Abstract
Reaching a comprehensive understanding of how nature solves the problem of degrading recalcitrant biomass may eventually allow development of more efficient biorefining processes. Here we interpret genomic and proteomic information generated from a cellulolytic microbial consortium (termed F1RT) enriched from soil. Analyses of reconstructed bacterial draft genomes from all seven uncultured phylotypes in F1RT indicate that its constituent microbes cooperate in both cellulose-degrading and other important metabolic processes. Support for cellulolytic inter-species cooperation came from the discovery of F1RT microbes that encode and express complimentary enzymatic inventories that include both extracellular cellulosomes and secreted free-enzyme systems. Metabolic reconstruction of the seven F1RT phylotypes predicted a wider genomic rationale as to how this particular community functions as well as possible reasons as to why biomass conversion in nature relies on a structured and cooperative microbial community.
Similar content being viewed by others
Introduction
Utilization of traditional fossil fuels is commonly associated with global environmental issues and supply concerns, emphasizing the need for sustainable biofuel alternatives. Cellulose, the Earth's most abundant renewable biomass substrate1 is gaining increased attention2,3, however its industrial-scale conversion to biofuel is impeded by structural recalcitrance and a paucity of available biocatalysts. Current research on enzyme systems for cellulose conversion focuses on the abilities of free-enzyme systems4 and/or cellulosomes5,6,7 within individual microorganisms. However, it is likely that cellulose turnover in nature benefits from the concerted actions within a microbial consortium or “microbiome”8,9,10. This has been previously illustrated in the laboratory for a defined mixed culture constructed from five isolated strains (designated SF356), which collectively degraded cellulose whilst maintaining stable population structure and improved function over time11,12.
As cellulose-rich plant biomass is constantly recycled in soil environments, a plethora of efficient cellulose-degrading machineries should perceivably exist in its resident microbiomes, potentially including hitherto concealed degradation mechanisms. Recently, metagenomic sequencing has enabled in-depth description of complex microbiomes specialized in plant biomass conversion including digestive8,10 and free-living ecosystems9. What is more, the currently attainable depth of such sequencing efforts combined with the progression of taxonomic “binning” software has facilitated the reconstruction of genomes representative of the as-yet uncultured microbes that reside within a microbiome8,13. Based on these notions, we have utilized a compilation of culture-independent “omics” techniques to analyze the genomic potential and functional synergies within a proficient low-complexity cellulose-degrading microbial consortium enriched from a naturally occurring soil sample.
Results
The discovery and structural depiction of the F1RT consortium
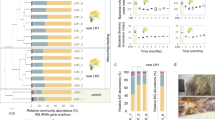
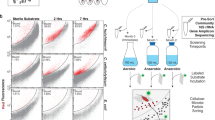
Five humified soil samples were collected from different locations in Shaoguan, China and their cellulolytic abilities were analyzed using degradation of filter paper as an indication (Supplementary Fig. 1). To circumvent the high levels of species complexity inherent to soil microbiomes, an enrichment strategy was applied to the best performing samples using serial-dilution, plating and repetitive subculturing on selective aerobically prepared media. Despite these extensive efforts, pure cultures were not obtained. Rather, we ended up with a consortium (hereafter termed F1RT) that initially appeared as one colony on a selective plate (Supplementary Fig. 2). F1RT was functionally stable during dozens of subsequent passages and was capable of completely degrading filter paper (Supplementary Fig. 3). 16S rRNA gene sequencing analysis revealed that it was a mixed consortium containing seven individual phylotypes that were affiliated at varying identity values (92.4–99.8 ID%) to both aerobic and anaerobic species (rrs clusters: RC1–7, Table 1). Given the phylogenetic and putative phenotypic variation in F1RT a metagenomic and genome binning approach was employed to recover near-complete genomes so that a holistic interpretation could be made on the metabolic capabilities for all seven phylotypes. Using Illumina technology we recovered 12.8 Gb of F1RT metagenomic reads, with rarefaction curves (Supplementary Fig. 4) suggesting that the sequencing depth was sufficient for de novo assembly. Approximately 90.1% of high-quality reads (10.23 Gb) were assembled into a total of 3,930 scaffolds (length ≥500 bp, average length 6.7 kb), totalling 26.51 Mb (Supplementary Fig. 5). Using GeneMark 27,532 open reading frames (ORFs) with length ≥100 bp were predicted.
To enable genomic analysis of F1RT phylotypes a combinatory binning approach based on read-coverage of conserved single-copy genes14, Markov model-based tetranucleotide frequencies15 and a naïve Bayesian classifier16 was used to reconstruct seven uncultured microbial genomes from the assembled metagenome scaffolds (FC1–7; Table 1, Supplementary Note 1, Supplementary Table 3). We applied the same workflow as described previously8 to assess the coverage of binned genomes when compared to complete genomes available in GenBank (Supplementary Note 2). Completeness estimations of the binned genomes indicated that all seven were near-complete draft genomes with low redundancy of conserved single-copy genes (i.e. high “authenticity”), although for FC7 the data indicated slightly lower levels of completeness and authenticity (Supplementary Fig. 9; Supplementary Table 4). Genome assembly coverage data indicated that FC1, FC2 and FC3 are the most abundant phylotypes in F1RT (Table 1, (Supplementary Fig. 7).
Draft genome bins FC1-7 were found to lack 16S rRNA genes, presumably due to the repetitive nature of 16S rRNA operons, which often leads to assembly issues in bacterial genomes. TETRA, ANI and AAI measurements17 were used to identify the taxonomy of the FC genomes and failed to designate species affiliations, implying that all phylotypes in this community have not been sequenced to date (Table 1, Supplementary Table 8). Therefore we compared the different taxonomic affiliations collected for both the FC and the RC datasets to enable putative associations between the F1RT draft genome bins (FC) and operational taxonomic units (OTUs; RC) (Table 1, Supplementary Note 3). Both FC and RC comparisons identified that all seven clusters were affiliated to the Firmicutes phylum with five clusters affiliated to the order Clostridiales and two clusters affiliated to the Bacillales (Table 1). Additional phylogenetic analysis of Clostridiales-affiliated 16S rRNA sequences from RC phylotypes, the closest RC relatives and the closest FC relatives (determined by TETRA matches) demonstrated distinct phylogenetic clustering, which was used to identify “direct” RC-FC associations (Supplementary Note 3). FC2 exhibited closest genome similarity to Clostridium thermocellum (TETRA coefficient = 0.95) and was putatively affiliated with Clostridium straminisolvens (RC3: 16S rRNA identity 99.6%), an anaerobic cellulolytic soil bacterium. FC1 also exhibited closest genome similarity to C. thermocellum (TETRA coefficient = 0.92), but was putatively affiliated with Clostridium thermosuccinogenes (RC2: 16S rRNA identity 99.1%; Table 1) as well as Clostridium sp. FG4 (16S rRNA identity 99.3%). Interestingly, the latter species is an oligosaccharide fermenting anaerobe that was found to co-exist with C. straminisolvens in SF356 enrichments12. FC5 was putatively affiliated with Clostridium sporogenes (RC1). The low TETRA coefficients displayed by FC6 and incoherent clustering of FC7 prevented direct associations with RC clusters and their affiliate species. By the process of elimination FC6 and FC7 were “non-directly” associated to either one of the remaining Clostridiales RC clusters, which were affiliated to Cellulosilyticum lentocellum (RC5: 16S rRNA identity 92.4%) and Tissierella praeacuta (RC6: 16S rRNA identity 94.1%) (Table 1, Supplementary Note 3). Similarly, the two F1RT phylotypes that were putatively affiliated with the order Bacillales (FC3 and FC4) were non-directly associated with either one of the RC clusters that were affiliated to the aerobe Brevibacillus borstelensis of the same order (RC4 and RC7, Table 1). For non-direct associations the best available clustering was used to speculate on a predictive association (Table 1).
Omics-based interpretation of F1RT's cellulolytic capacity
The cellulolytic capabilities exhibited by the F1RT consortium and the affiliations of some of its constituent phylotypes to cellulolytic clostridia (Table 1) directed our initial attention towards F1RT's potential to encode and utilize cellulosome structures. Cellulosomes, first described in C. thermocellum5,7, are elaborate multi-enzyme machines that are assembled by protein-protein interactions between type I cohesin modules in a scaffoldin and type I dockerin modules attached to each enzyme-subunit. Interactions between type II cohesins and type II dockerins, encoded within particular scaffoldins and/or cell-surface proteins, may link cellulosomal structures to cell surfaces and/or to other cellulosomes (polycellulosomes)6. We detected 74 putative dockerins that consisted of 69 type I and 5 type II domains (domain architecture: Supplementary Tables 9–11, domain alignment: Supplementary Fig. 11). A total of 27 putative cohesin domains arranged into 13 scaffoldin-encoding ORFs (domain architecture: Fig. 1A, Supplementary Table 12, domain alignment: Supplementary Fig. 12). All cellulosomal ORFs were affiliated to FC2, whose genome was most similar to the genome of the known cellulosomal bacteria C. thermocellum. Notably, the TETRA coefficients for genome comparisons were 0.95, i.e. well below the cut-off value (0.9917) that would indicate species affiliation (Table 1). Given the incomplete status of the FC2 genome, a detailed prediction of the structural assemblies of F1RT cellulosomes was not attempted. Regardless, the large modular variability observed in FC2 scaffoldins (Fig. 1A) and the identification of both type I and type II cohesin/dockerin modules, suggests a large number of diverse (poly)cellulosome assemblies.
Schematic representation of cellulosomal subunits identified in the F1RT consortium.
(A) Dockerin catalog and modular architecture of the various scaffoldins identified in the F1RT metagenome. Putative scaffoldins were affiliated to the draft genome bin FC2. Note that this is a partial genome bins, meaning that some proteins are incomplete and that we only have access to a subset of cellulosomal proteins. Domain architecture for all cellulosomal GH domains is summarized in Supplementary Table 10. Acronyms: GH, glycoside hydrolase; CBM, carbohydrate-binding module; Cu, copper; aa, amino acid. (B) Hypothetical example of a polycellulosome assembly that was predicted via metaproteomic analysis of soluble protein-protein complexes produced by the F1RT consortium (see Supplementary Fig. 14, Supplementary Table 21, “Band 10”). The shaded structure (corresponding to GL0023450) was not detected in the proteomics experiment. However, comparisons of ORF GL0026964 (lacking the 3′end), with the vanguard CipA scaffoldin of C.themocellum would indicate that it is likely that this protein encodes the necessary type-II dockerin that is found in GL0023450 (lacking the 5′end); see text.
To unearth the carbohydrate-active enzymes encoded within F1RT, we used the CAZy database18 as reference to identify 348 putative glycoside hydrolase (GH) domains (GH summary: Supplementary Table 13) and 227 putative carbohydrate-binding modules (CBMs) (Supplementary Table 9). A total of 50 GH domains were identified within 46 individual ORFs that also encoded a type-1 dockerin domain and these GHs were thus considered cellulosomal; 27 of these GH domains were predicted to be cellulases (GH5, GH9, GH44, GH48, GH74; Fig. 1A). Additional cellulosomal hydrolytic (GH) and binding (CBMs) abilities seem directed towards xylans (GH10, GH43; CBM6, CBM22), mannans (GH26; CBM35), xyloglucans (GH16, GH74) and beta-glucans (GH16, GH55, GH81) (Fig. 1A). Approximately 86% of all GH domains were considered non-cellulosomal, however in the high coverage cellulosome-containing FC2 77% of all predicted cellulases (including endoglucanases and exoglucanases) were predicted to be cellulosomal, in accordance with the expected importance of cellulosomes in cellulose degradation19. Several phylotypes contained large GH profiles, including cellulases but devoid of any cellulosomal domains, which suggested their saccharolytic capabilities extended via employment of a free-enzyme system (Supplementary Table 13). FC6 and the high-coverage FC1 phylotype encoded comprehensive (53 and 123 GH domains, respectively) yet slightly variant glycoside hydrolase catalogues that included inferred activity against hemicellulose (e.g. GH8, GH26, GH43), suggesting that F1RT has the capability to degrade heterogeneous and complex hemicellulosic substrates (Fig. 2, Supplementary Table 13). FC1 was also found to encode putative endoglucanases (GH5 and GH9) inferring a possible role in cellulose conversion as well. FC3, FC4 and FC5 were predicted to be incapable of degrading cellulose and hemicellulose, but were found to encode oligosaccharide-degrading GH domains such as GH3 beta-glucosidases and GH96 cellobiose phosphorylases.
Hypothetical network model summarizing putative inter-species cooperation in the F1RT enrichment.
Pathway analysis (Supplementary Table 24) and structural comparisons with the C. straminisolvens-containing SF356 enrichment12 were used to generate predictions regarding saccharification and downstream metabolism in the F1RT consortium. Phylotypes FC3 and FC4 were predicted to be (facultative) aerobes and are thus likely to consume oxygen by utilizing substrates contained in PCS media. Crucially this provides and subsequently maintains, the anoxic environment (shaded grey) that is required for FC1-2 and FC6 to degrade cellulose and various hemicellulose substrates, through the use of different glycoside hydrolases (GH) arranged in cellulosomes and/or free-enzyme systems. Metabolic reconstruction suggests that all anaerobic phylotypes are capable of fermenting the various sugars produced via cellulose and hemicellulose hydrolysis. The major fermentation end-products were predicted to be acetate, ethanol and succinate (bold text) as well as formate and lactate. FC3 and FC4 are able to utilize ethanol and acetate (glyoxylate cycle) as well as succinate (succinate to cytochrome bo oxidase electron transfer). Selected GH families are indicated for FC1-2 and FC6; for complete GH catalogs refer to Supplementary Table 13. For a detailed pathway analysis of FC1-7 refer to Supplementary Table 24.
Metaproteomic analysis of the F1RT consortium was performed to illustrate which phylotypes were actively producing cellulolytic machineries and other polysaccharide degrading enzymes. This approach confirmed a plethora of both extracellular and cell-wall associated cellulosomal proteins and free-enzyme GHs (Supplementary Note 4). Moreover, it showed that different consortium members contribute in different ways to the enzymatic potential of the consortium (Fig. 2, Supplementary Tables S15–18). Free-enzyme GHs were detected from all seven draft genomes however non-cellulosomal GHs affiliated to FC1 and FC2 were most abundant and included putative cellulases (GH48, GH9, GH5) and hemicellulases (GH10, GH11, GH29, GH51). (Supplementary Tables S17–18). Cellulosomal proteins were also detected in the extracellular fraction including a multiple-type II cohesin-encoding scaffoldin (GL0004812, Fig. 1A) that does not encode any cell-anchoring domains (i.e. SLH20), suggesting its involvement in cell-free cellulosomes. The domain structure of GL0004812 (seven type-II cohesins) is comparable to the Cthe_0736 scaffoldin from C. thermocellum, whose functional role is relatively unknown although it has been suggested to form cell-free extracellular polycellulosome complexes6. NanoLC-MS/MS analysis of extracellular cellulosome complexes that were isolated using native PAGE gels, frequently co-detected GL0004812 with a multiple-type I scaffoldin (GL0026964) that has resemblance to the vanguard CipA scaffoldin from C. thermocellum (Supplementary Fig. 14, Supplementary Tables 21–22). While the incomplete GL0026964 seems to lack the type-II dockerin required to interact with GL0004812, we predict it is likely to encode the necessary domain on the missing 3′ region of the ORF, similar or perhaps identical to the domain encoded on the also incomplete GL0023450 (Fig. 1A). This prediction is supported by the observation that GL0026964 and GL0023450 align with the N- and C-terminal regions, respectively, on the CipA scaffoldin in C. thermocellum (acc: YP_005687158). Collating this information we hypothesize that GL0004812 and GL0026964 could conceivably be interconnected, forming a polycellulosome assembly that is not attached to the cell wall (Fig. 1B). Enzymatic subunits detected within the native PAGE gel bands were dominated by cellulosomal cellulases belonging to families GH48 and GH9, whereas cellulosomal GH5, GH8, GH10, GH18, GH26, GH44 and GH74 domains were also detected (Supplementary Tables 21–22).
Metabolic reconstruction of F1RT predicts a genomic-derived rational for inter-species cooperation
The most distinctive feature of F1RT upon comparison with phylogenetically related cellulolytic bacteria C. straminisolvens, C. thermocellum and C. lentocellum was its ability to degrade cellulose in aerobically-prepared media, whereas these known individual cellulolytic clostridia require strict anaerobic media for growth and cellulose conversion. In their pioneering work on cellulose-degrading communities (enrichment SF356), Kato et al.12 demonstrated that the cellulolytic efficiency of C. straminisolvens is stimulated in the presence of other aerobic and non-cellulolytic anaerobic phylotypes11,12. Despite being enriched from independently collected samples, SF356 and F1RT show structural similarities, including FC2's affiliation to C. straminisolvens, FC1's affiliation to Clostridium sp. FG4 (99.3% identity) and FC3 and FC4's affiliation to the genus Brevibacillus.
Our F1RT metagenomic dataset allowed for a broad genomic analysis of these intriguing observations on bacterial co-existence (summarized in Fig. 2). Genome-scale reconstruction of metabolic pathways revealed contrasting functional roles for the F1RT phylotypes and suggested several syntrophic mechanisms. (Fig. 2, Supplementary Table 24). Similar to SF356, analysis of oxygen requirements distinguished Brevibacillus-affiliated phylotypes FC3 and FC4 as (facultative) aerobes that are likely to consume oxygen while utilizing nutrients present or generated in the F1RT enrichment medium (Fig. 2, Supplementary Table 24). Despite the reputation of C. straminisolvens (similar to FC2) being the sole polysaccharide-degrading microbe in SF356, our data suggested that three of the five anaerobic phylotypes in F1RT contribute to hydrolysis of plant polysaccharides by utilizing both cellulosome (e.g. FC2) and free-enzyme systems (e.g. FC1 and FC6) (Fig. 2). In particular, GH5 endoglucanases affiliated to FC1 were detected in F1RT metaproteomic analysis (Supplementary Table 17). This suggests a greater saccharolytic contribution by FC1 than its corresponding SF356 counterpart Clostridium sp. FG4, which was only deemed capable of fermenting oligosaccharides12.
The identification of putative transporters (Supplementary Table 25), beta-glucosidases (i.e. GH1, GH3; Supplementary Table 13) and cellobiose phosphorylases (GH94; Supplementary Table 13) indicates that all F1RT phylotypes are capable of “down-stream” metabolic utilization of liberated cellodextrins. Since glucose and cellobiose are known to inhibit cellulases and cellulosome formation21, their utilization by non-cellulolytic phylotypes such as FC5 and FC6 is likely to help maintain F1RT's capability to continuously deconstruct cellulose (Fig. 2). Analysis of the reconstructed pathways further showed that the anaerobic phylotypes differ with respect to which sugars they are capable of fermenting to end-products such as ethanol, acetate and succinate (Fig. 2, Supplementary Table 24). The Brevibacillus-affiliated phylotypes FC3 and FC4 seem capable of converting ethanol and acetate to acetyl-CoA, the major input to the glyoxylate cycle commonly utilized by facultative anaerobic and aerobic bacteria (Supplementary Tables 24). In addition to providing a nutritional rationale for microbial co-existence, the predicted consumption of acids acetate and succinate (Fig. 2, Supplementary Table 24) by these non-cellulolytic bacteria is expected to alleviate the negative pH affects of acid accumulation which are known to adversely affect cellulose degradation by anaerobic clostridia22.
Discussion
Our cumulative understanding regarding microbial plant biomass degradation is built from laboratory performance-based assessment of singular cellulolytic isolates and their enzymes. However this does not reflect the true capabilities of naturally occurring cellulolytic ecosystems, which utilize syntropic interactions within a consortium to envelop both polysaccharide hydrolysis and downstream metabolism. Here we have utilized the recent progression in DNA sequencing and computational biology technologies to describe both the genomes and proteomes for all the as-yet uncultured members within a cellulolytic consortium. Predictive metabolic “blue-print” analysis of all F1RT phylotypes presented a genomic rationale as to how the F1RT consortium functions, why it resisted initial aerobic culturing efforts and why it displayed structural similarity to the independently generated SF356 enrichment. In addition, our “omics” analyses allowed for the first time, the reconstruction of cellulosomes from uncultured bacteria and indications for the presence of extracellular (poly)cellulosomes, a poorly understood facet of the cellulosome “paradigm”. Moreover, our analysis demonstrated the putative operation of both cellulosome and free-enzyme systems by different phylotypes within a consortium. Indeed, recent in vitro studies indicate synergism between recombinant cellulosomes and free-enzymes, due to the two enzyme systems using different mechanisms for interacting with microcrystalline surfaces19,23. Our findings suggest that such a multi-system strategy for cellulose-degradation conceivably exists within naturally occurring microbial consortia. The functional capabilities of F1RT combined with the observed structural similarities with the previously described SF356 consortium would suggest that F1RT-like populations play crucial roles in natural cellulose-degrading soil ecosystems. Verification of this hypothesis would create significant insights towards understanding the impact of such populations on a broader scale.
Methods
Soil samples were collected from Shaoguan, China and their cellulose-degrading capabilities tested and compared. An optimal enrichment was domesticated and simplified utilizing filter paper-containing selective medium and approximately 64 continuous subcultures were performed prior to identification of a specific enrichment that demonstrated efficiency in cellulose-hydrolysis (termed F1RT). F1RT's status as a consortium with functional stability eluded further simplification via culture-based methods. Therefore we applied a metagenomic approach to decode and reconstruct the genetic components of F1RT, as well as observe the community-wide cellulosome repertoire in silico. In addition, metaproteomics was employed to analyze protein-protein complexes for validation of predictive annotations regarding F1RT's cellulolytic capabilities. Finally, a more extensive metabolic pathway analysis was performed to predict syntrophic relationships within the F1RT consortium. Full Methods and associated references are available in the Supplementary Information.
Additional information
Data are available at the NCBI Short Read Archive under accession number SRA065216. Metaproteomic and metagenomic data can also be accessed from the GigaScience database (http://dx.doi.org/10.5524/100049).
References
Leschine, S. B. Cellulose degradation in anaerobic environments. Annual review of microbiology 49, 399–426, 10.1146/annurev.mi.49.100195.002151 (1995).
Horn, S. J., Vaaje-Kolstad, G., Westereng, B. & Eijsink, V. G. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 5, 45 (2012).
Rubin, E. M. Genomics of cellulosic biofuels. Nature 454, 841–845 (2008).
Merino, S. & Cherry, J. Progress and challenges in enzyme development for biomass utilization. Adv. Biochem. Eng. Biotechnol 108, 95–120 (2007).
Bayer, E. A., Kenig, R. & Lamed, R. Adherence of Clostridium thermocellum to cellulose. J. Bacteriol. 156, 818–827 (1983).
Fontes, C. M. & Gilbert, H. J. Cellulosomes: highly efficient nanomachines designed to deconstruct plant cell wall complex carbohydrates. Annu. Rev. Biochem. 79, 655–681 (2010).
Lamed, R., Setter, E. & Bayer, E. A. Characterization of a cellulose-binding, cellulase-containing complex in Clostridium thermocellum. J. Bacteriol. 156, 828–836 (1983).
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467 (2011).
Allgaier, M. et al. Targeted Discovery of Glycoside Hydrolases from a Switchgrass-Adapted Compost Community. Plos one 5, e8812 (2010).
Warnecke, F. et al. Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450, 560–565 (2007).
Kato, S., Haruta, S., Cui, Z. J., Ishii, M. & Igarashi, Y. Effective cellulose degradation by a mixed-culture system composed of a cellulolytic Clostridium and aerobic non-cellulolytic bacteria. FEMS Microbiol. Ecol. 51, 133–142 (2004).
Kato, S., Haruta, S., Cui, Z. J., Ishii, M. & Igarashi, Y. Stable coexistence of five bacterial strains as a cellulose-degrading community. Appl. Environ. Microbiol. 71, 7099–7106 (2005).
Pope, P. B. et al. Isolation of Succinivibrionaceae implicated in low methane emissions from Tammar wallabies. Science 333, 646–648 (2011).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2012).
Teeling, H., Meyerdierks, A., Bauer, M., Amann, R. & Glockner, F. O. Application of tetranucleotide frequencies for the assignment of genomic fragments. Environmental microbiology 6, 938–947, 10.1111/j.1462-2920.2004.00624.x (2004).
Sandberg, R. et al. Capturing whole-genome characteristics in short sequences using a naive Bayesian classifier. Genome research 11, 1404–1409, 10.1101/gr.186401 (2001).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl Acad. Sci. USA 106, 19126–19131 (2009).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, 233–238, 10.1093/nar/gkn663 (2009).
Resch, M. G. et al. Fungal cellulases and complexed cellulosomal enzymes exhibit synergistic mechanisms in cellulose deconstruction. Energy Environ. Sci. 6, 1858–1867 (2013).
Bayer, E. A., Lamed, R., White, B. A. & Flint, H. J. From cellulosomes to cellulosomics. Chem. Rec. 8, 364–377 (2008).
Blair, B. G. & Anderson, K. L. Regulation of cellulose-inducible structures of Clostridium cellulovorans. Can. J. Microbiol. 45, 242–249 (1999).
Kato, S. et al. Clostridium straminisolvens sp. nov., a moderately thermophilic, aerotolerant and cellulolytic bacterium isolated from a cellulose-degrading bacterial community. Int. J. Syst. Evol. Microbiol. 54, 2043–2047 (2004).
Ding, S. Y. et al. How does plant cell wall nanoscale architecture correlate with enzymatic digestibility? Science 338, 1055–1060 (2012).
Lovitt, R. W., Kell, D. B. & Morris, J. G. The physiology of Clostridium sporogenes NCIB 8053 growing in defined media. J. Appl. Bacteriol. 62, 81–92 (1987).
Sridhar, J. & Eiteman, M. A. Metabolic flux analysis of Clostridium thermosuccinogenes: effects of pH and culture redox potential. Appl. Biochem. Biotechnol. 94, 51–69 (2001).
Miller, D. A. et al. Complete genome sequence of the cellulose-degrading bacterium Cellulosilyticum lentocellum. J. Bacteriol. 193, 2357–2358 (2011).
Collins, M. D. & Shah, H. N. Reclassification of Bacteroides praeacutus Tissier (Holdeman and Moore) in a new genus, Tissierella, as Tissierella praeacuta comb. nov. Int. J. Syst. Bacteriol. 36, 461–463 (1986).
Shida, O. et al. Proposal of Bacillus reuszeri sp. nov., Bacillus formosus sp. nov., nom. rev. and Bacillus borstelensis sp. nov., nom. rev. Int. J. Syst. Bacteriol. 45, 93–100 (1995).
Acknowledgements
We are indebted to the faculty and staff of Beijing Genomics Institute at Shenzhen, whose names were not included in the author list but who contributed to sequencing. We also thank Prof. Ming from the School of Bioscience and Bioengineering, South China University of Technology, who assisted us with comparative experiments of C. thermocellum. PBP and VGHE were funded by Norwegian Research Council projects 214042 and 221568.
Author information
Authors and Affiliations
Contributions
Jian W proposed this project. J.W. and R.Y. designed the experiments and supervised the project. Y.Z. performed the bioinformatical analysis. S.L., Q.S., C.Z., J.Z., S.C., J.C., J.Y., X.L., T.J., X.Z., S.Y. and F.L. did the experiments for screening the cellulolytic consortium, assembly accuracy assessment by PCR and other relevant assays. Y.Z., B.W. and F.T. did the metaproteomics analysis. Y.Z., P.B.P., V.G.H.E., R.Y., J.W., G.Z., H.Y., Jian W and X.D. contributed to analysis of the results and paper writing.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Table 9
Supplementary Information
Supplementary Table 10
Supplementary Information
Supplementary Table 24
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Zhou, Y., Pope, P., Li, S. et al. Omics-based interpretation of synergism in a soil-derived cellulose-degrading microbial community. Sci Rep 4, 5288 (2014). https://doi.org/10.1038/srep05288
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05288
This article is cited by
-
Role of metagenomics in prospecting novel endoglucanases, accentuating functional metagenomics approach in second-generation biofuel production: a review
Biomass Conversion and Biorefinery (2023)
-
Constructing metagenome-assembled genomes for almost all components in a real bacterial consortium for binning benchmarking
BMC Genomics (2022)
-
Effect of liquid volume and microflora source on degradation rate and microbial community in corn stover degradation
AMB Express (2021)
-
Comparative metagenomic discovery of the dynamic cellulose-degrading process from a synergistic cellulolytic microbiota
Cellulose (2021)
-
A completeness-independent method for pre-selection of closely related genomes for species delineation in prokaryotes
BMC Genomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.