Abstract
Lysine acetylation is a reversible post-translational modification, playing an important role in cytokine signaling, transcriptional regulation and apoptosis. To fully understand acetylation mechanisms, identification of substrates and specific acetylation sites is crucial. Experimental identification is often time-consuming and expensive. Alternative bioinformatics methods are cost-effective and can be used in a high-throughput manner to generate relatively precise predictions. Here we develop a method termed as SSPKA for species-specific lysine acetylation prediction, using random forest classifiers that combine sequence-derived and functional features with two-step feature selection. Feature importance analysis indicates functional features, applied for lysine acetylation site prediction for the first time, significantly improve the predictive performance. We apply the SSPKA model to screen the entire human proteome and identify many high-confidence putative substrates that are not previously identified. The results along with the implemented Java tool, serve as useful resources to elucidate the mechanism of lysine acetylation and facilitate hypothesis-driven experimental design and validation.
Similar content being viewed by others
Introduction
Lysine acetylation is an important type of reversible post-translational modification (PTM) that takes place in the ε-amino group of lysine residues in proteins. Regulation of lysine acetylation is activated by a highly balanced enzyme system. In this system, lysine acetyltransferases (KATs) transfer the acetyl group to the ε-amino group of lysine, while lysine deacetylases (KDACs) or histone deacetylases (HDACs) remove these acetyl groups1. Around 50 years ago, lysine acetylation of nuclear histones was discovered2,3,4, followed by the successive identification of several acetylation sites in histones. Research over the past five years has shown that this reversible covalent modification is strongly related to cell regulation. During this period, more than 2,000 proteins, including kinases, transcription factors, ubiquitin ligases, structural proteins, ribosomal proteins and metabolic enzymes, have been identified as acetylated, not only in histones but also in the cytoplasm of mammalian cells5,6,7. These proteins are critical for a variety of cellular activities, ranging from the DNA damage checkpoint, cell cycle control and cytoskeleton organization to metabolism and endocytosis. Lysine acetylation is crucial for both nuclear and cytoplasmic processes8. Most major enzymes involved in the tricarboxylic acid cycle (TCA) cycle, nitrogen metabolism, fatty acid oxidation, urea cycle, glycolysis, gluconeogenesis and glycogen metabolism undergo lysine acetylation5.
Our understanding of the regulatory roles of lysine acetylation remains nebulous. Identification of acetylation sites is an essential first step towards elucidation of the mechanism underlying protein acetylation. A number of experimental methods have been accordingly developed to determine potential acetylation sites, including the radioactive chemical method9, mass spectrometry10 and chromatin immunoprecipitation (ChIP)11. However, these conventional experimental techniques are laborious, time-consuming and usually expensive12. Several high-throughput experimental methods such as mass spectrometry-based proteomics also provide a better and larger coverage of proteome-wide acetylation sites13. As an alternative approach, computational prediction methods are more efficient and applicable for large-scale high-throughput screening of novel acetylation substrates.
A variety of computational approaches have been developed to predict lysine acetylation sites12,14,15,16,17,18,19,20,21,22,23,24. For example, Basu et al.12 developed a prediction method, PredMod, that combines experimental approaches with clustering analysis to predict protein acetylation, based on the characteristics of residues surrounding acetylated lysines. Clustering of sequences in histones and nonhistones was used to represent a local amino acid sequence composition. Xu and colleagues18 developed a novel approach, Ensemble-Pail, which implemented an ensemble support vector machine (SVM) classifier with encoded features based on positional weight matrices (PWMs). A two-stage SVM-based classifier, N-Ace, proposed by Lee et al.19, was applied to identify protein acetylation sites based on features combining the physicochemical properties of proteins with accessible surface area. Suo et al.16 developed a position-specific SVM-based method, PSKAcePred, with features that included information on amino acid composition, evolutionary similarity and physicochemical properties to predict lysine acetylation sites. In their study, entropy values were used to select or exclude residues around the acetylation sites. Although significant progress has been achieved in predicting acetylation sites, the existing methods have certain drawbacks: (i) The regulation mechanism of lysine acetylation differs among species, especially between prokaryotes and eukaryotes25. Therefore, sequences or structural patterns around the acetylation sites may significantly differ in different organisms. However, the majority of existing studies disregarded the differences between species by considering all species-specific acetylation sites as generic sites to build a simplified model; (ii) Most existing models are established using machine learning techniques, such as SVM. However, not all features are equivalently important for the performance of the trained model; redundant features will reduce the performance of the model. Accordingly, feature selection is generally required for removing redundant features and improving prediction performance. However, limited studies have involved this procedure to gain insights into the relative significance and contributory effects of various features; (iii) Most earlier studies only extracted features based on the sequence environment around the acetylated lysine, but failed to consider those descriptive of the whole protein that play a decisive role in determining the fate of a protein in terms of lysine acetylation, especially for those involved in different cell processes. The next generation of computational methods thus needs to address the above drawbacks in order to generate more accurate models for the efficient identification of species-specific lysine acetylation sites.
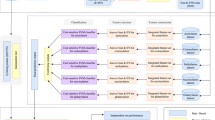
Here, we present a novel approach to predict species-specific lysine acetylation sites, based on the random forest (RF) algorithm, termed SSPKA (Species-Specific Prediction of lysine (K) Acetylation). In particular, our method incorporates various informative features, including sequence-derived features, predicted secondary structure and relevant functional features at both amino acid residue and protein levels, coupled with a two-step effective feature selection method, to assemble an optimal feature set for building the prediction model. SSPKA is benchmarked with other existing methods using both 5-fold cross-validation and independent tests. A user-friendly web server and the local Java tool of SSPKA are freely accessible at http://www.structbioinfor.org/Lab/SSPKA for the wider scientific community. A flowchart of the developed SSPKA approach is given in Supplementary Fig. S1.
Results
Analysis of sequence-level determinants of acetylation site specificity
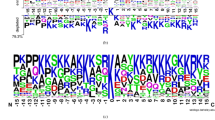
Based on the curated datasets, we analyzed the sequence surrounding the lysine acetylation sites and plotted a sequence logo (Fig. 1) for the six different species using IceLogo26, with the aim of identifying distinct patterns or conserved sequence motifs between acetylation and background sites26.
The sequence logo indicates the existence of distinct sequence patterns between the six species. In Fig. 1, the large ‘K’ represents the centered acetylation site. Apparently, a primary feature of the site specificity across all six species is the requirement that other lysine residues are located proximal to the centered acetylation site. In particular, ‘K’ is preferred at position +4 in all six species and this is more pronounced in H. sapiens and S. typhimurium, where ‘K’ tends to appear across all positions following the centered acetylation site, i.e., from +1 to +5. In addition, ‘K’ is favored at positions −5, −4 and −1 in S. cerevisiae and S. typhimurium. Other residue types in addition to ‘K’ are also observed. One example is residue ‘R’, favored at position +1 in H. sapiens and M. musculus, −1 in S. cerevisiae and −1 and −2 in R. norvegicus, respectively. Residue ‘G’, as another example, is favored at position −1 in eukaryotes such as H. sapiens, M. musculus and S. cerevisiae. On the other hand, we observe several residues that are disfavored using IceLogo. For instance, residue ‘L’ is disfavored at positions −6, −4, 3, 5 and 6 in E. coli and position −4 in R. norvegicus, respectively, whereas residue ‘D’ is not favored at position −2 in S. cerevisiae. Altogether, these results highlight the necessity and significance of addressing the task of precise lysine acetylation site recognition by developing species-specific predictors.
Two-step feature selection via random forest
As heterogeneous features are often noisy and redundant27, leading to an adverse impact on model training, such as decreasing performance, we performed feature selection to remove redundant features and assess those important for prediction performance. In particular, a two-step feature selection method was applied in our study. We have recently applied this two-step feature selection approach to address the task of protease-specific cleavage target prediction28.
In the first-step feature selection, we estimated the relative importance of each feature using the minimum-redundancy maximum-relevance (mRMR)29 approach, which ranked each input feature according to its relevance to the classification variable as well as redundancy among all features. Features that are ranked highly by mRMR generally have an appropriate balance between the maximum relevance and minimum redundancy. This step is based on the benchmark dataset. We obtained the top 100 features as the optimal candidates (OFCs) after this step. As observed from Fig. 2 and Supplementary Fig. S2, functional features had the highest ranking of importance scores. AAindex and PSSM features additionally had a relatively high importance score. The predicted secondary structures, such as those predicted by SABLE and SpineX, had relatively lower ranking values. These results indicate that the contributive features in our method are predominantly sequence-derived and functional.
mRMR results of the top 50 features (classified by feature type) for H. sapiens.
Each group of features is denoted by different colors. See Supplementary Table S1 for the full list.
The second step was a stepwise feature selection, i.e. incremental feature selection (IFS) based on the RF classifier. At each round of stepwise feature selection, the next feature from the mRMR-ranked feature list was added to the model and the resulting performance of the model calculated. To evaluate the performance, 5-fold cross-validation tests were applied, whereby the benchmark dataset was randomly divided into five subsets. Each subset in turn was used as a holdout set. For each holdout set the remaining four subsets were merged to form the training set for the RF model, while the holdout set was used as the testing set for validation of the model. The corresponding feature subset with which the RF classifier achieved the highest AUC score was considered the final optimal feature subset and used to build the prediction model. By iteratively adding informative features from the initial OFCs, the prediction performance of the model was gradually increased during this procedure (Fig. 3). The best performance in Fig. 3 is the final AUC for the corresponding species. And the feature set for that performance was the final optimal feature set. At the same time the corresponding model was the final model based on the benchmark datasets. Fig. 3 shows the whole process of second feature selection.
This two-step feature selection, which combines mRMR feature ranking and stepwise feature selection, provides a practical approach for selecting a useful subset of informative features and has been adopted by other prediction tasks28,30,31,32. Finally, we obtained a more compact informative feature subset that improved the prediction performance of RF classifiers for each species (A complete list of the final selected optimal features for each species is provided in Supplementary Table S1).
Feature importance and contribution
As mentioned previously, the optimal feature subset was selected with a two-step selection procedure. After this procedure, ten different feature types were retained in the respective optimal subsets for each species, including functional features, AAindex, functional annotation, physicochemical properties, PSSM, conservation score, Disopred, Sable and SpineX. The number of selected optimal features differed, depending on the species of interest. For example, H. sapiens had the largest subset of optimal features (a total of 58), while R. norvegicus had the smallest subset with only 8 features used to build its specific model. In addition, the number of AAindex features was greater than that of other features for all species, presumably because the proportion of initial AAindex features prior to selection is extremely high, relative to other features (7280 to 7973). In contrast, the numbers of physicochemical properties, conservation score, Disorder and Sable features were relatively small. More importantly, functional features (only 7 in total) were entirely selected for optimal subsets of H. sapiens and M. musculus, suggesting that protein functional features play significant roles in determining the prediction performance of the model. Moreover, both AAindex and PSSM features were included in the optimal feature subsets for all six species, indicating that sequence-derived features represent a critical factor in determining the predictive power of the model.
Sequence-derived features have been extensively used in model training and reported as crucial for acetylation site prediction in a number of previous studies. In our investigation, sequence-derived features, such as the PSSM profile, AAindex and physicochemical properties, were found to be indispensable for improving the prediction performance of lysine acetylation sites across all six species. To our knowledge, functional features were applied for the first time to build accurate models for predicting lysine acetylation sites. They have contributed significantly to performance improvement of our model, along with other complementary feature sets (see “Comparison with other tools” section for details).
Prediction performance was evaluated based on the AUC score. Firstly, we compared the mean values of the selected optimal features in positive and negative datasets using the statistical unpaired two-sample t-test to verify whether the two datasets were significantly different. The given P value was used to estimate statistical significance between the two datasets for a specific feature. Results are illustrated in Supplementary Fig. S3 and Fig. S4. For most selected optimal subset features, P values were lower than 0.01/n (where n is the number of tests performed, in this case, the number of features) according to the Bonferroni adjustment, indicating that the positive and negative datasets are significantly different from each other. This finding highlights the discriminative power of these features for prediction.
We continued to evaluate the importance and individual contribution of each feature type to the prediction performance of the model. For each species, all features of the feature type were taken out of the optimal feature set in turn and the remaining feature types used to build the corresponding model for predicting acetylation sites. The prediction performance of the resultant model was evaluated using the AUC measure. A feature is considered to contribute significantly to performance if the AUC score of the model in its absence decreases considerably, compared with that of the original model built using all the optimal features, as presented in Fig. 4. Taking H. sapiens as an example, there are eight types of features in its model. The AUC score for functional features was the lowest and this suggests that when removing this type of features from all eight types of features, the performance would considerably decrease, implying that functional features make a more important contribution than other features for predicting the acetylation sites of H. sapiens.
We additionally quantified the contribution of each specific feature by examining the difference between the AUC score of the model to that using only the examined feature as input and the AUC score using all other optimal features but excluding that particular feature as input. This analysis facilitated determination of the individual features that had contributed more significantly to the prediction performance of the model. Our results are presented in Supplementary Fig. S5. Taking H. sapiens as an example, the 7973rd feature was functional feature_7 (Protein-protein interaction score) (See Supplementary Table S1 for a full list of all the final features). The red bar in Supplementary Fig. S5 denotes the AUC score of the model that was trained using this particular feature only, which was over 0.6, whereas the blue bar indicates the AUC of the model that was trained using all other optimal features but excluding this particular feature, which was nearly 0.8. Altogether, these results indicate that this feature is relatively important.
Prediction performance of SSPKA based on benchmark datasets
We evaluated the prediction performance of the SSPKA models based on the final optimal features, using 5-fold cross-validation tests based on the benchmark datasets. The results are presented in Table 1 and Supplementary Table S2. RF models for all six species displayed relatively good performance with AUC scores ranging from 0.746 to 0.883. Among these species, the performance of the model for S. typhimurium was the worst with an AUC of 0.746, while that for R. norvegicus was optimal with an AUC of 0.883. The models trained using the selected optimal features for H. sapiens, M. musculus, E. coli and S. cerevisiae achieved AUC scores of 0.794, 0.788, 0.782 and 0.795, respectively.
A significant feature of this work, distinct from previous lysine acetylation site prediction studies, is the characterization and incorporation of statistically over-represented functional features by performing hypergeometric tests on the background protein datasets33. While previous studies mostly focused on the extraction of useful sequence or sequence-derived features, such as PSSM and AAindex, our current model took into consideration other important, relevant functional features that could be used in combination with sequence-derived features to improve accuracy.
Indeed, several key functional features, including KEGG, GO CC, BP, MF and PPI, contributed significantly to improvement of lysine acetylation site prediction. Supplementary Tables S3–S8 provide complete lists of the significantly enriched functional feature terms with P < 0.01/n (n is the number of tests performed and in this case, the number of terms) according to the Bonferroni adjustment) for all six species. The enrichment or depletion of these functional features reflects a specific inclination or preference of the functional requirements of different acetylated proteins, such as those for cellular compartments, related pathways and protein-protein interactions. Consequently, inclusion and encoding of the informative functional features helped improve prediction performance.
Here, we further characterized significantly enriched terms at the functional level. First, over-represented functional features were identified by hypergeometric tests. Significantly enriched functional feature terms for each species are shown in Supplementary Tables S3–S8 (only the top ten terms are listed). Taking H. sapiens as an example, there were 2233 enriched protein interaction partners in terms of PPI features. With regard to KEGG pathway features, acetylated H. sapiens proteins were enriched in metabolic pathway terms, such as “Valine, leucine and isoleucine degradation”, “Glycolysis/Gluconeogenesis” and “Pyruvate metabolism”. In addition, these proteins were enriched in certain disease pathway terms, including “Viral carcinogenesis”, “Systemic lupus erythematosus” and “Pathogenic Escherichia coli infection” (Supplementary Table S3). The results suggest that acetylated H. sapiens proteins are involved in metabolic processes related to disease pathways. Similar pathway terms were found in M. musculus (Supplementary Table S4). In terms of Biological Process (BP) terms, again, acetylated proteins were associated with transcriptional processes, including “translational termination”, “translational initiation”, “translational elongation”, “viral transcription” and “mRNA splicing, via spliceosome” (Supplementary Table S3). In terms of Molecular Function (MF) terms, acetylated proteins were enriched in functions related to nucleotide binding, such as “RNA binding”, “DNA binding”, “chromatin binding” and “nucleotide binding”.
Sequence-derived features are additionally useful for predicting lysine acetylation sites. As shown in Supplementary Table S1, the selected optimal features derived from sequences mainly include AAindex and PSSM features of residues at positions surrounding potential lysine acetylation sites. Supplementary Table S9 displays an overall statistical analysis of all the selected optimal features for each species. We did not elaborate on these features in this section, since their utility has been established in previous studies. In addition, the application of powerful feature selection techniques, such as those used in this study, allowed quantification of the relative importance and contribution of each feature type to lysine acetylation site prediction. Our findings collectively provide critical insights into the key determinants of lysine acetylation sites at both sequence and functional levels.
Comparison with other existing tools based on independent test datasets
Both 5-fold cross-validation and independent tests were conducted to compare the performance of our method with other previously published methods, including Phosida23, BRABSB24, PLMLA17, LysAcet20, ensemblePail18 and PSKAcePred16. Phosida23 and PSKAcePred16 used the binary encoding features of amino acids as input features of the model. BRABSB24 was a SVM-based human-specific lysine acetylation predictor that was developed using a novel bi-relative adapted binomial score Bayes (BRABSB) feature extraction method. PLMLA17, LysAcet20 and ensemblePail18 utilized position-weighted matrix or position-weighted amino acid properties, similar to the PSSM profile, as part of the input features to build the models. Moreover, PLMLA17 employed the secondary structure predicted by PSIPRED, while PSKAcePred16 combined solvent accessible surface area and KNN scores to train the model. Our SSPKA method incorporated not only the sequence-derived features previously shown to be useful for prediction but also the over-represented protein functional features, which made a significant contribution to the prediction power of the model. Another major difference between our method and other techniques lies in the fact that all earlier tools built the prediction model using SVM, whereas our method used RF based on decision trees to train and build the model.
We initially compared the performance of our method with other methods using the benchmark datasets based on 5-fold cross-validation tests. ROC curves of all methods are shown in Fig. 5, which describe the true positive rate as a function of false positive rate for different trade-offs between the sensitivity and specificity. Our method clearly outperformed the other five techniques for all six species. AUC scores of 0.794, 0.788, 0.746, 0.782, 0.795 and 0.883 were achieved for H. sapiens, M. musculus, S. typhimurium, E. coli, S. cerevisiae and R. norvegicus, respectively. These results indicate that our model provides a better predictive power than existing tools on benchmark datasets.
We further performed two independent tests by considering two different situations of negative sample selection to further compare the performance of our method (the first situation is to randomly select negative samples from proteins that contain positive samples and proteins in the background and the second is to select negative samples only from proteins that contain positive samples, excluding the background dataset). The corresponding ROC curves for these two situations are displayed in Supplementary Fig. S6 and S7, respectively.
In the first situation, our methods still outperformed the majority of other methods for all the species examined. One exception was the case of R. norvegicus for which SSPKA achieved an AUC of 0.696, which was slightly lower than that of LysAcet (AUC = 0.729), indicating slightly weaker performance of our method. The second situation represents a more difficult scenario for lysine acetylation site prediction, since both positive and negative samples were obtained from the same proteins, thus representing a subtler situation and confounding the prediction of positives and negatives. In the second situation, our method still outperformed others for three species, M. musculus, S. typhimurium and E. coli. However, our method performed worse than PLMLA for S. cerevisiae, LysAcet for R. norvegicus and PLMLA and BRABSB for H. sapiens, respectively. The main reason is that the contribution of functional features in this situation became marginal, due to the selection of samples only from proteins containing both negative and positive sites, reducing the predictive power of models relying on global functional features. The performance comparison results of our method with other existing methods based on different datasets are shown in Table 1 and Supplementary Table S2. To examine the statistical significance, we further performed pairwise t-test using the prediction outputs from different methods. The results showed that the performance differences between our method SSPKA and other methods were, in most cases, statistically significant (Supplementary Table S10). Finally, we would like to emphasize that the testing results on the independent tests were more reliable and hence should be given higher weights when evaluating the performance of different methods.
Cross-species performance evaluation
To examine whether each of the species-specific models could perform better for its original species than other species, we performed cross-species performance evaluation for each of the models by testing and comparing its performance on all other species. Here, the model's performance was tested by independent test datasets. As shown in Supplementary Fig. S8, the original model consistently performed the best when being applied to predict acetylation sites for the original species than other species. For example, the human-specific model achieved an AUC score of 0.756. In contrast, it achieved lower AUC scores of 0.606–0.698 for predicting acetylation sites of the other five species. This is also the case for other species-specific models, which consistently achieved the higher AUC scores when being applied to their own species than other species. In summary, these results justify the necessity and importance of developing species-specific models to improve the prediction of lysine acetylation sites.
Influence of the selection of negative datasets on the performance
As the selection of negative data might influence the final prediction performance of the model, we examine this aspect by re-training the models with randomly selected negative data and testing the resulting model's performance on the independent test dataset. This procedure was repeated five times and we generated the corresponding ROC curves in Supplementary Fig. S9. As can be seen, the red curve corresponds to the original model, which achieved an AUC value of 0.756, while all the other five blue curves correspond to the re-trained models with randomly generated negative data, with close AUC values ranging from 0.759 to 0.765. Therefore, these results show that the selection of negative datasets has little or minor influence on the performance of the model.
Proteome-wide prediction and implementation of online web server and local Java applet of SSPKA
We have implemented an online web server and local Java tool of SSPKA to facilitate high-throughput in silico prediction of lysine acetylation sites, which can be freely accessed at http://www.structbioinfor.org/Lab/SSPKA/. After demonstrating SSPKA's ability to predict lysine acetylation sites using both benchmark and independent datasets, we applied our model to screen the entire human proteome (a total of 84,919 proteins), with the aim of identifying high-confidence novel lysine acetylation sites that have been overlooked with other experimental techniques. To generate high-confidence prediction results, SSPKA models were trained using the final optimal features based on the whole training dataset. Proteins containing predicted lysine acetylation sites (adopting a prediction threshold of 0.517, which corresponds to the upper-left-most point in the ROC curve for human) were classified as high-confidence acetylated substrates. Consequently, our high-throughput in silico analysis identified 17,464 acetylated substrates with 66,255 predicted lysine acetylation sites. A complete list of the predicted acetylation substrates with detailed annotations of substrate protein IDs, acetylated lysine positions and amino acid sequences is available on the SSPKA website (http://www.structbioinfor.org/Lab/SSPKA/). The proteome-wide predictions represent a valuable resource for experimental validation of novel human acetylation substrates and generation of useful hypotheses.
The implemented online web server, local Java Applet and user instructions are also available at the website. In particular, an important advantage of the Java applet is that it provides a user-friendly interface and allows high-throughput in silico screening analysis of putative acetylation substrates (see Supplementary Fig. S10 for a screenshot of the interface and an example output of the implemented Java tool).
Analysis of proteome-wide prediction results in the sense of mimicking the distribution of acetylation sites
We further analyzed the proteome-wide prediction results to examine whether there was a correlation in the number of predicted acetylation sites to protein sequence length and whether there was a correlation in the number of predicted acetylation sites to the number of lysines in the protein sequence. We thus plotted these two distributions in Supplementary Fig. S11. We can see that the two distributions were not similar to each other, with varying correlation coefficients of less than 0.5, indicating no significant correlations between the two types of distributions.
Discussion
The performance of some competing methods (PSKAcePred and Phosida) seemed to be lower than their original published results. Here, we provide our explanations. Firstly, many machine learning algorithms work better on balanced datasets than on imbalanced datasets. In this study, the number of negative samples (i.e. non-acetylation sites) is much larger than that of positive samples (i.e. acetylation sites). A widely adopted strategy in model training is to select a balanced number of positives versus negatives with a ratio of 1:1, 1:2 and 1:3. This strategy has been used by many tools in protein post-translational modification (PTM) site prediction, including acetylation site prediction (ref. 18, 19, 20, 22); Secondly, use of different benchmarking datasets might lead to the lower performance of these methods. We selected the positive datasets with high credibility (according to PubmedMS2 or CstMS2 values from PhosphoSitePlus datasets). Proteins that were previously considered as non-acetylated might be acetylated under certain conditions. Thus, the benchmark datasets have to be updated in a timely manner to reflect the current annotation status of acetylated proteins; Thirdly, the negative datasets were selected not only from lysine residues excluding known lysine acetylation sites on the same protein, but also from other lysine residues on non-acetylated proteins (proteins not shown to be acetylated to date), while other methods only selected negative sites on the same proteins. In the latter case, it would be difficult to collect a group of proteins that can be strictly regarded as non-acetylated proteins. In such case, methods that did not consider the background proteins relative to acetylated proteins would not perform well on the benchmark datasets that not only included acetylated proteins but also incorporated vast numbers of background proteins; Lastly, some tools (e.g. BRABSB) only provided a valid model for H. sapiens. As it was trained using acetylation data only from H. sapiens, it might not work well when applied to predict the acetylation sites for other species.
In this study, we have developed a novel integrative approach, termed SSPKA, that has significantly improved the prediction performance of species-specific lysine acetylation sites across six different species, i.e., H. sapiens, M. musculus, S. typhimurium, E. coli, S. cerevisiae and R. norvegicus, by combining a variety of sequence-derived and functional features from multiple sources. SSPKA employs an efficient two-step feature selection framework to characterize the sequence and function-level features that are significant and relevant for the determination of true acetylation sites. Benchmarking experiments indicate that SSPKA is able to perform competitively, compared with existing tools. Moreover, a user-friendly webserver and local java program that suit the purposes of various biological users for the high-throughput in silico prediction of lysine acetylation substrates and sites have been made freely available. We anticipate that SSPKA will be used as a powerful tool for hypothesis-driven experimental studies on novel acetylation substrates and their biological functions.
Methods
Datasets
Annotations of lysine acetylation sites were extracted from multiple public resources. These include CPLA34 (http://cpla.biocuckoo.org/), N-ACE19 (http://N-Ace.mbc.NCTU.edu.tw/), Phosida35 (http://www.phosida.com/), ASEB15 (http://cmbi.bjmu.edu.cn/huac) and PhosphoSitePlus36 (http://www.phosphosite.org). Amongst these, CPLA is a lysine acetylation database that integrates abundant protein annotations, while PhosphoSitePlus and Phosida are comprehensive databases for post-translational modifications, including lysine acetylation data. N-ACE and ASEB are two lysine acetylation prediction tools that provide training datasets for their model15,19. To extract annotations from UniProtKB/Swiss-Prot37, all protein data were mapped to the UniProt database to retrieve the corresponding Uniprot IDs. After removing all identical sequences among the seven initial databases, we finally collected 27,075 experimentally verified acetylation sites from 10,713 protein sequences. Large amounts of protein acetylation data resulted from the rapid development of high-throughput proteomic technologies. However, in many cases, the probability scores of MS2 (mass spectrum) were substantially low, indicating a low likelihood of true acetylation sites. Accordingly, we removed acetylation sites from our datasets with PubmedMS2 or CstMS2 values smaller than 10 in PhosphoSitePlus. Acetylation sites were grouped according to the corresponding species. As homologous sequences lead to overestimation of the prediction accuracy of built models, we clustered protein sequences at the 30% identity level using CD-HIT38 software. Finally, species containing more than 40 acetylation sites were included in our positive datasets, predominantly because datasets of fewer samples are not sufficiently large to generate a valid machine learning model.
Table 2 shows the statistics of the final species-specific datasets curated. In total, our final datasets contained 1,936 proteins with 3,956 acetylation sites from six species, including both prokaryotes and eukaryotes. The datasets can be downloaded at the website http://www.structbioinfor.org/Lab/SSPKA. Negative samples were randomly selected, not only from lysine residues (excluding known lysine acetylation sites on the same protein) but also other lysines of non-acetylated proteins (proteins that were not shown to be acetylated to date), with a ratio of 1:2 of positive versus negative sites (i.e., random sampling of one positive sample accompanied with one negative sample on the same protein, as well as one negative sample from non-acetylated protein). In addition, 20% of each final dataset was randomly singled out as an independent test dataset to evaluate and compare performance between our method and other previously published protocols, while the rest was used as the training dataset to optimize the parameters, train the models and assess performance in 5-fold cross-validation tests.
Feature extraction
The extracted features were classified into four major categories: sequence-derived features, predicted secondary structures, functional annotations and functional features. In keeping with earlier studies17,19,20,22, a local sliding window approach with 13 residues centered on the lysine of interest was employed to extract the sequence-derived features of each candidate residue. In total, 7,973 features from different feature types were extracted (Table 3).
Sequence-derived features
Position-specific scoring matrix
Multiple sequence alignment containing the evolutionary information of a sequence in the form of position-specific scoring matrix (PSSM) has been shown to significantly improve prediction performance. Each element in PSSM indicates the probability of the individual residue at that specific position in the multiple sequence alignment31,39. The PSSM profile of each sequence was generated from PSI-BLAST40 and a local sliding window approach adopted to encode the matrix of a given sequence fragment surrounding potential acetylation sites. The parameters for running PSI-BLAST were set as the default E-value cutoff and three iterations used to search against the non-redundant NCBI NR database.
AAindex
We employed the AAindex41 database to extract various biochemical and physicochemical properties of amino acids, which are major features. Three sections are included in the AAindex, specifically, AAindex1 for the 20 numerical amino acid values, AAindex2 for the amino acid mutation matrix and AAindex3 for statistical protein contact potential.
Evolutionary conservation score
Evolutionary conservation is commonly employed as an important feature for prediction. A more conserved residue within a protein is indicative of higher importance for protein function. Evolutionary conservation features are extracted from the PSSM profile generated by PSI-BLAST. A lower conservation score means higher conservation at a specific position.
The conservation score is defined as:

where pi,j is the frequency of amino acid type j at position i.
Predicted secondary structure
Protein secondary structure is a useful feature to predict lysine acetylation sites. However, due to the limited number of protein substrates with available structural information, we predicted the secondary structure from amino acid sequences using SABLE42. For each residue of the query sequence, SABLE outputs three secondary structure types, H, E and C, denoting alpha-helix, beta-strand and coil, respectively. We encoded the predicted secondary structure in our model using 3-bit binary encoding43.
Predicted solvent accessibility
Solvent accessibility is another important feature for acetylation site prediction. SpineX44 was employed to predict the solvent accessibility information for each protein, which provided a quantitative score representing the extent of relative solvent accessibility of a residue from fully buried to fully exposed.
Disordered region
A protein disordered region lacks a well-defined tertiary structure and is either fully or partially unfolded. Earlier researchers suggested that disordered regions were ‘useless’. However, over recent years, disordered regions have been shown to be involved in several important biological functions45. For instance, many phosphorylation sites are located in disordered, rather than non-disordered regions46,47. As such, disorder information contributes to phosphorylation site prediction48. Here, we extracted predicted disorder information calculated using DISOPRED2, which was also added as the input to our models49.
Functional annotation
Functional annotation of a protein in UniProt can be found in the “FT” line of the annotation37. Several different types of functional annotations were used as features, including DOMAIN, NP_BIND, DISULFID, MOD_RES, CARBOHYD, ACT_SITE, VARIANT, METAL and BINDING, which represent domain, nucleotide binding, disulfide bond, post-translational modified residue (acetylation removed), glycosylation, active site, natural variant, metal ion binding site and binding site, respectively. Within the sliding window, the amino acid is encoded as “1” if that site has the annotation of a specific function. On the other hand, an amino acid without the functional annotation is encoded as “0”. In total, there are 13 (window size) × 9 (annotation types) = 117-dimensional encoded features for this type.
Functional features
Inclusion of functional features of a whole protein and assessment of their contribution to performance is a crucial aspect of this work. To address this, we included protein functional features from the Gene Ontology database50 and other biological databases, including Biological Process features (BP), Cellular Component features (CC), Molecular Function features (MF), functional domain features from InterPro51, pathway features from KEGG52, functional domain features from Pfam53 and protein-protein interactions from PPI54.
Random forest classifier
We employed a machine learning approach- random forest to generate models for lysine acetylation site prediction. RF is an ensemble learning method based on the classification tree55, which “votes” for one of the two classes, either positive (acetylation sites) or negative (non-acetylation sites). The experimentally verified acetylation sites in the datasets were labeled ‘1’ and all other lysine residues labeled ‘−1’. As described above, the physicochemical properties of a lysine residue of interest were represented by a series of input feature vectors and encoded into RF classifiers to identify whether or not the residue was an acetylation site. RF is considered as one of the most accurate machine learning algorithms available that produce highly accurate classification results. In addition, it can be used to select more important variables and efficiently handle large datasets56. Owing to these advantages, RF has been frequently used to address classification problems in bioinformatics, such as prediction of DNA-binding sites, RNA-binding sites, residue-residue contacts, functional sites, disease-causing non-synonymous SNPs and metal-binding sites30,31,57,58,59,60.
Performance evaluation
We used six measures, Matthews Correlation Coefficient (MCC), Accuracy (ACC), Sensitivity (SEN), Specificity (SPE), Precision (PRE) and Area Under the Receiver-Operating Characteristic Curve (AUC), to evaluate performance. For the six species-specific datasets, an under-sampling strategy with a 1:2 ratio between positive and negative samples was adopted. It is not reasonable to assess performance using Accuracy (i.e., the proportion of true positives and true negatives on the dataset) based on an imbalanced dataset. AUC is the area under the receiver-operating characteristic (ROC) curve, presented as a plot of true positive rate (TPR i.e. SEN) against false positive rate (FPR). The AUC value of a ROC curve summarizes the overall performance of a corresponding model or method. An AUC value of 1.0 indicates perfect prediction, while 0.5 signifies complete random prediction. We consider AUC a more appropriate measure for comprehensively evaluating the overall quality of the RF-based classifier performance.
MCC is defined as:

ACC is defined as:

SEN is defined as:

SPE is defined as:

PRE is defined as:

FPR is defined as:

where TP, TN, FP and FN represent the numbers of true positives, true negatives, false positives and false negatives, respectively.
References
Sadoul, K., Wang, J., Diagouraga, B. & Khochbin, S. The tale of protein lysine acetylation in the cytoplasm. J. Biomed. Biotechnol. 2011, 970382 (2011).
Allfrey, V. G., Pogo, B. G., Littau, V. C., Gershey, E. L. & Mirsky, A. E. Histone acetylation in insect chromosomes. Science 159, 314–316 (1968).
Allfrey, V. G., Faulkner, R. & Mirsky, A. E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl. Acad. Sci. USA 51, 786–794 (1964).
Phillips, D. M. The presence of acetyl groups of histones. Biochem. J. 87, 258–263 (1963).
Zhao, S. et al. Regulation of cellular metabolism by protein lysine acetylation. Science 327, 1000–1004 (2010).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Kim, S. C. et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell 23, 607–618 (2006).
Xiong, Y. & Guan, K. L. Mechanistic insights into the regulation of metabolic enzymes by acetylation. J. Cell Biol. 198, 155–164 (2012).
Welsch, D. J. & Nelsestuen, G. L. Amino-terminal alanine functions in a calcium-specific process essential for membrane binding by prothrombin fragment 1. Biochemistry 27, 4939–4945 (1988).
Medzihradszky, K. F. Peptide sequence analysis. Methods Enzymol. 402, 209–244 (2005).
Umlauf, D., Goto, Y. & Feil, R. Site-specific analysis of histone methylation and acetylation. Methods Mol. Biol. 287, 99–120 (2004).
Basu, A. et al. Proteome-wide prediction of acetylation substrates. Proc. Natl. Acad. Sci. USA 106, 13785–13790 (2009).
Choudhary, C. & Mann, M. Decoding signalling networks by mass spectrometry-based proteomics. Nat. Rev. Mol. Cell Biol. 11, 427–439 (2010).
Suo, S. B. et al. Proteome-wide analysis of amino acid variations that influence protein lysine acetylation. J. Proteome Res. 12, 949–958 (2013).
Wang, L., Du, Y., Lu, M. & Li, T. ASEB: a web server for KAT-specific acetylation site prediction. Nucleic Acids Res. 40, W376–379 (2012).
Suo, S. B. et al. Position-specific analysis and prediction for protein lysine acetylation based on multiple features. PLoS One 7, e49108 (2012).
Shi, S. P. et al. PLMLA: prediction of lysine methylation and lysine acetylation by combining multiple features. Mol. Biosyst. 8, 1520–1527 (2012).
Xu, Y., Wang, X. B., Ding, J., Wu, L. Y. & Deng, N. Y. Lysine acetylation sites prediction using an ensemble of support vector machine classifiers. J. Theor. Biol. 264, 130–135 (2010).
Lee, T. Y. et al. N-Ace: using solvent accessibility and physicochemical properties to identify protein N-acetylation sites. J. Comput. Chem. 31, 2759–2771 (2010).
Li, S. et al. Improved prediction of lysine acetylation by support vector machines. Protein Peptide Lett. 16, 977–983 (2009).
Cai, Y. D. & Lu, L. Predicting N-terminal acetylation based on feature selection method. Biochem. Biophys. Res. Commun. 372, 862–865 (2008).
Li, A., Xue, Y., Jin, C., Wang, M. & Yao, X. Prediction of Nepsilon-acetylation on internal lysines implemented in Bayesian Discriminant Method. Biochem. Biophys. Res. Commun. 350, 818–824 (2006).
Gnad, F., Ren, S., Choudhary, C., Cox, J. & Mann, M. Predicting post-translational lysine acetylation using support vector machines. Bioinformatics 26, 1666–1668 (2010).
Shao, J. et al. Systematic analysis of human lysine acetylation proteins and accurate prediction of human lysine acetylation through bi-relative adapted binomial score Bayes feature representation. Mol. Biosyst. 8, 2964–2973 (2012).
Jones, J. D. & O'Connor, C. D. Protein acetylation in prokaryotes. Proteomics 11, 3012–3022 (2011).
Colaert, N., Helsens, K., Martens, L., Vandekerckhove, J. & Gevaert, K. Improved visualization of protein consensus sequences by iceLogo. Nat. Methods 6, 786–787 (2009).
Saeys, Y., Inza, I. & Larranaga, P. A review of feature selection techniques in bioinformatics. Bioinformatics 23, 2507–2517 (2007).
Wang, M. et al. Cascleave 2.0, a new approach for predicting caspase and granzyme cleavage targets. Bioinformatics 30, 71–80 (2014).
Peng, H., Long, F. & Ding, C. Feature selection based on mutual information: criteria of max-dependency, max-relevance and min-redundancy. IEEE Trans. Pattern Anal. Mach. Intell. 27, 1226–1238 (2005).
Zheng, C. et al. An integrative computational framework based on a two-step random forest algorithm improves prediction of zinc-binding sites in proteins. PLoS One 7, e49716 (2012).
Wang, M. et al. FunSAV: predicting the functional effect of single amino acid variants using a two-stage random forest model. PLoS One 7, e43847 (2012).
Song, J. et al. PROSPER: an integrated feature-based tool for predicting protease substrate cleavage sites. PLoS One 7, e50300 (2012).
Li, T., Du, P. & Xu, N. Identifying human kinase-specific protein phosphorylation sites by integrating heterogeneous information from various sources. PLoS One 5, e15411 (2010).
Liu, Z. et al. CPLA 1.0: an integrated database of protein lysine acetylation. Nucleic Acids Res. 39, D1029–1034 (2011).
Gnad, F., Gunawardena, J. & Mann, M. PHOSIDA 2011: the posttranslational modification database. Nucleic Acids Res. 39, D253–260 (2011).
Hornbeck, P. V. et al. PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res. 40, D261–270 (2012).
The Uniprot Consortium. Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucleic Acids Res. 40, D71–75 (2012).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Song, J., Tan, H., Wang, M., Webb, G. I. & Akutsu, T. TANGLE: two-level support vector regression approach for protein backbone torsion angle prediction from primary sequences. PLoS One 7, e30361 (2012).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Kawashima, S. et al. AAindex: amino acid index database, progress report 2008. Nucleic Acids Res. 36, D202–205 (2008).
Wagner, M., Adamczak, R., Porollo, A. & Meller, J. Linear regression models for solvent accessibility prediction in proteins. J. Comput. Biol. 12, 355–369 (2005).
Song, J., Burrage, K., Yuan, Z. & Huber, T. Prediction of cis/trans isomerization in proteins using PSI-BLAST profiles and secondary structure information. BMC Bioinformatics 7, 124 (2006).
Faraggi, E., Zhang, T., Yang, Y., Kurgan, L. & Zhou, Y. SPINE X: improving protein secondary structure prediction by multistep learning coupled with prediction of solvent accessible surface area and backbone torsion angles. J. Comput. Chem. 33, 259–267 (2012).
Dunker, A. K. & Obradovic, Z. The protein trinity--linking function and disorder. Nat. Biotechnol. 19, 805–806 (2001).
Dunker, A. K. et al. The unfoldomics decade: an update on intrinsically disordered proteins. BMC Genomics 9 Suppl 2, S1 (2008).
Iakoucheva, L. M. et al. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res. 32, 1037–1049 (2004).
Gnad, F. et al. PHOSIDA (phosphorylation site database): management, structural and evolutionary investigation and prediction of phosphosites. Genome Biol. 8, R250 (2007).
Ward, J. J., McGuffin, L. J., Bryson, K., Buxton, B. F. & Jones, D. T. The DISOPRED server for the prediction of protein disorder. Bioinformatics 20, 2138–2139 (2004).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Hunter, S. et al. InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res. 40, D306–312 (2012).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–301 (2012).
Jensen, L. J. et al. STRING 8--a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 37, D412–416 (2009).
Liaw, A. & Wiener, M. Classification and regression by random forest. R news 2, 18–22 (2002).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Segura, J., Jones, P. F. & Fernandez-Fuentes, N. A holistic in silico approach to predict functional sites in protein structures. Bioinformatics 28, 1845–1850 (2012).
Wang, X. F. et al. Predicting residue-residue contacts and helix-helix interactions in transmembrane proteins using an integrative feature-based random forest approach. PLoS One 6, e26767 (2011).
Liu, Z. P., Wu, L. Y., Wang, Y., Zhang, X. S. & Chen, L. Prediction of protein-RNA binding sites by a random forest method with combined features. Bioinformatics 26, 1616–1622 (2010).
Wu, J. et al. Prediction of DNA-binding residues in proteins from amino acid sequences using a random forest model with a hybrid feature. Bioinformatics 25, 30–35 (2009).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (61202167, 61303169 and 11250110508), Hundred Talents Program of the Chinese Academy of Sciences (CAS), Knowledge Innovation Program of CAS (KSCX2-EW-G-8), National Health and Medical Research Council of Australia (NHMRC) (490989) and Australian Research Council (ARC) (LP110200333). GIW is a recipient of the ARC Discovery Outstanding Researcher Ward (DORA). JS is an NHMRC Peter Doherty Fellow and a recipient of the Hundred Talents Program of CAS.
Author information
Authors and Affiliations
Contributions
Y.L. collected the data, performed the experiments, wrote the programs and analyzed the data. M.W., H.W. and H.T. analyzed the data and contributed to the writing of the programs. Z.Z., G.I.W. and J.S. designed the algorithm and analyzed the data. Y.L. and J.S. wrote the manuscript. All authors discussed and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Li, Y., Wang, M., Wang, H. et al. Accurate in silico identification of species-specific acetylation sites by integrating protein sequence-derived and functional features. Sci Rep 4, 5765 (2014). https://doi.org/10.1038/srep05765
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05765
This article is cited by
-
IRC-Fuse: improved and robust prediction of redox-sensitive cysteine by fusing of multiple feature representations
Journal of Computer-Aided Molecular Design (2021)
-
SulSite-GTB: identification of protein S-sulfenylation sites by fusing multiple feature information and gradient tree boosting
Neural Computing and Applications (2020)
-
Analysis and prediction of human acetylation using a cascade classifier based on support vector machine
BMC Bioinformatics (2019)
-
Computational identification of microbial phosphorylation sites by the enhanced characteristics of sequence information
Scientific Reports (2019)
-
PhosContext2vec: a distributed representation of residue-level sequence contexts and its application to general and kinase-specific phosphorylation site prediction
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.