Abstract
Heading date is a critical trait for adaptation of rice to different cultivation areas and cropping seasons. We evaluated the heading dates of 1,123 chromosome segments substitution lines (CSSLs) in the genetic background of an elite rice variety Huajingxian74 (HJX74). A CSSL with the substituted segments from Zihui100 exhibited late heading under both natural long-day (NLD) and natural short-day (NSD) conditions and the late heading phenotype was controlled by two novel epistatic loci on chromosome 8 and chromosome 3, respectively, termed LH8 and EH3. The function of EH3 was dependent on the LH8 genotype through epistatic interaction between EH3Zihui100 and LH8Zihui100 alleles. Genetic and molecular characterization revealed LH8 encodes a CCAAT-box-binding transcription factor with Heading date1 (Hd1)-binding activity and may delay flowering by repressing the expression of Early heading date1 (Ehd1). Our work provides a solid foundation for further study on gene interaction in heading date and has application in breeding rice with greater adaptability.
Similar content being viewed by others
Introduction
Rice is a short-day (SD) plant and is becoming an increasingly important model monocot plant for molecular biological study on agronomic important traits such as heading date. Heading date, flowering time in cereal crops, is a critical agronomical determinant for adaptation to specific cropping locations and growing seasons for current varieties of cultivated rice and is an important consideration for rice breeders. The molecular and developmental determinants of heading date have thus particularly important genetic targets for domestication and for breeding new varieties of rice.
The timing of flower initiation is controlled by both environmental and internal signals1,2. By taking advantage of pioneering works in long-day (LD) plant Arabidopsis, many molecular components of rice flowering process initiated from photoperiod and light signals have been identified through natural variations, quantitative trait locus (QTL) analyses and by screening mutant populations (as summarized and reviewed in3). Two independent flowering pathways, the conserved Hd1-dependent and unique Ehd1-dependent pathways, control heading time in rice4,5,6,7. The two pathways function redundantly under SD conditions but antagonistically under LD conditions8. Hd1, a rice ortholog of Arabidopsis CONSTANS (CO) that is regulated by OsGI (a rice ortholog of Arabidopsis GIGANTEA), activates flowering under SD conditions and represses it strongly under LD conditions through regulating the expression of Heading date3a (Hd3a), a rice ortholog of Arabidopsis florigen FLOWERING LOCUS T (FT)4,9. Ehd1 encodes a B-type response regulator that is highly conserved in cultivated rice, but has no homolog in Arabidopsis6. Ehd1 can promote flowering by activating the expression of Hd3a and RICE FLOWERING LOCUS T1 (RFT1), the closest paralog of Hd3a6,7,10. The expression of Ehd1 is promoted by a number of positive regulators, including RID1 (also known as OsID1 and Ehd2)11,12,13, Early heading date3 (Ehd3)14, Early heading date4 (Ehd4)15, OsMADS5116 and OsMADS50 (also known as DTH3)17,18,19. On the other hand, several factors repress the expression of Ehd1, thereby delaying flowering, such as Ghd720, DTH8 (also known as Ghd8 and LHD1)21,22,23, SE524, OsCOL425 and OsLFL126,27. Therefore, Ehd1 serves as a critical convergence point of regulation by multiple signaling pathways. The flowering signals are collected from Hd1 and Ehd1 and are transducted to a pair of florigen genes, Hd3a and RFT1, promoting heading under SD and LD conditions, respectively7,10. After production in leaves, these florigens move to the shoot apical meristem and promote floral transition by activating the expression of OsMADS14 and OsMADS157. Although these studies have revealed much insight into the flowering control of rice, the underlying molecular mechanisms, especially the genetic interactions among these factors, are still not well understood.
The advantages of CSSLs in precise and detailed phenotype evaluation, minor QTL detection and validation of gene-gene and gene-environment interactions facilitated both genetic studies and rice improvement28,29,30. Therefore, we had constructed a library of 1,123 CSSLs in rice using HJX74, an elite indica variety from south China, as recipient and 24 accessions, including 14 indica and 10 japonica, collected worldwide as donors to understand the molecular mechanism of agronomic traits including heading date31,32. Each CSSL contains a very small number of well-characterized chromosome segments from one of the 24 donor varieties.
In this study, we found a CSSL with the substituted segments from Zihui100 exhibited late heading under both NLD and NSD conditions and the late heading phenotype was controlled by two novel epistatic loci on chromosome 8 and chromosome 3, respectively, termed LH8 and EH3. Map-based cloning of LH8 revealed it encodes a CCAAT-box-binding transcription factor and is allelic to DTH8. Molecular function of LH8 was further studied by expression analysis and yeast-two-hybrid experiment.
Results
Identification of a CSSL with late heading
The procedure for the development of the CSSLs was summarized in Fig. 1. HJX74 (as the female parent) was crossed with one of the 24 donors (Zihui100 was taken as an example) and the F1 plants were backcrossed with HJX74 to develop the BC1F1 generation. Polymorphic SSR markers were used in the selection of the donor chromosomal segments. Using the same method, BC6F1 plants were obtained and were self-crossed to produce BC6F2 lines in which the majority of genomic regions were homozygous for HJX74 alleles. Totally, 1,123 such CSSLs in the HJX74 genetic background were developed31,32. To further understand how flowering in rice was controlled, we evaluated the heading date of the 1,123 CSSLs under natural growth conditions. A CSSL, CSSL5, with substituted segments from Zihui100 exhibited late heading compared with the recipient HJX74 under both NLD (114.4 ± 1.4 d for CSSL5, 100.1 ± 0.5 d for HJX74) and NSD conditions (84.5 ± 1.6 d for CSSL5, 76.7 ± 1.4 d for HJX74) in 2009 (Fig. 2a and 2b).
The heading dates of HJX74, CSSL5 and their F2 population.
(a). Phenotypes of HJX74 (left) and CSSL5 (right). Yellow bar = 15 cm. (b). Days-to-heading in HJX74 and CSSL5 in different photoperiodic conditions. NLD, natural long-day conditions; NSD, natural short-day conditions. Each column represents mean ± s.d. (n = 30). (c). Frenquency distributions of days-to-heading in an F2 population from HJX74 x CSSL5 under NLD conditions. Black arrows indicate the mean DTH of HJX74 and CSSL5 (n = 30), respectively.
To analyze the genetic basis for the late heading of CSSL5, we crossed CSSL5 with the recipient HJX74 (Fig. 1). 73 out of 326 F2 plants showed heading date later than that of CSSL5 (113.0 ± 1.3 d) under NLD conditions in 2010 and the whole population exhibited trimodal distribution of heading date fitting 4:9:3 segration ratio (χ2 = 4.67 < χ20.05,2 = 5.99) (Fig. 2c). These results indicate that the late heading phenotype was controlled by two genes with negative epistasis.
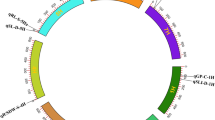
Confirmation of the two genes by resequencing
To delimit the heading date genes on the substituted segments, CSSL5 and HJX74 were high-throughput genotyped by whole-genome resequencing and an ultrahigh-quality physical map was constructed (Fig. 3). 11 substituted segments with a total length of 20.64 Mb derived from Zihui100 were found to be distributed over 6 chromosomes (Fig. 3 and Supplementary Table S1 online). Molecular markers were subsequently designed on the substituted segments and were used to analyze their linkage with heading date phenotype of the F2 population. Two QTLs were identified on chromosome 3 and 8, respectively, designated as EH3 and LH8, respectively. EH3 was mapped in the 750 kb genomic region between the Id32 and Id33 markers and had the contributions to phenotypic variation by 14.0%, the allele from Zihui100 could shorten the heading date by 2.32d and the dominant effect was −3.24. LH8 was mapped in the 460 kb genomic region between the Id83 and Id82 markers and had the contributions to phenotypic variation by 65.3%. The allele from Zihui100 could delay the heading date by 6.62d and the dominant effect was 3.44 (Table 1).
Epistatic interaction between EH3 and LH8
To examine the genetic interactions between EH3 and LH8, we scored heading dates in the F2 population under NLD and NSD conditions in 2011 (Fig. 4). The F2 plants were classified into nine genotype classes based on the EH3 and LH8 alleles and we compared the average heading date among the nine classes. Under NLD conditions, two-way ANOVA revealed the existence of digenic interaction between EH3 and LH8 (P = 5.96e-06). Orthogonal contrast test showed that AA (Additive by additive), but not AD (Additive by dominance), DA (Dominance by additive) or DD (Dominance by dominance) type of interaction was significant in the interaction of EH3 and LH8 (P < 0.0001). The effect of the Zihui100 allele at LH8 (i.e., increased days-to-heading (DTH)) was regardless of the allele status of EH3. In comparison, the effect of the Zihui100 allele at EH3 (i.e., shortened DTH) was observed in two genotype classes, homozygous for the Zihui100 allele at the LH8 locus and heterozygous, but not in the class homozygous for the HJX74 allele at the LH8 locus (Fig. 4a). Under NSD conditions, the Zihui100 allele at LH8 was also observed to delay heading regardless of the allele status of EH3. However, the two-way ANOVA suggested the digenic interaction between EH3 and LH8 was not significant under NSD conditions (P = 0.883) (Fig. 4b).
Epistatic interaction between LH8 and EH3.
Differences in days-to-heading for nine genotype classes for combinations of LH8 and EH3 under NLD (a) and NSD (b) conditions in the F2 populations. Genotypes were determined using the closely linked markers Id83-Id82 (LH8) and Id32-Id33 (EH3). Z, Hetero and H indicate homozygous for Zihui100 allele, heterozygous and homozygous for the HJX74 allele, respectively. PAA, PAD, PDA and PDD are significant probalities for AA, AD, DA and DD interactions between LH8 and EH3, respectively.
Map-based cloning of LH8
To fine map the LH8 gene, we selected F2 plants with homozygous EH3 allele from HJX74 and heterozygous LH8 allele from Zihui100 to generate F3 population (Fig. 1). F3 population showed bimodal distribution for DTH fitting 3:1 segregation ratio under both NLD and NSD conditions in 2012 (χ2 = 0.04 < χ20.05,1 = 3.84 and χ2 = 0.03 < χ20.05,1 = 3.84 for NLD and NSD, respectively), indicating the Zihui100 allele at LH8 increased DTH in a dominant manner (Fig. 5a and 5b). 7 new markers were developed in the marker interval of Id83-Id82 and were used to analyze a total of 2,159 F3 plants. 6 recombinants were identified between markers RM22475 and RM25 (Fig. 5c). The self-pollinated progeny (F4 lines) of those 6 plants were used to determine the genotypes of LH8 (Fig. 1). The recombinants r2 and r5 restricted LH8 to the 28.3 kb genomic region between the Id87 and Id811 markers, including 3 putative genes, a transferase family protein (LOC_Os08g07730), a SERK-family receptor-like protein kinase (LOC_Os08g07760) and a CCAAT-box-binding transcription factor (LOC_Os08g07740) which had been reported to function in heading date control in rice (Fig. 5c). The sequencing of LH8 genomic regions from CSSL5 and HJX74 revealed two GGC insertion and a 1,116-bp deletion in LOC_Os08g07740 of HJX74 (Fig. 6a), resulting in a 2-glycine insertion in the middle and a big alternation in the C-terminal region of the translated protein, respectively (Fig. 6b). These results suggest that LOC_Os08g07740 corresponds to LH8 controlling the late heading in CSSL5.
Map-based cloning of the LH8 gene in rice.
(a and b). Frenquency distributions of days-to-heading in an F3 population from HJX74 x CSSL5 under NLD (a) and NSD (b) conditions. Black arrows indicate the mean DTH of HJX74 and NIL-LH8 (n = 30), respectively. (c). High-resolution mapping of LH8. The genetic map of LH8 was based on recombinant events among 2,159 F3 plants from HJX74 x CSSL5. The number of recombinants between adjacent markers is shown above the bar. White, shadow and black regions shown in the recombinants indicate homozygous regions for HJX74 allele, heterozygous and homozygous regions for the Zihui100 allele, respectively.
Sequence comparision of LH8 from HJX74 and CSSL5.
(a). Structure of LH8 in HJX74 and CSSL5. Black and gray regions represent the ORF and UTR regions of LH8, respectively. The dotted line represents the 1,116 bp deletion in HJX74. (b). Protein alignment of LH8 from the predicted protein of HJX74 and CSSL5 using the EMBL software ClustalW2 multiple sequence alignment tool.
Characterization of LH8 in rice
To examine whether the sequence mutations can alter the expression of LH8, we examined the expression levels of LH8 in CSSL5, HJX74 and NIL-LH8 (a line selected from LH8 F2 segragating population with homozygous EH3 allele from HJX74 and homozygous LH8 allele from Zihui100) under NSD conditions in 2013. No significant difference in the LH8 expression levels could be detected among the three materials, suggesting that late heading did not result from the LH8 transcription. The expression levels of other heading date genes were also analyzed. For Hd1 and OsMADS50, no difference could be detected among the three materials under NSD conditions. The expression levels of Ehd1, Hd3a and RTF1 were less in NIL-LH8 than those in HJX74 under NSD conditions, indicating that LH8 may delay heading through suppressing the expressions of these floral transition activators. In contrast, the expression levels of Ehd1 and RTF1 were higher in CSSL5 than those in HJX74. The different expression levels of Ehd1 and RFT1 between CSSL5 and NIL-LH8 may result from the function of different EH3 alleles (Fig. 7a).
Characterization of the LH8 gene in rice.
(a). Expression of LH8, Hd1, OsMADS50, Ehd1, Hd3a and RFT1 in HJX74 (white pillars), CSSL5 (gray pillars) and NIL-LH8 (black pillars) at dawn (for LH8, OsMADS50, Ehd1, Hd3a and RFT1) and dusk (for Hd1) under NSD conditions. The analysis was performed in two independent experiments. The mean values of the relative expression levels of genes in CSSL5 and NIL-LH8 compared with HJX74 are shown by cloumns and standard deviations are indicated by the error bars. (b). Interaction of LH8 and Hd1 by yeast-two-hybrid assay. The yeast cells were grown on SD/-Leu/-Trp/-His/-Ade medium. The interaction between SE and HYL1 was served as positive control, while the combination of empty prey pGADT7 and that of the empty bait vector pGBKT7 with/without LH8 from Zihui100 were served as negative controls.
To study the involvement of the 2-glycine indel and the C-terminal region of the LH8 protein in its function, we tested the interaction between LH8 and Hd1 by yeast-two-hybrid method. Yeast cells that coexpressed the LH8Zihui100 bait and Hd1 prey fusion proteins were able to grow on media lacking histidine and adenine, indicating that these proteins can bind each other in yeast cells to promote the expression of the HIS3 and ADE2 reporter genes. However, the similar interaction could not be detected between LH8HJX74 and Hd1, indicating the 2-glycine indel and the C-terminal region of LH8 might be important for its interaction with Hd1 (Fig. 7b).
Discussion
Until now, a total of 734 rice heading date QTLs have been reported (http://www.gramene.org/qtl) and some of them have been molecularly characterized3. Besides understanding the biological function of single gene/QTL, clarification of genetic interactions among these genes/QTLs is also important. Actually, several examples of epistatic interactions among heading date genes/QTLs have been reported33,34,35,36,37. By eliminating genetic background noises, CSSLs offer a practical solution to bridge the huge gap of knowledge between the genotype and phenotype and therefore, are especially useful in studying complicated traits including heading date. In this study, a novel epistatic interaction between LH8 and EH3 involved in flowering control in rice was detected. Under NLD conditions and HJX74 genetic background, in plants homozygous for the LH8HJX74 allele, difference in DTH between EH3Zihui100 and EH3HJX74 alleles was small (about 1 day), as shown in HJX74 and NIL-EH3 (a line selected from LH8 F2 segragating population with homozygous LH8 allele from HJX74 and homozygous EH3 allele from Zihui100) (Fig. 8a). A large effect of EH3Zihui100 allele on heading promotion (about 7 days) occurred with the LH8Zihui100 allele, indicating EH3Zihui100 allele could function to shorten heading date only when LH8Zihui100 allele was concurrently present and therefore, the function of EH3 was dependent on the LH8 genotype through epistatic interaction between EH3Zihui100 and LH8Zihui100 alleles (Fig. 8a). This result suggests that EH3 might function as a modifier of LH8. Genetic interaction between LH8 and EH3 suggested that QTL pyramiding would be an effective method for the development of varieties with desirable heading dates under different growth conditions.
Schematic representation of the LH8 and EH3 mediated flowering pathway under natural growth conditions.
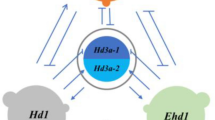
(a). Epistatic model of rice heading controlled by LH8 and EH3. (b). A proposed model for the flowering pathway controlled by LH8 and EH3 in rice under natural growth conditions. LH8 could down-regulate Ehd1, which could act on Hd3a and RFT1, thus delaying flowering in rice under natural growth conditions. EH3 might function as a modifier of LH8 to shorten heading date only when specific LH8 background was present. The interaction between LH8 and Hd1 depended on different alleles presented in different genetic backgrounds, possibly resulting in varied functions in flowering control.
To facilitate the identification of the location of LH8 and EH3, the high-throughput genotyping technology was exploited. The recipient HJX74 and CSSL5 were resequenced at the sequencing depth of 66- and 1-fold, respectively. Besides a large substituted segment from the short arm end to RM282 on chromosome 3 which was detected through traditional marker-assisted selection during CSSLs construction31, 10 additional substituted segments were identified distributing over 6 chromosomes, including some small introgressed segments that might be difficult to be detected by SSR markers (see Supplementary Table S1 online). Indeed, LH8 was found on untargeted chromosomal region on chromosome 8. These results showed that the high-throughput genotyping method by resequencing has a higher accuracy than marker-assisted selection. Owning to the deep sequencing coverage of the recipient HJX74, resequencing the CSSL5 with genome coverage as low as 1-fold was enough to rapidly validate and delimit the QTL, suggesting the efficiency of this mapping-by-sequencing method in discovering the genes responsible for quantitative trait loci. Our results presented a good example of combining the advanced material with uniform genetic background and high-throughput resequencing technology in economically accelerating gene identification.
The sequencing of LH8 genomic regions from CSSL5 and HJX74 revealed two GGC insertion and a 1,116-bp deletion in LOC_Os08g07740 of HJX74, resulting in a 2-glycine insertion in the middle and a big alternation in the C-terminal region of the translated protein, respectively (Fig. 6). Such sequence variations in the coding region of LOC_Os08g07740 between CSSL5 and HJX74 suggested that LOC_Os08g07740 might correspond to LH8 controlling the late heading in CSSL5 and NIL-LH8. LOC_Os08g07740 encodes a CCAAT-box-binding transcription factor which had been reported to strongly inhibit floral transition with names of DTH8, Ghd8 and LHD121,22,23. Similar to DTH8 and Ghd8, LH8 increased plant height and grain number per panicle under NLD conditions (Fig. 2 and Supplementary Table S2 online), indicating LH8 is allelic to DTH8, Ghd8 and LHD1. Indeed, the two GGC insertion and one 1,116-bp deletion in the coding region of LOC_Os08g07740 of HJX74 had also been found in the ZS97 allele at Ghd822. However, unlike DTH8 and LHD1 delaying flowering only under NLD/LD conditions21,23 and Ghd8 functioning oppositely under SD and LD conditions22, LH8 delays the heading date under both NSD and NLD conditions. These differences in function may come from different gene-background interactions or different growth conditions. Therefore, genes controlling important agronomic traits are necessary to be further studied in different genetic backgrounds and under different natural field conditions to give a full understanding of its effects and to offer practical instructions in rice breeding.
Expression analysis revealed the transcripts of several flowering promoting factors including Ehd1, Hd3a and RFT1 were fewer in NIL-LH8 than those in HJX74 under NSD conditions, suggesting LH8 might delay flowering by down-regulating the expression of Ehd1, Hd3a and RFT1 (Fig. 8b). The expressions levels of Hd1 and OsMADS50 showed no significant differences among HJX74, CSSL5 and NIL-LH8 under NSD conditions, suggesting they might function upstream of LH8 or parallel to LH8. Furthermore, no significant difference of LH8 expression were found among HJX74, CSSL5 and NIL-LH8 under NSD conditions, indicating the reason for the change of heading date was not caused by LH8 expression, but by nucleotide changes of the coding sequence. The 6-bp and 1,116-bp indels in the coding region of LH8 resulted in a big sequence alternation of the translated protein, especially in the C-terminal region.
The protein encoded by LH8 is a putative HAP3 subunit of the HAP complex, which binds to the CCAAT box, a cis-acting element present in approximately 25% of eukaryotic gene promoters38,39. HAP proteins always form a HAP2/HAP3/HAP5 trimeric complex with DNA binding activity in mammals40,41. In Arabidopsis, CO could replace AtHAP2 in the HAP complex to form a trimetric complex, CO/AtHAP3/AtHAP5, as the CCT domain of CO and the DNA-binding domain of AtHAP2 are similar to each other42. In LD conditions, overexpression of AtHAP2 or AtHAP3 could delay flowering by impairing formation of the CO/AtHAP3/AtHAP5 complex, leading to decreased expression of FT42. As a homologue of CO, Hd1 may be able to replace OsHAP2 in the HAP complex in rice. Our yeast-two-hybrid results revealed that Hd1HJX74 could indeed interact with LH8Zihui100, but not with LH8HJX74, indicating the sequence difference, especially in the C-terminal region, between LH8Zihui100 and LH8HJX74 might be important for its interaction with Hd1HJX74 and therefore for the regulation of rice heading (Fig. 8b). The domains responsible for the interaction between LH8HJX74 and Hd1HJX74 were needed to be further investigated. However, the interaction between LH8Zihui100 and Hd1Nip (Hd1 from Nipponbare) could not be detected (data not shown), in consistent with the previous study that effects of Ghd8 depend on the genetic background22. Consequently, whether the HAP complex could be formed or not was determined by the different subunits presented in different genetic backgrounds, possibly resulting in varied functions in flowering control.
Until now, 4 genes (Ehd4, OsMADS50, OsDof12, OsPhyB) on the short arm of chromosome 3 had been identified to be involved in flowering control in rice15,17,18,19,43,44. In our study, EH3 was mapped in the 750 kb genomic region between the Id32 and Id33 markers with 123 putative ORFs inside, including OsMADS50. OsMADS50, an ortholog of ArabidopsisSUPPRESSOR OF OVEREXPRSSION OF CONSTANS1 (SOC1), functions upstream of Ehd1 to indirectly induce RFT1 transcription17,18,19 and promote flowering. However, no epistatic interaction had been found between LH8 and OsMADS50. Furthermore, no significant difference of the expression of OsMADS50 was found between HJX74 and CSSL5. Therefore, whether EH3 is allelic to OsMADS50 remains to be investigated. Further fine mapping and molecular identification of EH3 will be necessary to provide molecular evidence for the epistatic interaction between LH8 and EH3 in regulating heading date in rice.
Methods
Plant materials and growth conditions
CSSL5 is one of a set of 1,123 chromosome segment substitution lines, which were developed from backcross progenies (BC6F2) derived from a cross between an elite indica variety from south china, HJX74, as the recurrent parent and another indica variety Zihui100 as the donor parent.
Plants were grown under natural short-day (from July to November) and natural long-day (from March to July) conditions in a paddy field in Guangzhou (23°07′N, 113°15′E), China. Heading date was defined as the time when the first panicle emerged.
High-throughput genotyping using whole-genome resequencing
DNA was extracted from 100 mg fresh rice leaves using the DNeasy Plant Mini Kit (QIAGEN Sciences). DNA was quantified using NanoDrop 1000 spectrophotometer (Thermo Scientific) and about 5 μg of DNA was used for preparation of libraries for Illumina sequencing according to the protocol for the Paired-End DNA Sample Prep kit (Illumina). The libraries were used for cluster generation on a flow cell and sequenced for 76 cycles on an Illumina Genome Analyzer IIx. Base calling and filtering of low-quality bases were done using sequence control software real-time analysis, Base calling (BCL) converter and the GERALD module (Illumina).
DNA extraction and PCR amplification
Fresh leaves were collected at the seedling stage and then ground in liquid nitrogen. Microquantities of DNA were extracted from fresh leaves of each individual using a previously reported method45. Amplification was carried out on the program for the initial denaturing step with 94°C for 3 min, followed by 35 cycles for 30 s at 94°C, 30 s at 55°C, 30 s at 72°C, with a final extension at 72°C for 5 min. PCR products were separated on 6% non-denaturing polyacrylamide gel and detected using the silver staining method46.
Linkage map and QTL analysis
We constructed the linkage map using Mapmaker/Exp3.047. The Kosambi function was used to calculate genetic distances. QTL for heading date were estimated by Mapmaker/QTL 1.148. Putative QTL were identified in regions exceeding 3.0 LOD (log-likelihood value).
Evaluation of genetic interactions
To analyze genetic interactions between LH8 and EH3, we classified a total of 359 and 225 F2 plants grown under NLD and NSD conditions in 2011, respectively, into nine classes according to the genotypes of SSR markers tightly linked to the QTLs (Id83-Id82 interval for LH8 and Id32-Id33 interval for EH3). We compared the mean DTH between the nine genotypes using two-way analysis of variance (ANOVA). Testing for each genetic interaction types (AA, AD, DA and DD) was done by orthogonal contrast test in STAR software (http://bbi.irri.org/).
Mapping for LH8
F2 plants with homozygous EH3 allele from HJX74 and heterozygous LH8 allele from Zihui100 were selected to generate F3 population. A total of 559 and 1,600 F3 plants were grown to fine map LH8 gene under NLD and NSD conditions in 2012, respectively. Each plant was genotyped using molecular markers and phenotyped by visual examination of the first panicle emergence. The F4 recombinant lines derived from the F3 population were used for progeny testing.
RNA Extraction and QRT-PCR
Total RNA from 30d-old leaves were isolated using TRIZOL reagent (Invitrogen) following the manufacturer's instruction. First-strand cDNA was reverse transcribed from DNaseI-treated RNA with oligo-dT as the primer using ReverTra Ace kit (Toyobo). Gene expression was measured by QRT-PCR using the ABI 7500 system (Life technologies). The QRT-PCR was carried out in a total volume of 20 μl containing 1X SYBR Green Master Mix (Life technologies). We normalized the expression levels by using UBQ5 gene as internal control. Each set of experiments was repeated three times and the relative standard curve quantification method was used to evaluate quantitative variation. The QRT-PCR procedure was conducted at 94°C for 3 min, followed by 40 cycles at 94°C for 30 s, 58°C for 30 s and 72°C for 30 s. The QRT-PCR primers were listed in Supplemental Table S3 online.
Yeast two-hybrid assay
The cDNA fragments encoding the entire putative LH8 and Hd1 proteins were PCR-amplified, verified by sequencing and cloned into NdeI–EcoRI restriction sites of the MATCHMAKER two-hybrid prey vector pGADT7 or bait vector pGBKT7 (Clontech), respectively. The bait and prey constructs were co-transformed into the yeast strain AH109 and the transformed cells were plated on synthetic defined medium containing all essential amino acids except leucine and tryptophan (SD/-Leu/-Trp) medium and incubated at 30°C for 3 d and then diluted and applied onto SD/-Leu/-Trp/-His/-Ade medium and SD/-Leu/-Trp medium (as loading control) and cultured at 30°C for 2–3 days. The interaction between HYL1 and SE was used as a positive control49. The primers used in vector construction were listed in Supplemental Table S3 online.
References
Kobayashi, Y. & Weigel, D. Move on up, it's time for change--mobile signals controlling photoperiod-dependent flowering. Genes Dev 21, 2371–84 (2007).
Putterill, J., Laurie, R. & Macknight, R. It's time to flower: the genetic control of flowering time. Bioessays 26, 363–73 (2004).
Tsuji, H., Taoka, K. & Shimamoto, K. Regulation of flowering in rice: two florigen genes, a complex gene network and natural variation. Curr Opin Plant Biol 14, 45–52 (2011).
Yano, M. et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12, 2473–2484 (2000).
Hayama, R., Yokoi, S., Tamaki, S., Yano, M. & Shimamoto, K. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 422, 719–22 (2003).
Doi, K. et al. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 18, 926–36 (2004).
Komiya, R., Yokoi, S. & Shimamoto, K. A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development 136, 3443–50 (2009).
Izawa, T. Adaptation of flowering-time by natural and artificial selection in Arabidopsis and rice. J Exp Bot 58, 3091–7 (2007).
Izawa, T., Takahashi, Y. & Yano, M. Comparative biology comes into bloom: genomic and genetic comparison of flowering pathways in rice and Arabidopsis. Curr Opin Plant Biol 6, 113–20 (2003).
Komiya, R., Ikegami, A., Tamaki, S., Yokoi, S. & Shimamoto, K. Hd3a and RFT1 are essential for flowering in rice. Development 135, 767–74 (2008).
Wu, C. et al. RID1, encoding a Cys2/His2-type zinc finger transcription factor, acts as a master switch from vegetative to floral development in rice. Proc Natl Acad Sci USA 105, 12915–20 (2008).
Park, S. J. et al. Rice Indeterminate 1 (OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod. Plant J 56, 1018–29 (2008).
Matsubara, K. et al. Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1. Plant Physiol 148, 1425–35 (2008).
Matsubara, K. et al. Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J 66, 603–12 (2011).
Gao, H. et al. Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice. PLoS Genet 9, e1003281 (2013).
Kim, S. L., Lee, S., Kim, H. J., Nam, H. G. & An, G. OsMADS51 is a short-day flowering promoter that functions upstream of Ehd1, OsMADS14 and Hd3a. Plant Physiol 145, 1484–94 (2007).
Lee, S., Kim, J., Han, J. J., Han, M. J. & An, G. Functional analyses of the flowering time gene OsMADS50, the putative SUPPRESSOR OF OVEREXPRESSION OF CO 1/AGAMOUS-LIKE 20 (SOC1/AGL20) ortholog in rice. Plant J 38, 754–64 (2004).
Ryu, C. H. et al. OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plant Cell Environ 32, 1412–27 (2009).
Bian, X. F. et al. Heading date gene, dth3 controlled late flowering in O. Glaberrima Steud. by down-regulating Ehd1. Plant Cell Rep 30, 2243–54 (2011).
Xue, W. et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40, 761–7 (2008).
Wei, X. et al. DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol 153, 1747–58 (2010).
Yan, W. H. et al. A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height and heading date in rice. Mol Plant 4, 319–30 (2011).
Dai, X. et al. LHD1, an allele of DTH8/Ghd8, controls late heading date in common wild rice (Oryza rufipogon). J Integr Plant Biol 54, 790–9 (2012).
Andres, F., Galbraith, D. W., Talon, M. & Domingo, C. Analysis of PHOTOPERIOD SENSITIVITY5 sheds light on the role of phytochromes in photoperiodic flowering in rice. Plant Physiol 151, 681–90 (2009).
Lee, Y. S. et al. OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB. Plant J 63, 18–30 (2010).
Peng, L. T., Shi, Z. Y., Li, L., Shen, G. Z. & Zhang, J. L. Ectopic expression of OsLFL1 in rice represses Ehd1 by binding on its promoter. Biochem Biophys Res Commun 360, 251–6 (2007).
Peng, L. T., Shi, Z. Y., Li, L., Shen, G. Z. & Zhang, J. L. Overexpression of transcription factor OsLFL1 delays flowering time in Oryza sativa. J Plant Physiol 165, 876–85 (2008).
Fukuoka, S., Ebana, K., Yamamoto, T. & Yano, M. Integration of Genomics into Rice Breeding. Rice 3, 131–137 (2010).
Yamamoto, T., Yonemaru, J. & Yano, M. Towards the understanding of complex traits in rice: substantially or superficially? DNA Res 16, 141–54 (2009).
Fukuoka, S., Nonoue, Y. & Yano, M. Germplasm enhancement by developing advanced plant materials from diverse rice accessions. Breeding Sci 60, 509–517 (2010).
Zhang, G. Q. et al. The construction of a library of single segment substitution lines in rice (Oryza sativa L.). Rice Genetics Newsletter 21, 85–87 (2004).
Xi, Z. Y. et al. Development of a wide population of chromosome single-segment substitution lines in the genetic background of an elite cultivar of rice (Oryza sativa L.). Genome 49, 476–84 (2006).
Yu, S. B. et al. Identification of quantitative trait loci and epistatic interactions for plant height and heading date in rice. Theor Appl Genet 104, 619–625 (2002).
Lin, H. X., Liang, Z. W., Sasaki, T. & Yano, M. Fine mapping and characterization of quantitative trait loci Hd4 and Hd5 controlling heading date in rice. Breeding Sci 53, 51–59 (2003).
Gu, X. Y. & Foley, M. E. Epistatic interactions of three loci regulate flowering time under short and long daylengths in a backcross population of rice. Theor Appl Genet 114, 745–54 (2007).
Uwatoko, N. et al. Epistasis among the three major flowering time genes in rice: coordinate changes of photoperiod sensitivity, basic vegetative growth and optimum photoperiod. Euphytica 163, 167–175 (2008).
Liu, S. et al. Genetic analysis and fine mapping of LH1 and LH2, a set of complementary genes controlling late heading in rice (Oryza sativa L.). Breeding Sci 62, 310–319 (2012).
Maity, S. N. & de Crombrugghe, B. Role of the CCAAT-binding protein CBF/NF-Y in transcription. Trends Biochem Sci 23, 174–8 (1998).
Mantovani, R. The molecular biology of the CCAAT-binding factor NF-Y. Gene 239, 15–27 (1999).
Goda, H. et al. Nuclear translocation of the heterotrimeric CCAAT binding factor of Aspergillus oryzae is dependent on two redundant localising signals in a single subunit. Arch Microbiol 184, 93–100 (2005).
Tuncher, A., Sprote, P., Gehrke, A. & Brakhage, A. A. The CCAAT-binding complex of eukaryotes: evolution of a second NLS in the HapB subunit of the filamentous fungus Aspergillus nidulans despite functional conservation at the molecular level between yeast, A. nidulans and human. J Mol Biol 352, 517–33 (2005).
Wenkel, S. et al. CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 18, 2971–84 (2006).
Takano, M. et al. Distinct and cooperative functions of phytochromes A, B and C in the control of deetiolation and flowering in rice. Plant Cell 17, 3311–25 (2005).
Li, D. et al. Functional characterization of rice OsDof12. Planta 229, 1159–69 (2009).
Shan, J. X., Zhu, M. Z., Shi, M., Gao, J. P. & Lin, H. X. Fine mapping and candidate gene analysis of spd6, responsible for small panicle and dwarfness in wild rice (Oryza rufipogon Griff.). Theor Appl Genet 119, 827–36 (2009).
Li, W. T., Zeng, R. Z., Zhang, Z. M. & Zhang, G. Q. Mapping of S-b locus for F1 pollen sterility in cultivated rice using PCR based markers. Acta Botanica Sinica 44, 463–467 (2002).
Lander, E. S. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–81 (1987).
Lincoln, S., Daly, M. & Lander, E. Mapping genes controlling quantitative traits with MAPMAKER/QTL1.1. 2nd edition (Whitehead Institute Technical Report, Cambridge, Massachusetts, 1992).
Yang, L., Liu, Z., Lu, F., Dong, A. & Huang, H. SERRATE is a novel nuclear regulator in primary microRNA processing in Arabidopsis. Plant J 47, 841–50 (2006).
Acknowledgements
This study was finacially supported by grants from the National Natural Science Foundation of P.R. China (31200905), the Guangdong Provincial Natural Science Foundation of China (10451064201005483), the Specialized Research Fund for the Doctoral Program of Higher Education (20104404120005) and the 40th Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry.
Author information
Authors and Affiliations
Contributions
Z.Q.L. and G.Q.Z. designed the experiments. Z.Q.L. wrote the manuscript. J.B.C. and M.Q. performed QTL analysis. J.B.C. and X.Y.L. performed map-based cloning. C.C. and H.T.Z. performed sequencing analysis. Y.H.W. performed real-time RT-PCR. R.Z.Z. and X.L.F. performed yeast-two-hybrid analysis. All authors commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Chen, J., Li, X., Cheng, C. et al. Characterization of Epistatic Interaction of QTLs LH8 and EH3 Controlling Heading Date in Rice. Sci Rep 4, 4263 (2014). https://doi.org/10.1038/srep04263
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04263
This article is cited by
-
Dynamic analysis of QTLs on plant height with single segment substitution lines in rice
Scientific Reports (2022)
-
GW10, a member of P450 subfamily regulates grain size and grain number in rice
Theoretical and Applied Genetics (2021)
-
Unconditional and conditional analysis of epistasis between tillering QTLs based on single segment substitution lines in rice
Scientific Reports (2020)
-
Identification and fine mapping of a major QTL, qHD19, that plays pleiotropic roles in regulating the heading date in rice
Molecular Breeding (2020)
-
Genotyping-by-sequencing and multilocation evaluation of two interspecific backcross populations identify QTLs for yield-related traits in pigeonpea
Theoretical and Applied Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.