Abstract
The dipeptidyl aminopeptidase BII (DAP BII) belongs to a serine peptidase family, S46. The amino acid sequence of the catalytic unit of DAP BII exhibits significant similarity to those of clan PA endopeptidases, such as chymotrypsin. However, the molecular mechanism of the exopeptidase activity of family S46 peptidase is unknown. Here, we report crystal structures of DAP BII. DAP BII contains a peptidase domain including a typical double β-barrel fold and previously unreported α-helical domain. The structures of peptide complexes revealed that the α-helical domain covers the active-site cleft and the side chain of Asn330 in the domain forms hydrogen bonds with the N-terminus of the bound peptide. These observations indicate that the α-helical domain regulates the exopeptidase activity of DAP BII. Because S46 peptidases are not found in mammals, we expect that our study will be useful for the design of specific inhibitors of S46 peptidases from pathogens.
Similar content being viewed by others
Introduction
Peptidases (proteases or proteinases) catalyse the hydrolysis of peptide bonds. These enzymes are widely distributed in nature and are involved in a wide variety of functions1. For example, viral peptidases are processing enzymes that cleave precursors of their coat proteins, many extracellular bacterial peptidases degrade proteins in their surroundings and higher organisms use peptidases for functions such as food digestion, blood coagulation, development2 (e.g. Nudel, Gastrulation Defective, Snake and Easter proteases are involved in dorsoventral axis formation in the Drosophila embryo3), immunity (e.g. C3 protein is cleaved and activated by C3-convertase to C3a and C3b in the complement system4) and apoptosis5. Peptidases can be grouped according to catalytic type; aspartic, cysteine, metallo- and serine peptidases have been extensively studied, whereas glutamic and threonine peptidases have only been identified. In the MEROPS database6, each peptidase is assigned to a “family” on the basis of amino acid sequence similarity and families that are considered homologous are grouped into a “clan”. Among the various peptidases, serine peptidases have been extensively studied7,8 and are found in all kingdoms of life. Structural studies have frequently been conducted of clan PA, family S1 enzymes, such as chymotrypsin9. Peptidases can also be grouped according to the pattern of proteolysis as either endo- or exo-peptidases. Exopeptidases catalyse the removal of amino acids (or short peptides) from the end of a polypeptide chain, whereas endopeptidases cleave a peptide bond between nonterminal amino acids. Exopeptidases are divided into aminopeptidases, dipeptidases, dipeptidyl-peptidases, tripeptidyl-peptidases, carboxypeptidases and omega peptidases by the IUBMB nomenclature.
Dipeptidyl aminopeptidase BII (DAP BII) from Pseudoxanthomonas mexicana WO24 was originally isolated from the waste produced in a bean curd (tofu) factory and possesses the dipeptidyl peptidase (DPP, EC 3.4.14) activity, that catalyses the removal of dipeptides from the amino terminus of peptides10. The bacterium prefers oligopeptides as a carbon and nitrogen source (Suzuki et al., unpublished result), indicating that DAP BII plays an important role in feeding by digesting oligopeptides absorbed into the periplasm. We recently determined the nucleotide sequence of DAP BII and the predicted amino acid sequence revealed that the enzyme is a new class of 79-kDa (722-a.a.) serine proteases that has been classified as belonging to the clan PA, family S4611. Only three dipeptidyl peptidases, DPP7 from Porphyromonas gingivalis12 and DPP11s from P. gingivalis and P. endodontalis13, have been placed in family S46. The clan PA contains over seven hundred identified peptidases, mainly endopeptidases such as trypsin and chymotrypsin, the best known of all peptidases and forms the largest of all peptidase clans7,8. The only exception in the clan PA is the S46 family in that the S46 enzymes are exopeptidases11,12,13. To act as an exopeptidase, the clan PA peptidase unit (the chymotrypsin fold) must be covered by a regulatory domain to spatially restrict the access of substrate peptides and proteins. Clear examples of exopeptidase regulatory mechanism have been known for clan SC, family S9 peptidases (e.g. dipeptidyl peptidase IV), in which a catalytic α/β hydrolase fold is covered by a regulatory β-propeller domain14,15,16,17. Similar regulatory mechanisms have been reported for clan SC, family S28 peptidases (e.g. prolylcarboxypeptidase and dipeptidyl peptidase 7), in which a catalytic α/β hydrolase fold is covered by a regulatory α-helical SKS domain, rather than a β-propeller domain observed in the family S9 peptidases18,19. Although many structural studies of clan PA peptidases, such as chymotrypsin9 and trypsin20, have been reported, no structural example representing exopeptidase activity by a chymotrypsin fold associated with a regulatory domain has yet been reported.
In this study, the crystal structures of DAP BII have been experimentally identified for the first time for a family S46 peptidase and the present study also shows the first structural example of an exopeptidase in clan PA. The structures revealed that DAP BII contains a peptidase domain with a double β-barrel fold and an unusual, previously unreported α-helical domain. The structures clearly explain the molecular basis of the exopeptidase activity of DAP BII, which is most likely the common mechanism among S46 peptidases. The residues directly involved in recognition of the N-terminus of the substrate peptide are Asn215, Asn330 and Asp674. These residues are completely conserved among family S46 peptidases and absent from clan PA endopeptidases. In addition, structural snapshots of the sequential digestion of peptide substrates by DAP BII were obtained through crystal structure analyses of wild-type and mutant DAP BIIs complexed with angiotensins II or IV as model peptides.
Results
Overall structure of DAP BII
In its mature form, DAP BII is a homodimer, each subunit comprises 698 residues (Gly25-Lys722) with a molecular weight of approximately 76 kDa (Fig. 1). The crystal structures of wild-type and mutant DAP BIIs in the peptide-free form (ground- and space-grown crystals), as well as the dipeptide- (Val-Tyr), hexapeptide- (Val-Tyr-Ile-His-Pro-Phe; angiotensin IV) and octapeptide- (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe; angiotensin II) bound complexes were determined at 2.2- to 1.74-Å resolutions (Tables S1 and S2). Representative electron density maps of the bound peptides are shown in Fig. S1. The subunit of DAP BII consists of two domains separated by a deep cleft (Fig. 2A). One domain, containing the chymotrypsin fold, is responsible for catalysis; the second, α-helical domain is the regulatory domain necessary for exopeptidase activity.
Amino acid sequences of clan PA S46 family peptidases.
The His-Asp-Ser catalytic triad and the N-terminus recognition residues are shown in red and blue, respectively. The signal sequence of DAP BII (residues 1–24) is shown in italics. Secondary structural elements are shown on top. The abbreviations used (UniProt accession numbers) are as follows: PmDAPBII (V5YM14), Pseudoxanthomonas mexicana WO24 DAP BII; PgDPP7 (Q7MWU6), Porphyromonas gingivalis DPP7; PgDPP11 (B2RID1), Porphyromonas gingivalis DPP11; PeDPP7 (C3JAQ3), Porphyromonas endodontalis DPP7; PeDPP11 (F8WQK8), Porphyromonas endodontalis DPP11; and SmDPP7 (B4SLK2), Stenotrophomonas maltophilia DPP7.
Three-dimensional structure of DAP BII.
(A) Stereoview of the DAP BII subunit. Helices (1 to 30) and strands (N1 to N7 and C1 to C6 in the N- and C-terminal barrels, respectively) are coloured and labeled as in Figure 1.(B) Dimeric structure of DAP BII in orthogonal views. One subunit is coloured blue and orange and the other is coloured pink and green, for the catalytic and α-helical domains, respectively. (C) Close view of the intermolecular four-stranded antiparallel β-sheet observed at the dimer interface of DAP BII.
In the present crystal structure, two subunits of an asymmetric unit form a homodimer (Fig. 2B). The dimer formation is consistent with gel-filtration data showing that the apparent molecular weight of DAP BII is approximately 150 kDa in solution10. Inter-subunit interactions in DAP BII occur primarily along protruding β-hairpins of the catalytic domain (Fig. 2B, left); thus, a four-stranded antiparallel β-sheet is formed from the β-hairpins from the catalytic domains of each subunit (Fig. 2C). The α-helical domains are not involved in dimer formation (Fig. 2B, right). Because this type of inter-subunit interaction is found among different crystal forms (Table S2), it is likely that dimer formation is the natural state of DAP BII and that it also exists in solution.
Catalytic domain
The catalytic domain comprises residues 25 to 276 and residues 574 to 722 (Figs. 1, 3A and S2) and forms a double β-barrel structure, which is characteristic of clan PA peptidases, consisting of 15 α-helices (α1–α10 and α25–α29) and 15 β-strands (βN1-βN7 and βC1-βC6). The domain's structural core is formed by two six-stranded antiparallel β-barrels. Although the catalytic domain of DAP BII exhibits low sequence identity to other clan PA peptidases, the cores of their three-dimensional structures are superimposable. For example, the structure of one of the best characterised clan PA enzymes, chymotorypsin (Fig. 3B), can be superimposed onto the catalytic domain of DAP BII (Fig. 3C) with a root mean square (rms) deviation between the two structures of 1.84 Å for 173 structurally equivalent Cα atoms, which had only 15% sequence identity for that region (Fig. S3). The serine peptidase catalytic triads, His86, Asp224 and Ser657 in DAP BII11 and His57, Asp102 and Ser19521 in chymotrypsin, are almost completely superimposable (Figs. 3C, 3D and S2). The binding mode of peptides will be presented later.
The catalytic domain of DAP BII.
Ribbon diagrams of the catalytic domain of DAP BII (A) and that of chymotrypsin (PDB ID: 1YPH) (B). The catalytic triads of these enzymes are shown in ball-and-stick format. (C) Superposition of DAP BII and chymotrypsin, which are shown in cyan and purple, respectively. (D) Domain structures of DAP BII and chymotrypsin. The N- and C-terminal barrels of the catalytic domains of the two enzymes are coloured blue and red, respectively. The α-helical domain of DAP BII is coloured green. (E) The catalytic triad and the α-helical domain of DAP BII. The figure corresponds to an enlarged detail of Figure 2A.
The α-helical domain
The α-helical domain comprises residues 277 to 573 (Figs. 1, 2A and S2) and is conserved among S46 peptidases (Fig. 1). This domain represents a completely new structure because no structural homologue of this domain is found in the DALI server22. The α-helical domain packs against the catalytic domain and the catalytic triad (His86, Asp224 and Ser657) of the catalytic domain lies at the interface between the two domains (Fig. 3E). A comparison of the crystal structure of peptide-free DAP BII with those of peptide (di, hexa, or octa)-bound forms of DAP BII shows that the large cleft between the catalytic and α-helical domains is closed upon peptide binding (Fig. 4A). A closed conformation was observed for both crystallographically independent subunits of the peptide-bound complexes. The largest change in Cα position (approx. 12.7 Å) was observed for Gln313. Interestingly, although both subunits of the DAP BII dimer in the asymmetric unit of the peptide-free crystal exhibit a relatively open conformation, the conformations of the two subunits are slightly different (Fig. S4). These observations indicate that the link between the two domains of the DAP BII subunit is flexible.
Substrate-binding mode of peptide-bound DAP BII.
(A) Comparison of the subunit structure of peptide-free DAP BII (orange) and that of peptide-bound DAP BII (purple). N330 in each of the α-helical domains is indicated using arrows. (B) Substrate N-terminus recognition mode observed in the Val-Tyr-bound DAP BII. The peptide-free structure (orange) is shown for comparison. (C) Schematic diagram showing the binding mode of the Val-Tyr dipeptide with DAP BII. (D) Surface drawing of subsites S2-S1 of DAP BII. (E) The catalytic triad and oxyanion hole of DAP BII.
Recognition of the substrate N-terminus
Peptide complex structures can clearly explain the molecular basis for the dipeptidyl peptidase activity of DAP BII. For simplicity, the following description refers primarily to subunit A of the Val-Tyr complex of DAP BII (dipeptide complex I, Table S1). DAP BII hydrolyses peptides from the N-terminus of oligopeptides and small proteins, cleaving dipeptide units (NH3-P2-P1-) when the second (P1) residue is Ala, Leu, Ile, Phe, Tyr, Arg, or His (but not Pro)10. To act as a dipeptidyl peptidase, DAP BII must be able to fix the N-terminus in position. This task is performed by the side chain of Asn330 in helix α13 of the α-helical domain (Figs. 4A and 4B); a bifurcated hydrogen bond is formed between the amide and carbonyl groups of the Asn330 side chain and the carbonyl and amino groups, respectively, of the main chain of the N-terminal P2 residue (Fig. 4B). In the open conformation observed in peptide-free DAP BII, Asn330 is located away from the catalytic triad. A structural difference of approx. 5.6 Å in the Cα position of Asn330 is observed between the closed and open conformations (Figs. 4A and 4B). The side chains of Asn215, Trp216 and Asp674 are also involved in the recognition of the N-terminus of the substrate (Figs. 4B and 4C). The side chain of Asp674 is conformationally restrained by hydrogen-bond interactions with the side chain of Arg220. The bulky side chain of Trp216 appears to function as a wall to restrict a part of the active site at the S3 subsite and is also involved in a cation-pi interaction with the N-terminus of the bound peptide. A similar interaction is observed in a cathepsin C/inhibitor complex (between Phe278 of the enzyme and the N-terminus of the bound inhibitor)23. Asn215 and Trp216 are located in an insertion (between strands βN6 and βN7, see Figs. S2 and S3; this insertion does not exist in chymotrypsin) of the N-terminal barrel of DAP BII, whereas Asp674 is located at a β-strand of the C-terminal barrel; the corresponding residue in chymotrypsin is Trp215, which contributes to a hydrophobic core. Individual Ala-substitution of Asn215, Trp216, Asn330 and Asp674 resulted in a drastic loss of enzymatic activity (Table S3), indicating that these residues play an important role in the recognition of the N-terminus of the substrate peptide in DAP BII. Among 661 amino acid sequences of S46 peptidases found in 306 species11, Asn215, Asn330 and Asp674 in DAP BII are completely conserved (e.g. Fig. 1). With respect to Trp216 in DAP BII, the tryptophan residue is conserved in 537 sequences (81.24%), while conservative substitutions are observed in the rest of 124 sequences [replaced by phenylalanine in 94 sequences (14.22%) and tyrosine in 30 sequences (4.54%)].
Recognition of P1 and P2 residues
The S1 subsite is sufficiently deep to accommodate the bulky Tyr side chain of the bound dipeptide (Fig. 4D). This is consistent with the relatively loose P1 preference of DAP BII for synthetic substrates (Xaa-Yaa-p-nitroanilide, Xaa-Yaa-pNA, where Xaa is any amino acid and Yaa is Ala, Leu, Ile, Phe, Tyr, Arg, or His). The pocket is lined by residues Asp651-Gly654, Phe673-Ser682 and Thr691-Ile694 in the C-terminal barrel. The residues in the S1 pocket of DAP BII are structurally equivalent to Ser189-Met192, Ser214-Cys220 and Thr224-Val227, respectively, of chymotrypsin24,25 (Figs. 5A and 5B and Table S4). The NH group of the P1 residue of the bound peptide is recognized by a hydrogen bond (3.0 Å) with the carbonyl oxygen of Phe673 (Fig. 4E). This interaction explains why DAP BII cannot cleave a substrate with Pro at the P1 position because such substrates could not provide a favourable hydrogen-bond interaction between NH(P1) and O(Phe673) and would add an unfavourable steric clash between CD(P1) and O(Phe673). The carbonyl oxygen of Phe673 in DAP BII is structurally equivalent to that of Ser214 in chymotrypsin (Figs. 5A and 5B and Table S4). The S2 subsite is also sufficiently deep to accommodate a larger side chain and is lined by the side chains of Lys208, Arg220 and Thr222 from the N-terminal barrel and Phe673 from the C-terminal barrel. As described above, the main-chain atoms of the P2 residue are recognized by the side chains of DAP BII (Asn215, Asn330 and Asp674). However, the S2 subsite appears to be nonspecific for the substrate side chain because intermolecular interaction between the S2 subsite and the side chain of P2 residue is limited. This result is also consistent with the lack of specificity of DAP BII for P2 observed with synthetic substrates.
The active site of clan PA peptidases.
The catalytic triad, the Gly residue providing the NH group for the oxyanion hole and the determinants of the strict P1 specificities are coloured red, blue and green, respectively. The bound ligands are shown in ball-and-stick format. The P1 specificities of the presented peptidases are shown at the bottom of each figure. (A) DAP BII complexed with Val-Tyr (present study). (B) Chymotrypsin complexed with a 14-residue peptide (PDB ID: 1OXG). (C) Trypsin complexed with an inhibitor, AERUGINOSIN 98-B (PDB ID: 1AQ7). (D) Elastase complexed with a tripeptide, acetyl-Ala-Pro-Ala (PDB ID: 1NES). (E) V8 protease (PDB ID: 2O8L). (F) Superposition of the five enzymes. The catalytic triads are shown in ball-and-stick format. Each molecule is coloured as described in (A) to (E).
The catalytic triad and oxyanion hole
The catalytic triad of DAP BII, His86, Asp224 and Ser65711 is located in the loop regions contributed by each barrel of the catalytic domain and is covered by the α-helical domain. The catalytic triad of DAP BII spans approximately 600 amino acids, whereas the catalytic triads of most clan PA peptidases are distributed among approximately only 200 amino acids, e.g., His57, Asp102 and Ser195 in chymotrypsin21 (Table S4). This is due to the insertion of the α-helical domain into the C-terminal barrel of the catalytic domain of DAP BII (Fig. 3D). The hydroxyl group of Ser657 is hydrogen bonded to the NE atom of the imidazole ring of His86 (2.7 Å). One of the oxygen atoms of the carboxylate group of Asp224 is hydrogen bonded to the ND of His86 (2.6 Å) and the other oxygen atom of Asp224 is fixed by the main chain NH group of His85 and a water molecule that is hydrogen bonded to the NH group of His86.
During serine peptidase catalysis, a tetrahedral intermediate is formed, which includes the negatively charged oxyanion generated from the carbonyl oxygen of the scissile bond; this oxyanion is stabilised by two hydrogen bonds within the active site26,27. In peptide-bound DAP BII, the carbonyl oxygen of the P1 residue lies in the oxyanion hole and forms hydrogen bonds to the main chain NH groups of the catalytic Ser657 residue (3.1 Å) and to another residue, Gly655 (2.8 Å) (Fig. 4E). These residues are structurally equivalent to Ser195 and Gly193, respectively, in chymotrypsin. Thus, the geometry of the catalytic triad and the oxyanion hole are well conserved among clan PA peptidases28,29,30,31 (Fig. 5).
Recognition of P1′-P2′-P3′ residues
In the present crystal structure analysis, we successfully determined the crystal structures of DAP BII (H86A single mutant) complexed with a hexapeptide (Val-Tyr-Ile-His-Pro-Phe) (hexapeptide complex I, Table S1) and DAP BII (H86A/D224A/S657A triple mutant) complexed with an octapeptide (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe) (octapeptide complex, Table S1). These structures revealed the substrate recognition mode of the peptide (at P1′-P2′-P3′) downstream of the scissile bond.
Hexapeptide complex I was obtained by co-crystallisation of DAP BII (H86A) with angiotensin II as a model peptide (Table S1). Clear, continuous electron density was observed for the region from third to the seventh residues of the peptide (Fig. S1B). Thus, the N-terminal dipeptide (Asp-Arg) appeared to be cleaved. Five of the remaining six residues form molecular interactions with the active site residues of DAP BII (Figs. 6A and 6B). No clear electron density was observed for the last residue (Phe) because it is exposed to solvent. The recognition mode of the P2 and P1 residues is equivalent to that observed in the dipeptide (Val-Tyr) complex described above. The carbonyl oxygen of the P1 residue is accommodated in the oxyanion hole and is hydrogen bonded to the main-chain NH groups of Ser657 (2.9 Å) and Gly655 (2.8 Å). Although a direct hydrogen bond (3.2 Å) is observed between the NH group of the P2′ residue (His) and the main-chain carbonyl group of Gly69, the P1′ residue (Ile) exhibits limited interaction with the active site of DAP BII; a distant hydrophobic contact is observed between the side chain of P1′-Ile and that of Cys70, which is involved in a disulphide bridge with Cys87. The carbonyl oxygen of Gly69 in DAP BII is structurally equivalent to that of Phe41 in chymotrypsin (Table S4). Another distant hydrophobic contact is observed between the side chain of P2′-His and that of Leu577. Electron density for the P3′ residue (Pro) was well defined despite the lack of tight interaction with the active site because the neighbouring P2′ residue (His) is fixed to the active site by a hydrogen bond as described above.
The recognition of P1′-P2′-P3′ residues.
(A) The P1′-P2′-P3′ recognition mode observed in hexapeptide (Val-Tyr-Ile-His-Pro-Phe)-bound DAP BII. Disordered residues are shown in italics. (B) Schematic diagram showing the binding mode of the hexapeptide in DAP BII. (C) The P1′-P2′-P3′ recognition mode observed in octapeptide (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe)-bound DAP BII. (D) Schematic diagram showing the binding mode of the octapeptide in DAP BII. (E) Superposition of the active sites of the hexapeptide complex (orange) and the octapeptide complex (yellow).
To reproduce the peptide-binding mode observed in hexapeptide complex I described above, DAP BII (H86A) and DAP BII (H86A/D224A/S657A) were crystallised with intact angiotensin IV (hexapeptide complexes II and III, Table S1). These complexes exhibited essentially equivalent enzyme-peptide interactions to those seen in the DAP BII (H86A)-hexapeptide (digested angiotensin II) complex (hexapeptide complex I).
An octapeptide (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe) complex was obtained by co-crystallisation of DAP BII (H86A/D224A/S657A triple mutant) with angiotensin II as a model peptide (octapeptide complex, Table S1). Clear continuous electron density was observed for the first five residues of the octapeptide (Fig. S1C) and no clear electron density was observed for the last three residues, which are exposed to solvent. In this complex, the bound peptide was intact, i.e., the N-terminus of the bound peptide was Asp because all residues of the catalytic triad were mutated to Ala. The peptide main-chain recognition mode for the P2-P1-P1′-P2′ residues of the octapeptide complex (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe) is essentially equivalent to that observed for the hexapeptide complex (Val-Tyr-Ile-His-Pro-Phe) (Figs. 6C and 6D). A clear structural difference was observed downstream of P3′ (Fig. 6E) due to the structural restriction caused by the presence of Pro at the P3′ position of the hexapeptide complex.
Discussion
The series of peptide-complex structures presented here provide insights into the catalytic mechanism of DAP BII, which may be common to clan PA peptidases inasmuch that the mechanism involves two tetrahedral intermediates32 (Fig. 7). In the first step, the hydroxyl oxygen atom of Ser657 attacks the P1-carbonyl carbon of the peptide substrate due to the action of the catalytic triad His86 as a general base to extract the proton of the Ser657 OH group (Fig. 7, arrow 1). The oxyanion-containing tetrahedral intermediate is stabilised by the backbone NH groups of Gly655 and Ser657, which generate a positively charged pocket known as the oxyanion hole26,27. When the peptide bond is cleaved, one peptide product attaches to the enzyme forming the acyl-enzyme intermediate (Ser657-P1-P2) and the exit of the N-terminus of the other, newly created peptide (P1′-P2′-…) is mediated by His86 and the peptide product rapidly diffuses away (Fig. 7, arrows 2 to 5). In a second step, a water molecule attacks the acyl-enzyme intermediate (Fig. 7, arrow 6). The oxyanion hole also stabilises the second tetrahedral intermediate. Then second peptide product (P2-P1) containing a complete C-terminus is released and the OH group of Ser is restored by His86 (Fig. 7, arrows 7 to 8).
The side chain of Asn215, Trp216 and Asp674 from the catalytic domain and the side chain of Asn330 from the α-helical domain fix the N-terminus of the substrate in the correct position. The relative arrangement between these side chains and the catalytic Ser657 limits the active site and leaves room for only two amino acids before the scissile bond reaches the catalytic Ser657, making DAP BII a dipeptidyl peptidase. In the case of another dipeptidyl peptidase, dipeptidyl peptidase IV (clan SC, family S9), in which a catalytic α/β hydrolase fold is covered by a regulatory β-propeller domain, the N-terminus of the substrate is held precisely in position by strong interactions with the side chains of a double Glu motif (Glu205-Glu206) from an insertion helix of the regulatory β-propeller domain14,16 (Fig. S5).
The structure of the dipeptide Val-Tyr complex, which was obtained by co-crystallisation of DAP BII (WT) with angiotensin II (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe), (dipeptide complex II, Table S1) is consistent with the above catalytic mechanism and provides insight into the sequential digestion mechanism of peptide substrates by DAP BII. Because the DAP BII enzyme reaction occurs in the solution used for crystallisation, the first substrate is the octapeptide (Fig. 8, State 1-1), the reaction products are Asp-Arg (P2-P1) and Val-Tyr-Ile-His-Pro-Phe (P1′-P2′-…) and both peptides dissociate from the active site of DAP BII. The dissociated Val-Tyr-Ile-His-Pro-Phe peptide then rebinds to the active site (Fig. 8, State 2-1) and the second reaction products are Val-Tyr (P2-P1) and Ile-His-Pro-Phe (P1′-P2′-…). At this time, while the N-terminal product Val-Tyr remains at the active site, the C-terminal product dissociates from the active site. Thus, the reaction appears to be trapped after hydrolysis of the acyl-enzyme intermediate by a water molecule (Fig. 8, State 2-2). Similarly, the structure of the dipeptide Val-Tyr complex obtained by co-crystallisation of DAP BII (WT) with angiotensin IV (Val-Tyr-Ile-His-Pro-Phe) (dipeptide complex III, Table S1) is consistent with the above catalytic mechanism. In this case, the first substrate is the hexapeptide (Fig. 8, State 1-1) and the reaction products are Val-Tyr (P2-P1) and Ile-His-Pro-Phe (P1′-P2′-…). At this time, while the N-terminal product Val-Tyr remains at the active site (Fig. 8, State 1-2), the C-terminal product dissociates from the active site.
Sequential peptide digestion of DAP BII.
Snapshots of states 1-0, 1-1, 1-2, 2-1 and 2-2 were obtained from the present crystal structure analyses (see Table S1).
The dipeptide Val-Tyr complexes II and III described above indicate that the dipeptide Val-Tyr can act as a competitive inhibitor of DAP BII. Thus we examined the inhibitory activities of the dipeptides Val-Tyr and Asp-Arg against the hydrolysis of the synthetic substrate Gly-Phe-pNA (Km value of 1.4 mM) by DAP BII (WT). The IC50 and Ki values of the dipeptide Val-Tyr were measured as 0.50 ± 0.02 mM and 0.41 ± 0.02 mM, respectively. In contrast, the dipeptide Asp-Arg, corresponding to the N-terminal dipeptide of angiotensin II (Table S1), exhibited no inhibitory activity against the hydrolysis of Gly-Phe-pNA by DAP BII (WT) (Table S5). Interestingly, however, the dipeptide Val-Tyr can be a portion of substrate of DAP BII. DAP BII sequentially releases Asp-Arg, Val-Tyr, Ile-His, Pro-Phe and His-Leu from the N terminus of angiotensin I to degrade it completely into dipeptides10. The S1 site of DAP BII has a tendency to prefer a large hydrophobic side chain11, although strict specificity is not observed.
The structure of the hexapeptide complex obtained by the co-crystallisation of DAP BII (H86A) with angiotensin II (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe) (hexapeptide complex I, Table S1) represents a snapshot of DAP BII complexed with the second substrate (Fig. 8, State 2-1). In this reaction, the first substrate is the octapeptide (Fig. 8, State 1-1), the reaction products are Asp-Arg (P2-P1) and Val-Tyr-Ile-His-Pro-Phe (P1′-P2′-…) and both peptides dissociate from the active site of DAP BII. The dissociated Val-Tyr-Ile-His-Pro-Phe peptide then rebinds to the active site as the second substrate (Fig. 8, State 2-1). Fortunately, the second substrate was not digested by the mutant enzyme and the hexapeptide remained in the active site. To test the resistance of the hexapeptide (corresponding to angiotensin IV) against proteolysis by DAP BII (H86A), the hexapeptide was co-crystallised with DAP BII (H86A). The crystallographic evidence clearly showed that the hexapeptide was not cleaved by DAP BII (H86A) (hexapeptide complex II, Table S1).
The present crystal structure analyses revealed that DAP BII substrate recognition is non-specific, with the exception that Pro is not suitable as a P1 residue. The S2 subsite forms a large space and the S1 subsite is sufficiently deep to accommodate any amino acids (Fig. 4D). In regard to the S1′, S2′, … subsites, few interactions were observed between the peptide side chain and the enzyme active site (Fig. 6). These observations imply that the P2 and/or P1 specificity of DAP BII may be altered by modification of the residues in the S2 and/or S1 subsites. Among the S46 peptidases, a novel dipeptidyl peptidase, DPP11, from P. gingivalis (PgDPP11) has been reported to have specificity for Asp- and Glu- at the P1 residue13. Biochemical studies of PgDPP11 revealed that Arg670 is responsible for the Asp/Glu specificity of PgDPP1113,33. Arg670 in PgDPP11 corresponds to Gly675 in DAP BII (Table S4). Because Gly675 is located at the bottom of a wall of the S1 subsite (Fig. 4E and 5A), mutation of Gly675 would alter the P1 specificity of DAP BII. Similarly, the P2 specificity of DAP BII could be altered by limiting the space available at the S2 subsite by introducing a mutation at this site. Mutagenesis studies revealed that Phe664 of PgDPP7 and Phe671 of PgDPP11, which correspond to Phe673 of DAP BII, contribute to the preference for hydrophobic residues at the substrate P2 position34. Dipeptides, which are absorbed more rapidly than amino acids, have a number of commercial and industrial uses and play an important role in living organisms. Therefore, we expect the present structures to be useful guides for the design of engineered forms of DAP BII that can produce custom dipeptides.
The DAP BII structure is also useful for drug design. As described below, S46 peptidases are not found in mammals but do exist in some pathogenic organisms, such as P. gingivalis (a periodontal pathogen)35, Stenotrophomonas maltophilia (an opportunistic pathogen)36 and Xylella fastidiosa (a plant pathogen)37. For example, three S46 peptidases have been characterised in Porphyromonas species12,13. These organisms are asaccharolytic, do not ferment glucose, cellobiose, lactose, or sucrose and require peptide/protein substrates as carbon and energy sources. In addition, it has recently been reported that P. gingivalis accelerates inflammatory atherosclerosis38. Therefore, DPP7 and DPP11 from P. gingivalis may be good targets for atherosclerosis therapy.
To clarify the origin of the α-helical domain found in DAP BII, we searched for amino acid sequence homologues of the α-helical domain. The resulting α-helical domain homologues are absolutely restricted to S46 peptidases. The α-helical SKS domains observed in S28 peptidases18,19 are not related to the α-helical domain of S46 peptidases. The search also showed that S46 peptidases are restricted to prokaryotes and do not exist in eukaryotes or archaea. Phylogenetic analysis of S46 peptidase genes (Fig. S6) revealed that S46 peptidases exist widely in Proteobacteria and Bacteroidetes. These observations indicate that Proteobacteria and Bacteroidetes acquired the S46 peptidase gene via horizontal gene transfer from an ancestral S46 gene, although the timing of the appearance of the ancestral gene is uncertain. Unlike the α-helical domain of DAP BII, homologues of the catalytic domain of DAP BII (the chymotrypsin fold) are ubiquitous. Given that homologues of the α-helical domain of DAP BII exist only in S46 peptidases, gene fusion of the α-helical domain and catalytic domain to form the two-domain architecture of S46 peptidases can be excluded.
In this paper, we present the structures of the peptide-free and peptide-bound forms of DAP BII, a member of the S46 peptidase family, which is a unique member of the clan PA peptidases. This is the first structural example of a catalytic chymotrypsin fold-containing exopeptidase that also contains a regulatory domain. The structures of peptide-bound complexes reveal the mode of recognition of the substrate N-terminus by the active site residues of DAP BII and clearly explain the molecular basis of the exopeptidase activity of DAP BII. In addition, a series of peptide-complex structures provide insight into the sequential peptide digestion mechanism of DAP BII. We expect that the present structures could act as useful guides for the design of specific inhibitors of S46 peptidases produced by pathogenic organisms.
Methods
Overexpression and purification of DAP BII
DAP BII was expressed and purified as described elsewhere11. DNA encoding DAP BII from Pseudoxanthomonas mexicana WO24 (accession: AB889525) was obtained by plaque hybridization with the chromosomal DNA library of the organism. The target sequence corresponding to mature DAP BII (Gly25-Lys722) containing the signal peptide of DAP BIII from Pseudoxanthomonas mexicana WO24 was amplified using PCR and cloned into the pET22b expression plasmid. E. coli Rosetta2(DE3)pLacI cells harbouring the pET22b expression plasmid were grown in YT media at 298 K to an OD600 of 0.6. Overexpression of DAP BII was induced by adding 0.2 mM IPTG for 16 h at 298 K. After this period, the cells were harvested by centrifugation at 8,000 × g. Cells were disrupted using BugBuster Protein Extraction Reagent (Novagen). The cell extract was obtained by centrifuging the lysate at 27,000 × g. DAP BII was purified using 35 to 70% ammonium sulphate precipitation and hydrophobic column chromatography using a HiPrep 16/10 Butyl column (GE Healthcare), desalted using a HiPrep 26/10 desalting column (GE Healthcare) and finally subjected to anion-exchange column chromatography using a Mono Q 5/50 GL column (GE Healthcare). The fractions containing DAP BII were pooled, buffer-exchanged to 80 mM Tris/HCl pH 8.5 using a Vivaspin 20 concentrator (GE Healthcare) and concentrated to 10 mg/ml using the concentrator.
Se-Met-substituted DAP BII was expressed using the Overnight Express Autoinduction System 2 (Novagen). The Se-Met derivative was purified in a manner similar to that used for wild-type DAP BII described above.
Crystallisation
To obtain peptide-free DAP BII crystals39, the samples were crystallised using the hanging-drop method, in which 1 μl of protein solution (10 mg/ml DAP BII in 80 mM Tris-HCl, pH 8.5) was mixed with the same volume of reservoir solution (18% (w/v) PEG8000, 20% (v/v) glycerol and 2 mM zinc chloride in 0.08 M CHES-NaOH, pH 9.5) and incubated at 293 K. The drops were suspended over 500 μl of reservoir solution in 24-well plates. Peptide-free crystals were also obtained using a counter-diffusion crystallisation method40 under a microgravity environment in the Japanese Experimental Module “Kibo” at the International Space Station (ISS)41. Although autolysis was not observed for DAP BII, a mutant (S657A, the catalytic serine was mutated to alanine) was used just to be safe for the prolonged (4 to 6 months) crystallisation experiment at the ISS. For peptide-free DAP BII, higher resolution data were obtained from the space-grown crystal (1.95 Å resolution) than from the ground-grown crystal (2.20 Å resolution). To obtain crystals of the peptide complex, protein solution (11.1 mg/ml DAP BII in 80 mM Tris-HCl, pH 8.5) was mixed with solution of each of the peptides (30 mM peptide in 80 mM Tris-HCl, pH 8.5) at a volume ratio of 9:1. The DAP BII constructs used to prepare the peptides complex crystals were the wild type, a single mutant (H86A) and a triple mutant (H86A/D224A/S657A). The peptides used were the dipeptide Val-Tyr, angiotensin II (Asp-Arg-Val-Tyr-Ile-His-Pro-Phe) and angiotensin IV (Val-Tyr-Ile-His-Pro-Phe). The samples were crystallised using the hanging-drop method, in which 1 μl of peptide complex solution (10 mg/ml DAP BII and 3 mM peptide in 80 mM Tris-HCl, pH 8.5) was mixed with the same volume of reservoir solution (18% (w/v) PEG8000, 20% (v/v) glycerol, 2 mM zinc chloride and 160 mM magnesium acetate) and incubated at 293 K. The drops were suspended over 500 μl of reservoir solution in 24-well plates. An outline of the crystallisation of DAP BII in the absence and presence of the peptides is provided in Table S1.
X-ray data collection
Because the crystallisation conditions of DAP BII described above included 20% (v/v) glycerol in the reservoir solution, X-ray data could be collected under cryogenic conditions without the need for any further addition of cryoprotectant. Crystals obtained from the hanging drop were directly mounted in nylon loops and flash-cooled in a cold nitrogen gas stream at 100 K immediately before data collection. Data were collected by the rotation method at 100 K using an ADSC Q315r CCD detector with synchrotron radiation [λ = 1.000 Å on beamline 17A of the Photon Factory] for most of the crystals and using a Rayonix MX225HE CCD detector with synchrotron radiation [λ = 0.8000 Å on beamline 41XU of SPring-8] for the space-grown crystal. The Laue group and unit-cell parameters were determined using the HKL2000 package42. The resulting cell parameters and data-collection statistics are summarised in Table S2.
Structure determination
Initial phases were determined for the peptide-free DAP BII using the single-wavelength anomalous diffraction (SAD) method. Se-Met-substituted DAP BII was crystallised in the space group P43212. Se-SAD phases were determined using the program SHARP/autoSHARP43. Automatic model building and refinement were performed using the programs ARP/wARP44 and REFMAC545 and further iterative manual model building and refinement were performed using the programs Coot46 and REFMAC5. The refined peptide-free model was used for structural determination of the dipeptide (Val-Tyr) complex of DAP BII by the molecular replacement method using the program Phaser47 from the CCP4 suite48. The Val-Tyr complex of DAP BII was refined at 1.74 Å resolution. The refined Val-Tyr complex model was used as the starting model for the structural determination of the remaining peptide complexes. The refinement statistics for the peptide-free and peptide-bound complexes of DAP BII are summarised in Table S2. Figures were produced using the programs UCSF Chimera49 and LIGPLOT50. Least-squares comparisons of the molecular models were carried out using the program UCSF Chimera and XtalView51. Multiple sequence alignment of S46 peptidases was performed using the program ClustalW52 and a phylogenetic tree was produced using the program FigTree (http://tree.bio.ed.ac.uk/software/figtree/).
Enzymatic activity assay of mutant DAP BIIs
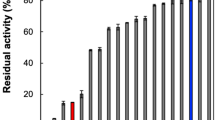
Asn215, Trp216, Asn330 and Asp674 of DAP BII, which are involved in the recognition of the N-terminus of the substrate peptide, were individually mutated to Ala. The mutants were expressed using the pET system and purified using the same method as that described for wild-type DAP BII above. The enzymatic activities of the mutant enzymes were measured based on the hydrolysis of a synthetic substrate, Gly-Phe-p-nitroanilide (Gly-Phe-pNA). The reaction mixture, comprising 100 μl 3 mM Gly-Phe-pNA substrate, 100 μl appropriately diluted enzyme solution, 500 μl 100 mM Tris-HCl buffer (pH 8.0) and 300 μl water, was incubated at 303 K for 120 min. The reaction was stopped by the addition of 50 μl of 100% trichloroacetic acid and centrifuged at 20,000 × g for 5 min; the extent of hydrolysis was measured by detecting the absorbance at 385 nm. The activity of the incubation mixtures was measured at a final substrate concentration of 0.3 mM and expressed as the mean of three different experiments. The relative activities of the mutant DAP BIIs are summarised in Table S3. kcat and Km values could not be obtained due to the low activity of the mutant enzymes.
Product inhibition of wild-type DAP BII
The inhibitory activities of the dipeptides Val-Tyr and Asp-Arg were determined based on the hydrolysis of 0.3 mM Gly-Phe-pNA as described above. The inhibitory activities were measured at dipeptide concentrations from 0.01 to 4 mM and expressed as the means of three different experiments (Table S5).
Additional information
Accessions codes: Atomic coordinates for the reported structures have been deposited with the Protein Data Bank under accession codes 3WOI (Se-Met DAP BII), 3WOJ (peptide-free, ground-grown crystal), 3WOK (peptide-free, space-grown crystal), 3WOL (dipeptide complex I), 3WOM (dipeptide complex II), 3WON (dipeptide complex III), 3WOO (hexapeptide complex I), 3WOP (hexapeptide complex II), 3WOQ (hexapeptide complex III) and 3WOR (octapeptide complex).
References
Neurath, H. Evolution of proteolytic enzymes. Science 224, 350–357 (1984).
Antalis, T. M., Buzza, M. S., Hodge, K. M., Hooper, J. D. & Netzel-Arnett, S. The cutting edge: membrane-anchored serine protease activities in the pericellular microenvironment. Biochem. J. 428, 325–346 (2010).
Moussian, B. & Roth, S. Dorsoventral axis formation in the Drosophila embryo--shaping and transducing a morphogen gradient. Curr. Biol. 15, R887–899 (2005).
Sahu, A. & Lambris, J. D. Structure and biology of complement protein C3, a connecting link between innate and acquired immunity. Immunol. Rev. 180, 35–48 (2001).
Marino, G., Niso-Santano, M., Baehrecke, E. H. & Kroemer, G. Self-consumption: the interplay of autophagy and apoptosis. Nat. Rev. Mol. Cell Biol. 15, 81–94 (2014).
Rawlings, N. D., Barrett, A. J. & Bateman, A. MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 40, D343–350 (2012).
Botos, I. & Wlodawer, A. The expanding diversity of serine hydrolases. Curr. Opin. Struct. Biol. 17, 683–690 (2007).
Page, M. J. & Di, Cera E. Serine peptidases: classification, structure and function. Cell. Mol. Life. Sci. 65, 1220–1236 (2008).
Matthews, B. W., Sigler, P. B., Henderson, R. & Blow, D. M. Three-dimensional structure of tosyl-alpha-chymotrypsin. Nature. 214, 652–656 (1967).
Ogasawara, W., Kobayashi, G., Okada, H. & Morikawa, Y. Two types of novel dipeptidyl aminopeptidases from Pseudomonas sp. strain WO24. J. Bacteriol. 178, 6288–6295 (1996).
Suzuki, Y. et al. Identification of the catalytic triad of family S46 exopeptidases, closely related to clan PA endopeptidases. Sci. Rep. 4, 4292 (2014).
Banbula, A. et al. Porphyromonas gingivalis DPP-7 represents a novel type of dipeptidylpeptidase. J. Biol. Chem. 276, 6299–6305 (2001).
Ohara-Nemoto, Y. et al. Asp- and Glu-specific novel dipeptidyl peptidase 11 of Porphyromonas gingivalis ensures utilization of proteinaceous energy sources. J. Biol. Chem. 286, 38115–38127 (2011).
Rasmussen, H. B., Branner, S., Wiberg, F. C. & Wagtmann, N. Crystal structure of human dipeptidyl peptidase IV/CD26 in complex with a substrate analog. Nat. Struct. Biol. 10, 19–25 (2003).
Hiramatsu, H. et al. The structure and function of human dipeptidyl peptidase IV, possessing a unique eight-bladed beta-propeller fold. Biochem. Biophys. Res. Commun. 302, 849–854 (2003).
Thoma, R. et al. Structural basis of proline-specific exopeptidase activity as observed in human dipeptidyl peptidase-IV. Structure. 11, 947–959 (2003).
Engel, M. et al. The crystal structure of dipeptidyl peptidase IV (CD26) reveals its functional regulation and enzymatic mechanism. Proc. Natl. Acad. Sci. U. S. A. 100, 5063–5068 (2003).
Soisson, S. M. et al. Structural definition and substrate specificity of the S28 protease family: the crystal structure of human prolylcarboxypeptidase. BMC Struct. Biol. 10, 16 (2010).
Bezerra, G. A. et al. Structures of human DPP7 reveal the molecular basis of specific inhibition and the architectural diversity of proline-specific peptidases. PLoS One 7, e43019 (2012).
Bode, W. & Schwager, P. The refined crystal structure of bovine beta-trypsin at 1.8 A resolution. II. Crystallographic refinement, calcium binding site, benzamidine binding site and active site at pH 7.0. J. Mol. Biol. 98, 693–717 (1975).
Polgar, L. The catalytic triad of serine peptidases. Cell. Mol. Life Sci. 62, 2161–2172 (2005).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–549 (2010).
Molgaard, A. et al. The crystal structure of human dipeptidyl peptidase I (cathepsin C) in complex with the inhibitor Gly-Phe-CHN2. Biochem. J. 401, 645–650 (2007).
James, M. N., Sielecki, A. R., Brayer, G. D., Delbaere, L. T. & Bauer, C. A. Structures of product and inhibitor complexes of Streptomyces griseus protease A at 1.8 A resolution. A model for serine protease catalysis. J. Mol. Biol. 144, 43–88 (1980).
Perona, J. J. & Craik, C. S. Structural basis of substrate specificity in the serine proteases. Protein Sci. 4, 337–360 (1995).
Henderson, R. Structure of crystalline alpha-chymotrypsin. IV. The structure of indoleacryloyl-alpha-chyotrypsin and its relevance to the hydrolytic mechanism of the enzyme. J. Mol. Biol. 54, 341–354 (1970).
Robertus, J. D., Kraut, J., Alden, R. A. & Birktoft, J. J. Subtilisin; a stereochemical mechanism involving transition-state stabilization. Biochemistry 11, 4293–4303 (1972).
Singh, N. et al. Detection of native peptides as potent inhibitors of enzymes. Crystal structure of the complex formed between treated bovine alpha-chymotrypsin and an autocatalytically produced fragment, IIe-Val-Asn-Gly-Glu-Glu-Ala-Val-Pro-Gly-Ser-Trp-Pro-Trp, at 2.2 angstroms resolution. FEBS J. 272, 562–572 (2005).
Meyer, E. F., Jr, Radhakrishnan, R., Cole, G. M. & Presta, L. G. Structure of the product complex of acetyl-Ala-Pro-Ala with porcine pancreatic elastase at 1.65 A resolution. J. Mol. Biol. 189, 533–539 (1986).
Prasad, L., Leduc, Y., Hayakawa, K. & Delbaere, L. T. The structure of a universally employed enzyme: V8 protease from Staphylococcus aureus. Acta Crystallogr. D Biol. Crystallogr. 60, 256–259 (2004).
Sandler, B., Murakami, M. & Clardy, J. Atomic Structure of the Trypsin-Aeruginosin 98-B Complex. J. Am. Chem. Soc. 120, 595–596. (1998).
Kraut, J. Serine proteases: structure and mechanism of catalysis. Annu. Rev. Biochem. 46, 331–358 (1977).
Rouf, S. M. et al. Discrimination based on Gly and Arg/Ser at position 673 between dipeptidyl-peptidase (DPP) 7 and DPP11, widely distributed DPPs in pathogenic and environmental gram-negative bacteria. Biochimie 95, 824–832 (2013).
Rouf, S. M. et al. Phenylalanine 664 of dipeptidyl peptidase (DPP) 7 and Phenylalanine 671 of DPP11 mediate preference for P2-position hydrophobic residues of a substrate. FEBS Open. Bio. 3, 177–183 (2013).
Bostanci, N. & Belibasakis, G. N. Porphyromonas gingivalis: an invasive and evasive opportunistic oral pathogen. FEMS. Microbiol. Lett. 333, 1–9 (2012).
Brooke, J. S. Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clin. Microbiol. Rev. 25, 2–41 (2012).
Lambais, M. R., Goldman, M. H., Camargo, L. E. & Goldman, G. H. A genomic approach to the understanding of Xylella fastidiosa pathogenicity. Curr. Opin. Microbiol. 3, 459–462 (2000).
Hayashi, C. et al. Porphyromonas gingivalis accelerates inflammatory atherosclerosis in the innominate artery of ApoE deficient mice. Atherosclerosis 215, 52–59 (2011).
Sakamoto, Y. et al. Crystallization and preliminary X-ray crystallographic studies of dipeptidyl aminopeptidase BII from Pseudoxanthomonas mexicana WO24. Acta Crystallogr. F Struct. Biol. Commun. 70, 221–224 (2014).
Garcia-Ruiz, J. M. & Morena, A. Investigations on protein crystal growth by the gel acupuncture method. Acta Crystallogr. D Biol. Crystallogr. 50, 484–490 (1994).
Takahashi, S. et al. JAXA protein crystallization in space: ongoing improvements for growing high-quality crystals. J. Synchrotron. Radiat. 20, 968–973 (2013).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods in Enzymol. 276, 307–326 (1997).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
Lamzin, V. S. & Wilson, K. S. Automated refinement of protein models. Acta Crystallogr. D Biol. Crystallogr. 49, 129–147 (1993).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta. Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Collaborative, Computational Project & Number The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995).
McRee, D. E. XtalView/Xfit--A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Acknowledgements
We thank Drs. Y. Yamada and N. Matsugaki of the Photon Factory and Dr. K. Hasegawa of SPring-8 for their help with data collection at the synchrotron facilities. This study was supported in part by the Platform for Drug Discovery, Informatics and Structural Life Science (to N.T.), the Program for the Strategic Research Foundation at Private Universities from the MEXT of Japan (to N.T. & Y.Sa) and a Grant-in-Aid for Scientific Research(C) (to Y.Sa.). This study was also supported in part by “High-Quality Protein Crystal Growth Experiment on JEM” promoted by JAXA (Japan Aerospace Exploration Agency) (to Y.Sa.). The Russian Spacecraft “Progress” and “Soyuz” provided by the Russian Federal Space Agency were used for space transportation. Some of the space crystallisation technology was developed by ESA (the European Space Agency) and the University of Granada.
Author information
Authors and Affiliations
Contributions
Y.Sa. crystallised the protein, collected X-ray data, analysed the data and wrote the paper; Y.Su. cloned the construct, performed biochemical analyses and wrote the paper; I.I., C.T., S.R. and M.F. crystallised the protein, collected X-ray data and analysed the data; K.I., H.T., M.M. and K.O. supported the crystallisation of the protein under microgravity; H.O., T.N., Y.M. and K.T.N. were involved in the study design; W.O. and N.T. designed the study, analysed the data and wrote the paper. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sakamoto, Y., Suzuki, Y., Iizuka, I. et al. S46 Peptidases are the First Exopeptidases to be Members of Clan PA. Sci Rep 4, 4977 (2014). https://doi.org/10.1038/srep04977
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04977
This article is cited by
-
Bioprocess optimization platform for valorization of poly(lactic)-based bioplastic waste using PLA-degrading actinobacteria, Saccharothrix sp. MY1 cultured in silk wastewater as low-cost nutrient source
Biomass Conversion and Biorefinery (2022)
-
Structural basis for an exceptionally strong preference for asparagine residue at the S2 subsite of Stenotrophomonas maltophilia dipeptidyl peptidase 7
Scientific Reports (2021)
-
Fragment-based discovery of the first nonpeptidyl inhibitor of an S46 family peptidase
Scientific Reports (2019)
-
Bacterial protease uses distinct thermodynamic signatures for substrate recognition
Scientific Reports (2017)
-
Structural and mutational analyses of dipeptidyl peptidase 11 from Porphyromonas gingivalis reveal the molecular basis for strict substrate specificity
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.