Abstract
While there is a rich literature on transcription dynamics during the development of many organisms, protein data is limited. We used iTRAQ isotopic labeling and mass spectrometry to generate the largest developmental proteomic dataset for any animal. Expression dynamics of nearly 4,000 proteins of Xenopuslaevis was generated from fertilized egg to neurula embryo. Expression clusters into groups. The cluster profiles accurately reflect the major events that mark changes in gene expression patterns during early Xenopus development. We observed decline in the expression of ten DNA replication factors after the midblastula transition (MBT), including a marked decline of the licensing factor XCdc6. Ectopic expression of XCdc6 leads to apoptosis; temporal changes in this protein are critical for proper development. Measurement of expression in single embryos provided no evidence for significant protein heterogeneity between embryos at the same stage of development.
Similar content being viewed by others
Introduction
Xenopus laevis has a long history as a model organism for studying early vertebrate development. Landmark advances in cell, molecular and developmental biology using this animal include nuclear transfer experiments that demonstrated the totipotency of the nucleus from a differentiated cell1, the first isolation of a gene from any organism2, the first complete nucleotide sequence of a gene3 and the purification of the first eukaryotic transcription factor4.
Xenopus has been especially important for studies of early vertebrate development because in vitro fertilization yields synchronized embryos that mature outside the mother, so that embryogenesis can be easily monitored in real time. Fate maps for organ development have been determined and major regulators of these processes have been identified and characterized, providing an abundance of tissue- and organ-specific markers to track embryo formation. Development of embryos is rapid (the gastrula stage is reached 9 hours post fertilization and a nearly fully developed nervous system at 4 days). The relatively large size of the egg (>1.2 mm) allows for facile microinjection of material of all types and for microsurgery.
A number of techniques have been used to monitor changes in transcription expression during early stages of development in Xenopus5,6,7,8. These studies have revealed a number of subtle mechanisms by which the organism controls protein expression. Only maternal mRNAs are present at fertilization and during the first seven stages of development. Post-transcriptional regulation controls protein translation, both by the destruction of mRNA and by activating “masked” or dormant mRNA through polyadenylation. Zygotic transcription begins at the mid-blastula transition (MBT - developmental stage 8); only then does the embryo begin to translate its own mRNA into protein.
Although there is a rich literature on transcript analysis, the analysis of Xenopus protein expression changes during development is much more poorly documented. Many researchers have applied western blotting and other traditional methods to study protein expression changes during development9,10,11,12. However, these methods are only able to characterize expression changes for a small number of proteins and are limited to the available antibodies.
Large-scale qualitative and quantitative proteomic technologies have matured over the past two decades13,14,15,16. The vast majority of proteomic studies employ a bottom-up protocol, wherein proteins are digested to peptides. These peptides undergo one or more stages of chromatographic separation, are detected using electrospray ionization-tandem mass spectrometry and are identified through database searching algorithms17,18,19. A number of quantitative proteomic techniques have been developed based on stable isotope labeling17,20,21,22,23,24. Isobaric tags for relative and absolute quantitation (iTRAQ) is an isotopic labeling method that produces protein identification from peptide fragments and protein quantitation from low mass reporter ions at the MS/MS level22. It has several benefits. First, it can analyze up to eight pools of peptides in a single analysis (iTRAQ 8plex)24, significantly speeding comparisons of protein abundance across many samples. Second, iTRAQ labeling does not increase the complexity of parent ion spectra because the mass increase produced by each label is the same. Since its introduction, iTRAQ has been widely used for quantitative proteomics25,26.
There have been only a few reports determining protein expression changes in model systems of animal developmental biology27, including Drosophila28, Danio29 and Xenopus30,31 embryos and mouse oocytes32. In this work, we employed iTRAQ-8plex chemistry to monitor protein expression kinetics of Xenopuslaevis embryos at six early stages of development. We generated quantitative data on the expression changes of nearly 4,000 proteins during four or more stages of Xenopus development. These are by far the largest Xenopus proteomic dataset and the largest dataset on developmental proteomics for any organism. All the data related to this work was deposited in Peptide Atlas33, http://www.peptideatlas.org/PASS/PASS00436.
The mRNA expression of roughly 5,000 genes has been detected in Xenopuslaevis early development (stages 2–33)8. Our observation of expression changes of nearly 4,000 proteins suggests that our dataset includes a large fraction of the proteome of the organism at early stages of development.
This work included single zygotes in two iTRAQ experiments. The resulting datasets are by far the largest protein expression studies performed on single cells.
Results and discussion
Deep proteome analysis of Xenopus laevis embryos
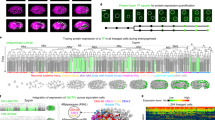
We performed three independent iTRAQ experiments on Xenopuslaevis embryos during early development. Micrographs of embryos at the stages of development employed in these experiments are shown in Fig. 1A and the experimental design is shown in Fig. 1B. Experiments I and II each employed single embryos and 8plex chemistry. In Experiment I (E1), single embryos at stages 1, 5, 8 and 11 were used; two embryos from each stage were analyzed in separate iTRAQ channels as biological duplicates. In Experiment II (E2), single embryos from stages 1, 5, 13 and 22 were used; again, two embryos from each stage were analyzed as biological duplicates. To provide more material for analysis and to average any embryo-to-embryo heterogeneity in protein expression, experiment III (E3) employed homogenates of four embryos each at stages 1, 8, 13 and 22, which were labeled using four channels of 8plex chemistry. Following digestion with trypsin and subsequent labeling using iTRAQ chemistry, the pooled peptides from each of the experiments were separated into 20 fractions by strong cation exchange liquid chromatography, followed by UPLC-ESI-MS/MS analysis with a Q-Exactive mass spectrometer. The .raw files were converted to .mgf files with RAW2MSM software34 using the default settings, followed by ProteinPilot™ 4.5 (AB Sciex) analysis with database searching as “Thorough”. Because RAW2MSM software filters out relatively low signal intensity fragment ions in tandem spectra, peptide identification is improved.
Experimental design.
Micrographs of Xenopus laevis embryos at developmental stages used for iTRAQ measurements (A). Design and workflow of three independent iTRAQ experiments (B). Three experiments were performed. In experiment 1 (E1), two embryos at stage 1, two embryos at stage 5, two embryos at stage 8 and two embryos at stage 11 were separately lysed and digested with trypsin. The first embryo at stage 1 was labeled with the iTRAQ reagent channel 113, the second embryo at stage 1 was labeled with the iTRAQ reagent channel 114, the two embryos at stage 5 were labeled with the iTRAQ reagents channels 115 and 116, the two embryos at stage 8 were labeled with the iTRAQ reagent channels 117 and 118 and the two embryos at stage 11 were labeled with the iTRAQ reagent channels 119 and 121. These labeled peptides were pooled, subjected to strong cation exchange chromatography fractionation and each fraction was analyzed using reversed-phase liquid chromatography and detection with a Q-Exactive mass spectrometer. Tandem mass spectra were analyzed both to identify the peptide and to quantitate the abundances of each peptide from each of the eight embryos. A similar procedure was performed in experiment 2 (E2), except that the biological duplicates consisted of single embryos taken from stages 1, 5, 13 and 22. Finally, experiment 3 (E3) employed four pools of four embryos, where each pool was taken embryos at stages 1, 8, 13 and 22.
Roughly 1 million MS/MS spectra were acquired over 150 hours in the three experiments, identifying an average of 3,403 ± 157 proteins per experiment, with a protein-level false discovery rate (FDR) < 1% (Supplementary Fig. S1A). The union of the three experimental results generated a total of 4,455 non-redundant protein features, which is the largest Xenopuslaevis proteome dataset generated to date. The mRNA from ~5,000 genes has been detected at early stages of X. laevis development8. Our dataset captures proteins translated from 90% of the genes listed in the published mRNA dataset of Xenopus embryos at these early developmental stages.
When we filtered the peptide identification with 95% confidence (peptide-level FDR as ~0.5%), 23,618 (E1), 21,956 (E2) and 29,422 (E3) distinct peptides were identified; the union of three experiments produced 36,977 peptides with confidence higher than 95%. Peptides with 95% or higher confidence are candidates for use in single/multiple reaction monitoring experiments (SRM/MRM); the large peptide datasets generated in this work can be used as a library for further targeted proteomic analysis35. SRM/MRM analysis can replace western blotting, which is particularly valuable for high throughput quantitation of selected protein and for quantifying proteins for which no antibody is available. Protein and peptide lists are provided in supplementary material II.
We analyzed the proteins using the DAVID Bioinformatics Resource 6.736. The official gene symbols were obtained for 4,065 proteins; 390 putative proteins have not yet been annotated. We determined the biological process, molecular function and cellular localization of these proteins (Supplementary Fig. S1 B–D). The highly enriched biological processes (Supplementary Fig. S1B) are related to DNA replication, transcription, translation, energy generation and post-translational operations (i.e. protein transport, protein localization and protein folding). The highly ranked molecular functions (Supplementary Fig. S1C) are related to nucleotide, nucleoside and RNA binding; helicase activity; ATP binding and ATPase activity; ribosome structural constituents; and protein transporter activities. Biological processes and molecular functions reflect cell division, transcription and protein expression and organization during the early stages of development. Cellular localization information of the proteins (Supplementary Fig. S1D) mirrors the biological processes; many proteins are associated with ribonucleoprotein (RNP) complexes, ribosome, mitochondrion, nuclear pore, Golgi-associated vesicle, translation initiation factors and proton-transporting ATP-synthase complex. The DAVID bioinformatics results are also included in supplementary material II.
Expression kinetics of 3,983 proteins during early Xenopus development
Nearly 90% of the identified proteins were also quantified in the three iTRAQ experiments. The high quantitation percentage is a result of high iTRAQ labeling efficiency and high specificity (100% at lysine and 99.33% at N-termini for the first 20,000 peptide matches per experiment). There were no identified labels on Tyr, His, Ser, or Thr residues in the 20,000 peptide sets. Quantitation was further enhanced by the Q-Exactive mass spectrometer, which generated high intensity iTRAQ reporter ion signals (Supplementary Fig. S2). For charge 2 and 3 parent ions, the iTRAQ reporter ions were quite intense (Supplementary Fig. S2A); the median reporter ion signal intensities range from 5.2E + 04 (channel 116) to 9.6E + 04 (channel 121) for E1 data, Supplementary Fig. S2B, which helps ensure high quality quantitation estimates.
2,889 (E1), 2,948 (E2) and 3,259 (E3) proteins were quantified in the three independent iTRAQ experiments, with identification FDRs of less than 1%. The union of the results for the three experiments yields quantitation data for 3,983 proteins, where each protein generated expression information at four or more embryonic stages (supplementary material III). This is by far the largest database of protein expression kinetics during the embryonic development of any organism. We were able to calculate pValues for more than 80% of our reported quantitations. Using a target-decoy approach by comparing ratios on duplicate iTRAQ channels, the calculated FDR for our quantitation data is less than 20%.
Biological replicates provide insight on the precision of the protocol. E1 and E2 generate an iTRAQ replicate based on the ratios between stage 5 and stage 1 from the two experiments (Supplementary Fig. S3A). The log2 ratio distribution is very narrow with a standard deviation of 0.19. E1 and E3 generate another biological replicate based on stage 8 embryos; E2 and E3 yield two more replicates based on stages 13 and 22. The excellent correlations between those replicates (slope 0.91–0.99; r 0.96–0.99) further confirm the method's reproducibility (Supplementary Fig. S3B).
The distributions of iTRAQ ratios are plotted in Supplementary Fig. S4. In all cases, the means of the log2 protein ratio distributions are very close to 0, which demonstrates that total protein abundances vary little during early development. These results are consistent with our BCA data (Supplementary Table S1). The protein abundance ratio distributions become broader at later stages (stage 11 (E1) and 22 (E2, E3)) as more proteins undergo a significant expression change.
Cluster analysis
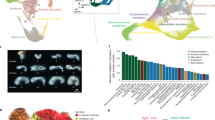
We performed cluster analysis based on abundance changes during development for all the quantified proteins from the three experiments (E1–E3), segregating proteins into those with no significant expression change (cluster 0) and into six others that showed significant changes (clusters 1–6). Fig. 2 presents the clustering results from experiment E3; a total of 3,259 proteins were quantified in this experiment. Biological process information for the proteins in clusters 1–6 is presented in Supplementary Fig. S5. Results for E1 and E2 are presented in Supplementary Fig. S6–S9 and are also listed in supplementary material III.
Cluster analysis of quantified proteins.
Proteins with significant change of abundance are grouped into one of six clusters according to the changes in expression as a function of developmental stage. Log2(protein abundance ratio) from experiment 3 (E3) was used for cluster analysis. The number of clusters was fixed at 6. The upper regulation threshold for the log2 data was 0.26 and lower threshold was −0.32 corresponding to the original ratios of 1.2 and 0.8.
Proteins with expression ratios higher than 1.2 or lower than 0.8 are considered as significantly up- and down-regulated, corresponding to log2 values of regulation thresholds as 0.26 and −0.32, respectively. 2,028 proteins showed no significant expression changes in experiment E3 (cluster 0), whereas 1,231 proteins were recognized as significantly up- or down-regulated during early development (clusters 1–6).
The cluster profiles accurately reflect the three major events that mark changes in gene expression patterns during early Xenopus development. Owing to the rapid cell divisions that occur immediately following fertilization, protein synthesis in early stage embryos relies on maternal transcripts that accumulated during oogenesis. After 12 cell divisions, the cell cycle lengthens and zygotic transcription begins during stage 8 at what is known as the midblastula transition (MBT). Many mRNAs synthesized in the oocyte are dormant or “masked” and become activated by cytoplasmic polyadenylation upon hormone-dependent maturation of the egg or upon fertilization37. Organ and system specific proteins appear following the transcriptional reprogramming that occurs at stage 13.
Clusters 1 and 2
Proteins in clusters 1 and 2 are down-regulated during early development. These proteins are maternal in origin and their functions are mostly limited to oocytes and early stage embryos. The difference between these two groups is the time at which levels begin to decline. Proteins in cluster 1 decrease immediately after fertilization; whereas, those in cluster 2 remain constant up to the MBT and then decline. Proteins found in cluster 1 include those that bind to and mediate translational repression of maternal mRNAs in Xenopus oocytes such as the Y box proteins (frgy 2 a/b) and Zygote arrest protein38,39 as well as proteins such as Maskin and RAP55 that are constituents of the protein complex that binds to the cytoplasmic polyadenylation element and controls activation of masked mRNA through lengthening the poly(A) tail10,40. Since this mechanism of translational control is limited to oocytes, we can easily rationalize the loss of these proteins following fertilization.
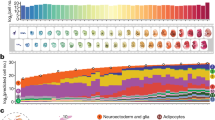
Recently, it was demonstrated that the lengthening of the cell cycle and the resulting asynchronous cell division that begins at the MBT is due, at least in part, to a decline in the amounts of four replication factors (Cut5, RecQ4, Treslin and Drf1) at the MBT41. Overexpression of these four proteins in Xenopus embryos resulted in a two-fold increase in DNA replication that could be correlated with increased initiation. We, likewise, detect a decrease in the amounts of RecQ4 and Cut5 at the MBT that places them in cluster 2; Treslin and Drf1 were not detected, most likely due to their low abundance. However, we also observe a much more pronounced decline during this period in the level of the replication licensing factor, XCdc6 (Fig. 3A). We confirmed the changes in XCdc6 by western blot (Fig. 3B). Cdc6 acts early in the initiation of replication by binding to the origin recognition complex and recruiting Mcm2–7 to form the licensing or prereplicative complex; the other four factors are active later during kinase-dependent formation of the replisome.
Expression levels of DNA replication factors and histones.
The expression levels of several DNA replication factors and histones change at the mid-blastula transition. (A) Xcdc6 exhibits a marked decline after the mid-blastula transition relative to other replication factors. (B) Levels of Cdc6 protein measured by western blot for control embryos and embryos injected with Cdc6 mRNA (1 ng). (C) Overexpression of Cdc6 triggers increased levels of apoptosis. Apoptosis was detected by staining late blastula (stage 9) embryos with PSS-380 for the fluorescent detection of phosphatidylserine. Left panel is water-injected control embryo; center panel is Cdc6 mRNA (1 ng) injected embryo; right panel is embryo incubated with 0.1 μM staurosporine to induce apoptosis. (D) Levels of maternal histone H1 (H1M/B4) decline while levels of adult H1 and core histones increase. Data from experiments E1 and E3 were used to generate (A) and (D) and the error bars for stage 8 were based on the results from experiments E1 and E3.
We questioned whether the marked decline in XCdc6 also accounts for lengthening of the cell cycle at the MBT. Injection of mRNA encoding XCdc6 into one-cell embryos results in elevated levels of the factor that persist beyond stage 13 (Fig. 3B). However, overexpression of Xcdc6 did not lead to a measureable increase in replication (results not shown). Caspase cleavage of Cdc6 can generate truncated forms of the factor that are incorporated into the prereplicative complex, but cannot support the recruitment of the Mcm complex42. This incorporation, in turn, activates p53-mediated apoptosis. We asked whether overexpression of the licensing factor could be triggering a similar response in Xenopus embryos. Notably, the combined overexpression of Cut5, RecQ4, Treslin and Drf1 inhibits gastrulation and also leads to cell death at stage 1741. Embryos injected with XCdc6 mRNA were compared to water injected controls and to embryos incubated with the pro-apoptotic drug, staurosporine43. Stage 9 embryos were incubated with a stain (PSS-380) for phosphatidylserine, an early marker of apoptosis (Fig. 3C). Embryos overexpressing XCdc6 exhibit an appreciable amount of apoptosis that would mask detection of any increase in replication. The immediate effect of XCdc6 overexpression relative to the other replication factors may reflect their different roles (licensing vs. initiation) in replication or the means through which they signal replication defects. Notwithstanding this point, iTRAQ accurately identified a critical change in the amounts of a licensing factor that triggers apoptosis when perturbed (stage 8).
We note that two XCdc6 isoforms (XCdc6A and XCdc6B) have been identified and XCdc6A is down-regulated after the MBT44. The two isoforms, which are named as XCdc6 protein and Cdc6B protein in the protein database, can be placed into two different protein groups in the protein identification lists (supplementary material II). However, in the protein quantitation list (supplementary material III), only XCdc6 protein (isoform A) was confidently quantified, which is most likely due to the two different data analysis pipelines employed for protein identification and quantitation in this work. In addition, the XCdc6A expression kinetics discovered in this work also agree with results from the Coleman group44.
Clusters 3 and 5
Clusters 3 and 5 are comprised of over 300 proteins, including G10 protein and Stat1, encoded by maternal mRNAs that become activated upon fertilization. These proteins are classic examples of the discordance between transcript and protein expression. This discordance arises due to an appreciable amount of post-transcriptional regulation that controls expression of maternal mRNAs beyond the MBT.
The notable difference between cluster 3 and 5 is that zygotic transcription appears to sustain protein levels in the former; whereas, a marked decline in expression of proteins in cluster 5 is clearly coincident with the resumption of transcription and is presumably due to the decay of maternal transcripts. The diminished protein levels in cluster 5 coincide with progression of the embryo into the gastrula stage when organization of the three primary germ layers occurs, so the reduced levels of proteins in cluster 5 can also be a consequence of restricted, tissue-specific expression as more cells become differentiated, which is indeed the case for Stat145.
Cluster 6
Cluster 6 is by far the largest in our dataset (351 proteins). Expression increases at the MBT for proteins in this cluster, which would indicate that this heightened expression results from the initiation of zygotic transcription at the MBT. For many of these proteins, this simple scenario appears correct, ranging from very large increases in mRNA levels (e.g., alkaline phosphatase, lin28a) to more moderate changes (e.g., fructose-1,6-bisphosphatase, fus, YAP, U2 auxiliary factor 2).
However, comparison of protein and mRNA levels reveal a far greater complexity for this cluster that, in some cases, reflects contributions from both maternal and zygotic transcripts. Unlike clusters 3 and 5, some members of cluster 6 appear to be expressed initially from maternal transcripts whose activation is delayed until the MBT. For the BMP signaling agonist, Twisted gastrulation (xTsg), maternal transcripts decrease and zygotic transcripts increase well after the MBT at late gastrulation46; yet, we measure a continuous increase of xTsg protein from the MBT through stage 22. Thus, there must be a coordinated transition from one pool of mRNA to the other that likely reflects the increasingly restricted, tissue-specific expression of the protein following gastrulation. Fibronectin, which is also a member of cluster 6, provides a particularly good example of delayed activation of a maternal transcript. In accord with our data, Lee et al. measured a pronounced increase in the synthesis of fibronectin beginning around the MBT that is unaffected by inhibition of zygotic transcription using α-amanitin47, demonstrating that some maternal mRNAs are activated even as zygotic transcription is initiated.
Several members of cluster 6 exhibit a disparity between mRNA and protein levels that provide additional evidence for post-transcriptional regulation. In these instances there is little change in mRNA levels that can account for the elevated levels of protein (i.e., RAC1, aldehyde dehydrogenase 1, Ssbp2, Snap29, chromobox homolog 3). An especially important lesson from these data is that increased protein expression that commences at the MBT should not be immediately attributed to zygotic transcription. Regulation of maternal transcripts at or even beyond the MBT can be used to transition into zygotic gene expression patterns. While there appears to be little regulation of protein expression by miRNAs in Xenopus oocytes and early embryos48, it would not be surprising if some members of cluster 6 are controlled by this mechanism as well.
The amount of RNA-binding protein VgRBP71 shows a pronounced increase beginning at the MBT. VgRBP71 is associated with localized mRNAs in Xenopus oocytes; in somatic cells isoforms of the protein function in alternative mRNA splicing, mRNA decay and in transcriptional regulation49. The prominent increase during gastrulation (from stage 8 to 13) is confirmed in a western blot analysis of staged embryos (Supplementary Fig. S10).
Cluster 4
Major changes in gene expression occur at the gastrula-neurula transition (stage 13), which immediately precedes organogenesis50,51. This period should mark the appearance of tissue specific proteins, which is clearly reflected in cluster 4 where a constant level of protein through the first 15 hours of embryogenesis is followed by a marked increase beginning at stage 13. Proteins in this cluster that reflect the imminent anatomical changes include: globin Y, multiple skeletal troponins, skeletal myosins, myosin 10 (neural cells), neurolfilament protein, SPARC/osteonectin (cilia cells) and Na+-K+-ATPase (kidney).
Histone variants
We also identified eight histone variants that show a significant expression change during development, Fig. 3D. Two variants, H1M (B4 protein) and H2A.X-F52 (histone cluster 1, H2aa), undergo a similar down-regulation at stage 11. Six histone variants (H1A, H1B, H2A, H2B, H3 and H4) show an increase in expression beginning at stage 8. These data suggest that H2A.X-F is a maternal histone H2 subtype due to its similar expression profile to H1M, which has been shown to be a maternal histone H1 subtype53. These data also suggest that histone H1M and H2A.X-F are the main histone variants before the MBT and that they are gradually replaced by H1A, H1B and H2A, at later stages of development. Because H1M binds to nucleosomal core and linker DNA with much lower affinity than somatic H1 variants54, replacement of H1M with somatic linker histones fixes the position of nucleosomes on the DNA55, resulting in chromatin compaction and formation of higher-order structure56. The data support the hypothesis that the higher-order chromatin structure established during embryogenesis56 restricts the binding of initiation factors to the DNA, limiting origin use57.
Heterogeneity of protein expression in single Xenopuslaevis embryos
Experiments E1 and E2 employed a single embryo for each iTRAQ channel and biological duplicates were included for each embryonic stage, Fig. 1B. These results allow us to investigate stochastic variation in protein expression between embryos at the same stage of development at the single embryo level.
We first calculated the Expression Ratio for proteins quantitated in two embryos in the same experiment at the same stage of development, eq. 1:

where the ratio is calculated for each protein at the nth stage of development. The distributions of log2(Expression Ratio) for embryos are presented in Supplementary Fig. S11. These distributions are centered at zero, corresponding to equal average expression in the two embryos, at each stage of development. The standard deviations of the log2 distributions are ~0.25, corresponding to a ~15% relative standard deviation in protein expression; embryo-to-embryo differences in protein expression at the same stage of development are very low in Xenopuslaevis.
The small variation that was observed in the expression of some proteins could either be due to noise in the experimental protocol or due to noise in protein expression. If the variation is due to experimental noise, then the variation should not be correlated between experiments E1 and E2. We used the standard deviation of the absolute value of log2(Expression ratio) for each protein in each experiment as our measure of variation (Supplemental figure S12). This arithmetic procedure generates the standard deviation of the relative change and ensures that expression ratio always has the larger value in the numerator. The standard deviations are clustered at very low levels and are essentially uncorrelated (r = 0.40), which supports the interpretation that experimental noise rather than biological processes dominate any embryo-to-embryo variations in protein expression.
Single cell proteomics
Embryos at stage 1 are zygotes that have not undergone division; they are single cells. While there have been efforts to characterize protein expression in single cells, the data presented here for Experiments E1 and E2 include the largest dataset for single cell proteomics58,59,60. These amphibian zygotes are roughly three orders of magnitude larger than mammalian zygotes and six orders of magnitude larger than somatic cells. Significant instrumental advances will be required to obtain a similar proteomic depth on those cells. Recent advances in electrospray technology and on-column digestion provide some hope that comprehensive single cell proteomics may be extended to those cells61,62.
Methods
Materials and reagents
Bovine pancreas TPCK-treated trypsin, urea, ammonium bicarbonate (NH4HCO3), dithiothreitol (DTT) and iodoacetamide (IAA) were purchased from Sigma–Aldrich (St. Louis, MO). Acetonitrile (ACN) and formic acid (FA) were purchased from Fisher Scientific (Pittsburgh, PA). Water was deionized by a Nano Pure system from Thermo scientific (Marietta, OH). iTRAQ 8-plex kits were purchased from AB Sciex (Foster City, CA).
Xenopuslaevis was purchased from Nasco (Fort Atkinson, WI). Mammalian Cell-PE LB™ buffer for embryo lysis was purchased from G-Biosciences (St. Louis, MO). Complete, mini protease inhibitor cocktail (provided in EASYpacks) was purchased from Roche (Indianapolis, IN).
Xenopuslaevis embryo culture and collection
All animal procedures were performed according to protocols approved by the University of Notre Dame Institutional Animal Care and Use Committee. Female Xenopuslaevis were induced to lay eggs by injection with 600 units of human chorionic (C1063, Sigma-Aldrich, St. Louis, MO) 12–15 hours prior to spawning. Testes were isolated from anesthetized males at the time of spawning. Eggs and minced testis were combined in 1/3 MMR (Marc's Modified Ringers) for fertilization and embryos were maintained at room temperature and throughout development. Embryos were collected at different stages based on the information from references 63 and 64. Eight eggs were collected separately into eight different Eppendorf tubes at stage 1. Then, six embryos were collected at stages 5, 8, 11, 13 and 22. A single embryo was collected into one Eppendorf tube.
Embryo lysis and protein sample preparation
Each collected embryo was suspended in 50 μL of mammalian cell-PE LB™ buffer containing complete protease inhibitor. After shaking for 30 s at room temperature, the tubes were sonicated for 3 min on ice with a Branson Sonifier 250 (VWR Scientific, Batavia, IL) to lyse the embryos completely. The lysates were then kept on ice for half an hour. The tubes were next centrifuged at 12,000 g for 10 min. Finally, the supernatants were collected into fresh Eppendorf tubes and stored at −80°C before use.
The embryo lysate supernatants were precipitated with 300 μL of cold acetone at −20°C for 6 h. After centrifugation, the supernatants were removed and another 300 μL of cold acetone was added to each tube to wash the pellets again. After further centrifugation, the supernatants were removed and protein pellets were dried at room temperature.
The pellet in each tube was dissolved in 30 μL of 8 M urea, 100 mM NH4HCO3 (pH 8.0) buffer via vortex with sonication. After centrifugation, an 8 μL aliquot of protein solution was taken from each tube and further diluted to 24 μL with 100 mM NH4HCO3 (pH 8.0), followed by protein concentration measurement using the bicinchoninic acid (BCA) method65. For experiment I (E1), two embryos from stage 1, 5, 8 and 11 were used and each embryo lysate was further prepared individually. For experiment II (E2), two embryos from stage 1, 5, 13 and 22 were used and each embryo lysate was also further prepared individually. For experiment III (E3), the mixtures of four embryos from stage 1, 8, 13 and 22 were used for further preparation. All the samples were denatured at 37°C for 1 h, followed by protein reduction in 20 mM DTT at 56°C for 1 h and alkylation in 50 mM IAA at room temperature for 30 min in dark. The treated samples were further diluted four times with 100 mM NH4HCO3 (pH 8.0) to reduce the urea concentration to about 2 M, followed by trypsin digestion at 37°C overnight with trypsin/protein mass ratio as 1/25.
The digests were acidified with formic acid to terminate the tryptic digestion, followed by peptide desalting with C18 spin columns (Pierce Biotechnology, Rockford, IL) for E1 and E2 samples and with Sep-Pak C18 1 cc Vac Cartridge (Waters Corporation, Milford, MA) for E3 samples. After lyophilization, the peptides were labeled by ‘isobaric tags for relative and absolute quantitation’ (iTRAQ) 8-plex reagents according to the manufacturer's protocols (AB Sciex, Foster City)22 with the following minor modifications. The lyophilized digests for E1 and E2 were dissolved in 12.5 μL of dissolution buffer and the digests for E3 were dissolved in 35 μL of dissolution buffer. After addition of 50 μL of isopropanol to each iTRAQ reagent vial, 25 μL of iTRAQ reagent was added to the digest for E1 and E2 and an entire vial of iTRAQ reagent was used to label the digests for E3. After labeling at room temperature for 2 hours, 35 μL of 100 mM Tris-HCl buffer (pH 8.0) was added to the samples for E1 and E2 and incubated at room temperature for 40 min to block the residual iTRAQ reagents. For E3 samples, 100 μL of 100 mM Tris-HCl buffer (pH 8.0) was used. Then, the labeled samples in each experiment were mixed and three tubes of labeled digests (E1, E2 and E3) were obtained. After lyophilization, the samples were dissolved in 500 μL (E1 and E2) or 800 μL (E3) of 2% ACN and 0.1%FA solution, followed by desalting with Sep-Pak C18 1 cc Vac Cartridge (Waters). The digests were lyophilized again and then redissolved in 250 μL of 0.1% FA, followed by strong cation exchange (SCX) liquid chromatography fractionation.
SCX fractionation
The labeled samples from three experiments (E1, E2 and E3) were fractionated by SCX liquid chromatography using a Waters Alliance HPLC system (Waters, Milford, MA, USA) at a flow rate of 0.25 mL/min. About 150 μL of the labeled peptide samples was loaded onto an SCX guard column (4.6 mm i.d. × 12.5 mm length, Agilent Technologies, Wilmington, DE, USA) and then separated with a Zorbax 300-SCX column (2.1 mm i.d. × 50 mm length, 5 μm particles, Agilent Technologies). The mobile phase gradient was generated using buffer A (10 mM KH2PO4, 20% ACN, pH 2.85) and buffer B (1 M KCl in A, pH 2.85). The samples in 0.1% FA were loaded, followed by 10 min washing with 100% A to remove excess iTRAQ reagent. Then, the peptides were separated by a 25 min linear gradient from 100% A to 100% B. Finally, the column was washed by 100% B for 5 min, followed by column equilibration with 100% A. Fractions were collected from 12 min to 42 min as follows. Eluate from 12 min to 18 min was collected to one fraction and from 36 min to 42 min as one fraction. From 18 min to 36 min, the eluate was collected as 1 min/fraction. In total, 20 fractions were collected from each sample.
Peptide desalting
The collected SCX fractions were first lyophilized. Then, the fractions from E1 and E2 were redissolved in 30 μL of 2% ACN and 0.1% FA, followed by peptide desalting with a C18 ZipTip (ZTC18S096, Millipore, Bedford, MA, USA). The fractions from E3 were redissolved in 30 or 50 μL of 2% ACN and 0.1% FA, followed by desalting with a C18 ZipTip (Millipore) or a C18 spin column (Pierce Biotechnology) based on the sample amount in each fraction. The eluates were lyophilized and redissolved in 6 μL or 8 μL (E1 and E2) and 12 μL (E3) of 2% ACN and 0.1% FA solution, followed by ultra-performance liquid chromatography (UPLC)-ESI-MS/MS analysis.
UPLC-ESI-MS/MS analysis
A nanoACQUITY UltraPerformance LC® (UPLC®) system (Waters, Milford, MA, USA) was used for peptide separation. Buffer A (0.1% FA in water) and buffer B (0.1% FA in ACN) were used as mobile phases for gradient separation. Peptides were automatically loaded onto a commercial C18 reversed phase column (Waters, 100 μm × 100 mm, 1.7 μm particle, BEH130C18, column temperature 40°C) with 2% buffer B for 10 min at a flow rate of 1 μL/min, followed by 3-step gradient separation, 2 min from 2% to 8%, 114 min to 28% B, 3 min to 85% B and maintained at 85% B for 13 min. The column was equilibrated for 12 min with 2% B before analysis of the next sample. The eluted peptides from the C18 column were pumped through a capillary tip for electrospray and analyzed by a Q-Exactive mass spectrometer (Thermo Fisher Scientific). For each sample, 2 μL of peptides were used for analysis.
The electrospray voltage was 1.6 kV and the ion transfer tube temperature was 280°C. The S-Lens RF level was 50.00. The data acquisition was programmed in data dependent acquisition (DDA) mode. A top 12 method was used. Full MS scans were acquired in Orbitrap mass analyzer over m/z 380–1800 range with resolution of 70,000 (m/z 200) and the number of microscans set to 1. The target value was 1.00E + 06 and maximum injection time was 250 ms. For MS/MS scans, the twelve most intense peaks with charge state ≥ 2 were sequentially isolated and further fragmented in the higher-energy-collisional-dissociation (HCD) cell following one full MS scan. To reduce the interference of peptide co-fragmentation to the iTRAQ quantitation, the isolation window was set as 1.0 m/z. The normalized collision energy was 33% and tandem mass spectra were acquired in the Orbitrap mass analyzer with resolution 35,000 (m/z 200). The fixed first mass was m/z 100.0. The target value was 1.00E + 06 and maximum injection time was 120 ms. The number of microscans was 1 and the ion selection threshold was 5.0E + 04 counts. Peptide match and exclude isotopes were turned on. Dynamic exclusion was set as 30 s.
Data analysis
The .raw files were converted to Mascot generic format (.mgf) files via RAW2MSM software34 with default settings for deep proteome analysis and via Proteome Discoverer 1.3 (Thermo Fisher Scientific) with default settings for protein quantitation analysis. Protein Pilot™ 4.5 (AB Sciex, Foster City, CA, USA) was used for deep proteome analysis and protein quantitation analysis with .mgf files as input. Paragon™ algorithm (v. 4.5.0.0, 1654)66 integrated in the Protein Pilot™ 4.5 was used for database searching. The Xenopuslaevis database (12/28/2012 version) downloaded from the Xenbase website (ftp://ftp.xenbase.org/pub/Genomics/Sequences/). The Xenopus database was then combined with common contaminants and used for database searching. The parameters for database searching were as follows: iTRAQ 8plex (peptide labeled) was set as sample type. Iodoacetamide was set as the cysteine alkylation, trypsin as the digestion enzyme, Orbitrap MS (1–3 ppm) and Orbitrap MS/MS as instrument and urea denaturation as special factors. In addition, search effort was set as “thorough” for deep proteome analysis and “rapid” for protein quantitation. The database searching for the reversed database was also performed in order to evaluate FDRs at the peptide and protein levels67,68.
For deep proteome and protein quantitation analysis, the peptide confidence was filtered to produce a peptide level global FDR of less than 1%. On protein group level, the protein unused score was used to filter the protein identification to produce a protein level FDR less than 1%. For protein quantitation, only ratios from the spectra that were unique to each protein were used for calculation of protein ratio and only “Auto” peptides were used for protein quantitation. Bias correction was applied for protein quantitation results, which determines the median average protein ratio and corrects it to unity and then applies this factor to all quantitation results. Proteins with iTRAQ ratio higher than 20 or lower than 0.05 were not considered as quantified and only proteins with reasonable ratios across all channels were recognized as quantified ones. We obtained the final protein quantitation information based on normalization to the mean of channels 113 and 114 (biological replicate of stage 1) for E1 and E2 and the mean of iTRAQ protein ratios of biological replicates for each embryonic stage was used for further data analysis. We calculated the FDR for our quantitative data using a target-decoy approach by comparing ratios on duplicate iTRAQ channels.
DAVID Bioinformatics Resources 6.736 was used to generate the gene symbols of the Xenopuslaevis proteome, biological process, molecular function and cellular component information.
Open source software, GProX69 used to visualize protein quantitation data including histograms and clustering analysis. The log2 ratios were used for analysis. For histograms analysis, default settings were used. For clustering analysis, the number of clusters was set to 6 and fixed regulation threshold (upper limit as 0.26 and lower limit as −0.32, corresponding to the original ratios of about 1.2 and 0.8) was used. The minimal membership for plot was set as 0.5. Other parameters were default settings.
Western blot analysis
To confirm protein quantitation determined by mass spectrometry, we also performed western blot analysis for several proteins. Following SDS-PAGE, proteins were transferred to a nitrocellulose membrane overnight at 4°C and developed with antibody, produced in rabbit, specific to Xenopuslaevis VgRBP71 (generated in the Huber laboratory), actin (AHP1629, AbD Serotec, Raleigh, NC) and Cdc670 (kindly provided by the laboratory of Dr. William G. Dunphy at the California Institute of Technology).
Xcdc6 mRNA injection experiments
One cell embryos were injected with 1 ng capped Xcdc6 mRNA or an equivalent volume of H2O and allowed to developed at room temperature in 1/3 MMR solution. At stage 9, embryos were incubated for one hour in a 200 μM solution of PSS-380 (a gift from Dr. Bradley Smith, University of Notre Dame), a fluorescent probe that binds to phosphatidyl serine exposed on the surface of apoptotic cells. Apoptosis was induced in control embryos by incubation with staurosporine (0.1 μM) for one hour before staining with PSS-380. Fluorescence images were acquired using a Nikon TE-2000U epifluorescence microscope equipped with the appropriate UV filter set (ex: 340/80, em: 435/485) for blue fluorescence and edited using the Nikon Imaging Software (NIS) Elements v. 4.0.
References
Gurdon, J. B. The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J. Embryol. Exp. Morphol. 10, 622–640 (1962).
Wallace, H. & Birnstiel, M. L. Ribosomal cistrons and the nucleolar organizer. Biochim. Biophys. Acta 114, 296–310 (1966).
Miller, J. R., Cartwright, E. M., Brownlee, G. G., Fedoroff, N. V. & Brown, D. D. The nucleotide sequence of oocyte 5S DNA in Xenopuslaevis. II. The GC-rich region. Cell 13, 717–725 (1978).
Engelke, D. R., Ng, S.-Y., Shastry, B. S. & Roeder, R. G. Specific interaction of a purified transcription factor with an internal control region of 5S RNA genes. Cell 19, 717–728 (1980).
Altmann, C. R. et al. Microarray-based analysis of early development in Xenopuslaevis. Dev. Biol. 236, 64–75 (2001).
Paranjpe, S. S., Jacobi, U. G., van Heeringen, S. J. & Veenstra, G. J. A genome-wide survey of maternal and embryonic transcripts during Xenopus tropicalis development. BMC Genomics 14, 762 (2013).
Tan, M. H. et al. RNA sequencing reveals a diverse and dynamic repertoire of the Xenopus tropicalis transcriptome over development. Genome Res. 23, 201–216 (2013).
Yanai, I., Peshkin, L., Jorgensen, P. & Kirschner, M. W. Mapping gene expression in two Xenopus species: evolutionary constraints and developmental flexibility. Dev. Cell 20, 483–496 (2011).
Ginsberg, A. M., King, B. O. & Roeder, R. G. Xenopus 5S gene transcription factor, TFIIIA: Characterization of a cDNA clone and measurement of RNA levels throughout development. Cell 39, 479–489 (1984).
Stebbins-Boaz, B., Cao, Q., de Moor, C. H., Mendez, R. & Richter, J. D. Maskin is a CPEB-associated factor that transiently interacts with elF-4E. Mol. Cell 4, 1017–1027 (1999).
Hara, M. et al. Greatwall kinase and cyclin B-Cdk1 are both critical constituents of M-phase-promoting factor. Nat. Commun. 3, 1059 (2012).
Miyamoto, K. et al. Nuclear Wave1 is required for reprogramming transcription in oocytes and for normal development. Science 341, 1002–1005 (2013).
Ong, S. E. & Mann, M. Mass spectrometry-based proteomics turns quantitative. Nat. Chem. Biol. 1, 252–262 (2005).
Gstaiger, M. & Aebersold, R. Applying mass spectrometry-based proteomics to genetics, genomics and network biology. Nat. Rev. Genet. 10, 617–627 (2009).
Zhang, Y., Fonslow, B. R., Shan, B., Baek, M.-C. & Yates, J. R., III Protein analysis by shotgun/bottom-up proteomics. Chem. Rev. 113, 2343–2394 (2013).
Zhou, F. et al. Genome-scale proteome quantification by DEEP SEQ mass spectrometry. Nat. Commun. 4, 2171 (2013).
Gygi, S. P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Washburn, M. P., Wolters, D. & Yates, J. R., III Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Mann, M. & Kelleher, N. L. Precision proteomics: The case for high resolution and high mass accuracy. Proc. Natl. Acad. Sci. USA 105, 18132–18138 (2008).
Yao, X., Freas, A., Ramirez, J., Demirev, P. A. & Fenselau, C. Proteolytic 18O labeling for comparative proteomics: model studies with two serotypes of adenovirus. Anal. Chem. 73, 2836–2842 (2001).
Ong, S. E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Ross, P. L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
McClatchy, D. B., Liao, L., Park, S. K., Venable, J. D. & Yates, J. R., III Quantification of the synaptosomal proteome of the rat cerebellum during post-natal development. Genome Res. 17, 1378–1388 (2007).
Ow, S. Y. et al. Quantitative shotgun proteomics of enriched heterocysts from Nostoc sp. PCC 7120 using 8-plex isobaric peptide tags. J. Proteome Res. 7, 1615–1628 (2008).
Wolf-Yadlin, A., Hautaniemi, S., Lauffenburger, D. A. & White, F. M. Multiple reaction monitoring for robust quantitative proteomic analysis of cellular signaling networks. Proc. Natl. Acad. Sci. USA 104, 5860–5865 (2007).
Phanstiel, D. H. et al. Proteomic and phosphoproteomic comparison of human ES and iPS cells. Nat. Methods 8, 821–827 (2011).
Reintsch, W. E. & Mandato, C. A. Deciphering animal development through proteomics: requirements and prospects. Proteome Sci. 6, 21 (2008).
Zhai, B., Villén, J., Beausoleil, S. A., Mintseris, J. & Gygi, S. P. Phosphoproteome analysis of Drosophila melanogaster embryos. J. Proteome Res. 7, 1675–1682 (2008).
Lucitt, M. B. et al. Analysis of the Zebrafish proteome during embryonic development. Mol. Cell. Proteomics 7, 981–994 (2008).
McGivern, J. V., Swaney, D. L., Coon, J. J. & Sheets, M. D. Toward defining the phosphoproteome of Xenopus laevis embryos. Dev. Dyn. 238, 1433–1443 (2009).
Wang, R., Liu, X., Küster-Schöck, E. & Fagotto, F. Proteomic analysis of differences in ectoderm and mesoderm membranes by DiGE. J. Proteome Res. 11, 4575–4593 (2012).
Wang, S. et al. Proteome of mouse oocytes at different developmental stages. Proc. Natl. Acad. Sci. USA 107, 17639–17644 (2010).
Desiere, F. et al. The PeptideAtlas Project. Nucleic Acids Res. 34, D655–D658 (2006).
Olsen, J. V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Picotti, P. & Aebersold, R. Selected reaction monitoring–based proteomics: workflows, potential, pitfalls and future directions. Nat. Methods 9, 555–566 (2012).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protoc. 4, 44–57 (2009).
Villalba, A., Coll, O. & Gebauer, F. Cytoplasmic polyadenylation and translational control. Curr. Opin. Gene. Dev. 21, 452–457 (2011).
Yurkova, M. S. & Murray, M. T. A translation regulatory particle containing the Xenopus oocyte Y box protein mRNP3+4. J. Biol. Chem. 272, 10870–10876 (1997).
Wu, X., Wang, P., Brown, C. A., Zilinski, C. A. & Matzuk, M. M. Zygote Arrest 1 (Zar1) Is an Evolutionarily Conserved Gene Expressed in Vertebrate Ovaries. Biol. Reprod. 69, 861–867 (2003).
Tanaka, K. J. et al. RAP55, a Cytoplasmic mRNP Component, Represses Translation in Xenopus Oocytes. J. Biol. Chem. 281, 40096–40106 (2006).
Collart, C., Allen, G. E., Bradshaw, C. R., Smith, J. C. & Zegerman, P. Titration of Four Replication Factors Is Essential for the Xenopuslaevis Midblastula Transition. Science 341, 893–896 (2013).
Yim, H. et al. Cleavage of Cdc6 by caspase-3 promotes ATM/ATR kinase–mediated apoptosis of HeLa cells. J. Cell Biol. 174, 77–88 (2006).
Chae, H. J. et al. Molecular mechanism of staurosporine-induced apoptosis in osteoblasts. Pharmacol. Res. 42, 373–381 (2000).
Tikhmyanova, N. & Coleman, T. R. Isoform switching of Cdc6 contributes to developmental cell cycle remodeling. Dev. Biol. 260, 362–375 (2003).
Turpen, J. B., Carlson, D. L. & Huang, C. Cloning and developmental expression of Xenopus Stat1. Dev. Comp. Immunol. 25, 219–229 (2001).
Oelgeschlager, M., Larrain, J., Geissert, D. & De Robertis, E. M. The evolutionarily conserved BMP-binding protein Twisted gastrulation promotes BMP signalling. Nature 405, 757–763 (2000).
Lee, G., Hynes, R. & Kirschner, M. Temporal and spatial regulation of fibronectin in early Xenopus development. Cell 36, 729–470 (1984).
Lund, E., Sheets, M. D., Imboden, S. B. & Dahlberg, J. E. Limiting Ago protein restricts RNAi and microRNA biogenesis during early development in Xenopuslaevis. Genes Dev. 25, 1121–1131 (2011).
Kroll, T. T., Zhao, W. M., Jiang, C. & Huber, P. W. A homolog of FBP2/KSRP binds to localized mRNAs in Xenopus oocytes. Development 129, 5609–5619 (2002).
Andrews, M. T., Loo, S. & Wilson, L. R. Coordinate inactivation of class III genes during the Gastrula-Neurula Transition in Xenopus. Dev. Biol. 146, 250–254 (1991).
Howe, J. A., Howell, M., Hunt, T. & Newport, J. W. Identification of a developmental timer regulating the stability of embryonic cyclin A and a new somatic A-type cyclin at gastrulation. Genes Dev. 9, 1164–1176 (1995).
Shechter, D. et al. A distinct H2A.X isoform is enriched in Xenopus laevis eggs and early embryos and is phosphorylated in the absence of a checkpoint. Proc. Natl. Acad. Sci. U S A 106, 749–754 (2009).
Dworkin-Rastl, E., Kandolf, H. & Smith, R. C. The maternal histone H1 variant, H1M (B4 protein), is the predominant H1 histone in Xenopus pregastrula embryos. Dev. Biol. 161, 425–439 (1994).
Ura, K., Nightingale, K. & Wolffe, A. P. Differential association of HMG1 and linker histones B4 and H1 with dinucleosomal DNA: structural transitions and transcriptional repression. EMBO J. 15, 4959–4969 (1996).
Ura, K., Hayes, J. J. & Wolffe, A. P. A positive role for nucleosome mobility in the transcriptional activity of chromatin templates: restriction by linker histones. EMBO J. 14, 3752–3765 (1995).
Wolffe, A. P. Dominant and specific repression of Xenopus oocyte 5S RNA genes and satellite I DNA by histone H1. EMBO J. 8, 527–537 (1989).
Stillman, B. Cell cycle control of DNA replication. Science 274, 1659–1664 (1996).
Whittal, R. M., Keller, B. O. & Li, L. Nanoliter chemistry combined with mass spectrometry for peptide mapping of proteins from single mammalian cell lysates. Anal. Chem. 70, 5344–5347 (1998).
Neupert, S., Rubakhin, S. S. & Sweedler, J. V. Targeted single-cell microchemical analysis: MS-based peptidomics of individual paraformaldehyde-fixed and immunolabeled neurons. Chem. Biol. 19, 1010–1019 (2012).
Abiko, M. et al. Identification of proteins enriched in rice egg or sperm cells by single-cell proteomics. PLoS One 8, e69578 (2013).
Sun, L., Zhu, G. & Dovichi, N. J. Integrated capillary zone electrophoresis-electrospray ionization tandem mass spectrometry system with an immobilized trypsin microreactor for online digestion and analysis of picogram amounts of RAW 264.7 cell lysate. Anal. Chem. 85, 4187–4194 (2013).
Sun, L. et al. Ultrasensitive and Fast Bottom-up Analysis of Femtogram Amounts of Complex Proteome Digests. Angew. Chem. Int. Ed. Engl. 52, 13661–13664 (2013).
Nieuwkoop, P. D. & Faber, J. Normal Table of Xenopuslaevis (Daudin) (Garland Publishing Inc, New York) (1994).
Khokha, M. K. et al. Techniques and probes for the study of Xenopus tropicalis development. Dev. Dyn. 225, 499–510 (2002).
Smith, P. K. et al. C. Measurement of protein using bicinchoninic acid. Anal. Biochem. 150, 76–85 (1985).
Shilov, I. V. et al. The Paragon Algorithm, a next generation search engine that uses sequence temperature values and feature probabilities to identify peptides from tandem mass spectra. Mol. Cell. Proteomics 6, 1638–1655 (2007).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Rigbolt, K. T. G., Vanselow, J. T. & Blagoev, B. GProX, a user-friendly platform for bioinformatics analysis and visualization of quantitative proteomics data. Mol. Cell. Proteomics 10, O110.007450 (2011).
Coleman, T. R., Carpenter, P. B. & Dunphy, W. G. The Xenopus Cdc6 protein is essential for the initiation of a single round of DNA replication in cell-free extracts. Cell 87, 53–63 (1996).
Acknowledgements
We thank Leon Peshkin of Marc Kirshner's group at Harvard, who provided abstract 732 (Quantitative molecular embryology: coupled proteomic and transcriptomic analysis) at the 2013 ASCB conference. We thank Dr. William Boggess in the Notre Dame Mass Spectrometry and Proteomics Facility for his help with this project. We thank Dr. William G. Dunphy at the California Institute of Technology for kindly providing the antibody specific to Cdc6 and Dr. Bradley Smith (Notre Dame) for the sample of PSS-380. This project was supported by a grant from the National Institutes of Health (Grant R01GM096767).
Author information
Authors and Affiliations
Contributions
L.S., M.M.B., P.W.H. and N.J.D. designed research. L.S., M.M.B. and G.Z. performed research. L.S., M.M.B., M.M.C., P.W.H. and N.J.D. analyzed data. L.S., P.W.H. and N.J.D. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information I
Supplementary Information
Supporting material II
Supplementary Information
Supporting material III
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sun, L., Bertke, M., Champion, M. et al. Quantitative proteomics of Xenopus laevis embryos: expression kinetics of nearly 4000 proteins during early development. Sci Rep 4, 4365 (2014). https://doi.org/10.1038/srep04365
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04365
This article is cited by
-
Comparative antler proteome of sika deer from different developmental stages
Scientific Reports (2021)
-
SSRP1-mediated histone H1 eviction promotes replication origin assembly and accelerated development
Nature Communications (2020)
-
The neglected part of early embryonic development: maternal protein degradation
Cellular and Molecular Life Sciences (2020)
-
An integrated genome-wide multi-omics analysis of gene expression dynamics in the preimplantation mouse embryo
Scientific Reports (2019)
-
Dynamics of Non-Canonical Amino Acid-Labeled Intra- and Extracellular Proteins in the Developing Mouse
Cellular and Molecular Bioengineering (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.