordon, A. J., Satory, D., Halliday, J. A. & Herman, C. Heritable change caused by transient transcription errors. PLoS Genet. 9, e1003595 (2013).
Satory, D., Gordon, A. J., Halliday, J. A. & Herman, C. Epigenetic switches: can infidelity govern fate in microbes? Curr. Opin. Microbiol. 14, 212–7 (2011).
Morreall, J. F., Petrova, L. & Doetsch, P. W. Transcriptional mutagenesis and its potential roles in the etiology of cancer and bacterial antibiotic resistance. J. Cell Physiol. 228, 2257–61 (2013).
Bregeon, D. & Doetsch, P. W. Transcriptional mutagenesis: causes and involvement in tumour development. Nat. Rev. Cancer 11, 218–27 (2011).
Saxowsky, T. T., Meadows, K. L., Klungland, A. & Doetsch, P. W. 8-Oxoguanine-mediated transcriptional mutagenesis causes Ras activation in mammalian cells. Proc. Natl. Acad. Sci. U S A 105, 18877–82 (2008).
Bregeon, D., Peignon, P. A. & Sarasin, A. Transcriptional mutagenesis induced by 8-oxoguanine in mammalian cells. PLoS Genet. 5, e1000577 (2009).
Wu, S. C. & Zhang, Y. Active DNA demethylation: many roads lead to Rome. Nat. Rev. Mol. Cell. Biol. 11, 607–20 (2010).
Liutkeviciute, Z. et al. Direct decarboxylation of 5-carboxylcytosine by DNA C5-methyltransferases. J. Am. Chem. Soc. 136, 5884–7 (2014).
Guo, J. U., Su, Y., Zhong, C., Ming, G. L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–34 (2011).
Bomgarden, R. D. et al. Opposing effects of the UV lesion repair protein XPA and UV bypass polymerase eta on ATR checkpoint signaling. EMBO J. 25, 2605–14 (2006).
Ji, D., Lin, K., Song, J. & Wang, Y. Effects of Tet-induced oxidation products of 5-methylcytosine on Dnmt1- and DNMT3a-mediated cytosine methylation. Mol. Biosyst. 10, 1749–52 (2014).
Yuan, B. et al. The roles of DNA polymerases κ and ι in the error-free bypass of N2-carboxyalkyl-2′-deoxyguanosine lesions in mammalian cells. J. Biol. Chem. 286, 17503–11 (2011).
Yuan, B., O'Connor, T. R. & Wang, Y. 6-Thioguanine and S6-methylthioguanine are mutagenic in human cells. ACS Chem. Biol. 5, 1021–7 (2010).
Baker, D. J. et al. Nucleotide excision repair eliminates unique DNA-protein cross-links from mammalian cells. J. Biol. Chem. 282, 22592–604 (2007).
You, C., Dai, X., Yuan, B. & Wang, Y. Effects of 6-thioguanine and S6-methylthioguanine on transcription in vitro and in human cells. J. Biol. Chem. 287, 40915–23 (2012).
Delaney, J. C. & Essigmann, J. M. Assays for determining lesion bypass efficiency and mutagenicity of site-specific DNA lesions in vivo. Methods Enzymol. 408, 1–15 (2006).
Acknowledgements
We thank Prof. Timothy R. O'Connor for providing the initial pTGFP-Hha10 vector and Prof. Karlene A Cimprich for kindly providing XPA-complemented (GM15876A) and XPA-deficient (XP12RO) lines. This work was supported by the National Institutes of Health (R01 CA101864).
Author information
Authors and Affiliations
Contributions
C.Y. and Y.W. designed the research, analyzed the data and wrote the paper; C.Y., D.J. and X.D. performed the research; Y.W. conceived and supervised the study.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Materials
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
You, C., Ji, D., Dai, X. et al. Effects of Tet-mediated Oxidation Products of 5-Methylcytosine on DNA Transcription in vitro and in Mammalian Cells. Sci Rep 4, 7052 (2014). https://doi.org/10.1038/srep07052
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07052
This article is cited by
-
Quantitative measurement of transcriptional inhibition and mutagenesis induced by site-specifically incorporated DNA lesions in vitro and in vivo
Nature Protocols (2015)
-
5-Formylcytosine alters the structure of the DNA double helix
Nature Structural & Molecular Biology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.