Abstract
The emergence and rapid spread of New Delhi Metallo-beta-lactamase-1 (NDM-1)-producing Klebsiella pneumoniae strains has caused a great concern worldwide. To better understand the mechanisms underlying environmental adaptation of those highly drug-resistant K. pneumoniae strains, we took advantage of the China's Shenzhou 10 spacecraft mission to conduct comparative genomic and transcriptomic analysis of a NDM-1 K. pneumoniae strain (ATCC BAA-2146) being cultivated under different conditions. The samples were recovered from semisolid medium placed on the ground (D strain), in simulated space condition (M strain), or in Shenzhou 10 spacecraft (T strain) for analysis. Our data revealed multiple variations underlying pathogen adaptation into different environments in terms of changes in morphology, H2O2 tolerance and biofilm formation ability, genomic stability and regulation of metabolic pathways. Additionally, we found a few non-coding RNAs to be differentially regulated. The results are helpful for better understanding the adaptive mechanisms of drug-resistant bacterial pathogens.
Similar content being viewed by others
Introduction
Klebsiella pneumoniae is a highly important bacterial pathogen that causes a wide range of community- and hospital-acquired infections, such as pneumonia, urinary tract infections, intra-abdominal infections and respiratory tract infections1,2. The emergence and rapid spread of carbapenem-resistant K. pneumoniae isolates such as the New Delhi Metallo-beta-lactamase-1 (NDM-1)-producing strains has caused great concern worldwide3,4,5. The NDM-1, an Ambler class B metallo-β-lactamase (MBL), is capable of hydrolyzing all β-lactams (including carbapenems) except monobactams. It was first identified in K. pneumoniae and Escherichia coli in India in 2008. To date, NDM-1 producing pathogens have been reported in more than 40 countries6,7,8. The successful persistence and transmission of this “superbug” remain largely unknown, but could be partly due to the ability of this pathogen to sense and react to environmental and host stress signals, which allow it to persist and disseminate in harsh conditions such as in various medical settings and inside the human host. Thus, understanding the ability of those pathogens to adapt to various stressors they encountered during environmental persistence could facilitate the comprehension of their pathobiology. In addition, identification of specific genomic variations and transcriptome patterns critical for drug resistance and environmental adaptation of the drug-resistant pathogenic bacteria is important for the development of more effective pathogen control strategies.
During spaceflight, a variety of physiological stressors associated with the space environment and spacecraft conditions could potentially contribute to detrimental alterations in the human immune system. At the same time, the bacteria introduced into the extraterrestrial environments by the space crew members inhabiting the stressful airspace environments might develop resistance traits which could be important to spaceflight missions and the general public medicine. Indeed, the recurrent isolation of various extremotolerant bacteria from spacecrafts emphasizes the possibility of those microorganisms gaining access to spacecrafts and being transferred to extraterrestrial environments9,10,11. Furthermore, it has been demonstrated that following spaceflight the gene expression pattern of Salmonella enterica serotype Typhimurium changed and the virulence of those bacteria increased12. However, the microbial mechanisms of resistance in those pathogenic bacteria during spaceflight remained largely unknown. The extreme conditions encountered by the microorganisms in air space could facilitate the occurrence of the environmental adaptation-associated genomic variations and transcriptomic changes in them and that information could be useful for taking pre-emptive measures to manage medically important drug-resistant pathogens on earth. The Shenzhou-10 spacecraft, which was launched in June 11, 2013 and returned to earth after completion of its 15-day spaceflight, is China's fifth manned space mission and its longest so far, thus it provided us a unique opportunity to explore the underlying mechanisms important for bacterial adaptation to extreme environments.
In this study, we conducted comparative whole genome sequencing and RNA sequencing (RNA-Seq) analysis of a NDM-1- producing K. pneumoniae strain (ATCC BAA- 2146). The K. pneumoniae strains were cultivated in semisolid medium and the culture placed in the following conditions: on the ground (D strain), in simulated space condition (M strain) and in the Shenzhou 10 spacecraft (T strain). Comparative genomic analysis revealed 3 and 2 strain-specific mutations in M and T strains, respectively. The comparative transcriptomic analysis revealed different transcriptome patterns among the strains, especially with respect to the regulation of multiple metabolic pathways. Additionally, we found a few non-coding RNAs (ncRNAs) to be differentially regulated. Phenotype comparisons showed that the T strain exhibited elongated forms, reduced hydrogen peroxide (H2O2) tolerance and increased biofilm formation ability. Our results provide new insights into the environmental adaptability of drug-resistant K. pneumoniae. These findings could contribute to understanding the persistence and widespread of those highly drug-resistant pathogenic bacteria.
Results
Phenotypic characteristics of K. pneumoniae strains
The K. pneumoniae ATCC BAA-2146 was an ESBL-positive and NDM-1-producing strain. According to multilocus sequence typing (MLST), this strain belonged to sequence type 11 (ST11). Drug susceptibility testing results showed that the reference strain ATCC BAA-2146 was resistant to all antibiotics tested and the drug resistance profiles of the recovered D, M and T strains remained unchanged (Supplementary Table S1). No notable differences in colony morphology of the strains were observed under light microscopy. Field emission scanning electron microscopy (SEM) was further used to monitor single cell morphology of the strains and the results showed that the bacterial cell walls of all strains were intact. But interestingly, some of the cells of the T strain turned into elongated forms and adhered to each other under SEM (Fig. 1). In the H2O2 sensitivity assay, the 1 mM and 5 mM H2O2 exposure resulted in slightly reduced survival for the T strain in comparison with the BAA-2146 control strain (P < 0.001), whereas, the D and M strains showed no obvious differences in sensitivity to H2O2 when compared to the BAA-2146 strain (Fig. 2a). The ability of K. pneumoniae to form biofilms is thought to be an important phenotype with respect to adaptability and virulence traits such as host colonization, antibiotic resistance and environmental persistence, etc.13,14. We therefore performed biofilm assays to determine the biofilm formation abilities of the strains and the results showed that the T strain showed enhanced biofilm-forming capacity compared to the BAA-2146 strain (P < 0.001) while the D strain and the M strain showed similar biofilm-forming capacity with the reference strain BAA-2146 (Fig. 2b). The fitness of the K. pneumoniae strains was further assessed by determining the growth rates of the strains in a nutrient LB medium as well as in chemically defined media (CDM), as this parameter also could contribute to environmental adaptability. All strains showed similar growth curves in LB medium (Fig. 3a) and in CDM (Fig. 3b). Furthermore, the growth curves were also similar when the bacteria were cultivated in LB medium at different temperatures including 30°C, 37°C and 40°C (Fig. 3c, 3a and 3d). To better understand the phenotypic changes in the strains, we also performed carbon source utilization assays and chemical sensitivity assays using the 96-well Biolog GEN III MicroPlate. Among the 71 carbon source utilization assays, the α-D-Lactose utilization ability was found to be defective in all three strains (including the D, M and T strains) as compared to the reference strain BAA-2146. Interestingly, the M strain was shown to gain the ability to use D-Mannose. Among the 23 chemical sensitivity assays, no obvious changes were observed for all three strains compared to the reference strain (Supplementary Table S2).
Oxidative stress tolerance and biofilm formation ability of K. pneumoniae strains.
(a) Survival of K. pneumoniae strains in oxidative stressed environment. The ATCC BAA-2146, D, M and T strains were incubated with different concentrations of H2O2 in sodium phosphate buffer and then tested for their survival on LB plates. Growth is expressed as relative survival after 2 h incubation with H2O2. (b) Biofilm assays of K. pneumoniae strains. OD595 readings were measured for each strain to determine the amount of biofilm formed. Data represent one of three independent experiments.
Growth curves of the K. pneumoniae strains in LB medium (a, c and d) or CDM (b).
Overnight-grown bacteria were diluted 100 folds in fresh LB medium or CDM and were cultivated at designated temperature with shaking (180 rpm). OD600 readings were measured at designated time points for each strain. The data represent one of two independent experiments performed in triplicate, with s.d. indicated by error bars.
Whole genome sequencing statistics
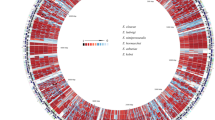
The basic whole genome sequencing statistics were shown in Table 1. The sequencing depth ranged between 127× and 128× and read mapping results reported nearly complete genome coverage (~99.99%) for all three strains. Fig. 4 showed the circular representation of the genome features including the COG annotated coding sequences, KEGG enzymes, RNA genes, GC content, GC skew, strain-specific SNPs, etc. As shown in Supplementary Fig. S1, all three samples had similar mapping coverage across the reference genome and the coverage was higher for the regions closer to the origin of replication, probably because of a higher speed of the DNA replication than that of cellular doubling during exponential growth.
Circular representation of the genome features.
Genome sequences (ring 1), COG Annotated coding sequences (rings 2 + 3), KEGG enzyme (ring 4), RNA genes (ring 5: red, rRNA; blue, tRNA), GC content (rings 6), GC skew (ring 7), strain-specific SNPs (ring 8) are shown. All strain-specific SNPs from D (yellow), M (red) and T (blue) are shown on ring 8. Very short features were enlarged to enhance visibility. Clustered genes and SNPs, such as several rRNA genes, may appear as one line due to space limitations. The image was created by using the software Circos.
Identification of the mutations in D, M and T strains compared to reference strain ATCC BAA-2146
A total of 34 mutations (excluding the synonymous mutations) were identified in all strains (Supplementary Tables S3), among which, 25 mutations were shared by all three strains (Supplementary Tables S4). No strain-specific mutation was identified in the D strain, while 3 and 2 strain-specific mutations were identified in the T and M strains, respectively. In addition, a few mutations were shared by T and M, T and D, or M and D strains (Table 2). PCR and sequencing analysis confirmed that these mutations were not sequencing or assembly errors.
Drug resistance determinants of K. pneumoniae strains
Electronic PCR was conducted for ATCC BAA-2146, D, M and T strains to analyze drug resistance determinants conferring resistance to carbapenems, folate pathway inhibitors, fluoroquinolones, aminoglycosides, etc. The drug resistance-associated genes and gene mutations detected in the D, M and T strains were identical to those of the reference strain ATCC BAA-2146 (Supplementary Table S5).
RNA-Seq mapping statistics
Approximately 96% of the sequencing reads could be mapped to ATCC BAA-2146 (CP006659) reference genome (Table 3). The uniquely mapped reads for D, M and T were 96.0%, 95.9% and 96.3% respectively. The number of reads mapped each gene ranged from 1 to 449,669 with a median of 206, 209 and 214 for D, M and T, respectively. In addition, the RPKM for D, M and T was 48.8, 49.5 and 52.3 for D, M and T respectively. The distribution of the number of mapped reads and RPKM values across all three samples was displayed in Supplementary Fig. S2. The expression of total genes and the differentially expressed genes identified among D, M and T strains (fold changes >2) were shown in Fig. 5.
Expression of total genes (a) and the differentially expressed genes (b) identified among D, M and T strains (Fold changes >2).
The heatmap was generated from hierarchical cluster analysis of genes. The RPKM values were mean-centered and normalized across samples, with each row representing a different gene.
Comparative transcriptomic analysis
By using the KEGG orthology based annotation system to identify metabolic pathways, we identified extensive changes in the transcriptomes of the M and T strains in comparison to the D strain. Expression of the differentially expressed genes identified among D, M and T strains (including: T strain vs. D strain, M strain vs. D strain and T strain vs. M strain) were shown in Supplementary Tables S6–S8. Compared to the D strain transcriptome, the T strain transcriptome was characterized by regulation of a number of genes involved in fatty acid degradation (P = 0.0464189), microbial metabolism in diverse environments (P = 0.0464189) and phenylalanine metabolism (P = 0.0464189). Among those three categories, the majority of the genes involved in fatty acid degradation and phenylalanine metabolism were down-regulated, while the genes involved in microbial metabolism in diverse environments include many up-regulated and many down-regulated genes (Supplementary Table S6). Compared to the D strain transcriptome, the M strain transcriptome showed that many genes associated with tyrosine metabolism (P<0.001), degradation of aromatic compounds (P = 0.003804639) and microbial metabolism in diverse environments (P = 0.005292205) were regulated (Supplementary Table S7) and the majority of those genes were up-regulated. Genes altered in their expression common to both M strain and T strain transcriptomes included a variety of genes involved in microbial metabolism in diverse environments. In comparison to the M strain, the T strain transcriptome was further characterized by regulation of a number of genes involved in citrate cycle (P = 0.0179529) and phenylalanine metabolism (P = 0.0404427). In addition, many more genes involved in microbial metabolism in diverse environments (P = 0.0404427) were regulated in the T strain than in the M strain. Additionally, we found a few ncRNAs to be differentially regulated in the T and M strains as compared to the D strain. Some of those ncRNAs (including GlmZ RNA activator of glmS mRNA and RtT RNA) were similarly regulated in both M and T strains. But interestingly, sroC RNA was down-regulated in the T strain but up-regulated in the M strain (Table 4). The number of mapped reads and estimation of gene expression (RPKM) values between the D, M and T strains were displayed in a boxplot as shown in Supplementary Fig. S2. Ten genes were selected for quantitative RT-PCR analysis in parallel with the RNA-Seq analysis and the results validated that gene expression fold changes from RNA-Seq analysis for specific genes were well correlated with transcript levels measured using real-time quantitative RT-PCR (qRT-PCR) analysis, supporting that RNA-Seq provides reliable quantitative estimates of transcript levels (Supplementary Tables S9 and S10). The Orthologus Groups (COG) enrichment analysis was also determined to show the distribution of differentially expressed genes (DEGs) in COG functional categories (Fig. 6). A large number of the genes that were differentially expressed in the T strain vs. D strain (T/D) play putative roles in the following two COG categories: amino acid transport and metabolism (P = 0.01814) and carbohydrate transport and metabolism (P = 0.04659). In addition, many genes that were differentially expressed in the M strain vs. D strain (M/D) belonged to the following three COG categories: Cell motility, Secondary metabolites biosynthesis (P = 0.004304), transport and catabolism (P = 0.007086) and Inorganic ion transport and metabolism (P = 0.0346).
Discussion
The explanation for the epidemiological success of highly drug-resistant bacterial pathogens, such as the NDM-1-producing K. pneumoniae strains, is complex. In general, the antibiotic resistance phenotype of bacteria is attributed to their capacity to acquire and express genes associated with a wide range of antibiotic resistance functions, as well as their intrinsic properties such as porins and gene mutations, etc. In this study, the reference strain ATCC BAA-2146 was shown to possess a large number of previously reported genes associated with resistance to β-lactams, fluoroquinolones, aminoglycosides and folate pathway inhibitors15,16,17,18,19. Among the more recently reported carbapenemases genes (such as blaIMP, blaVIM, blaNDM-1, blaKPC and blaOXA-48, etc.), only NDM-1 was detected in ATCC BAA-2146. We also identified a gyrA mutation T247A (Ser83Ile) which was shown to be associated with fluoroquinolone resistance16. In this study, the D, M and T strains all remained their drug resistance phenotype and maintained all those drug resistance-associated genes detected in the reference strain ATCC BAA-2146, suggesting that this highly drug-resistant K. pneumoniae strain did not lose resistance to those drugs while grown in extreme conditions. Based on MLST genotyping analysis, ATCC BAA-2146 belonged to ST11, which was reported to be the most dominant clone among the carbapenem-resistant K. pneumoniae isolates in China20. Thus this drug-resistant K. pneumoniae strain could be intrinsically highly transmissible.
Morphological analysis revealed that some cells of the T strain were elongated and adhered to each other under SEM, suggesting that certain genes involved in maintaining bacterial morphology and adherence ability could be affected or regulated. Normally, the function and morphology of the bacteria are maintained by well-coordinated equilibrium of their fusion and fission activities. Therefore, the explanations for the mechanism underlying the formation of elongated bacteria could be an enhanced fusion process, a blocked fission process, or a combined action of the two. Since we did not identify mutations in genes directly involved in bacterial fission or fusion function in the T strain, we thus hypothesize that certain mutations which occurred in the intergenic regions or ncRNAs could play regulatory roles on genes associated with these morphological changes in the T strain. We also predict that the increased bacterial fusion and adherence capabilities might be beneficial for the pathogen to form biofilm under stress conditions in order to promote bacterial survival. The elongated forms of the bacteria have previously been observed in bacteria being treated by certain antibiotics. For example, in one study, the scanning electron microscopy analysis demonstrated that Pseudomonas aeruginosa became elongated after exposure to ciprofloxacin21. Another study also showed that most of the P. aeruginosa cells turned into elongated forms and adhere to each other when exposed to nalidixic acid22. Thus, this morphological change could be a typical response of pathogenic bacteria towards various stressors.
Bacterial cells are constantly challenged by various environmental stressors from their natural habitats. Similar to many pathogens, K. pneumoniae faces several challenges during infection and colonization of the human body. Among these, the ability of the pathogen to tolerate oxidative stress is critical for their survival since the transition from aerobic to microaerophilic conditions or the transition from a microaerophilic to oxidative stress environment is frequently encountered by the pathogen during infection. Oxidative stress on aerobic bacteria is mainly mediated by partially reduced oxygen species, or reactive oxygen species, most notably superoxide and H2O2, which are by-products of aerobic metabolism. These reactive oxygen species can cause damage to DNA, proteins and membranes. As a result, all aerobic bacteria possess various mechanisms to scavenge superoxide and H2O2, thus protecting the cells from being damaged by these reactive oxygen species. Thus, the response of drug-resistant bacteria under oxidative stress conditions is of particular interest. Our study revealed that the T strain exhibited a slightly reduced tolerance toward H2O2, this could be partly explained by the decreased expression of the universal stress response gene as well as a few ncRNAs such as sroC RNA, 6S/SsrS RNA and the bacterial small signal recognition particle RNA in this strain, since those genes and ncRNAs were shown to be involved in stress tolerance or bacterial fitness23,24,25,26,27. Interestingly, it was demonstrated that S. Typhimurium inside macrophages led to repression of the expression of the sroC and 6S/SsrS RNA genes, suggesting that those ncRNAs may also direct pathogen adaptation to a non-proliferative state inside the host cell27. Taken together, we hypothesize that those ncRNAs could also play critical roles during environmental adaptation of K. pneumoniae.
The bacterial biofilm consists of an aggregate of cells contained within a matrix of surface polysaccharides, proteins and DNA. The ability to produce a biofilm results in enhanced resistance to host defense factors, antimicrobials, as well as various environmental stressors, thus it has been increasingly recognized as an important virulence property. An observation by Wu et al. showed that certain K. pneumoniae strains such as hvKP strains produced more biofilm than cKP strains, suggesting that biofilm formation may be a contributing factor to the increased virulence of the hvKP strains28. In this study, though the T strain showed slightly decreased tolerance towards H2O2, it exhibited increased biofilm production. The similar growth rates of the reference strain ATCC BAA-2146 as well as the D, M and T strains in LB medium and CDM ruled out the possibility that the increased biofilm production in the T strain was due to a change in growth. The mechanism underlying the increase biofilm formation in the T strain remains currently unclear, but could be associated with increased expression of certain regulatory genes involved in surface polysaccharide formation as suggested by the increased adherence between bacterial cells under SEM. We also noticed that in the T strain a large number of up-regulated genes were involved in the citrate cycle, which is a central metabolic pathway that facilitates the adaption of bacteria to environmental stress. This finding is also consistent with a previous study which showed an increased abundance of genes involved in the citrate cycle in E. coli when grown as a biofilm29. The results from that study and ours suggest that pathogens grown as a biofilm could still be energetically viable by using amino acids as an indirect carbon source through citrate cycle.
Using the 96-well Biolog GEN III MicroPlate for further phenotypic characterization of the strains, we observed that the α-D-Lactose utilization ability was defective in all three strains as compared to the reference strain BAA-2146 and the M strain gained the ability to use D-Mannose. Regulation of metabolic pathways has been demonstrated to be an important stress response of pathogenic bacteria and a variety of transcriptional regulators such as LysR, RutR and Atf1 have been shown to play central roles during this process30,31,32,33,34. For example, LysR was shown to be an important regulatory component of P. aeruginosa adaptation during oxidative stress35. RutR was demonstrated to be involved in glutamate-dependent acid resistance of E. coli for survival under acidic conditions34. In addition, many regulatory networks of the pathogen contain ncRNAs, which could also regulate the expression of key protein-coding genes during the adaptive response to environmental stimuli36,37. We identified a non-synonymous mutation in Kpn2146_2018 (P184S), which belongs to the transcriptional regulator LysR family. In addition, a couple of mutations were also identified in the intergenic regions near Kpn2146_2018 and Kpn2146_5387, the latter of which is also a transcriptional regulator, RutR. We also identified two M strain-specific mutations in the intergenic regions between Kpn2146_3273 (RtT RNA) and Kpn2146_3274 (tRNA-Tyr) (T-39A and C-37T). Those mutations within or nearby those transcriptional regulators and non-coding RNA regions could be associated with the regulation of metabolic pathways through up-regulating or down-regulating their targets in the K. pneumoniae strains during pathogen environmental adaptation.
The study of genomic variations as a function of pathogen adaptation to a specific niche could reveal important insights into how they sense and respond to varied habitats. Our comparative genomic analysis revealed only a few mutations specific to each strain, but a large number of mutations were shared by the D, M and T strains, suggesting the all three strains underwent adaptation genomic variations while being cultivated in different environments, though each strain were exposed to different combination of stressors and the strength of each stressors they encounter might be different. We also noticed that a large proportion of the mutations occurred in the intergenic regions or ncRNAs, suggesting that those mutations could play important regulatory roles on their down-stream genes or more distantly located genes, which could be involved in stress response and environmental adaptation. There were also some mutations occurring in hypothetical proteins such as Kpn2146-1328, Kpn2146-2411 and Kpn2146-2412, the roles of which in pathogen environmental adaption await further studies.
Comparative transcriptomic analysis revealed that the gene expression patterns of the three strains were significantly different from each other. The transcriptomes of both the M and T strains indicated significant changes in microbial metabolism compared to the D strain. Many more genes were regulated in the T strain compared to the M strain, suggesting that the T strain experienced more stress during spaceflight than the M strain which was cultivated in a simulated space condition with 10−3 g microgravity. Furthermore, a large proportion of the differentially expressed genes in the T strain, as compared to both D and M strains, were involved in a variety of different metabolic pathways including the following COG categories: amino acid transport and metabolism and carbohydrate transport and metabolism. These results suggest that in response to environmental stress, pathogenic bacteria including highly drug-resistant ones exhibit a great flexibility and adaptability to survive successfully through regulation of multiple physiological functions and cellular pathways, among which the regulation of metabolism pathways seems to play a very important role. Using RNA-Seq, we also identified a few previously reported or putative ncRNAs, which could play critical roles during the environmental adaptation of K. pneumoniae. As the RNA isolation procedure used in this study selected against small RNA molecules, it is likely that additional small ncRNAs not detected here could also be transcribed during stress responses of the K. penumoniae. Nevertheless, the results from this study provide powerful evidence showing that RNA-Seq allow quantitative characterization of bacterial transcriptomes and provide a useful tool for exploring transcriptional regulatory networks in bacteria.
In summary, our comparative genomic and transcriptomic analysis of a NDM-1 K. pneumoniae strain being cultivated in different conditions support the possible resistance or adaptation of the highly drug-resistant pathogenic bacteria towards the extreme environments such as the conditions in spacecrafts, which could have important impacts on the microbial ecology of the extraterrestrial space. The present study could also serve as a basis for future studies examining the complex network systems that regulate bacterial adaptation to extreme environments. Furthermore, the insights we gained from the observations of persistence and pathogenesis-related genomic variations and transcriptomic changes of the NDM-1 K. pneumoniae strain in response to spaceflight could also facilitate the development of novel or alternative therapeutic methods needed to treat the recalcitrant infections caused by those highly drug-resistant K. pneumoniae strains. Finally, the data obtained from this study could also be important for future infectious disease risk assessment and prevention during spaceflight missions and in general public health as well.
Methods
Bacterial strains and culture conditions
The reference K. pneumoniae strain KP ATCC BAA- 2146 was used in the study. K. pneumoniae was cultivated in semisolid medium (with 0.5% agar) for 15 days at 21°C and the culture were placed in the following conditions: on the ground (D strain), in simulated space condition with rotation at 30 rpm and 10−3 g microgravity (M strain) and in the Shenzhou 10 spacecraft (T strain). For phenotypic analysis, the K. pneumoniae strains were routinely grown in Luria-Bertani (LB) broth. Agar was added to a final concentration of 1.5% when necessary. For growth curve measurement, the strains were grown at designated temperature (30°C, 37°C or 40°C) with shaking (180 rpm). The CDM used for growth curve measurement was described previously38.
Drug susceptibility testing
Drug susceptibility testing (DST) for the K. pneumoniae strains was performed using the bioMe'rieux VITEK-2 AST-GN13 system following manufacturer's instructions as described previously16. The following 18 drugs were tested: ampicillin (AMP), piperacillin/tazobactam (TZP), ampicillin/sulbactam (SAM), cefazolin (CFZ), ceftriaxone (CRO), ceftazidime (CAZ), cefepime (FEP), cefotetan (CTT), ertapenem (ETP), imipenem (IMP), aztreonam (ATM), ciprofloxacin (CIP), levofloxacin (LVX), gentamicin (GM), tobramycin (TOB), amikacin (AMK), trimethoprim-sulfamethoxazole (SXT), furadantin (FD). The ESBLs were detected by the bioMe'rieux VITEK-2 AST-GN13 test. E. coli strains ATCC 25922 and ATCC 35218, K. pneumoniae strain ATCC 700603 and P. aeruginosa strain ATCC 27853 were used as quality control strains for the DST.
H2O2 sensitivity assays
To measure the susceptibility of K. pneumoniae to oxidative stress, H2O2 sensitivity assays were conducted according to the method reported by Cumley et al. with minor modifications39. Briefly, overnight-grown bacteria were diluted 100 folds in fresh LB medium and grown to early stationary phase at 37°C with vigorous shaking. Bacteria were resuspended in 0.1 M sodium phosphate buffer (pH 7.4) with increasing concentrations of H2O2 (0 mM, 1 mM, 5 mM) without agitation. After 2 h incubation at 37°C, cultures were diluted and plated onto LB plates. All experiments were repeated at least three times.
Biofilm formation assay
Bacteria were diluted 100 folds in LB broth and was then inoculated into each well of a 96-well polystyrene flat-bottom microtiter plates and statically incubated at 37°C for 24 h. Then the wells were washed with phosphate-buffered saline (PBS) to remove unattached cells. Crystal violet (0.1% w/v; Sigma) was used to stain the attached cells for 30 min at 25°C. The plates were then washed with PBS and left to dry for a further 10 min and the stained biomass was solubilized in 1% (w/v) SDS. The absorbance of each well was determined at OD595 nm. All experiments were repeated at least three times.
Scanning electron microscopy
Bacterial cells, grown in LB medium, were fixed with 2.5% (vol/vol) glutaraldehyde in phosphate-buffered saline (PBS, pH 7.4) and subsequently post-fixed in 1% (wt/vol) osmium tetroxide. The samples were then dehydrated in acetone, critical-point dried and coated with gold-palladium. The specimens were finally examined with a FEI Quanta 200 SEM scanning electron microscope (USA).
Carbon source utilization assays and chemical sensitivity assays
The Biolog GEN III MicroPlate was used to analyze the K. pneumoniae strains in 94 phenotypic tests, which including 71 carbon source utilization assays and 23 chemical sensitivity assays. Briefly, the bacterial culture were picked up from the surface of the BUG+B agar plate (Biolog, CA, USA) using a sterile cotton-tipped swab and inoculated into the IF-A Inoculum (Biolog, CA, USA). The target cell density of Inoculum was set to 90–98% T by turbidimeter (BioMérieux, Lyon, Fance). Then 100 µl of the inoculum was added into each well of the 96 GEN III MicroPlateTM (Biolog, CA, USA). After incubating the culture for 24 hours at 37°C, the OD590 readings were measured with a BIOLOG microplate reader automatically and further confirmed visually.
Genome sequencing
Genome sequencing was performed by Beijing Genomics Institute (BGI, China). Briefly, the genomic DNA for each bacterium was prepared by conventional phenol-chloroform extraction methods. A 500 bp paired-end library was constructed for each purified DNA sample following the standard Illumina paired-end protocol with a low-cycle polymerase chain reaction during the fragment enrichment and sequencing was performed on the Illumina Hiseq 2000 with 90 cycles. Low quality reads were filtered using the DynamicTrim and LengthSort Perl scripts within SolexaQA. Short reads were assembled using SOAPdenovo version 2.0440 and the gaps were closed by GapCloser version 1.12.
Genetic mutations detection and phylogenetic analysis
First, the short reads were aligned onto the K. pneumoniae ATCC BAA -2146 genome reference using the SOAP2 program41. Second, SOAPsnp was used to score SNPs from aligned reads42. SOAPsnp results were filtered as follows: (1) The read coverage of the SNP site was more than three, (2) The Illumina quality score of either allele was more than 30 and (3) The count of all mapped best base was more than two times the count of all mapped second best base. In addition, BWA 0.6.2 and SAMtools 0.1.18 were used to confirm the SNP results. The Illumina reads were first aligned by BWA with default parameters for each sample. The aligned results were piped to SAMtools to perform SNP and Indel analysis.
Electronic PCR for MLST genotyping and identification of drug resistance-associated genes
MLST with seven genes (gapA, infB, mdh, pgi, phoE, rpoB and tonB) was performed on isolates according to the protocol described on the K. pneumoniae MLST website (www.pasteur.fr/mlst). STs were assigned by using the MLST database (www.pasteur.fr/mlst/Kpneumoniae.html). Electronic PCR was used to extract the drug resistance-associated genes, then the DNA sequences were annotated using the BLAST program at http://www.ncbi.nlm.nih.gov.Mutations in the gyrA and parC genes were identified by comparing the DNA sequences with gyrA and parC sequences of the K. pneumoniae (GenBank accession numbers DQ673325 and NC009648 for gyrA and parC respectively).
RNA-Seq and comparative transcriptomic data analysis
Total RNA isolation as well as construction and sequencing of cDNA libraries of the K. pneumoniae strains were conducted by BGI (China). Briefly, total RNA samples were isolated using a RNeasy Protect Bacteria Mini Kit (QIAGEN, Germany) according to the manufacturer's instructions. Sequencing was carried out by running 90 cycles on the Illumina Hiseq 2000. Paired-end reads were mapped to K. pneumoniae ATCC BAA -2146 (CP006659) reference sequence using Bowtie243. The number of reads mapped to each gene was counted by using HTSeq-count (http://www-huber.embl.de/users/anders/HTSeq). The edgeR package44 from R/BioConductor was used to normalize the mapped count data and for differential gene expression analysis. Fold changes with FDR ≤ 0.001 were considered to be statistically significant. RPKM (Reads Per kb per Million reads) values were provided to enable comparison of relative transcript abundance among difference samples.
Functional annotation and enrichment analysis
KOBAS 2.0 (KEGG Orthology Based Annotation System) was used to identify metabolic pathways and to calculate the statistical significance of each pathway45. The COG annotation was performed using the Blastall software against the Cluster of Orthologus Groups (COG) database. COG enrichment analysis was determined by comparing the prevalence of differentially expressed genes assigned to a specific COG category to the prevalence of genes in the whole genome assigned to that COG category with a Fisher's exact test.
Quantitative RT-PCR of selected targets
To validate whether RNA-Seq provides reliable quantitative estimates of transcript levels, qRT-PCR analysis was conducted for 10 randomly selected genes. cDNA samples were analyzed by quantitative PCR with KAPA SYBR FAST qPCR Kit (KAPA Biosystems) on ABI 7300 system (Applied Biosystems). Data were analyzed by the 2−ΔΔCT method and normalized to the gapdh. Each experiment was performed in triplicates and repeated at least three times.
References
Little, M. L., Qin, X., Zerr, D. M. & Weissman, S. J. Molecular epidemiology of colonizing and disease-causing Klebsiella pneumoniae in paediatric patients. J Med Microbiol 63, 610–616, 10.1099/jmm.0.063354-0 (2014).
Shon, A. S., Bajwa, R. P. & Russo, T. A. Hypervirulent (hypermucoviscous) Klebsiella pneumoniae: a new and dangerous breed. Virulence 4, 107–118, 10.4161/viru.22718 (2013).
Lippmann, N., Lubbert, C., Kaiser, T., Kaisers, U. X. & Rodloff, A. C. Clinical epidemiology of Klebsiella pneumoniae carbapenemases. Lancet Infect Dis 14, 271–272, 10.1016/S1473-3099(14)70705-4 (2014).
van Duin, D. et al. Surveillance of Carbapenem-Resistant Klebsiella pneumoniae: Tracking Molecular Epidemiology and Outcomes through a Regional Network. Antimicrob Agents Chemother, 10.1128/AAC.02636-14 (2014).
Munoz-Price, L. S. et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect Dis 13, 785–796, 10.1016/S1473-3099(13)70190-7 (2013).
Kim, S. Y., Rhee, J. Y., Shin, S. Y. & Ko, K. S. Characteristics of community-onset NDM-1-producing Klebsiella pneumoniae isolates. J Med Microbiol 63, 86–89, 10.1099/jmm.0.067744-0 (2014).
Bushnell, G., Mitrani-Gold, F. & Mundy, L. M. Emergence of New Delhi metallo-beta-lactamase type 1-producing enterobacteriaceae and non-enterobacteriaceae: global case detection and bacterial surveillance. Int J Infect Dis 17, e325–333, 10.1016/j.ijid.2012.11.025 (2013).
Dortet, L., Poirel, L. & Nordmann, P. Worldwide Dissemination of the NDM-Type Carbapenemases in Gram-Negative Bacteria. Biomed Res Int 2014, 249856, 10.1155/2014/249856 (2014).
Stieglmeier, M. et al. Abundance and diversity of microbial inhabitants in European spacecraft-associated clean rooms. Astrobiology 12, 572–585, 10.1089/ast.2011.0735 (2012).
Vaishampayan, P. A., Rabbow, E., Horneck, G. & Venkateswaran, K. J. Survival of Bacillus pumilus spores for a prolonged period of time in real space conditions. Astrobiology 12, 487–497, 10.1089/ast.2011.0738 (2012).
Ghosh, S., Osman, S., Vaishampayan, P. & Venkateswaran, K. Recurrent isolation of extremotolerant bacteria from the clean room where Phoenix spacecraft components were assembled. Astrobiology 10, 325–335, 10.1089/ast.2009.0396 (2010).
Wilson, J. W. et al. Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq. Proc Natl Acad Sci U S A 104, 16299–16304, 10.1073/pnas.0707155104 (2007).
Lee, K. W. et al. Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J 8, 894–907, 10.1038/ismej.2013.194 (2014).
Andrade, L. N. et al. Expansion and Evolution of a Virulent, Extensively Drug-Resistant (Polymyxin B-Resistant), QnrS1-, CTX-M-2- and KPC-2-producing Klebsiella pneumoniae ST11 International High-risk Clone. J Clin Microbiol, 10.1128/JCM.00088-14 (2014).
Frye, J. G. & Jackson, C. R. Genetic mechanisms of antimicrobial resistance identified in Salmonella enterica, Escherichia coli and Enteroccocus spp. isolated from U.S. food animals. Front Microbiol 4, 135, 10.3389/fmicb.2013.00135 (2013).
Li, B. et al. Analysis of drug resistance determinants in Klebsiella pneumoniae isolates from a tertiary-care hospital in Beijing, China. PLoS One 7, e42280, 10.1371/journal.pone.0042280 (2012).
Robicsek, A., Sahm, D. F., Strahilevitz, J., Jacoby, G. A. & Hooper, D. C. Broader distribution of plasmid-mediated quinolone resistance in the United States. Antimicrob Agents Chemother 49, 3001–3003, 10.1128/AAC.49.7.3001-3003.2005 (2005).
Ramirez, M. S. & Tolmasky, M. E. Aminoglycoside modifying enzymes. Drug Resist Updat 13, 151–171, 10.1016/j.drup.2010.08.003 (2010).
Sangurdekar, D. P., Zhang, Z. & Khodursky, A. B. The association of DNA damage response and nucleotide level modulation with the antibacterial mechanism of the anti-folate drug trimethoprim. BMC Genomics 12, 583, 10.1186/1471-2164-12-583 (2011).
Qi, Y. et al. ST11, the dominant clone of KPC-producing Klebsiella pneumoniae in China. J Antimicrob Chemother 66, 307–312, 10.1093/jac/dkq431 (2011).
Reid, G. et al. Effects of ciprofloxacin, norfloxacin and ofloxacin on in vitro adhesion and survival of Pseudomonas aeruginosa AK1 on urinary catheters. Antimicrob Agents Chemother 38, 1490–1495 (1994).
Al Bahry, S., Sivakumar, N. & Al-Khambashi, M. Effect of nalidixic acid on the morphology and protein expression of Pseudomonas aeruginosa. Asian Pac J Trop Med 5, 265–269, 10.1016/S1995-7645(12)60037-6 (2012).
Kim, H., Goo, E., Kang, Y., Kim, J. & Hwang, I. Regulation of universal stress protein genes by quorum sensing and RpoS in Burkholderia glumae. J Bacteriol 194, 982–992, 10.1128/JB.06396-11 (2012).
Hasona, A. et al. Streptococcal viability and diminished stress tolerance in mutants lacking the signal recognition particle pathway or YidC2. Proc Natl Acad Sci U S A 102, 17466–17471, 10.1073/pnas.0508778102 (2005).
Cavanagh, A. T. & Wassarman, K. M. 6S RNA, A Global Regulator of Transcription in Escherichia coli, Bacillus subtilis, and Beyond. Annu Rev Microbiol, 10.1146/annurev-micro-092611-150135 (2014).
Steuten, B. et al. Regulation of transcription by 6S RNAs: Insights from the Escherichia coli and Bacillus subtilis model systems. RNA Biol 11 (2014).
Ortega, A. D., Gonzalo-Asensio, J. & Garcia-del Portillo, F. Dynamics of Salmonella small RNA expression in non-growing bacteria located inside eukaryotic cells. RNA Biol 9, 469–488, 10.4161/rna.19317 (2012).
Li, W. et al. Increasing occurrence of antimicrobial-resistant hypervirulent (hypermucoviscous) Klebsiella pneumoniae isolates in China. Clin Infect Dis 58, 225–232, 10.1093/cid/cit675 (2014).
Mukherjee, J., Ow, S. Y., Noirel, J. & Biggs, C. A. Quantitative protein expression and cell surface characteristics of Escherichia coli MG1655 biofilms. Proteomics 11, 339–351, 10.1002/pmic.201000386 (2011).
Kang, A., Tan, M. H., Ling, H. & Chang, M. W. Systems-level characterization and engineering of oxidative stress tolerance in Escherichia coli under anaerobic conditions. Mol Biosyst 9, 285–295, Doi 10.1039/C2mb25259g (2013).
Van Nguyen, T., Kroger, C., Bonnighausen, J., Schafer, W. & Bormann, J. The ATF/CREB Transcription Factor Atf1 Is Essential for Full Virulence, Deoxynivalenol Production and Stress Tolerance in the Cereal Pathogen Fusarium graminearum. Mol Plant Microbe In 26, 1378–1394, Doi 10.1094/Mpmi-04-13-0125-R (2013).
Hartmann, T. et al. Catabolite Control Protein E (CcpE) Is a LysR-type Transcriptional Regulator of Tricarboxylic Acid Cycle Activity in Staphylococcus aureus. J Biol Chem 288, 36116–36128, DOI 10.1074/jbc.M113.516302 (2013).
Dufour, V. et al. Inactivation of the LysR regulator Cj1000 of Campylobacter jejuni affects host colonization and respiration. Microbiol-Sgm 159, 1165–1178, Doi 10.1099/Mic.0.062992-0 (2013).
Shimada, T., Hirao, K., Kori, A., Yamamoto, K. & Ishihama, A. RutR is the uracil/thymine-sensing master regulator of a set of genes for synthesis and degradation of pyrimidines. Mol Microbiol 66, 744–757, 10.1111/j.1365-2958.2007.05954.x (2007).
Reen, F. J., Haynes, J. M., Mooij, M. J. & O'Gara, F. A non-classical LysR-type transcriptional regulator PA2206 is required for an effective oxidative stress response in Pseudomonas aeruginosa. PLoS One 8, e54479, 10.1371/journal.pone.0054479 (2013).
Huang, S. H. et al. Role of the small RNA RyhB in the Fur regulon in mediating the capsular polysaccharide biosynthesis and iron acquisition systems in Klebsiella pneumoniae. BMC Microbiol 12, 148, 10.1186/1471-2180-12-148 (2012).
Gerstle, K., Klatschke, K., Hahn, U. & Piganeau, N. The small RNA RybA regulates key-genes in the biosynthesis of aromatic amino acids under peroxide stress in E. coli. RNA Biol 9, 458–468, 10.4161/rna.19065 (2012).
Willett, N. P. & Morse, G. E. Long-chain fatty acid inhibition of growth of Streptococcus agalactiae in a chemically defined medium. J Bacteriol 91, 2245–2250 (1966).
Cumley, N. J., Smith, L. M., Anthony, M. & May, R. C. The CovS/CovR acid response regulator is required for intracellular survival of group B Streptococcus in macrophages. Infect Immun 80, 1650–1661, 10.1128/IAI.05443-11 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18, 10.1186/2047-217X-1-18 (2012).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967, 10.1093/bioinformatics/btp336 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res 19, 1124–1132, 10.1101/gr.088013.108 (2009).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, 10.1038/nmeth.1923 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, 10.1093/bioinformatics/btp616 (2010).
Xie, C. et al. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39, W316–322, 10.1093/nar/gkr483 (2011).
Acknowledgements
Financial support was provided by the National Basic Research Programs of China (2014CB744400 and 2012CB518700), National Natural Science Foundation of China (81371769) and the Ministry of Health and the Ministry of Science and Technology, China (2013ZX10003006) and the Chinese Academy of Sciences (KJZD-EW-L02). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.L., F.L. and Q.W. contributed equally to this paper. C.H.L. and C.L. designed and coordinated the project. J.L., Q.W. and P.G. performed laboratory experiments. C.H.L., F.L. and Q.W. performed the data analysis; C.H.L. wrote the manuscript with assistance from other authors. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Li, J., Liu, F., Wang, Q. et al. Genomic and transcriptomic analysis of NDM-1 Klebsiella pneumoniae in spaceflight reveal mechanisms underlying environmental adaptability. Sci Rep 4, 6216 (2014). https://doi.org/10.1038/srep06216
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06216
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.