Abstract
Status epilepticus (SE) is a life-threatening condition that can give rise to a number of neurological disorders, including learning deficits, depression and epilepsy. Many of the effects of SE appear to be mediated by alterations in gene expression. To gain deeper insight into how SE affects the transcriptome, we employed the pilocarpine SE model in mice and Illumina-based high-throughput sequencing to characterize alterations in gene expression from the induction of SE, to the development of spontaneous seizure activity. While some genes were upregulated over the entire course of the pathological progression, each of the three sequenced time points (12-hour, 10-days and 6-weeks post-SE) had a largely unique transcriptional profile. Hence, genes that regulate synaptic physiology and transcription were most prominently altered at 12-hours post-SE; at 10-days post-SE, marked changes in metabolic and homeostatic gene expression were detected; at 6-weeks, substantial changes in the expression of cell excitability and morphogenesis genes were detected. At the level of cell signaling, KEGG analysis revealed dynamic changes within the MAPK pathways, as well as in CREB-associated gene expression. Notably, the inducible expression of several noncoding transcripts was also detected. These findings offer potential new insights into the cellular events that shape SE-evoked pathology.
Similar content being viewed by others
Introduction
Status epilepticus (SE) is defined as persistent, unremitting seizure activity lasting 5–30 minutes1. This potentially lethal bout of sustained excitatory synaptic activity is caused by a variety of conditions, including, hemorrhage, stroke, viral infection and the withdrawal of anticonvulsive medication. In rodent models, pilocarpine-evoked SE is commonly utilized to profile neuroprotective and cell death signaling pathways, as well as potential cellular and molecular processes that underlie epileptogenesis. With respect to epileptogenesis, pilocarpine initiates a very well characterized step-wise pattern of pathophysiological changes that ultimately lead to the development of spontaneous seizure activity. Hence, the acute pilocarpine-induced SE event manifests at a behavioral level as tonic-clonic seizures that persist for many hours; within this time period, actuation of cell death signaling pathways and reactive gliosis are observed (reviewed in Turski et al., 1989). Subsequent to the SE phase, mice enter a ‘seizure silent’ period that persists for several weeks. During this period, alterations in synaptic organization are observed, including the well-characterized formation of recurrent granule cell collaterals2,3. Following this quiescent phase, animals enter a chronic epileptic-like state. At the cellular level, marked changes in neuronal excitability, cellular morphology and synaptic reorganization are observed within the hippocampus. In addition, increased angiogenesis, granule cell dispersion and aberrant neurogenesis within the subgranular zone of the dentate gyrus are also observed2,4.
At a molecular level, SE triggers an array of changes in cell death, neuroprotective and plasticity-associated signaling pathways. Along these lines, there is also a rapid rise in the generation of reactive oxygen species (ROS) levels, resulting in oxidative stress and damage to DNA, RNA, proteins and lipids5. Likewise, SE stimulates the activation of apoptotic signaling pathways6. Paralleling this rise in cell death signaling, SE also drives a robust neuroprotective response, including the expression of phase II detoxifying enzymes and antiapoptotic signaling cascades. Finally, consistent with neuroanatomical changes resulting from epileptogenesis, a number of studies have shown that there is an increase in expression of genes involved in axonal outgrowth and synapse formation3.
Many of these molecular events are likely resulting from altered transcriptional activity. Consistent with this, seizure activity is associated with an increase in SRF-, ARE- and CREB-dependant transcription7. Further, the activation state of upstream kinase pathways, including the p42/44 MAPK, p38(MAPK), JNK2/3, AKT and PKC is increased at distinct phases of epileptogenesis and in a cell-type-specific manner8,9,10. Notably, in addition to changes in the activation state of kinase pathways, epileptogenesis has also been shown to alter kinase expression patterns11.
To gain a clearer picture of the transcriptional events that could contribute to SE-evoked brain pathology, a number of studies have utilized high-throughput, discovery-based approaches, such as proteomics and large-scale microarray transcriptome profiling12,13,14. The development of next-generation deep sequencing technologies provides a new approach to such profiling, potentially allowing for the discovery of novel genes, noncoding transcripts and methylation changes in a high-throughput manner15. Because of its relatively recent advent, the statistical tools available for RNA-Seq analysis are still evolving, but the insights garnered from next-generation sequencing make it a compelling method for identifying novel biomarkers. Thus, through its ability to probe expression changes on a transcriptome-wide basis, RNA-seq approaches may yield additional therapeutic targets by highlighting novel genes and cellular signaling networks.
Here, we took advantage of these strategies to examine the differential expression and molecular regulation characterizing SE-evoked hippocampal pathology across epileptogenesis. We hypothesize that the discrete and tightly-regulated phases of the epileptogenic process are mediated by the regulation of transcriptionally-inducible gene expression patterns that subsequently give rise to the next phase of pathogenesis, ultimately manifesting themselves in spontaneous seizure activity. To this end, we performed whole transcriptome profiling to identify differentially expressed mRNAs at 12 hrs, 10 days and 6 weeks post-SE. These time points where selected to approximate the distinct phases of the epileptogenic process (acute, seizure silent and spontaneous-seizure phases) as described above and parallel the timepoints chosen for previous transcriptomic examinations of epileptogenesis16,17. The data presented here identify distinct alterations in gene expression and functional analysis provides potential clues regarding how these changes may contribute to the epileptogenic process.
Results
Altered gene expression across phases of epileptogenesis
To determine the transcriptional regulation that occurs over the course of epileptogenesis, we used Illumina-based high-throughput sequencing. Hippocampi were dissected 12 hours, 10 days and 6 weeks after induction of SE, as well as from vehicle–treated controls (Fig. 1A, B). These time points were selected to profile each stage of the epileptogenic process and are consistent with timepoints used in previous studies (acute, seizure silent and spontaneous-seizure phases)16,17. Concordantly, animals sacrificed during the acute timepoint exhibited robust excitotoxic cell death within the hippocampus (Fig. 1C), followed later by evidence of substantial mossy fiber synaptic reorganization within the dentate gyrus (Fig. 1D). Further, animals sacrificed at the 6-week time point exhibited spontaneous seizure activity, occurring every 10–20 minutes and lasting several seconds each. Libraries were sequenced to an average depth of ~3.4 million reads per group across the three timepoints (Supplemental Table 1). Because our cDNA libraries are 3′ biased, this depth was adequate to detect genes with low levels of transcription (1–10 copies per cell). Approximately 2.2 million reads mapped to unique locations on the M. musculus UCSC mm9 reference genome, with ~50% of these mapping to exons of known RefSeq genes (Supplemental Fig. 1).
Differential expression of mRNA across phases of epileptogenesis.
(A) Timeline depicts tissue collection across the progression of epileptogenesis. Animals were sacked 12 hours, 10 days and 6 weeks after pilocarpine induction of status epilepticus (SE). These timepoints correspond to the acute, seizure-silence and spontaneous-seizure phases, respectively, paralleling the time course of pathogeneses. (B) Schematic outline of processing and analysis procedures. Hippocampal tissue was collected from each time point and RNA was isolated and cDNA libraries were created. Libraries were sequenced and subsequent datasets were filtered and compared to controls to yield fold changes across experimental groups. Genes showing changes in regulation were examined for differential functionality using the Database for Annotation, Visualization and Integrated Discovery (DAVID) v6.7. (C) At 2 days post-SE, Fluoro-Jade B labeling revealed cell death within CA1, CA3 subfields and the hilar region (Hil) of the hippocampus. GCL, granule cell layer. Scale bar: 200 μm. (D) Timm staining revealed reorganization of mossy fiber projections within the hippocampus 6 weeks after the induction of SE. Arrows indicate mossy fiber sprouting within the intermolecular layer. Mol, molecular layer; MF, mossy fibers. Scale bars: 200 μm (left); 100 μm (right). (E) Hierarchical cluster analysis of differentially expressed genes between control, 12-hour, 10-day and 6-week samples (q < 0.05). Increases in expression are indicated in red hues and decreases are in green hues. (F) Venn diagram showing the number of genes with significant upregulation (q < 0.05) in each time point. Color schemes representing each time point are used in all subsequent figures.
To examine temporal regulation of gene expression, we performed an unsupervised clustering analysis of the 300 most significantly-regulated genes (Fig. 1E). Hierarchical clustering (hclust) showed distinct patterns with significant clusters of genes showing an increase at 12 hr and 10 days. The 6 week timepoint showed a mixed pattern of gene expression with most gene blocks showing a decrease relative to the 10 day time point. Cell death resulting from SE may contribute to reports of down regulated transcripts. Given the potential difficulty that cell death poses for the interpretation of the downregulated transcript set, our subsequent analysis focused more heavily on those transcripts that were upregulated after SE induction relative to untreated animals (downregulated genes, their fold changes and subsequent analysis are included for reference in Supplemental Fig. 1 and Supplemental Fig. 4). Of the total list of upregulated genes, only 197 genes were upregulated in all three timepoints, whereas the majority of genes in each timepoint were unique to their respective group (Fig. 1F). Of note, data corresponding to the 12-hour, 10-day and 6-week timepoints are represented by magenta, yellow and cyan (respectively) throughout our report, with overlapping datasets denoted by the resultant mix of colors presented in Figure 1F.
The transcriptional patterns revealed by our analysis offer a good degree of overlap with previously reported datasets, as well as reveal new avenues for investigation. SE is known to trigger robust excitotoxicity and cell death throughout the hippocampus18. In response, a range of neuroprotective and reparatory mechanisms are activated, which was reflected in our transcriptional analysis of epileptogenesis (Supplemental Fig. 1 and Supplemental Fig. 4N). For example, metallothionein, a Golgi-associated metal binding protein with neuroprotective effects after seizure insult, was upregulated at both the 12 hour (Fold Change [FC] = 2.54, q < 0.05) and 10 day (FC = 3.03, q < 0.05) timepoints18. Similarly, upregulation of hypoxia response genes like Hmox2 during the seizure-silent phase (FC = 3.31, q < 0.05) contribute to recovery after excitotoxic insult19. We also saw an increase in neurotrophic and growth factors (e.g., Bdnf: FC = 3.03, q < 0.05; Fgf1: FC = 3.31, q < 0.05) that play a protective role after induction of SE20,21. Induced vascularization is highly associated with the post-SE response and numerous genes linked with blood vessel development were upregulated across all three epileptogenic phases, including Vegf (FC = 4.64 [12 hours], 5.36 [10 days], 4.00 [6 weeks]), q < 0.05) the expression of which has a protective effect in the hippocampus after seizure activity22. Furthermore, although cell death is widespread throughout pathogenesis, an upregulation of expression related to axonogenesis was also prevalent across all three timepoints (Supplemental Fig. 1E, largest changes presented and discussed in Fig. 4 below). Aberrant mossy fiber sprouting leads to recurrent excitatory circuits that exacerbate seizure activity. We found numerous genes associated with axonal growth that were upregulated across epileptogenesis. Some are known to be upregulated with seizure activity (e.g., Srf: FC = 5.74 [12 hours], 5.58 [10 days], 6.53 [6 weeks], q < 0.05; Cxcl12: FC = 15.77 [12 hours], 24.67 [10 days], 24.15 [6 weeks], q < 0.05)23,24, but several were yet to be described within the context of epileptogenesis up to this point (e.g., Dpysl5: FC = 11.94 [12 hours], 15.50 [10 days], 14.95 [6 weeks], q < 0.05; Cyfip1: FC = 2.13 [12 hours], 2.42 [10 days], 2.48 [6 weeks], q < 0.05). Given the heterogeneous nature of cell-types within hippocampal tissue, interpretation of altered pathways should be undertaken with caution. Nevertheless, these genes sets may provide substantial insight when used in conjunctions with established literature on the signaling mechanisms underlying epileptogenesis.
Functional profiling of upregulated genes.
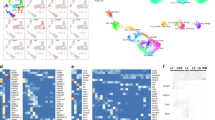
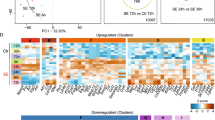
DAVID categories of upregulated genes were arranged into functional arrays to illustrate the proportions of four ontological domains (‘Molecular Functions’, ‘Cellular Components’, ‘Biological Processes' and ‘Signaling Pathways') dominated by each phase of epileptogenesis. Each category is represented by a single square and the color of the square denotes whether and when, its expression profile was affected following SE (using the color scheme established in Figure 1D). An interactive digital version of this figure that details each category is available in Supplemental Fig. 3 (Chrome web browser or iPad tablet recommended for greatest compatibility). A single square (GO:0007409 Axonogenesis), has been magnified and presented as an example here. Each square contains the total number of genes upregulated at each time point, followed by the names of those genes. In Supplemental Fig. 3, each square can be zoomed in to view its ontological term, as well as the names of all the genes upregulated in that category at each timepoint. The total number of upregulated genes at each timepoint is also provided. Genes upregulated at 6-weeks showed the broadest range of biological processes. In contrast, expression changes at the two earlier time points had more a focused range of processes. Relatively little overlap in function was observed between time points. http://obrietanlab.org.ohio-state.edu/.
Enriched ontological clusters across epileptogenesis.
The top 20 enriched clusters of DAVID categories are reported for the 12-hour, 10-day and 6-week timepoints. Clusters were sorted by enrichment score and the number of upregulated genes within each cluster is reported as bars corresponding to the lower axis. In addition, EASE Scores (DAVID's modified Fischer Exact P-value) for each cluster are depicted as a measure of the association strength of genes within each cluster with a line graph corresponding to the upper axis.
A subset of functionally-associated genes are upregulated at 12 hours, 10 days and 6 weeks post-SE.
DAVID analysis revealed an enrichment of upregulated genes associated with ‘blood vessel development’ across all phases of epileptogenesis. Here, relative fold changes are presented for the top five genes upregulated within this cluster.
The twenty unique genes with the greatest increases in expression at each timepoint are presented in Table 1, as well as the top twenty upregulated genes that were common to all three groups. Values reflect the average of statistically-significant fold changes across all replicates and splice-variant transcripts. Several genes, such as Cxcl12 and Notch3, had previously been reported to be involved in seizure-related neuronal activity and SE-affected tissue24,25. Again, several others had not been previously associated directly with seizure activity. These genes represent a wide array of functional categories, including transcriptional regulation (e.g., Atf7), regulation of cell survival (e.g., Cflar, Mul1) and neuronal outgrowth (e.g., Ltbp3, Dnaja3)26,27,28,29,30.
Pathogenic modification of molecular and cellular signaling
To investigate the functional effects of gene induction at each timepoint, we examined Gene Ontology (GO) terms across three domains: molecular functions, cellular components and biological processes (see Supplemental Fig. 2 for a glossary of terminology). In addition, we examined gene enrichment within Kyoto Encyclopedia of Genes and Genomes (KEGG) and BioCarta pathways. As detailed in the Methods section, we created an array of possible ontologies for five functional domains of interest (‘Molecular Functions’, ‘Cellular Components’, ‘Biological Processes' and ‘Signaling Pathways'), with each ontology represented as cube within the array, to produce a colorimetric profile of the gene functionalities upregulated at each timepoint. In addition to the static overview presented in Figure 2, Supplemental Fig. 3 offers an interactive digital version, which allows for dynamic exploration of each category. Each square can be zoomed in to view its ontological term, as well as the names of all the genes upregulated in that category at each timepoint. The total number of upregulated genes at each timepoint is also provided. The greatest diversity of changes appeared at the 6-week timepoint, whereas dysregulated genes at the earlier two timepoints showed more focused functionality. Interestingly, the greatest overlap in functional categories appeared between the 12-hour and 6-week timepoints. This pattern is reflective of the temporally distinct pathological changes that occur across epileptogenesis and highlights the unique nature of the molecular signaling events within the quiescent period compared with the hyperexcitability that occurs during the acute and chronic phases. The ontological categories that correspond to each cube, as well as the associated genes from each timepoint and the statistical strength of each category are available and searchable in Supplemental Fig. 4.
As part of our functional analysis, we also identified the ontologies of the genes uniquely expressed at each timepoint. Mutually-exclusive lists of genes were used to produce clusters of enriched categories for each timepoint. The top 20 clusters of upregulated categories are presented for each timepoint in order of DAVID Enrichment Score, along with the number of genes within each cluster (Fig. 3). The overlaid line graphs depict the –log(EASE), which indicates the strength of gene associations within each cluster. Of note, the acute timepoint (12 hours) was enriched for genes associated with synaptic activity (e.g., ‘Synapse,’ ‘Synaptic Transmission,’ and ‘Protein Kinase Regulator Activity’), morphological modifications (e.g., ‘Neuron Projection,’ and ‘Microtubule’) and transcriptional regulation (e.g., ‘Chromatin Modification,’ ‘Transcription Factor Binding,’ ‘mRNA Processing’, ‘Methylation’). In contrast, the 10-hour timepoint showed heightened metabolic regulation (e.g., ‘Mitochondrion,’ ‘Oxidative Phosphorylation’) and homeostatic damage response signaling (e.g., ‘Response to Oxidative Stress,’ ‘Negative Regulation of Apoptosis,’ ‘Cell Redox Homeostasis’). By six weeks, the transcriptional responses appear to have reached a new equilibrium reflective of adaptive cellular function that show fewer significant cluster differences compared to control (Supplemental Fig. 4). However there remained evidence of heightened excitability and morphogenesis reflected in the transcriptome at the chronic phase (e.g. ‘Synapse,’ ‘Regulation of Cytoskeleton Organization,’ ‘Neuron Projection Morphogenesis,’). Once again, these timepoint-specific gene lists yielded greater functional similarity between the 12-hour and 6-week timepoints than with those from the 10-day timepoint. This pattern is likely reflective of the hyperexcitability observed at both acute and chronic epileptogenic phases that is absent during the quiescent seizure-silent phase. A detailed examination of the genes associated with these enriched ontologies may yield greater insight into the signals that drive the pathological progression along each phase of epileptogenesis.
In addition to a functional examination of those transcripts unique to each phase of pathogenesis, we probed the subset of genes that were upregulated across all three timepoints for functional patterns and commonalities (Supplemental Fig. 1E). Because of the limited number of genes that were upregulated, this analysis yielded significantly fewer enriched ontological categories. Nevertheless, among the resulting clusters ‘blood vessel development’ and ‘axonogenesis’ were both highly enriched. Given the massive cellular reorganization and vascularization that occurs during epileptogenesis, it is not surprising that these clusters would be strongly represented. Here, we present the relative fold changes across time for five angiogenesis cluster-associated genes upregulated across experimental groups (Fig. 4). Of note is the particularly robust enrichment of Cxcl12 (FC = 15.77 [12 hours], 24.67 [10 days], 24.15 [6 weeks], q < 0.05), a chemokine known to play a significant role in angiogenesis and progenitor cell migration acutely after seizure activity, though its role in chronic SE has yet to be fully examined24.
Among the signaling pathways identified from lists of upregulated genes, enrichment within the MAPK cascade (57 genes out of 118 in the KEGG representation of the JNK, p38 and p42/44 ERK1/2 pathways) was of particular interest. Activation of these pathways is known to mediate many of the cellular responses to seizure activity in a time-dependant manner10,31,32. Here, we characterized changes in expression across the entirety of the JNK, p38 and classical MAPK signaling pathways at all three timepoints (Fig. 5). Accompanying previously demonstrated increases in kinase phosphorylation are robust changes in transcriptional regulation within all three arms of the MAPK cascade. Positive regulation of these pathways was prominent at all three timepoints (Fig. 5, inset). Upregulation of upstream activators of Ras suggest a facilitation of subsequent Raf, ERK and CREB phospho-activation and all four of these signaling proteins are also upregulated transcriptionally. In addition, the induction of MAPK kinase (Mkk) 6 (FC = 42.29 [12 hours], 25.79 [6 weeks], q < 0.05) and Mkk3 (FC = 67.83 [12 hours], 67.44 [6 weeks], q < 0.05) at early and late timepoints is consistent with the idea that stress-mediated cellular responses persist throughout epileptogenesis and well into the chronic phase of SE. Activation of each of these pathways may be counterbalanced by the upregulation of inhibitors such as MKPs and protein tyrosine phosphatases. Key inhibitors, such as Pp2ca (Fig. 5, circled), that did not show concordant upregulation may serve as potential therapeutic targets to counter hyper-activation of inflammatory and apoptotic signaling33.
Functional enrichment of genes involved in the MAP Kinase signaling pathways.
Pathway analysis of upregulated genes indicated an enrichment within the MAPK signaling pathway. The relevant genes from each timepoint were mapped onto an adapted version of the Kyoto Encyclopedia of Genes and Genomes (KEGG) MAPK pathway. Signaling cascades associated with classical ERK1/2, JNK and p38 signaling are indicated. A Venn diagram (inset) indicates the proportions of MAPK-associated genes that were upregulated at each timepoint.
CREB mediates activity-dependent gene expression and is known to be phosphorylated following seizure activity34. Given the increased expression of both Creb and its upstream activators across multiple timepoints, we determined the proportions of upregulated genes across epileptogenesis that are potentially under control of CREB transcriptional regulation. To accomplish this, we used gene lists generated from a genome-wide screen of CREB occupancy in hippocampal neurons35. In this approach, chromatin immunoprecipitation (ChIP) high throughput sequencing was performed after CREB pull down and the resulting list of enriched genes was intersected with our datasets of upregulated transcripts from each timepoint. Thus, we generated outputs representing the subset of genes that are both CREB-associated and upregulated at each phase of epileptogenesis (1474 genes total; Fig. 6A, Supplemental Fig. 5). Altogether, 37% of upregulated genes were associated with CREB binding at each timepoint. Among these was found an enrichment of genes associated with neuronal projection and synaptic transmission during the acute phase that gave way to oxidative stress response, myelination and homeostatic regulation in the second epileptogenic phase (Supplemental Fig. 5). The chronic phase was characterized by genes regulating cytoskeletal reorganization and synapse formation, along with increased transcriptional regulation. Hence, these CREB-associated genes likely reflect the robust reorganization that occurs during epileptogenesis and provide insights into key molecular mediators of these processes.
Temporal and cell-type profile of SE-evoked, CREB-mediated, gene expression.
(A) Data from a genome-wide analysis of CREB occupancy by ChIP-Seq was used to determine the proportion of upregulated epileptogenic genes that are CREB-associated. Venn diagram depicts subsets of genes at each timepoint. (B) Representative immunohistochemical labeling of tissue from CRE-β-Gal transgenic mice reveals robust reporter gene expression within the GCL at 4 hrs post-SE onset. Mol: molecular cell layer; GCL: granular cell layer; Hil: hilus. Bar: 50 microns. (C) Double immunofluorescence labeling for β-Gal and NeuN or parvalbumin A (Parv A) revealed that the reporter gene was inducibly expressed in both excitatory and inhibitory neurons at the 4 hour time point. Limited transgene expression was also observed in astrocytes, as assessed via GFAP double labeling. At 2 days post-SE (right panels), limited β-gal was detected in GCL neurons; rather, the reporter was detected in astrocytes and microglia, as assessed via GFAP and CD11b double-labeling. (D) Representative data from animals that were rendered epileptic (sacrificed at 6 weeks post-SE) revealed marked CRE-mediated gene expression within the GCL.
Given that our RNA seq data strongly suggest that SE drives a lasting increase in CREB-mediated gene expression, we sought to profile the activation of the CRE pathway at multiple time points following SE. To this end, we utilized a well-described transgenic mouse model in which 6 tandem CRE motifs drive the expression of a β -galactosidase reporter gene41. Consistent with our prior work, we found that SE triggers a marked upregulation of CREB-dependent transcription in the dentate GCL at 4 hrs post SE onset (Fig. 6B). Double labeling with the neuronal-specific marker NeuN and the interneuron marker parvalbumin A revealed that CRE-transcriptional activity occurred in both excitatory glutamatergic neurons and inhibitory GABAergic neurons (Fig. 6C). Low level β -galactosidase expression was also detected in astrocytes (as assessed via GFAP double labeling). Of note, at the 2 day post-SE time point, the most pronounced labeling was observed in astrocytes and microglia (as assessed via CD11b double labeling), within the molecular layer and hilar region; limited labeling was observed in neurons of the GCL. Quantitative analysis of SE-evoked CRE-reporter expression within the hippocampus can be found in Lee et al, 2007. Interestingly, in mice that developed spontaneous seizure activity (6 week post-SE), marked CRE transgene expression was detected within the GCL (Fig. 6D). Together, the data are consistent with the RNA seq data; thus, 1) SE triggered a rapid and prolonged increase in CRE mediated gene expression and 2) robust expression was detected within a multitude of cell types.
Identification of a predicted noncoding transcripts upregulated after induction of SE
An advantage of unbiased high-throughput sequencing relative to more traditional profiling approaches is its capacity to detect unannotated non-protein-coding transcripts. To this end, we identified several predicted noncoding regions that showed transcriptional upregulation after the induction of SE. A sliding window approach was used to find clusters of repeat-masked sequence reads that did not overlap with annotated protein-coding exons. Sequence tags within these regions were counted and pair-wise comparisons were conducted. Here, we highlight two examples of predicted noncoding regions that exhibited transcriptional activity (Storey Q-test) and that did not have homology with RNA or DNA repeats (Fig. 7). First, we show upregulation of a long noncoding RNA (lncRNA) whose involvement in epileptic pathophysiology has been previous established. MALAT1 regulates synaptic density in neurons and was shown to be upregulated in high-activity areas of the human epileptic cortex36. We also show its increased expression across multiple phases of epileptogenesis (Fig. 7A). In addition to Malat1, we present a predicted noncoding transcript on chromosome 9 that was previously unassociated with epileptic activity. This short sequence showed robust upregulation predominantly 6 weeks after induction of SE (Fig. 7B). The predicted noncoding transcript is contained within a small EST that has only been detected in pre-implantation embryo cDNA libraries. No exonic sequences from this EST were detected in any of our hippocampal RNA-Seq data sets (unpublished observation). Thus, we are confident that the small transcript reported in Figure 7B is not associated with the EST sequence. These transcripts are representative examples of a wide array of noncoding transcriptional activity that occurs across the phases of epileptogenesis. The increased transcription of such predicted noncoding regions supports the possibility of a regulatory role for noncoding signaling over the course of epileptogenesis37.
Upregulation of predicted noncoding RNAs after induction of SE.
Representative predicted noncoding regions showing increased transcription across epileptogenesis. The UCSC Genome Browser was used to visualize tag counts from each timepoint. (A) Increased expression of the MALAT1 long noncoding RNA. (B) An intergenic transcript showing increased expression during the chronic phase of epileptogenesis.
Discussion
Here, we used high-throughput deep sequencing to profile the hippocampal transcriptome across three phases of SE-evoked epileptogenesis. Comparative analysis of samples derived from each timepoint revealed significant differences in transcriptomic profiles, reflecting temporal alterations underlying the progression of epileptic pathology. In addition, we performed functional analysis to determine the cellular processes associated with altered mRNA expression across epileptogenesis. Several patterns emerged from an ontological analysis of these datasets. Although some functional overlap is apparent, the induced functional profile at each timepoint was largely unique. Together, these data provided insight into both the temporally divergent and sustained cellular responses to pilocarpine-induced SE. Several of the cellular and molecular processes mentioned above are under the regulatory control of the MAPK-family signaling pathways. We demonstrate that the transcriptional expression of these signaling cascades is markedly altered at multiple phases of the epileptogenic process. The p42/44 MAPK cascade plays a prominent role in progenitor cell proliferation and neuronal survival following SE38,39,40. Given this, some of the upregulation of these signaling cascades may be due to the increase in neurogenesis known to occur following SE, presenting a potential confound to our data by biasing our results toward this population of cells. Nevertheless, the density of these developing neurons is relatively low and likely only accounts for a small subset of the transcriptional increases observed. Thus, given the robust activation of p42/44 MAPK signaling across epileptogenesis, an increase in CREB-dependant gene expression is not unexpected. Indeed, CREB-mediated gene expression is associated with SE in a cell-layer and time-dependant manner41,42. Concordantly, we showed that CREB-bound genes were highly enriched across timepoints, as demonstrated by robust intersectionality with CREB ChiP-seq gene lists. This subset of CREB-associated genes yields insight into the protective mechanisms of CREB phosphorylation.
Conversely, the JNK and p38 MAPK pathways are known to contribute to the apoptotic and inflammatory responses following seizure activity43,44. Within these pathways, genes have emerged out of our non-biased interrogation that may not have been emphasized a priori as having therapeutic potential. The serine/threonine protein phosphatase PP2CA is one such example. Research surrounding the regulatory potential of PP2CA in epilepsy is limited, though initial studies suggest that PP2CA/CACNB4 signaling is essential to activity-dependent gene expression and is impaired in juvenile myoclonic epilepsy33. PP2CA is a known regulator of cellular stress response, attenuating excitotoxic JNK/p38 signaling after ischemic injury45,46. Given the absence of compensatory upregulation of PP2CA in response to JNK/p38 activation after SE, therapeutic strategies to increase PP2CA activity may prove protective against epileptic pathology as well. Nevertheless, the scale of cascade-wide transcriptional dysregulation revealed by high-throughput technology suggests that therapies focused on a single target may be insufficient for meaningful protective benefit.
Though we employed high-throughput RNA sequencing to describe the transcriptional changes underlying the three phases of epileptogenesis, others have previously made use of microarray technology to profile the epileptogenic transcriptome14,16,17. These studies all exhibited the greatest changes in gene expression at the earliest timepoints, which is to be expected given the profoundly heightened neuronal excitability that occurs during the acute phase. Here, we too found the greatest number of genes expressed at the acute timepoint (using similar significance and fold-change criterion). Though there remains some level of variability between datasets due to both the methods of initial approach and subsequent analysis used in each, there remains remarkable overlap between studies that lends confidence to our new findings and the new datasets provided here that expand upon previous work. Consistent among each of these studies is the robust expression of morphogenesis, blood vessel development and stress-response signaling pathways throughout epileptogenesis12,14,47. Further, the MAPK family signaling cascades were a prominent feature of multiple studies and PI3K, TGF-beta, IGF-1, mTOR signaling were all frequent regulatory themes12,14.
Though high-throughput screening of transcription across multiple phases of epileptogenesis is problematic in the context of human studies, microarray assays of postmortem epileptic brain tissue have proven useful in characterizing the chronically-epileptic transcriptome48. Arion et al. (2006) demonstrated that transcriptional regulation (including of the MAPK signaling cascade) correlated with intracranial EEG activity measured immediately prior to partial lobectomies performed in patients with temporal lobe epilepsy. Additionally, transcriptional expression within the CA3 subfield of the epileptic hippocampus was shown to correlate with patients' initial precipitating injury49. That the transcriptome varied substantially between patients with differing precipitating injuries emphasizes the importance of the method of SE induction used in animal models and the need to validate transcriptional changes found in rodent models against those observed in human patients50.
Though still subject to false positives, RNA-Seq has been demonstrated to be a reliable method of transcriptional analysis51,52. Nevertheless, several factors should be considered when interpreting the data presented here. As mentioned, the heightened neurogenesis following SE should be considered when interpreting the increases in gene transcription reported here. In addition, not only will cell death associated with excitotoxicity lead to some bias with regard to down regulated genes, the remaining spared cells (as well as those migrating into the region as part of the pathogenic response) will also create some level of expression bias in our reports of upregulated genes in a cell-type specific manner (relevant genes with the heightened susceptibility to these biases are noted in Supplemental Fig. 4N). Furthermore, the current study was performed on whole-hippocampus homogenates, resulting in a transcriptional readout from a heterogeneous cell populations, including pyramidal neurons, basket cells, granule cells, astrocytes, microglia and oligodendrocytes. Beyond these cellular-level confounds, there is still a need for more advanced statistical tools that have a sophistication paralleling the high-throughput capacity of RNA-seq outputs. Further, some issues with regard to sample replication and normalization remain unresolved in the field.
Up to this point, RNA-Seq had only been employed to screen for specific, strain-dependent single-nucleotide polymorphisms (SNPs) across genetic animal models of epilepsy, as well as changes in transcription and methylation at the chronic phase15,53. By using RNA sequencing to profile the entire transcriptome across multiple phases of epileptogenesis, we were able to gain perspective on both the protein-coding and noncoding transcriptional changes that occur. While many of the functional gene associations reported here build upon previous studies of molecular signaling following induction of SE, we also reveal expression patterns that suggest new regulatory pathways for which there is untapped therapeutic potential. Collectively, the data presented here emphasize the need for a coordinated investigation into the treatment of repetitive seizure activity that takes into account both gene-by-gene and high-throughput approaches.
A final word regarding current methods of reporting next-generation sequencing studies: as the sheer volume of data output increases, better graphic representation will be essential for the effective interpretation and translation of results. To date, published figures are almost exclusively comprised of bar graph, Venn diagram and heatmap depictions of data, which are limited in their utility for meaningful therapeutic discovery. Though the authors have made some attempts here to further the endeavor for novel methods of representation, there remains a need for network-level visualization graphics to a degree that has not yet been achieved. The ability to overlay datasets and interacting pathways in scalable, digital representations would greatly enhance the insights to be garnered from high-throughput investigations and aid in the discovery of therapeutic models that more closely approximate biological reality. By these means, a more complete picture of gene expression and molecular signaling networks will emerge.
Methods
Pilocarpine-induced SE
Adult (~10 weeks old) C57Bl/6 mice were intraperitoneally injected with atropine methyl nitrate (1 mg/kg), followed 30 minutes later with an injection of pilocarpine hydrochloride (325 mg/kg)54. Only those animals that survived and achieved SE were included in this study. SE was defined as an animal exhibiting multiple stage 5 seizures (tonic-clonic seizure extending to all four limbs, resulting in loss of balance and falling) and subsequent persistent and spontaneous seizure activity55. SE was induced in 20% of animals, with a mortality rate of 40%. Pilocarpine acts as an agonist for muscarinic receptors in the entorhinal cortex and changes within the hippocampus are a result of downstream excitatory transmission, rather than any direct confounding effects of pilocarpine itself. Thus, given that sub-threshold pilocarpine still drives synaptic activity at higher than normal levels, pilocarpine-injected mice that did not develop seizures were not included in our RNA-seq analysis. Instead, control animals were injected with atropine, followed by an equal volume of saline (0.9% Sodium Chloride; B. Braun Medical Inc.) 30 minutes later. No systemic, adverse side-effects of pilocarpine administration, beyond those expected by the induction of SE, were observed in treated mice. The Ohio State University Lab Animal Use Committee approved all animal procedures and all experiments were conducted in accordance with the approved guidelines.
Fluoro-Jade B
Fluoro-Jade B staining was performed as previously described in Choi et al. (2007)8.
Timm staining
Timm staining was performed as previously described in Li (2010)13.
RNA isolation and library preparation
Twelve hours, ten days, or six weeks after induction of SE, animals were sacrificed by cervical dislocation and hippocampi were removed bilaterally and subjected to total RNA extraction by TRIzol (Invitrogen #15596-018) as per manufacturer instructions. Polyadenylated RNA was purified by Dynabeads® mRNA purification kit (Life Technologies #61006). As per manufacturer protocol, 1 μg of total RNA was incubated at 50°C for 2 min and placed on ice. The sample was added to Dynabeads® with 50 μL binding buffer, mixed thoroughly at room temperature for 5 min, washed twice and then eluted into 10 μL 10 mM Tris-HCl (pH 7.5) by heating to 70°C for 2 min. Purified mRNA was treated with DNase I (Life Technologies #18068015) at 37°C for 30 min followed by 70°C for 5 min. For first strand synthesis, 10 μL of the digestion product, 1 μL oligo(dT)18 (500 ug/ml) and 1 μl 10 mM dNTP mix were mixed. Samples were heated to 65°C for 5 minutes and incubated on ice. Four microliters of 5× First-Strand Buffer, 2 μl of 0.1 M DTT and 1 μl of RNaseOUT Recombinant RNase Inhibitor (Life Technologies #10777-019) were added and incubated at 37°C for 2 min. One microliter of MMLV reverse transcriptase (Life Technologies #26025) was added to samples, which were then incubated at 37°C for 50 minutes, followed by a five-minute inactivation at 70°. cDNA was then treated with NEBNEXT mRNA second strand synthesis mix (neb #e6112) and incubated for 2.5 hours at 16°C, followed by sonication for 10 seconds. Samples were purified on a PCR column (Fermentas #K0702) and eluted in 22 μl 10 mM Tris pH 8. Blunt ends were using the DNATerminator® End Repair Kit (Lucigen #40035). Samples were purified using AMPure xp beads (Beckman #A63880) and A-tailing of 3′ ends of eluted cDNA was performed using Exo-minus Klenow DNA polymerase (Epicentre #KL0810259) at 37°C for 30 min. Uniquely tagged Illumina genomic adopters were ligated by Enzymatics T4 DNA ligase (Invitrogen #15224) at RT for 2 hrs. Size visualization was performed using a 2% agarose gel, yielding cDNA libraries ranging in size from 200–250 bp, which were amplified by PCR (5 sec at 98°C; 10 sec at 72°C; 10–20 cycles) using Phusion High-Fidelity DNA Polymerase (ThermoScientific #F530).
RNA-Seq and bioinformatic analyses
cDNA from two animals was pooled into two independent biological replicates for each timepoint (ie. two sets of two animals per experimental group: control, 12 hours, 10 days, 6 weeks). Samples were sequenced using a Genome Analyzer II (GAII) at a concentration of 10pM in each lane. Base-calling was conducted with the standard Illumina Analysis Pipeline 1.0 (Firescrest-Bustard). Eight FASTQ sequence files (sequencing reads plus quality information) were generated and mapped to the mouse genome (UCSC mm9) using the Bowtie algorithm with default settings. A C++ program was used to count the number of uniquely mapped reads within exons of Ref-Seq genes (UCSC Genome Browser mm9 annotation). The sequence data have been submitted to the NCBI Short Read Archive with accession number in progress. RefSeq gene sequence counts were median normalized and overlapping gene IDs were discarded. Data was visualized using the UCSC Genome browser (NCBI37/mm9). All statistical analyses were performed in the R programming environment. RNA-Seq tag counts in Ref-Seq gene regions were scaled to a weighted trimmed mean that excluded the top 5% of genes. Genes with a normalized reads per million of less than 10 were excluded for differential expression analysis. Quality control analyses showed that less than 1% of exonic tags were common repetitive artifacts (e.g. satellite repeats and RNA repeats, Repeat masker4). Statistical comparisons were conducted using the likelihood–ratio statistic56 and the Storey Q-test was used to correct for multiple comparisons (Bioconductor qvalue package). Differentially regulated Ref-Seq genes with a q < 0.01 and fold change > 1.5 were considered significant. Because some differential expression calls were due to single samples, we required the chi-square statistic for both paired comparisons to have an FDR-adjusted q < 0.01. To best represent the dynamic range of this time course, we selected the 300 most significant genes (ranked q-values of all pairwise comparisons), which were then plotted as a hierarchically-clustered heatmap of log2- and median-scaled tag counts (R heatmap2 package and hclust R function—complete-linkage clustering). Software and scripts used for these analyses will be provided upon request. In addition, previously-published, genome-wide screens of CREB occupancy in hippocampal neurons were intersected with our complete datasets of upregulated transcripts at each timepoint using the R statistical programming environment35. These CREB ChiP data were prepared by the Wayman lab from hippocampal neurons and ChIP-Seq libraries were generated and sequenced using standard Illumina protocols. Areas of enrichment at an FDR of 0.001 were determined using a 500 bp sliding window approach (see Lesiak et al. for details)35.
Functional analysis of gene lists
Using the web-based annotation tool DAVID v 6.7 (Database for Annotation, Visualization and Integrated Discovery) to cluster genes overexpressed in each timepoint by their common functionality, enriched ontologies were identified and statistically clustered by comparison of their constituent genes57. DAVID-defined annotation selection was used, including analysis of BioCarta Pathways, Kyoto Encyclopedia of Genes and Genomes (KEGG)-pathways and GO terms (GOTERM_XX_FAT)58,59. Clustering enrichment thresholds required DAVID EASE scores such that the modified Fischer Exact P-value was less than 0.05. Gene lists were uploaded to the DAVID web interface and the provided M. musculus background was selected. Outputs were compared with those derived from pseudo-random sets of 2000 genes generated using the atmospheric noise algorithms at random.org60.
We next determined which of the remaining categories were unique to specific timepoints and which were common to multiple epileptogenic phases. Using a union of all categories of upregulated genes (to avoid the potentially confounding effect of SE-induced cell death), enriched in the random set of genes in conjunction with those appearing at each timepoint, template arrays were created for four domains of interest: molecular functions, cellular components, biological processes and signaling pathways. Each category from the DAVID output was represented in its appropriate array by a square box. Each box was then subsequently assigned a color that corresponded to the timepoints showing enrichment in that category according to the color scheme established in Figure 1. Category boxes were ordered in accordance with the experimental timepoints used, yielding a profile for the ontology of the original gene lists. Ontological arrays were presented for dynamic interactive viewing using Prezi (Prezi.com), such that each square may be expanded to view its ontological term, as well as the names of all the genes upregulated in that category at each timepoint. The total number of upregulated genes at each timepoint is also provided.
CRE-β-galactosidase transgenic mice
CRE-β-galactosidase transgenic mice were provided by Dr. Daniel Storm (University of Washington, Seattle WA, USA). Immunohistochemical and immunofluorescent labeling was performed using the methods and reagents outlined in Lee et al (2007). Bright field images were acquired using a Leica DMIR microscope and fluorescence images were acquired using a Zeiss 510 Meta confocal microscope (Oberkochen, Germany). All photomicrographs were digitally assembled using Photoshop (Adobe Systems).
References
Gaitanis, J. N. & Drislane, F. W. Status epilepticus: a review of different syndromes, their current evaluation and treatment. Neurologist 9, 61–76, 10.1097/01.nrl.0000051445.03160.2e (2003).
Turski, L., Ikonomidou, C., Turski, W. A., Bortolotto, Z. A. & Cavalheiro, E. A. Review: cholinergic mechanisms and epileptogenesis. The seizures induced by pilocarpine: a novel experimental model of intractable epilepsy. Synapse 3, 154–171, 10.1002/syn.890030207 (1989).
Okazaki, M. M., Evenson, D. A. & Nadler, J. V. Hippocampal mossy fiber sprouting and synapse formation after status epilepticus in rats: visualization after retrograde transport of biocytin. J Comp Neurol 352, 515–534, 10.1002/cne.903520404 (1995).
Rigau, V. et al. Angiogenesis is associated with blood-brain barrier permeability in temporal lobe epilepsy. Brain 130, 1942–1956, 10.1093/brain/awm118 (2007).
Frantseva, M. V. et al. Oxidative stress is involved in seizure-induced neurodegeneration in the kindling model of epilepsy. Neuroscience 97, 431–435, 10.1016/S0306-4522(00)00041-5 (2000).
Pollard, H. et al. Kainate-induced apoptotic cell death in hippocampal neurons. Neuroscience 63, 7–18, 10.1016/0306-4522(94)90003-5 (1994).
Lee, B. et al. The CREB/CRE transcriptional pathway: protection against oxidative stress-mediated neuronal cell death. J Neurochem 108, 1251–1265, 10.1111/j.1471-4159.2008.05864.x (2009).
Choi, Y. S. et al. Status epilepticus-induced somatostatinergic hilar interneuron degeneration is regulated by striatal enriched protein tyrosine phosphatase. J Neurosci 27, 2999–3009, 10.1523/JNEUROSCI.4913-06.2007 (2007).
Fuortes, M. G., Faria, L. C. & Merlin, L. R. Impact of protein kinase C activation on epileptiform activity in the hippocampal slice. Epilepsy Res 82, 38–45, 10.1016/j.eplepsyres.2008.07.002 (2008).
Lopes, M. W. et al. Time-Dependent Modulation of Mitogen Activated Protein Kinases and AKT in Rat Hippocampus and Cortex in the Pilocarpine Model of Epilepsy. Neurochem Res 37, 1868–1878, 10.1007/s11064-012-0797-y (2012).
Xi, Z. Q. et al. Gene expression analysis on anterior temporal neocortex of patients with intractable epilepsy. Synapse 63, 1017–1028, 10.1002/syn.20681 (2009).
Laurén, H. B., Lopez-Picon, F. R., Brandt, A. M., Rios-Rojas, C. J. & Holopainen, I. E. Transcriptome analysis of the hippocampal CA1 pyramidal cell region after kainic acid-induced status epilepticus in juvenile rats. PLoS One 5, e10733, 10.1371/journal.pone.0010733 (2010).
Li, A. et al. Proteomic profiling of the epileptic dentate gyrus. Brain Pathol 20, 1077–1089, 10.1111/j.1750-3639.2010.00414.x (2010).
Okamoto, O. K. et al. Whole transcriptome analysis of the hippocampus: toward a molecular portrait of epileptogenesis. BMC Genomics 11, 230, 10.1186/1471-2164-11-230 (2010).
Kobow, K. et al. Deep sequencing reveals increased DNA methylation in chronic rat epilepsy. Acta Neuropathol 126, 741–756, 10.1007/s00401-013-1168-8 (2013).
Gorter, J. A. et al. Potential new antiepileptogenic targets indicated by microarray analysis in a rat model for temporal lobe epilepsy. J Neurosci 26, 11083–11110, 10.1523/JNEUROSCI.2766-06.2006 (2006).
Lukasiuk, K., Kontula, L. & Pitkänen, A. cDNA profiling of epileptogenesis in the rat brain. Eur J Neurosci 17, 271–279, 10.1046/j.1460-9568.2003.02461.x (2003).
Meldrum, B. Excitotoxicity and epileptic brain damage. Epilepsy Res 10, 55–61 (1991).
Kim, Y. S. & Doré, S. Catalytically inactive heme oxygenase-2 mutant is cytoprotective. Free Radic Biol Med 39, 558–564, 10.1016/j.freeradbiomed.2005.04.009 (2005).
Cuevas, P., Revilla, C., Herreras, O., Largo, C. & Giménez-Gallego, G. Neuroprotective effect of acidic fibroblast growth factor on seizure-associated brain damage. Neurol Res 16, 365–369 (1994).
Biagini, G., Avoli, M., Marcinkiewicz, J. & Marcinkiewicz, M. Brain-derived neurotrophic factor superinduction parallels anti-epileptic–neuroprotective treatment in the pilocarpine epilepsy model. J Neurochem 76, 1814–1822, 10.1046/j.1471-4159.2001.00163.x (2001).
Nicoletti, J. N. et al. Vascular endothelial growth factor is up-regulated after status epilepticus and protects against seizure-induced neuronal loss in hippocampus. Neuroscience 151, 232–241, 10.1016/j.neuroscience.2007.09.083 (2008).
Herdegen, T. et al. Expression of activating transcription factor-2, serum response factor and cAMP/Ca response element binding protein in the adult rat brain following generalized seizures, nerve fibre lesion and ultraviolet irradiation. Neuroscience 81, 199–212, 10.1016/S0306-4522(97)00170-X (1997).
Hartman, N. W. et al. CXCL12-mediated guidance of migrating embryonic stem cell-derived neural progenitors transplanted into the hippocampus. PLoS One 5, e15856, 10.1371/journal.pone.0015856 (2010).
Martikainen, M. H. & Roine, S. Rapid improvement of a complex migrainous episode with sodium valproate in a patient with CADASIL. J Headache Pain 13, 95–97, 10.1007/s10194-011-0400-y (2012).
Dobolyi, A. & Palkovits, M. Expression of latent transforming growth factor beta binding proteins in the rat brain. J Comp Neurol 507, 1393–1408, 10.1002/cne.21621 (2008).
Liu, H. Y., MacDonald, J. I., Hryciw, T., Li, C. & Meakin, S. O. Human tumorous imaginal disc 1 (TID1) associates with Trk receptor tyrosine kinases and regulates neurite outgrowth in nnr5-TrkA cells. J Biol Chem 280, 19461–19471, 10.1074/jbc.M500313200 (2005).
Vilimanovich, U. & Bumbasirevic, V. TRAIL induces proliferation of human glioma cells by c-FLIPL-mediated activation of ERK1/2. Cell Mol Life Sci 65, 814–826, 10.1007/s00018-008-7513-8 (2008).
Bae, S. et al. Akt is negatively regulated by the MULAN E3 ligase. Cell Res 22, 873–885, 10.1038/cr.2012.38 (2012).
Hamard, P. J. et al. Sumoylation delays the ATF7 transcription factor subcellular localization and inhibits its transcriptional activity. Nucleic Acids Res 35, 1134–1144, 10.1093/nar/gkl1168 (2007).
Garrido, Y. C., Sanabria, E. R., Funke, M. G., Cavalheiro, E. A. & Naffah-Mazzacoratti, M. G. Mitogen-activated protein kinase is increased in the limbic structures of the rat brain during the early stages of status epilepticus. Brain Res Bull 47, 223–229, 10.1016/S0361-9230(98)00075-6 (1998).
Jeon, S. H., Kim, Y. S., Bae, C. D. & Park, J. B. Activation of JNK and p38 in rat hippocampus after kainic acid induced seizure. Exp Mol Med 32, 227–230, 10.1038/emm.2000.37 (2000).
Tadmouri, A. et al. Cacnb4 directly couples electrical activity to gene expression, a process defective in juvenile epilepsy. EMBO J 31, 3730–3744, 10.1038/emboj.2012.226 (2012).
Tanis, K. Q., Duman, R. S. & Newton, S. S. CREB binding and activity in brain: regional specificity and induction by electroconvulsive seizure. Biol Psychiatry 63, 710–720, 10.1016/j.biopsych.2007.08.003 (2008).
Lesiak, A. et al. A Genome-Wide Screen of CREB Occupancy Identifies the RhoA Inhibitors Par6C and Rnd3 as Regulators of BDNF-Induced Synaptogenesis. PLoS One 8, e64658, 10.1371/journal.pone.0064658 (2013).
Lipovich, L. et al. Activity-dependent human brain coding/noncoding gene regulatory networks. Genetics 192, 1133–1148, 10.1534/genetics.112.145128 (2012).
Qureshi, I. A. & Mehler, M. F. Epigenetic mechanisms underlying human epileptic disorders and the process of epileptogenesis. Neurobiol Dis 39, 53–60, 10.1016/j.nbd.2010.02.005 (2010).
Choi, Y. S., Cho, H. Y., Hoyt, K. R., Naegele, J. R. & Obrietan, K. IGF-1 receptor-mediated ERK/MAPK signaling couples status epilepticus to progenitor cell proliferation in the subgranular layer of the dentate gyrus. Glia 56, 791–800, 10.1002/glia.20653 (2008).
Choi, Y. S. et al. Mitogen- and stress-activated kinases regulate progenitor cell proliferation and neuron development in the adult dentate gyrus. J Neurochem 123, 676–688, 10.1111/jnc.12035 (2012).
de Araújo Herculano, B., Vandresen-Filho, S., Martins, W. C., Boeck, C. R. & Tasca, C. I. NMDA preconditioning protects against quinolinic acid-induced seizures via PKA, PI3K and MAPK/ERK signaling pathways. Behav Brain Res 219, 92–97, 10.1016/j.bbr.2010.12.025 (2011).
Lee, B., Dziema, H., Lee, K. H., Choi, Y. S. & Obrietan, K. CRE-mediated transcription and COX-2 expression in the pilocarpine model of status epilepticus. Neurobiol Dis 25, 80–91, 10.1016/j.nbd.2006.08.015 (2007).
Beaumont, T. L., Yao, B., Shah, A., Kapatos, G. & Loeb, J. A. Layer-specific CREB target gene induction in human neocortical epilepsy. J Neurosci 32, 14389–14401, 10.1523/JNEUROSCI.3408-12.2012 (2012).
Yang, D. D. et al. Absence of excitotoxicity-induced apoptosis in the hippocampus of mice lacking the Jnk3 gene. Nature 389, 865–870, 10.1038/39899 (1997).
Kim, S. W., Yu, Y. M., Piao, C. S., Kim, J. B. & Lee, J. K. Inhibition of delayed induction of p38 mitogen-activated protein kinase attenuates kainic acid-induced neuronal loss in the hippocampus. Brain Res 1007, 188–191, 10.1016/j.brainres.2004.02.009 (2004).
Cao, Q. et al. Negative feedback regulation of Raf/MEK/ERK cascade after sublethal cerebral ischemia in the rat hippocampus. Neurochem Res 36, 153–162, 10.1007/s11064-010-0285-1 (2011).
Budziszewska, B. et al. The decrease in JNK- and p38-MAP kinase activity is accompanied by the enhancement of PP2A phosphate level in the brain of prenatally stressed rats. J Physiol Pharmacol 61, 207–215 (2010).
Hunsberger, J. G., Bennett, A. H., Selvanayagam, E., Duman, R. S. & Newton, S. S. Gene profiling the response to kainic acid induced seizures. Brain Res Mol Brain Res 141, 95–112, 10.1016/j.molbrainres.2005.08.005 (2005).
Venugopal, A. K. et al. Transcriptomic Profiling of Medial Temporal Lobe Epilepsy. J Proteomics Bioinform 5, 10.4172/jpb.1000210 (2012).
Bando, S. Y. et al. Hippocampal CA3 transcriptome signature correlates with initial precipitating injury in refractory mesial temporal lobe epilepsy. PLoS One 6, e26268, 10.1371/journal.pone.0026268 (2011).
Becker, A. J. et al. Correlated stage- and subfield-associated hippocampal gene expression patterns in experimental and human temporal lobe epilepsy. Eur J Neurosci 18, 2792–2802, 10.1111/j.1460-9568.2003.02993.x (2003).
van Delft, J. et al. RNA-seq provides new insights in the transcriptome responses induced by the carcinogen benzo[a]pyrene. Toxicol Sci 130(2), 427–39. 10.1093/toxsci/kfs250 (2012).
Richard, H. et al. Prediction of alternative isoforms from exon expression levels in RNA-Seq experiments. Nucleic Acids Res 38, e112, 10.1093/nar/gkq041 (2010).
Hawkins, N. A. & Kearney, J. A. Confirmation of an epilepsy modifier locus on mouse chromosome 11 and candidate gene analysis by RNA-Seq. Genes Brain Behav 11, 452–460, 10.1111/j.1601-183X.2012.00790.x (2012).
Curia, G., Longo, D., Biagini, G., Jones, R. S. & Avoli, M. The pilocarpine model of temporal lobe epilepsy. J Neurosci Methods 172, 143–157, 10.1016/j.jneumeth.2008.04.019 (2008).
Racine, R. J. Modification of seizure activity by electrical stimulation. II. Motor seizure. Electroencephalogr Clin Neurophysiol 32, 281–294 (1972).
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res 18, 1509–1517, 10.1101/gr.079558.108 (2008).
Huang, d. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57, 10.1038/nprot.2008.211 (2009).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 27, 29–34, 10.1093/nar/28.1.27 (1999).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29, 10.1038/75556 (2000).
Horvath, J. W., Barnett, G. E., Jimenez, R. E., Young, D. C. & Povoski, S. P. Comparison of intraoperative frozen section analysis for sentinel lymph node biopsy during breast cancer surgery for invasive lobular carcinoma and invasive ductal carcinoma. World J Surg Oncol 7, 34, 10.1186/1477-7819-7-34 (2009).
Acknowledgements
We thank Heather Dziema, Andrea M. Hesse and Yen Lee Loh for technical support. Funding was provided by the National Institutes of Health, grant numbers: F31-MH096460-01, NS066345 and NS067409 and by an Epilepsy Foundation of America Post-doctoral Fellowship to K. Sakamoto.
Author information
Authors and Affiliations
Contributions
K.O. and S.I. supervised the project. K.O., S.I. and K.S., formulated the hypothesis and designed invivo experiments. K.F.H., C.P. and S.I. processed and analyzed the high throughput sequencing data and prepared the figures. K.S. performed animal induction of SE, isolation of RNA and RNA-sequencing of samples. K.O. and K.F.H. wrote the manuscript. All authors reviewed and edited the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemental Table 1 and Supplemental Figures 2 & 3
Supplementary Information
Supplemental Figure 1
Supplementary Information
Supplemental Figure 4
Supplementary Information
Supplemental Figure 5
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Hansen, K., Sakamoto, K., Pelz, C. et al. Profiling status epilepticus-induced changes in hippocampal RNA expression using high-throughput RNA sequencing. Sci Rep 4, 6930 (2014). https://doi.org/10.1038/srep06930
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06930
This article is cited by
-
Similar brain proteomic signatures in Alzheimer’s disease and epilepsy
Acta Neuropathologica (2024)
-
Highly dynamic inflammatory and excitability transcriptional profiles in hippocampal CA1 following status epilepticus
Scientific Reports (2023)
-
The Promising Epigenetic Regulators for Refractory Epilepsy: An Adventurous Road Ahead
NeuroMolecular Medicine (2023)
-
Long-term in vivo application of a potassium channel-based optogenetic silencer in the healthy and epileptic mouse hippocampus
BMC Biology (2022)
-
Abnormal expression profile of plasma-derived exosomal microRNAs in patients with treatment-resistant depression
Human Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.