Abstract
Hypoxia and reoxygenation are common characteristics of solid tumors, which lead to oxidative stress and activation of stress-response genes. Previously, we observed that N-myc downstream-regulated gene 1 (NDRG1) was strongly down-regulated after shifting to reoxygenation, but the regulatory mechanism of NDRG1 remained elusive. Here we focused on the regulation of NDRG1 by microRNAs (miRNAs). Breast cancer MCF-7 cells were cultured under hypoxia for 24 h followed by 24 h of reoxygenation. The miRNA profiles were examined by Nanostring nCounter assays. Forty-three miRNAs had significant changes upon reoxygenation. In silico analysis identified four oxygen-sensitive miRNAs whose seed regions perfectly matched the 3′-UTR of NDRG1. In particular, miR-769-3p was able to inhibit the expression of NDRG1, which caused a significant reduction of NDRG1 protein upon reoxygenation. Furthermore, overexpression of miR-769-3p significantly inhibited cell proliferation and enhanced apoptosis. Our results revealed that miR-769-3p can functionally regulate NDRG1 during changes in oxygen concentration.
Similar content being viewed by others
Introduction
Tumor populations need to overcome distinct microenvironmental barriers prior to metastasizing to other organs. It has been suggested that the tumor microenvironment is a unique setting for tumor progression, which requires genetic and adaptive changes in cancer cells for further survival and proliferation. Invasive cancers, therefore, could be viewed as a sequence of phenotypic adaptations to their microenvironment. The tumor microenvironment is characterized by nutrient deprivation, low pH and hypoxia1. These aberrations were linked to perfusion deficits in solid tumors, which came from rapid tumor growth and profoundly disorganized vasculature2.
Although regions of hypoxia are a common feature in solid tumors, the oxygen concentrations within a hypoxic region are highly variable. Because tumor vasculature is highly inefficient and unstable, masses of red blood cells irregularly flow to the hypoxic regions, resulting in reperfusion or reoxygenation3. Reoxygenation not only increases oxygen supply but also results in complex changes in cellular energy charge, oxidant generating systems and antioxidant defenses4. These changes lead to oxidative stress5, tissue damage and activation of stress-response genes6. These cellular responses are activated by several key signaling pathways, such as hypoxia-inducible factor (HIF-1), nuclear factor-κB (NF-κB), activator protein 1 (AP-1) and some mitogen-activated protein kinase (MAPK) pathways6.
Previously, we used genome-wide expression microarrays to investigate the dynamics of gene profiling during reoxygenation in breast cancer MCF-7 cells and demonstrated that N-myc downstream-regulated gene 1 (NDRG1) was involved in tumor adaptation to reoxygenation7. NDRG1 is a 43-kD protein composed of 394 amino acids and is highly conserved among multicellular organisms. It is expressed ubiquitously in tissues stimulated under a wide variety of stress and cell growth-regulatory conditions, such as DNA damage8, cellular differentiation9, proliferation and growth arrest10, or hypoxia11,12. NDRG1 was reported to be strongly up-regulated under hypoxic conditions or nickel (Ni2+) treatment via hypoxia inducible factor 1 (HIF-1), Egr-1, or Sp113,14,15. Yet, the regulatory mechanism of NDRG1 upon oxygen fluctuation is still unclear.
An oncogenic and tumor-promoting role of NDRG1 has been suggested, because it is overexpressed in various human cancers, including lung, brain, skin, kidney and breast cancers13,16. Since hypoxia is prevalent in many solid tumors, its regulation is governed by hypoxia-inducible factor 1 alpha (HIF-1α) and p53-dependent pathways17,18. NDRG1 can be induced to promote cell migration under intermittent hypoxia19. However, NDRG1 functions as a metastatic suppressor in prostate and colon cancers9,20. The contradictory roles of NDRG1 in different cancers remain to be clarified, although it might be explained by their multiple cellular localizations and complex regulation by diverse physiological and pathological factors.
MicroRNAs (miRNAs), 18–25 nucleotides in length, are a class of non-protein-encoding small RNAs21. MiRNAs originate from longer primary miRNA transcripts located in either intergenic or intronic regions22. Intronic miRNAs are co-expressed with the genes in which they are located and are regulated from the same promoters as their host genes22. It has been indicated that miRNAs regulate the expression of more than 30% of protein-coding genes23 and that miRNAs control multiple biological and metabolic processes, ranging from tissue development and signal transduction to the disease processes of tumorigenesis and cancer progression24,25. Regulation of gene expression by miRNAs is achieved by translation repression, direct mRNA degradation or miRNA-mediated mRNA decay26. In a majority of cases, miRNAs bind with imperfect complementarity, with the exception of their ‘seed region’, to their targeted mRNAs at the 3′-UTR by the RNA induced silencing complex (RISC). MiRNA-RISC complex may interact with various translation factors, such as eIF4F and inhibit the initiation and/or elongation of protein translation27. MiRNAs can also directly degrade their targeted mRNAs or destabilize mRNAs through de-adenylation and/or de-capping of targeted mRNAs28.
Although >2,000 miRNAs have been discovered in mammals, the functions of most miRNAs are not understood. Recent studies revealed that a significant proportion of hypoxia-regulated miRNAs were overexpressed in human cancers and could affect important processes, such as apoptosis, proliferation and angiogenesis29. For instance, miR-210, miR-199a and miR-34a-5p were reported to regulate hypoxia and angiogenesis30,31,32. MiR-182 promoted proliferation and invasion of human prostate cancer cells by directly suppressing NDRG133.
Since the regulatory mechanism of NDRG1 upon reoxygenation remains elusive, we investigated whether the down-regulation of NDRG1 upon reoxygenation was achieved partly through miRNA regulation. Thus, miRNA expression upon reoxygenation was examined using NanoString nCounter® miRNA Expression Assays and analyzed by in silico analysis. We discovered that, upon reoxygenation, miR-769-3p can directly inhibit NDRG1, resulting in reduced proliferation and enhanced apoptosis in MCF-7 breast cancer cells.
Results
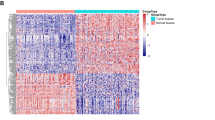
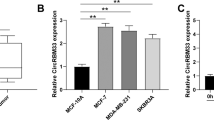
Previous studies have reported that the expression levels of miRNAs were changed under hypoxia34,35. To investigate whether NDRG1 is regulated by miRNA under different O2 conditions, we first used a hybridization-assisted, expressed sequence tag (EST)-based technique, the NanoString nCounter™ Analysis System, to identify differentially expressed miRNAs during reoxygenation. MCF-7 cells were grown in hypoxia for 24 h and harvested at 0, 1, 12 and 24 h after reoxygenation to measure the expression profiles of miRNAs. Fold change assessments (>1.5×) and t-tests (P < 0.05) of each time point as compared to the hypoxia control (time 0) were applied for selecting oxygen-responsive miRNAs. We identified 43 miRNAs passing these criteria, with 63% (n = 27) of the miRNAs up-regulated upon reoxygenation. The heat map of these differentially expressed miRNAs is shown in Fig. 1A. Because miRNAs usually negatively regulate their target genes and because NDRG1 was down-regulated upon reoxygenation, the up-regulated miRNAs were chosen for further investigation.
Identification of miRNAs that were differentially expressed during reoxygenation.
(A) Heatmap of the differentially expressed miRNAs during reoxygenation (re-O2) using NanoString nCounter® miRNA Expression Assays. Red: up-regulated in re-O2 as compared to hypoxia control (0 h); green: down-regulated in re-O2. (B–E) NanoString nCounter® miRNA Expression Assays and real-time PCR validation of miR-769-3p (B), miR-501-3p (C), miR-2276 (D) and miR-1282 (E) in re-O2. RNU44 was used as an internal control for real-time PCR. Data in the bar chart are the mean ± SD for three independent experiments.*, P < 0.05.
In order to examine which miRNAs directly regulate NDRG1, a bioinformatics algorithm, Target Scan (http://www.targetscan.org), was used to search the binding sites of miRNAs in the NDRG1 3′-UTR. Among the 27 up-regulated miRNAs, the seed regions of miR-769-3p, miR-501-3p, miR-2276 and miR-1282 matched perfectly the NDRG1 3′-UTR sequence. Therefore, we validated the expression of these four miRNAs by quantitative RT-PCR. As shown in Fig. 1B–E, the expression levels of miR-769-3p, miR-501-3p, miR-2276 and miR-1282 were all significantly up-regulated at 24 h under reoxygenation, showing that these miRNAs are responsive to reoxygenation.
Furthermore, molecular dynamics simulation and thermodynamics analysis were used to predict the strength of binding affinity36. The results showed that miR-769-3p had the highest alignment score and the most favorable thermodynamics (i.e., lowest G score) (Table 1). Therefore, we focused on miR-769-3p for the following experiments.
First of all, we examined the endogenous expression levels of miR-769-3p and its precursor in breast cancer MCF-7 cells at different oxygen concentrations. As shown in Fig. 2A, miR-769-3p was up-regulated during normoxia and reoxygenation as compared to hypoxia, indicating that miR-769-3p is an oxygen-dependent miRNA. However, the expression levels of precursor mir-769-3p in hypoxia were not significantly different from those in normoxia and reoxygenation (Fig. 2B).
MiR-769-3p is down-regulated in hypoxia and up-regulated upon re-oxygenation.
(A) Relative expression levels of miR-769-3p in normoxia (O2), hypoxia (N2) and reoxygenation (Re-O2) using quantitative RT-PCR. Breast cancer MCF-7 cells were cultured under normoxia, hypoxia for 24 h, or 24 h of hypoxia followed by 24 h of reoxygenation. The relative expression level of each condition was compared to hypoxia; 18s rRNA was used as an internal control. *: P < 0.05. (B) Relative expression levels of precursor mir-769-3p at different oxygen levels using quantitative RT-PCR.
Next, in order to explore whether miR-769-3p could directly bind and regulate NDRG1, we used computational algorithms37 to predict the potential binding sites in the NDRG1 3′-UTR and examined their interaction using luciferase assays. The location of the potential binding site was 1,014–1,036 bp downstream relative to the start site of NDRG1 3′-UTR (Fig. 3A). Because the seed region of the miRNA, which includes 2 to 8 nucleotides at the 5′-end of the miRNA, must be complementary to the 3′-UTR of target genes, we mutated five nucleotides in the binding site to validate their interaction (Fig. 3A). By co-transfecting the miR-769-3p plasmids with the reporter construct, which contained the NDRG1 3′-UTR behind the luciferase gene (Fig. 3A), to HEK 293 cells, we observed that, in normoxia, miR-769-3p was able to inhibit the luciferase activity as compared to the miR-empty vector control (Fig. 3B). When the binding site was mutated, the luciferase activity was recovered (Fig. 3B). These results suggest that NDRG1 is the target of miR-769-3p.
MiR-769-3p directly regulates NDRG1 activity.
(A) Schematic representation of the NDRG1 3′-UTR expression plasmid and the miR-769-3p target sequence. Firefly luciferase constructs contained the CMV promoter, luciferase coding region and a fragment of the NDRG1 3′-UTR wild-type (WT) or mutant (MUT). The mutation sites were underlined. (B) Luciferase assays of miR-769-3p co-transfected with firefly luciferase constructs containing NDRG1 3′-UTR wild-type or mutant into HEK 293 cells. Renilla luciferase was used as the transfection control. The relative luciferase activity represented the dual luciferase activity ratio (firefly/Renilla luciferase). *: P < 0.05.
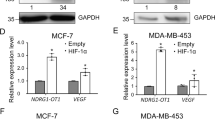
Afterwards, we overexpressed miR-769-3p in MCF-7 cells (Fig. S1) and examined the mRNA and protein expression of NDRG1 upon reoxygenation. As expected, overexpression of miR-769-3p led to a significant reduction of both NDRG1 mRNA (Fig. 4A) and protein (Fig. 4B) upon reoxygenation. These data indicate that down-regulation of NDRG1 is at least partly regulated by miR-769-3p under reoxygenation.
Overexpression of miR-769-3p suppresses NDRG1 expression.
(A) Relative expression levels of NDRG1 in MCF-7 cells overexpressing pri-mir-769-3p in normoxia using quantitative RT-PCR; 18S rRNA was used as a loading control. (B) Western blot analysis of NDRG1 in MCF-7 cells overexpressing pri-mir-769-3p in normoxia (O2), hypoxia (N2) and reoxygenation (Re-O2). GAPDH was used as an internal control. Full-length blots/gels are presented in Supplementary Figure 2 & 3.
Lastly, previous studies report that silencing NDRG1 can reduce proliferation. Therefore, we hypothesized that overexpression of miR-769-3p could also reduce proliferation through inhibiting NDRG1. As shown in Fig. 5A, cell proliferation was indeed suppressed under normoxia, hypoxia and reoxygenation in the presence of ectopic miR-769-3p using MTT assays. Furthermore, flow cytometry analysis showed that miR-769-3p-expressing cells exhibited more late apoptosis at 24 h of normoxia (O2), hypoxia (N2) and reoxygenation (Re-O2) than control cells using Annexin V binding assays (Fig. 5B & 5C). In addition, at 24 h of reoxygenation, there were higher proportions of miR-769-3p-expressing cells in sub-G1 phase than control cells (Fig. 5D & 5E). These results suggest that miR-769-3p can reduce proliferation and enhance apoptosis by inhibiting NDRG1 upon reoxygenation.
Overexpression of miR-769-3p suppresses proliferation and induces apoptosis of MCF-7 breast tumor cells.
(A) MTT assays of MCF-7 cells overexpressing miR-769-3p in normoxia (O2), hypoxia (N2) and reoxygenation (Re-O2). (B) A representative diagram of annexin V binding assay at 24 h of O2, N2 and Re-O2. Left panel: empty vector; right panel: MCF-7 cells overexpressing miR-769-3p. (C) Flow cytometry analysis for apoptosis at 24 h of O2, N2 and Re-O2. using annexin V and propidium iodide in MCF-7 cells overexpressing miR-769-3p. (D) A representative diagram of flow cytometry for analyzing cell cycle at 24 h of reoxygenation. Upper panel: empty vector; lower panel: MCF-7 cells overexpressing miR-769-3p. (E) Quantitative graph of flow cytometry results in panel (D) showing the percentage of cells in each phase of the cell cycle. *, P < 0.05.
Discussion
In this study, we used a genome-wide approach to screen all miRNAs and identified 43 oxygen-sensitive miRNAs in MCF-7 breast cancer cells. Among these oxygen-sensitive miRNAs, we demonstrated that miR-769-3p could directly inhibit NDRG1. Also, miR-769-3p could enhance apoptosis and suppress cell proliferation upon reoxygenation.
Previous studies have used miRNA microarrays to screen all miRNAs and identify hypoxia-related miRNA candidates34,38. However, in our pilot study, many previously reported hypoxia-related miRNAs could not be validated to regulate NDRG1. There are at least two possible reasons for this. First, since the previous microarrays used hybridization between fluorescent miRNAs and probes and the relative expression levels were quantified by intensity of fluorescence, the microarrays had low sensitivity and limited dynamic range. Therefore, the fluorescence signals were amplified using PCR/in vitro transcription, which may have generated non-linear amplification and led to misrepresentation of the quantitative transcript values39,40. A second possible reason why previous microarray approaches failed to identify more NDRG1-regulatory miRNAs is a lack of probes to detect newly discovered miRNAs. Therefore, we used a new version of the NanoString nCounter® Analysis System, a sequence-tag counting-based technology, to do a genome-wide screen in this study. The advantages of NanoString are that it has higher sensitivity and dynamic range and does not require amplification or labeling of miRNA41.
Through the screening of oxygen-responsive miRNAs, 43 miRNAs had dramatic changes as compared to 0 h upon reoxygenation. Among these 43 miRNAs, miR-210 and miR-34a-5p were known to be up-regulated in hypoxia signaling pathways30,42. Hence, miR-210 and miR-34a-5p should be down-regulated upon reoxygenation, which was consistent with our Nanostring results (Fig. 1A).
Based on the results of sequence analysis, molecular dynamics simulation and thermodynamics analysis, miR-769-3p was found to be the most likely miRNA to regulate NDRG1 and hence was chosen for further validation. MiR-769-3p was first discovered in 200643. The location of miR-769-3p is in the first intron of CCDC61. However, the role of miR-769-3p has not previously been reported. In this study, we demonstrated that miR-769-3p could down-regulate NDRG1 by degrading the mRNA of NDRG1 and inhibiting its protein levels when oxygen was available. This also implied that miR-769-3p was dissociated from NDRG1 under hypoxia. Yet, the detailed mechanism of the oxygen-sensitive association of miR-769-3p with NDRG1 needs further exploration. Furthermore, miR-769-3p not only regulated NDRG1 but was also involved in other hypoxia/reoxygenation-responsive pathways. According to the computational prediction of the function of miR-769-3p's target genes37, miR-769-3p may be involved in c-Myc's transcription factor network, the Rac-1signaling pathway, the MAPK pathway and the HIF-2α transcription factor network, via targeting to PGK1, MAPK14, CDKN1A (P21) and SP1, which are upstream or downstream of NDRG1 in these signaling pathways.
Overexpression of miR-769-3p led to a significant reduction of NDRG1 protein and mRNA upon reoxygenation, but not in hypoxia (Fig. 4B). This result suggested that miR-769-3p alone was not sufficient to reduce NDRG1 levels in hypoxia. We can envision two possible scenarios. First, up-regulation of NDRG1 under hypoxia may not be regulated solely by miRNAs, in which case the enormous expression of NDRG1 in hypoxia by other transcriptional regulations may offset the inhibitory effect of ectopic miR-769-3p on NDRG1. Second, since miR-769-3p may have many target genes in addition to NDRG1, the miR-769-3p-regulated genes may compete with NDRG1 for miR-769-3p. These RNAs containing the same miRNA binding sites in their 3′-UTR are referred to as competing endogenous RNAs (ceRNAs), which could regulate each other by competing for the shared miRNAs44,45. Therefore, when ceRNAs were up-regulated, more miR-769-3p bound to these ceRNAs, which led to less inhibition of NDRG1 and subsequently up-regulated NDRG1. According to computational predictions37, miR-769-3p could target MAPK14, PGK1 and SP1, which all have increased expression in hypoxia. Therefore, the up-regulation of MAPK14, PGK1 and SP1 could compete with NDRG1 for miR-769-3p and weaken its inhibition of NDRG1 in hypoxia.
Regarding the functions of miR-769-3p, we showed that overexpression of miR-769-3p could inhibit cell proliferation and induce cell apoptosis (Fig. 5). However, miR-769-3p might regulate hundreds of mRNAs. Whether the induction of apoptosis and inhibition of cell proliferation were solely through the NDRG1 signaling pathway is not currently known. Yet, we found that the patterns of proliferation and apoptosis were similar in MCF-7 cells transfected with NDRG1 wild-type 3′-UTR and mutant 3′-UTR in normoxia, hypoxia and reoxygenation (data not shown), implicating that NDRG1 was not the only target gene of miR769-3p. That is, the effects of miR-769-3p on cell proliferation and apoptosis were not solely dependent on NDRG1. In summary, this study is the first one to report the function of miR-769-3p and demonstrates that miR-769-3p can functionally regulate NDRG1 upon reoxygenation.
Methods
Cell culture
Human breast adenocarcinoma cell line MCF-7 was obtained from the Bioresource Collection and Research Center (Hsinchu, Taiwan). MCF-7 cells were maintained in Dulbecco's modified Eagle's medium (DMEM, Life Technologies, Carlsbad, CA, USA) supplemented with 10% (v/v) fetal bovine serum (Life Technologies) and 1% antibiotic solution (penicillin-streptomycin-amphotericin solution, Biological Industries, Beit-Haemek, Israel) at 37°C in a humidified incubator under 5% CO2. For hypoxia cultures, cells were incubated in a hypoxia chamber (InVivO2-200, Ruskinn Technology, Leeds, UK) for 24 h with 0.5% O2, 5% CO2 and residual N2. After 24 h of hypoxia, cells were shifted to a humidified incubator with 5% CO2 and 95% air and incubated at 37°C. The samples were harvested at selected time points after reoxygenation.
Nanostring data analysis
The nCounter® miRNA Expression Assays kit (Nanostring, Seattle, WA, USA) was utilized to perform miRNA expression profiling. Each miRNA was detected by ligating the unique digital tags which contain a specific digital barcode representing the counts of miRNAs. The workflow of normalization was according to the manufacturer's instructions. The samples were normalized to the geometric mean of the 100 miRNAs with the highest counts. The O2-responsive miRNAs were selected when their expression change was significantly (P < 0.05) greater than 1.5-fold at a given time point as compared to the hypoxia control (0 h).
Quantitative reverse transcription PCR
Total RNA was extracted using TRIZOL Reagent (Ambion, Carlsbad, CA, USA) according to the manufacturer's instructions. Reverse transcription of miRNA was performed using the TaqMan MicroRNA Reverse Transcription kit (Life Technologies). The resulting cDNA was detected using the FastStart Universal SYBR Green Master Mix (Roche, Branchburg, NJ, USA) or Universal ProbeLibrary Probe #21 (Roche) with a 7900HT Fast Real-Time PCR system (Life Technologies). RNU44 was used as an endogenous control to normalize the expression of miRNA. The following primers were used to detect miRNA expression: miR-769-3p: 5′- GCGGCGGCTGGGATCTCCGGGGTC -3′ (forward), 5′- GTGCAGGGTCCGAGGT -3′ (reverse); miR-1282: 5′- GCGGCGGTCGTTTGCCTTTTT -3′ (forward), 5′- GTGCAGGGTCCGAGGT -3′ (reverse); miR-501-3p: 5′- GCGGCGGAATGCACCCGGGCAAG -3′ (forward), 5′- GTGCAGGGTCCGAGGT -3′ (reverse); miR-2276: 5′- GCGGCGGTCTGCAAGTGTCAGAG -3′ (forward), 5′- GTGCAGGGTCCGAGGT -3′ (reverse); RNU44: 5′- TCGCGCCTGGATGATGATAGC -3′ (forward), 5′- GTGCAGGGTCCGAGGT -3′ (reverse).
Construct design and cell transfection
MiRNA-expressing vector
The primary sequence of miR-769-3p, including the flanking precursor sequence (150 bp long), was amplified from human leukocyte DNA. The following primers were used: miR-769-3p: 5′- CATGGATCCAGGATGCCTTTGGCTGTGT -3′ (forward) and 5′- CATAGATCTACACCTCCCCTCCTTCTCTC -3′ (reverse). The PCR product was gel-purified and cloned into the pcDNA6.2-GW/EmGFP-miR-neg vector (Life Technologies).
Luc-NDRG1 vector
The NDRG1 3′-UTR was amplified by PCR from genomic DNA isolated from human blood with the following primers: 5′- CATACTAGTGCGGCCTGCCCAGCTGCCGC -3′ (forward) and 5′- CATGCCGGCCCGCTGCAAAGTTACAA -3′ (reverse). The pMIR-NDRG1-3′-UTR construct was digested with SpeI and NaeI and the generated fragment was inserted into the SpeI-NaeI sites of the pMIR-REPORT miRNA Expression Reporter Vector (Life Technologies). The miR-769-3p binding site (1,014–1,036 bp downstream relative to the start site of 3′-UTR of NDRG1) was predicted using miRSystem37. Five nucleotide mutations of the miR-769-3p binding site were made in the NDRG1 3′-UTR as a negative control with no binding. The Luc-NDRG1 vector was amplified by PCR with KAPA HiFi HotStart polymerase (KAPA Biosystems, Woburn, MA, USA) with mutated primers. After amplification, the template plasmid DNA with methylation was eliminated by DpnI endonuclease (New England Biolabs, Inc., Ipswich, MA, USA). The mutated plasmids were then transformed into E. coli for amplification, extracted and sequenced for validation.
Cell transfection
MCF-7 cells were seeded in antibiotic-free medium at 70–80% confluence. The cells were transfected with the miR-769-3p-expression vector using TransIT®-2020 Transfection Reagent (Mirus Bio LLC, Madison, WI, USA) according to the manufacturer's instructions.
Luciferase reporter assay
HEK293 cells were co-transfected with 300 ng of miR-769-3p, 100 ng of the reporter vector containing the NDRG1 3′-UTR or the mutant 3′-UTR and 25 ng of the Renilla luciferase vector as a transfection control. After 48 h of transfection, the cells were collected and the luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Western blot
The cells were washed twice with phosphate-buffered saline (PBS) (Life Technologies) and lysed in RIPA lysis buffer (Sigma, St. Louis, MO, USA). Protein concentrations were determined using the Protein Assay Reagent (Bio Rad Laboratories, Hercules, CA, USA). Protein samples (30 μg) were loaded on 10% sodium dodecyl sulfate (SDS)-polyacrylamide gels. After electrophoresis, the proteins were transferred to polyvinylidene difluoride membranes (Bio Rad Laboratories, Inc.). Blocking was performed with 3% nonfat milk in a 1× mixture of Tris-buffered saline. The membranes were incubated overnight at 4°C with antibodies to NDRG1 (Abcam Inc., Cambridge, MA, USA) and GAPDH (GeneTex, Irvine, CA, USA). After washing and incubation with secondary antibodies for 1 h at room temperature, blotted proteins were detected using an enhanced chemiluminescence (ECL) system (Millipore, Billerica, MA, USA) with the BioSpectrum Imaging System (UVP, Upland, CA, USA).
MTT assay
Cells were seeded at a density of 3.0 × 104 cells/400 μl in a 24-well plate. After 12 h of incubation, cells were incubated with 3-(4,5-dimethylthiazol-2-yl)- 2,5-diphenyltetrazolium bromide (MTT) solution (Sigma) (1 ml per well of 5 mg/ml solution in PBS) for 3 h. Next, 400 μl/well of DMSO (Sigma) was added to dissolve the converted purple formazan and the absorbance of formazan was measured at 540 nm using an enzyme-linked immunosorbent assay (ELISA) reader (BioTek, Winooski, VT, USA).
Flow cytometry analysis of cell cycle and cell death
MCF-7 cells were seeded at a density of 4 × 105 cells/well in a 6-well plate and transfected with miR-769-3p plasmid. After transfection, cells were incubated in hypoxia for 24 h and shifted to normoxia for 24 h. Samples were trypsinized, resuspended in PBS (Life Technologies) and fixed with cold 70% ethanol overnight. The DNA contents were evaluated after staining with propidium iodide (PI) solution containing 50 μg/ml propidium iodide (PI) (Sigma), 0.1 mg/ml RNase A (Sigma), 0.05% Triton X-100 (Sigma) in PBS (Life Technologies). Cell cycle analysis was carried out using a Beckman Coulter FC500 (Beckman, Brea, CA, USA) and CellQuest software.
The FITC Annexin V Apoptosis Detection Kit (BD Pharmingen, San Jose, CA, USA) was used to detect apoptotic cells by flow cytometry. Annexin V binding buffer was used to resuspend cells and the cell suspensions were stained with FITC-annexin V and PI staining solution for 15 min at room temperature. The apoptotic/necrotic cell population was analyzed with a Beckman Coulter FC500 (Beckman).
References
Williams, K. J., Cowen, R. L. & Stratford, I. J. Hypoxia and oxidative stress. Tumour hypoxia--therapeutic considerations. Breast Cancer Res 3, 328–31 (2001).
Vaupel, P., Kallinowski, F. & Okunieff, P. Blood flow, oxygen and nutrient supply and metabolic microenvironment of human tumors: a review. Cancer Res 49, 6449–65 (1989).
Brown, J. M. & Giaccia, A. J. The unique physiology of solid tumors: opportunities (and problems) for cancer therapy. Cancer Res 58, 1408–16 (1998).
McCord, J. M. Oxygen-derived free radicals in postischemic tissue injury. N Engl J Med 312, 159–63 (1985).
Li, C. & Jackson, R. M. Reactive species mechanisms of cellular hypoxia-reoxygenation injury. Am J Physiol Cell Physiol 282, C227–41 (2002).
Harris, A. L. Hypoxia--a key regulatory factor in tumour growth. Nat Rev Cancer 2, 38–47 (2002).
Lai, L. C. et al. Down-Regulation of NDRG1 Promotes Migration of Cancer Cells during Reoxygenation. PloS one 6, e24375 (2011).
Kurdistani, S. K. et al. Inhibition of tumor cell growth by RTP/rit42 and its responsiveness to p53 and DNA damage. Cancer Res 58, 4439–44 (1998).
Guan, R. J. et al. Drg-1 as a differentiation-related, putative metastatic suppressor gene in human colon cancer. Cancer Res 60, 749–55 (2000).
Piquemal, D. et al. Differential expression of the RTP/Drg1/Ndr1 gene product in proliferating and growth arrested cells. Biochim Biophys Acta 1450, 364–73 (1999).
Salnikow, K., Costa, M., Figg, W. D. & Blagosklonny, M. V. Hyperinducibility of hypoxia-responsive genes without p53/p21-dependent checkpoint in aggressive prostate cancer. Cancer Res 60, 5630–4 (2000).
Salnikow, K. et al. The regulation of hypoxic genes by calcium involves c-Jun/AP-1, which cooperates with hypoxia-inducible factor 1 in response to hypoxia. Mol Cell Biol 22, 1734–41 (2002).
Cangul, H. Hypoxia upregulates the expression of the NDRG1 gene leading to its overexpression in various human cancers. BMC Genet 5, 27 (2004).
Wang, Q. et al. HIF-1alpha up-regulates NDRG1 expression through binding to NDRG1 promoter, leading to proliferation of lung cancer A549 cells. Mol Biol Rep 40, 3723–9 (2013).
Zhang, P., Tchou-Wong, K. M. & Costa, M. Egr-1 mediates hypoxia-inducible transcription of the NDRG1 gene through an overlapping Egr-1/Sp1 binding site in the promoter. Cancer Res 67, 9125–33 (2007).
Gomez-Casero, E. et al. Regulation of the differentiation-related gene Drg-1 during mouse skin carcinogenesis. Mol Carcinog 32, 100–9 (2001).
Salnikow, K., An, W. G., Melillo, G., Blagosklonny, M. V. & Costa, M. Nickel-induced transformation shifts the balance between HIF-1 and p53 transcription factors. Carcinogenesis 20, 1819–23 (1999).
Masuda, K. et al. Downregulation of Cap43 gene by von Hippel-Lindau tumor suppressor protein in human renal cancer cells. Int J Cancer 105, 803–10 (2003).
Toffoli, S. et al. NDRG1 and CRK-I/II are regulators of endothelial cell migration under Intermittent Hypoxia. Angiogenesis 12, 339–54 (2009).
Bandyopadhyay, S. et al. The Drg-1 gene suppresses tumor metastasis in prostate cancer. Cancer Res 63, 1731–6 (2003).
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–54 (1993).
Monteys, A. M. et al. Structure and activity of putative intronic miRNA promoters. RNA 16, 495–505 (2010).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–45 (2005).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–5 (2004).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6, 259–69 (2006).
Zhang, B., Wang, Q. & Pan, X. MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol 210, 279–89 (2007).
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5, 522–31 (2004).
Wu, L., Fan, J. & Belasco, J. G. MicroRNAs direct rapid deadenylation of mRNA. Proc Natl Acad Sci U S A 103, 4034–9 (2006).
Kulshreshtha, R., Davuluri, R. V., Calin, G. A. & Ivan, M. A microRNA component of the hypoxic response. Cell Death Differ 15, 667–71 (2008).
Camps, C. et al. hsa-miR-210 Is induced by hypoxia and is an independent prognostic factor in breast cancer. Clin Cancer Res 14, 1340–8 (2008).
Rane, S. et al. Downregulation of miR-199a derepresses hypoxia-inducible factor-1alpha and Sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ Res 104, 879–86 (2009).
Du, R. et al. Hypoxia-induced down-regulation of microRNA-34a promotes EMT by targeting the Notch signaling pathway in tubular epithelial cells. PloS one 7, e30771 (2012).
Liu, R., Li, J., Teng, Z., Zhang, Z. & Xu, Y. Overexpressed microRNA-182 promotes proliferation and invasion in prostate cancer PC-3 cells by down-regulating N-myc downstream regulated gene 1 (NDRG1). PloS one 8, e68982 (2013).
Kulshreshtha, R. et al. A microRNA signature of hypoxia. Mol Cell Biol 27, 1859–67 (2007).
Huang, X. et al. Hypoxia-inducible mir-210 regulates normoxic gene expression involved in tumor initiation. Mol Cell 35, 856–67 (2009).
Krek, A. et al. Combinatorial microRNA target predictions. Nat Genet 37, 495 (2005).
Lu, T. P. et al. miRSystem: An Integrated System for Characterizing Enriched Functions and Pathways of MicroRNA Targets. PloS one 7, e42390 (2012).
Guimbellot, J. S. et al. Correlation of microRNA levels during hypoxia with predicted target mRNAs through genome-wide microarray analysis. BMC Med Genomics 2, 15 (2009).
Nygaard, V. & Hovig, E. Options available for profiling small samples: a review of sample amplification technology when combined with microarray profiling. Nucleic Acids Res 34, 996–1014 (2006).
Murphy, D. Gene expression studies using microarrays: Principles, problems and prospects. Adv Physiol Educ 26, 256–270 (2002).
Pritchard, C. C., Cheng, H. H. & Tewari, M. MicroRNA profiling: approaches and considerations. Nat Rev Genet 13, 358–69 (2012).
Raitoharju, E. et al. miR-21, miR-210, miR-34a and miR-146a/b are up-regulated in human atherosclerotic plaques in the Tampere Vascular Study. Atherosclerosis 219, 211–7 (2011).
Berezikov, E. et al. Many novel mammalian microRNA candidates identified by extensive cloning and RAKE analysis. Genome Res 16, 1289–98 (2006).
Rubio-Somoza, I., Weigel, D., Franco-Zorilla, J. M., Garcia, J. A. & Paz-Ares, J. ceRNAs: miRNA target mimic mimics. Cell 147, 1431–2 (2011).
Su, X. et al. microRNAs and ceRNAs: RNA networks in pathogenesis of cancer. Chin J Cancer Res 25, 235–9 (2013).
Acknowledgements
This research was supported by a grant from the National Science Council (Grant No. NSC 101-2320-B-002-015). We thank Melissa Stauffer for editorial assistance.
Author information
Authors and Affiliations
Contributions
L.C.L. and Y.P.S. conceived and designed the experiments. E.C.L., Y.C.C. and W.Y.H. performed the experiments. E.C.L., Y.P.S., L.L.C. and Y.C.C. analyzed the data. Y.P.S., M.H.T., E.Y.C. and L.C.L. contributed reagents, materials, and/or analysis tools. E.C.L. and L.C.L. wrote the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Luo, EC., Chang, YC., Sher, YP. et al. MicroRNA-769-3p Down-regulates NDRG1 and Enhances Apoptosis in MCF-7 Cells During Reoxygenation. Sci Rep 4, 5908 (2014). https://doi.org/10.1038/srep05908
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05908
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.