Abstract
The paper shows how a table top superbright microfocus laboratory X-ray source and an innovative restoring-data algorithm, used in combination, allow to analyze the super molecular structure of soft matter by means of Small Angle X-ray Scattering ex-situ experiments. The proposed theoretical approach is aimed to restore diffraction features from SAXS profiles collected from low scattering biomaterials or soft tissues and therefore to deal with extremely noisy diffraction SAXS profiles/maps. As biological test cases we inspected: i) residues of exosomes' drops from healthy epithelial colon cell line and colorectal cancer cells; ii) collagen/human elastin artificial scaffolds developed for vascular tissue engineering applications; iii) apoferritin protein in solution. Our results show how this combination can provide morphological/structural nanoscale information to characterize new artificial biomaterials and/or to get insight into the transition between healthy and pathological tissues during the progression of a disease, or to morphologically characterize nanoscale proteins, based on SAXS data collected in a room-sized laboratory.
Similar content being viewed by others
Introduction
Small angle X-ray scattering (SAXS) is a method extensively used for the structural analysis of a wide range of materials, such as metals, alloys, polymers as well as biologic macromolecules in solution, porous materials, nanoparticles, etc1,2,3. Thousands of papers have been published to describe this X-ray based technique and acknowledge them all is a titanic effort. SAXS ability consists in retrieving a morphological (size/shape) information of the scattering objects, or - in case of ordered systems - in a structural analysis of nanoscale lattice periodicities4,5,6,7,8,9,10,11. Over the last decades, SAXS has been increasingly employed in the study of biological macromolecules thanks first of all to the availability of brighter synchrotron radiation sources with a time resolution down to sub-ms and, secondly, to the improvement of SAXS data analysis methods allowing reliable ab-initio shape and domain structure determination and detailed modeling of macromolecular complexes12.
SAXS technique has found interesting applications to medicine: i) for the analysis of healthy or pathologic tissues extracted from bones13,14,15,16,17,18,19,20,21,22; cornea23,24,25; breast26,27,28,29,30,31; brain32,33,34; ii) for the characterization of nano-materials employed as novel therapeutic vectors35,36; iii) for the investigation of biomaterials for tissue engineering37,38,39, to cite few references in the field. Indeed, SAXS has shown to be a promising tool to detect structural changes at supramolecular level and in some cases a clear relationship between tissue changes and disease development has been derived31,40. For example, characteristic parameters extracted from SAXS patterns were used to differentiate human breast tissues aimed to a disease-related classification26,29,30,31. A key issue in most of the cited experiments was the need for a high brilliance synchrotron radiation X-ray source to obtain high-quality scattering patterns, although a limited number of papers made use of laboratory systems mainly for bone and breast tissues analysis. The possibility to use a laboratory X-ray source for SAXS studies in nanomedicine is extremely important, because - in order to transfer this knowledge into a clinic - a room-sized system is mandatory.
Recently, a new class of home laboratory X-ray sources has shown brightness exceeding some second generation bending magnet synchrotron radiation beamlines (such as the beamline X33 of the European Molecular Biology Laboratory (EMBL) at the DORIS III storage ring41). Such remarkable advances in laboratory instrumentation has proved to be effective for in-house crystallographic research42, especially if empowered by original algorithms able to significantly improve the signal-to-noise ratio and to enhance fringe visibility and therefore virtually ameliorate source coherence43,44. As a result, interesting performances and flexibility have been already demonstrated in ex-situ Grazing Incidence SAXS45 on self-assembled nanoscale inorganic superstructures, as well as in ex-situ SAXS experiments on air-dried rat tail tendon44 or residues of cell exosomes46. The experiments were realized to quantitatively compare the performances of the actual table top instrumentation with European Synchrotron Radiation Facility (ESRF, ID01 beamline) and Synchrotron Light Source (SLS, cSAXS beamline) SAXS beamlines and proved its viability in detecting structural changes on a molecular/nanoscale level in inorganic matter43 as well as in fiber tissues44.
In this paper, the restoration algorithm developed for SAXS data44 was further improved with the aim to extend data treatment from 1D to 2D as well as to deal with data collected from biological low-scattering soft tissues and therefore with extremely noisy diffraction profiles/maps. The novel algorithm contains a denoising/deconvolution iterative procedure, working (if needed) directly on the 2D raw data, studied to deal with SAXS data collected from soft tissues with laboratory instrumentation, affected by severe noise and background problems. In particular, we will focus our attention on different nanomedicine-related cases; (i) residues of exosomes' drops from healthy epithelial colon cell line and colorectal cancer cells, as an example of SAXS used a diagnostic tool; (ii) collagen/human elastin scaffolds, to show a practical case of vascular tissue engineering applications; (iii) apoferritin protein, to prove the effectiveness of the algorithm on bioSAXS data. The paper is organized as follows: the original and restored SAXS profiles are described and discussed in the Results and Discussion sections respectively, the Methods section contains a description of the restoration algorithm, for maps (2D) and profiles (1D) SAXS data, to prove its efficiency for noisy SAXS signals embedded in a high background contribution. A brief description of the SAXS table-top facility used to collect the data is given in the Methods.
Results
In the following we present three examples of application of the algorithm, described in the Methods section, to very noisy or strongly background-affected SAXS data. The aim of these examples is to show the potentialities of this table top instrumentation, empowered by the restoration algorithm, to demanding cases of bio-matter.
SAXS analysis of exosomes
Exosomes are 30–100 nm diameter membrane vesicles released by most cells types, including tumor cells, that contribute in many aspects of physiology and disease and in intercellular communication47. In particular, tumor exosomes are gaining increasing interest in medicine and oncology as efficient markers for the delivery of defined signals, since they contain a great variety of bioactive molecules, such as proteins and RNA (mRNA and miRNA)48, that could promote tumor progression by means of other cells49. However, there is also increasing evidence that these nanovesicles may contribute to cancer by favoring metastatic niche onset50. Recent findings highlight the potential of exosomal profiles as diagnostic biomarkers of disease through a noninvasive blood test51.
Here, SAXS diffraction patterns have been collected from exosomes derived from two different colon cell lines: CCD841-CoN (healthy epithelial colon cell line) and HCT116 (human colorectal carcinoma cell line). An innovative procedure for isolation of exosomes and preparation of the sample for the SAXS measurements is described elsewhere46,52. With respect to the laboratory SAXS data already published in Accardo et al 201346, the SAXS data here presented were acquired on the same specimens but for a much lower time, down to 100 s. This was to have a controlled experimental test of the proposed algorithm, but at very reduced collection time and therefore reduced dose of X-ray exposure.
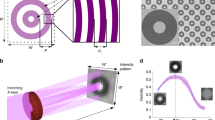
Figures 1a and 1c display the 2D SAXS frames from CCD and HCT collected at an acquisition time of 1200 and 100 s, respectively, whose azimuthal integrations are shown in Figures 1b and 1d (black curves). Starting from these raw data, both denoising, background subtraction and deconvolution have been performed and the results showed as red and blue profiles, respectively, in Figures 1b and 1d. For clarity, the deconvoluted profiles are properly scaled with respect to the original and denoised ones. The comparison clearly shows the efficiency of the restoration algorithm to extract relevant features from the raw data (see Discussion).
2D SAXS patterns from (a) CCD and (c) HCT residues, collected at the XMI-L@b at an acquisition time of 1200 s and 100 s, respectively; original (dotted black curve), denoised (red profile) and deconvolved (blue profile) 1D SAXS patterns, after restoration, are shown in (b) for CCD and (d) HCT residues.
SAXS studies on collagen-based three-dimensional scaffolds
In the second example described in this work, we report on the SAXS investigation of 3D collagen based scaffolds artificially produced as vessels. Three-dimensional biomimetic scaffolds have recently found extensive applications in biomedical tissue engineering, thanks to their micro-scaled design analogous to the native extracellular matrix53,54. In particular, natural polymers such as collagen and elastin, represent a promising alternative in creating three-dimensional scaffolds for vascular cell tissue engineering. Actually type I collagen combines suitable biomechanical properties as well as excellent biological and hematological properties and the use of recombinant Human Elastin-Like Polypeptides (HELPs) is a very promising complement able to significantly improve the biomechanical properties of three-dimensional collagen matrices in terms of tensile stress and elastic modulus55,56.
The artificial scaffolds have been prepared by extracting type I collagen from rat tail tendon according to a protocol described elsewhere57,58. Then, 50 mg/ml of Human Elastin-like Polypeptide (HELP), a bioinspired component that mimics human elastin, was added to the collagen blend and jellified with cells trapped within59. The samples have been also left in culture for 3, 7 and 21 days, in presence and absence of HELP. All the samples were fixed in formaldehyde 4%/water solution and maintained at 4°C until the analyses were performed. SAXS patterns were collected from tissues inserted into Lindemann glass capillary tubes of 0.7 mm diameter, to permanently keep them in formaldehyde 4%/water solution.
In Figure 2a we report the 2D SAXS data of porcine aorta, which has been collected as reference model for three-dimensional scaffolds, thanks to the high structural correlation between the human and porcine vessel walls. The raw 2D data, once radially integrated, are shown as 1D profile in Figure 2b (green curve). In Figure 2b SAXS patterns collected, for around 8 h, for collagen scaffolds, after 21 days of culture with (black) and without (red) HELP, are also displayed. Figure 2c compares the same profiles after application of the restoration algorithm, while the numbers in the upper side of the figure denote the collagen fiber diffraction orders.
(a) 2D SAXS of the aorta porcine benchmark material; (b) comparison between 1D raw data SAXS profiles of the collagen scaffolds, with (black) and without (red) HELP, after 21 days of culture and of the native porcine vessel (green). (b) comparison between 1D SAXS profiles after application of the restoring algorithm of the collagen scaffolds, with (black) and without (red) HELP, after 21 days of culture and of the native porcine vessel (green).
SAXS studies on apoferritin protein in aqueous solution
In Figure 3 the final test was performed on bioSAXS data from horse spleen apoferritin (ApoFt) purchased from Sigma60. Apoferritin is a globular nanosized cage-shaped protein composed by 24 subunits forming a stable and soluble hollow sphere. It derives from ferritin (450 kDa), a ubiquitous intracellular protein that stores iron and releases it in a controlled fashion.
(a) 1D SAXS of apoferritin: original (black) and denoised data (red); (b) comparison between laboratory (red) and synchrotron (black) data collected on the same sample (c) denoised data (dotted profile) compared with the best fit (red curve); (d) pair distribution function extracted by the fit in (c) and atomic model (inset) derived by fitting the denoised data with Dammif program.
Raw data - collected for 600 s and subtracted for the buffer contribution - are shown as black profile in panel (a) along with the denoised pattern (red curve), the latter compared with synchrotron data60,61 (black profile of Figure 3b) collected at the Austro SAXS beamline at ELETTRA facility on the same sample. Denoised laboratory data were fitted with Gnom62 (Figure 3c) and modeled with Dammin/Dammif62 (inset Figure 3d) programs, respectively and the pair distribution function was derived (Figure 3d).
Discussion
The first example holds as a controlled test as the same biological residues were extensively studied also with synchrotron radiation46. The application of the restoration algorithm improved the visibility of the diffraction peaks, beyond the first order and consequently the accuracy in the lattice periodicity determination. This allowed to determine the periodicity for the CCD and HCT exosomes of 14.6 nm ± 0.5 nm and 15.7 nm ± 0.5 nm, respectively, in perfect agreement with data acquired by micro-SAXS measurements at ESRF synchrotron (ID13 beamline)46. A lamellar morphology was also inferred from SAXS data, confirmed by SEM pictures of cross sections of the exosomes residues46.
The possibility to investigate the microstructure of healthy and diseased residues of exosomes proved the effectiveness of the SAXS technique even with a table-top X-ray microsource if combined with the restoration algorithm here described.
This first result shows how the algorithm properly works even for diffraction signals acquired only in 100 s (case of the HCT residues), with laboratory instrumentation, offering a broad range of applications for different radiation sensitive tissues, as often is the case with soft matter.
The second example, in Figure 2, shows how recovering structural information also for biomimetic engineered tissues. Also in this case, the SAXS profiles were heavily affected by the background, being the diffraction peaks hardly distinguishable from it. The restored profiles (Figure 2c) of all the investigated scaffolds (porcine aorta and collagen or collagen/HELP vessels), show many clear diffraction peaks, which allowed us to appreciate common features and almost the same periodicity among samples. In order to perform a quantitative comparison between the structural features of porcine vessel and collagen scaffolds, as a function of the days of culture (3-7-21 days) and to relate it to the presence or absence of HELP, the correlation coefficients between the collagen profiles and the one from the porcine vessel have been calculated over an interval of q ranging from 0.026 Å−1 to 0.100 Å−1.
Results, reported in Table 1, show that the correlation increases with the time of culture and in presence of HELP, reaching the highest correlation, cij = 0.72, for the longest culturing period (21days) in the presence of HELP, with a level of significance of 99%59.
The last example, in Figure 3, reports the results of the denoising procedure, applied to bioSAXS collected on the apoferritin protein in aqueous solution. Original raw data show a quite noisy pattern and therefore the denoising procedure is here a fundamental step before proceeding with data analysis. The results of the analysis was a gyration radius of Rg = 5.1 ± 0.2 nm, a pair distribution function and an empty sphere model (panels c and d of Figure 3) in perfect agreement with synchrotron derived findings60.
The reported results demonstrate the effective combination of a table top superbright microfocus X-ray source and a proper restoration algorithm, here employed for the investigation of bio-matter or soft natural or engineered tissues. The three examples here chosen - residues of exosomes' drops from healthy and pathologic tissues, collagen/human elastin artificial scaffolds developed for vascular tissue engineering applications and protein in solutions - are severe tests of the overall approach because are low scattering samples, whose SAXS diffraction signal is hardly detectable out of the matrix background or highly affected by noise.
The present equipment typology is fully compatible with real clinical investigations, where on site and room-sized equipment are mandatory. Therefore, a future could be envisioned where such a table top SAXS instrument could be considered also as novel diagnostic technique.
Methods
Algorithm description and application to extreme cases: denoising approach for 2D SAXS data
The algorithm reported in the paper is an evolution of the previous one (De Caro et al.44): here it has been introduced an iterative denoising/deconvolution procedure which can work, if needed, directly on the 2D raw data (not only on 1D profiles as for the previous one). The novelty therefore is two-fold: i) an original denoising algorithm was combined to the deconvolution procedure, where denoising is a fundamental requirement in case of lab SAXS data collected from soft tissue considering the poor signal to noise ratio; ii) the present algorithm allows to denoise and deconvolute directly the original 2D frames.
Following the same notation used in De Caro et al.44, the measured integrated intensity I(qi) at the scattering vector qi is related to the “ideal” profile Iid(qi) by means of the following equation:

where  denotes the convolution product. Here, the difference between Iid(qi) and the measured one I(qi) is caused by instrumental broadening effects like the finite beam divergence. The instrumental broadening is described by means of far-field source intensity angular distribution function Is(qi), experimentally determined. Background intensity Ibk(qi) and noise n(qi) are taken into account.
denotes the convolution product. Here, the difference between Iid(qi) and the measured one I(qi) is caused by instrumental broadening effects like the finite beam divergence. The instrumental broadening is described by means of far-field source intensity angular distribution function Is(qi), experimentally determined. Background intensity Ibk(qi) and noise n(qi) are taken into account.
We report in Figures 4a and 4b the 2D SAXS experimental data acquired on rat tail tendon and the corresponding denoised/deconvoluted one, as obtained by our algorithm, respectively.
The new denoising/deconvolution procedure here applied is iterative: each cycle is based on the alternation of a convolution with a Gaussian kernel of standard deviation σ and a deconvolution with a Gaussian kernel whose standard deviation is slightly smaller than σ. The final effect of this alternation is denoising/deconvolution of data. At the end of the n-th cycle each intensity value of the map In(qi) is updated by combining the filtered value Ifil,n-1(qi), obtained by the previous iteration, with raw data I(qi) in the following way:

where Ifil,n-1(qi) = Conv[Deconv[In-1(qi)]] is the denoised intensity, obtained by applying the convolution and the deconvolution steps on the map of the previous cycle; W(qi) = 1 − G(qi) is the weight function, being G(qi) a 2D Gaussian function having the maximum in the center of the SAXS 2D frame and the same full width at half maximum (FWHM) as the azimuthal average of the SAXS 2D frame. This weighting scheme leaves almost unchanged higher intensity signal, corresponding to smaller qi values, but allows the denoising of smaller intensity one (more affected by noise). The final result, shown in Figure 4b, leads to a great improvement of the visibility of smaller SAXS intensities, hidden by the high noise raw data, leaving unchanged higher intensity values. In particular the denoised 2D frame, shown in Figure 3b, has been obtained by putting the standard deviation of the convolving function equal to 4 pixels; the one of the deconvolving step equal to 3 pixel; the whole procedure was repeated for 10 cycles.
After denoising, higher diffraction orders become visible in the 2D SAXS map, even without any background subtraction.
Algorithm description and application to extreme cases: 1D profiles collected from low-scattering materials
Several noised and convoluted one-dimensional (1D) SAXS simulated profiles were computed, from Eq. (1), to reproduce, under control, typical SAXS experimental data collected from biomaterials, which were used to verify if the deconvolution algorithm was able to correctly extract the known Iid(qi) from I(qi). Before the deconvolution step, we applied the denoising iterative procedure described in the previous sub-section which acts directly on raw experimental data with the purpose to prevent noise enhancement during the subsequent signal restoration process.
We characterized the simulated deconvolution tests by means of the following quantities:
‐ the signal-to-background visibility V = <I(qi) − Ibk(qi)>/<I(qi)>, where <…> denotes averaging on the measured qi range;
‐ the relative error ε = (<w(qi)2(I(qi) − Ibk(qi))>)0.5/<I(qi) − Ibk(qi)> on the evaluation of Icon(qi) = I(qi) − Ibk(qi), calculated by means of the error propagation formula, where the function w(qi) can be evaluated by adding all points of the 2D map that would contribute to the same qi value of the 1D profile and by calculating the square root of the so-obtained sum  ;
;
‐ the residual error R = Σi|Iid(qi) − Idec(qi)|/ΣiIid(qi) between the deconvolved Idec(qi) and the input profile Iid(qi) in the qi-range characterized by the presence of less intense diffraction peaks.
Figure 5a shows a simulated 1D profile (Input 1 - black curve) obtained from Eq. (1) by properly convoluting the Iid(qi) spectra - plotted in red (Solution1) - with the assumed beam divergence and adding the contributions of the background Ibk(qi) - green profile - and the noise n(qi). Let us note that almost all features of the deconvoluted Idec(qi) profile (deconvoluted 1 - blue curve) were correctly reconstructed. The residual factor R = 0.18, calculated in a middle region from pixel 100 to 300 and the relative error ε = 2.5%, give a quantitative idea of the quality of the reconstructed profile. This test indicates that all the original information of Iid(qi) could be correctly extracted from the SAXS profile with similar characteristics of visibility of the structure peaks with respect to the background when the signal-to-background visibility V is around 55%. The level of added noise is obviously related to the statistics of the X-ray counts. In our tests we have chosen maximum scattered intensity values which range between 10 and 100 counts, typical of laboratory experimental data, at least for the specific microsource described in the Methods section. It is worth noting that even if the maximum scattered intensity is small – namely less than 100 counts – the relative error ε obtained after the restoration algorithm is quite lower than a Poisson-noise relative error, because the considered 1D profiles are obtained by the integration over circular regions of a 2D map, which reduces the statistical fluctuations of intensities. It is possible to verify that the 1D reduction of the 2D map reduces the relative error by about two orders of magnitude.
(a) Simulated SAXS 1D profiles I(qi) (black curve) affected by an overlapped background intensity Ibk(qi) (green curve), by the finite-size convolution effects, i.e., I(qi) = Icon(qi) + Ibk(qi) and by noise n(qi) (see main text). The red curve is the ideal intensity Iid(qi) that should be extracted by I(qi) by deconvolving the finite source size effects and subtracting the background intensity. The blue curve is the deconvolved Idec(qi) profile which is obtained by means of the new proposed algorithm. We have V = 55%; ε = 2.5%; R = 0.18; (b) Same notation as in Figure 5a. We have V = 23%; ε = 8.3%; R = 0.26.
In Figure 5b we report the results of another test in which we considered a worsened signal-to-background visibility V, of about 23%. We assumed a maximum intensity I(qi) equal to about 20 counts. In this test the relative error increases to about ε = 8.3% and the quality of the reconstructed Idec(qi), when compared with Iid(qi), is obviously worse than in the previous case, as confirmed by the obtained residual error value R = 0.26. Nevertheless almost all important features of Iid(qi) have been correctly reconstructed in the deconvoluted Idec(qi).
In Figure 6a we report the results of a third test in which we considered a quite low signal-to-background visibility V = 13%. In this case, the relative error increases to about ε = 16.7%. The zoom of Figure 6b gives an idea of the poor quality of the assumed I(qi) profile and of the noise level. It is interesting to verify the possibility to extract many useful information regarding the Iid(qi) profile by means of the proposed algorithm, as shown by the comparison between the blue and red curves in Figure 6b, even when starting from a very poor quality I(qi) profile. In fact, from a quantitative point of view, only the higher peaks of Figure 6b in the pixel range 125–250 are correctly reconstructed which, on the other hand, are the only peaks already clearly visible in the I(qi) profile. Nevertheless all other peaks which are almost completely hidden in I(qi) by background, convolution effects and noise, are sufficiently reconstructed in Idec(qi), giving much more information concerning the scattering sample than achievable from the convoluted profile. These results show that this version of the algorithm, here improved for the SAXS analysis of low scattering materials (such as soft tissues), could be a very useful tool to analyze SAXS 1D and 2D profiles, even in the unlucky case of a very low visibility (V) of the sample features.
(a) Same notation as in Figure 5. We have V = 13%; ε = 16.7%; R = 0.31; b) zoom of figure 6a with the red and blue profiles multiplied by a factor 10 to reduce the dynamical range of the plot to only two decades, thus showing the small features of the black profile from which the blue curve is reconstructed.
Experimental
A Fr-E+ SuperBright rotating anode copper anode microsource (Cu Kα, λ = 0.15405 nm, 2475W) is focused by a multilayer focusing optics (Confocal Max-Flux; CMF 15–105) to a SAXS three-pinhole camera (Rigaku SMAX-3000)45. The system is equipped with a Triton 20 gas-filled proportional counter (1024 × 1024 array, 195 µm pixel size) for SAXS acquisition, placed at a sample-detector distance of around 2200 mm. This distances give access to a range of scattering vector moduli (q = 4πsinθ/λ, where θ is half the scattering angle) of about 0.006–0.2 Å−1. A detailed description of the instrument performances can be found in Altamura et al.45.
References
Craievich, A. F. [Small-angle X-ray scattering by nanostructured materials]. Characterization of Sol-Gel Materials and Products, Kluwer Academic Publishers, Massachusetts, USA [vol. 2, 161–189] (2005).
Petoukhov, M. V. & Svergun, D. I. Applications of small-angle X-ray scattering to biomacromolecular solutions. Int. J. Biochem. Cell Biol. 45, 429–437 (2013).
Blanchet, C. E. & Svergun, D. I. Small-Angle X-Ray Scattering on Biological Macromolecules and Nanocomposites in Solution. Annu. Rev. Phys. Chem. 64, 37–54 (2013).
Glatter, O. & Kratky, O. Small-Angle X-ray Scattering (Academic Press, London, 1982).
Guinier, A. & Fournet, G. Small Angle Scattering of X-Rays (Wiley, New York, 1955).
Feigin, L. A. & Svergun, D. I. Structure analysis by small-angle X-ray and neutron scattering (Plenum Press, New York, 1987).
Wess, T. J., Hammersley, A. P., Wess, L. & Miller, A. A consensus model for molecular packing of type I collagen. J. Struct. Biol. 122, 92–100 (1998).
Koch, M. H. J., Vachette, P. & Svergun, D. I. Small-angle scattering: A view on the properties, structures and structural changes of biological macromolecules in solution. Q. Rev. Biophys. 36, 147–227 (2003).
Orgel, J. P. R. O., Irving, T. C., Miller, A. & Wess, T. J. Microfibrillar structure of type I collagen in situ. Proc. Natl. Acad. Sci. USA 103, 9001–9005 (2006).
Svergun, D. I., Shtykova, E. V., Volkov, V. V. & Feigin, L. A. Small-angle X-ray scattering, synchrotron radiation and the structure of bio- and nanosystems. Crystallogr. Rep. 56, 725–750 (2011).
Dorfs, D. et al. L. Comprehensive Nanoscience and Technology. (Elsevier B.V., Amsterdam, 2011), [vol. 1, 219–270].
Svergun, D. I. Small-angle scattering studies of macromolecular solutions. J. Appl. Cryst. 40, s10–s17 (2007).
Fratzl, P., Fratzl-Zelman, N. & Klaushofer, K. An x-ray scattering investigation of turkey leg tendon. Biophys. J. 64, 260–266 (1993).
Fratzl, P. et al. S. Fibrillar structure and mechanical properties of collagen. J. Struct. Biol. 122, 119–122 (1998).
Fratzl, P., Gupta, H. S., Paschalis, E. P. & Roschger, P. Structure and mechanical quality of the collagen–mineral nano-composite in bone. J. Mater. Chem. 14, 2115–2123 (2004).
Rinnerthaler, S. et al. P. Scanning Small Angle X-ray Scattering Analysis of Human Bone Sections. Calcif. Tissue Int. 64, 422–429 (1999).
Al-Jawad, M. et al. D.J. 2D mapping of Texture and Lattice Parameters of Dental Enamel. Biomaterials 28, 2908–2914 (2007).
Grabner, B. et al. Age-and genotype-dependence of bone material properties in the osteogenesis imperfecta murine model. Bone 29, 453–457 (2001).
Paris, O. From diffraction to imaging: New avenues in studying hierarchical biological tissues with x-ray microbeams. Biointerphases 3, FB16–FB26 (2008).
Roschger, P., Paschalis, E. P., Fratzl, P. & Klaushofer, K. Bone mineralization density distribution in health and disease. Bone 42, 456–466 (2008).
Giannini, C. et al. Correlative light and scanning X-ray scattering microscopy of healthy and pathologic human bone sections. Sci. Rep. 2, 435 (2012).
Giannini, C. et al. O. Scanning SAXS-WAXS microscopy on osteoarthritis-affected bone - an age-related study. J. Appl. Cryst. 47, 110–117 (2014).
Aghamohammadzadeh, H., Newton, R. H. & Meek, K. M. X-Ray Scattering Used to Map the Preferred Collagen Orientation in the Human Cornea and Limbus. Structure 12, 249–56 (2004).
Abahussin, M. et al. 3D collagen orientation study in human cornea using x-ray diffraction and femtosecond laser technology. Invest. Ophthalmol. Vis. Sci. 50, 5159–5164 (2009).
Boote, C. et al. Quantification of collagen organization in the peripheral human cornea at micron-scale resolution. Biophys J. 101, 33–42 (2011).
Avtandilov, G. et al. Human tissue analysis by small-angle X-ray scattering. J. Appl. Cryst. 33, 511–514 (2000).
Round, A. et al. A preliminary study of breast cancer diagnosis using laboratory based small angle x-ray scattering. Phys. Med. Biol. 50, 4159–4168 (2005).
James, V. J. Fiber diffraction of skin and nails provides an accurate diagnosis of malignancies. Int. J. Cancer 125, 133–138 (2009).
Sidhu, S. et al. X-ray scattering for classifying tissue types associated with breast disease. Med. Phys. 35, 4660–4670 (2008).
Sidhu, S. et al. Classification of breast tissue using a laboratory system for small-angle x-ray scattering (SAXS). Phys. Med. Biol. 56, 6779–6991 (2011).
Lewis, R. A. et al. D.R. Breast cancer diagnosis using scattered X-rays. J. Synchrotron Rad. 7, 348–352 (2000).
Malinchik, S. B., Inouye, H., Szumowski, K. E. & Kirschner, D. A. Structural analysis of Alzheimer's beta(1–40) amyloid: protofilament assembly of tubular fibrils. Biophys J. 74, 537–545 (1998).
De Felici, M. et al. Structural characterization of the human cerebral myelin sheath by small angle x-ray scattering. Phys. Med. Biol. 53, 5675–5688 (2008).
Jensen, T. H. et al. Brain tumor imaging using small-angle x-ray scattering tomography. Phys. Med. Biol. 56, 1717–1726 (2011).
Akiba, I. et al. Encapsulation of a Hydrophobic Drug into a Polymer-Micelle Core Explored with Synchrotron SAXS. Langmuir 26(10), 7544–7551 (2010).
Bigall, N. C. et al. Colloidal Ordered Assemblies in a Polymer Shell – A Novel Type of Magnetic Nanobeads for Theranostic Applications. Chem. Mater. 25, 1055–1062 (2013).
Guagliardi, A. et al. Toward the x-ray microdiffraction imaging of bone and tissue-engineered bone. Tissue Eng. B 15, 423–4429 (2009).
Guagliardi, A. et al. M. Debye function analysis and 2D imaging of nanoscaled engineered bone. Biomaterials 31, 8289–8298 (2010).
Campi, G. et al. Early stage mineralization in tissue engineering mapped by high resolution X-ray microdiffraction. Acta Biomater. 8, 3411–3418 (2012).
Fernandez, M. et al. P. Phys. Med. Biol. 47, 577–592 (2002).
Blanchet, C. E. et al. Instrumental setup for high-throughput small- and wide-angle solution scattering at the X33 beamline of EMBL Hamburg. J Appl Cryst. 45, 489–495 (2012).
Altamura, D. et al. Exploiting GISAXS for the Study of a 3D Ordered Superlattice of Self-Assembled Colloidal Iron Oxide Nanocrystals. Cryst.Growth Des. 12, 5505–5512 (2012).
De Caro, L. et al. A superbright X-ray laboratory microsource empowered by a novel restoration algorithm. J. Appl. Cryst. 45, 1228–1235 (2012).
De Caro, L. et al. Rat-tail tendon fiber SAXS high-order diffraction peaks recovered by a superbright laboratory source and a novel restoration algorithm. J. Appl. Cryst. 46, 672–78 (2013).
Altamura, D. et al. X-ray microimaging laboratory (XMI-LAB). J. Appl. Cryst. 45, 869–73 (2012).
Accardo, A. et al. Superhydrophobic surfaces allow probing of exosome self organization using X-ray scattering. Nanoscale 5, 2295–99 (2013).
Huotari, J. & Helenius, A. Endosome maturation. Embo J. 30, 3481–3500 (2011).
Filipazzi, P., Burdek, M., Villa, A., Rivoltini, L. & Huber, V. Recent advances on the role of tumor exosomes in immunosuppression and disease progression. Semin. Cancer Biol. 22, 342–349 (2012).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Nieuwland, R., van der Post, J. A., Lok, C. A., Kenter, G. & Sturk, A. Microparticles and exosomes in gynecologic neoplasias. Semin. Thromb. Hemost. 36, 925–929 (2010).
Simpson, R. J., Lim, J. W., Moritz, R. L. & Mathivanan, S. Exosomes: proteomic insights and diagnostic potential. Expert Rev. Proteomic 6, 267–83 (2009).
Tirinato, L. et al. Analysis On Exosomes Using Super-Hydrophobic Surfaces. Microelectron. Eng. 97, 337–340 (2012).
Dutta, C. R. & Dutta, A. K. Cell-interactive 3D-scaffold; advances and applications. Biotechnol. Adv., 27, 334–339 (2009).
Kim, B. S., Nikolovski, J., Bonadio, J., Smiley, E. & Mooney, D. J. Engineered smooth muscle tissues: Regulating cell phenotype with the scaffold. Exp. Cell. Res. 251, 318–328 (1999).
Bandiera, A. et al. Biotechnol. Expression and characterization of human-elastin-repeat-based temperature-responsive protein polymers for biotechnological purposes. Appl. Biochem. 42, 247–256 (2005).
Bandiera, A., Sisto, P. & Urbani, R. Comparison of Thermal Behavior of Two Recombinantly Expressed Human Elastin-Like Polypeptides for Cell Culture Applications. Biomacromolecules 11, 3256–3265 (2010).
Rajan, N., Habermehl, J., Cotè, M. F., Doillon, C. J. & Mantovani, D. Preparation of ready-to-use, storable and reconstituted type I collagen from rat tail tendon for tissue engineering applications. Nat. Protoc. 1, 2753–2758 (2006).
Boccafoschi, F., Bosetti, M., Mosca, C., Mantovani, D. & Cannas, M. The role of shear stress on mechanically stimulated engineered vascular substitutes: influence on mechanical and biological properties. J. Tissue Eng. Regen. M. 6, 60–67 (2012).
Boccafoschi, F. et al. Human elastin polypeptides (HELP) improve the biomechanical properties of three-dimensional matrices through the regulation of elastogenesis. J. Biomed. Mater. Res: A, Accepted Article (2014).
Ciasca, G. et al. Reconstitution of aluminium and iron core in horse spleen apoferritin. J Nanopart Res 13, 6149–6155 (2011).
Ciasca, G. et al. Transient state kinetic investigation of ferritin iron release. Appl. Phys. Lett. 100(7), 073703 (2012)
Data analysis software ATSAS 2.5.2 A program suite for small-angle scattering data analysis from biological macromolecules, http://www.embl-hamburg.de/biosaxs/software.html
Acknowledgements
The authors would like to thank the FIRB 2009/2010 project “Rete integrata per la Nano Medicina (RINAME)” – RBAP114AMK_006. Rocco Lassandro, Caterina Chiarella, Lucrezia Cassano, Brunella Aresta and Giovanni Filograsso are acknowledged for technical and administrative support in the XMI-LAB.
Author information
Authors and Affiliations
Contributions
T.S., D.A. and C.G. realized the SAXS experiments at the XMI-L@b, M.R. and F.B. prepared the collagen based vessels, L.T. and E.d.F. prepared the exosome residues and characterize them with microSAXS at ESRF; Gab.C. and Gae. C. prepared the apoferritin samples and measure the SAXS pattern at Elettra. L.D.C. developed the restoring algorithm. D.S. transformed the algorithm into a software which was used by T.S., D.A. and C.G. to analyze the SAXS data. T.S., L.D.C. and C.G. discussed the results and wrote the paper in close collaboration with all the authors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Sibillano, T., De Caro, L., Altamura, D. et al. An Optimized Table-Top Small-Angle X-ray Scattering Set-up for the Nanoscale Structural Analysis of Soft Matter. Sci Rep 4, 6985 (2014). https://doi.org/10.1038/srep06985
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06985
This article is cited by
-
Fabrication of fluorescent nanospheres by heating PEGylated tetratyrosine nanofibers
Scientific Reports (2021)
-
X-ray Fokker–Planck equation for paraxial imaging
Scientific Reports (2019)
-
Cross-beta nanostructures based on dinaphthylalanine Gd-conjugates loaded with doxorubicin
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.