Abstract
Gene set enrichment analysis (GSEA) associates gene sets and phenotypes, its use is predicated on the choice of a pre-defined collection of sets. The defacto standard implementation of GSEA provides seven collections yet there are no guidelines for the choice of collections and the impact of such choice, if any, is unknown. Here we compare each of the standard gene set collections in the context of a large dataset of drug response in human cancer cell lines. We define and test a new collection based on gene co-expression in cancer cell lines to compare the performance of the standard collections to an externally derived cell line based collection. The results show that GSEA findings vary significantly depending on the collection chosen for analysis. Henceforth, collections should be carefully selected and reported in studies that leverage GSEA.
Similar content being viewed by others
Introduction
With the advent of high-throughput gene expression profiling using microarrays and RNA sequencing, researchers are now able to quantify the expression of a cell's genes in response to various environments, stimuli or other controlled experimental factors1. It has thus become common practice to refer to a cell's expression profile, meaning the complete set of its gene expression levels for a specific experimental condition. Among numerous applications of gene expression profiling, the identification and quantification of differential gene expression have been shown to be informative and reproducible across different teams and technology platforms2,36.
Differential expression of individual genes have led to critical discoveries in numerous diseases such as the genes ESR1, ERBB2 and AURKA used in breast cancer molecular subtyping3. However it is now well established that it is generally not individual genes but sets of genes (and sets of gene products) that collectively define phenotypes such as targeted cancer therapy response. This suggests that the association of a set of expression levels with phenotype may be more robust than biomarkers consisting of individual gene expression levels. To this end Gene Set Enrichment Analysis (GSEA) has been developed to associate gene sets with sample phenotypes4,5.
In the context of cancer therapy decision-making, it is important to understand the mechanism of action of anticancer agents and to identify efficient drug response biomarkers. However given the rapid development of many new compounds, it is neither sustainable nor ethical to test all of them in clinical trials6. Therefore several research groups investigated the use of large panels of cell lines to effectively screen the therapeutic potential of numerous compounds7,8,9. In particular Garnett and colleagues at the Welcome Trust Sanger Institute recently published the results of a large panel of 727 unique cancer cell lines screened with 138 drugs (the resulting dataset is referred to hereafter as the Cancer Genome Project's dataset or CGP).
Developing therapeutic strategies based on such studies is an elusive and attractive target. Much of the difficulty in establishing reliable predictors lies in the genetic diversity of human cancers and so gene set association is a natural investigative avenue. Cell line drug response trials offer a vast array of quantifiable phenotypes and so GSEA can be utilized to find gene sets associated with a particular drug response10,11.
GSEA requires a pre-defined collection of gene sets as input then provides a score to each gene set's association with a phenotype. We refer to the distribution of these scores attributed to a particular collection by GSEA as that collection's enrichment profile. In this work we hypothesize that different gene set collections yield heterogeneous enrichment profiles when investigating the biological mechanisms of drug response in cell lines. We therefore tested the use of various collections within GSEA and compared their enrichment profiles within the context of drug response in human cancer cell lines. We analyzed the CGP pharmacogenomic dataset to compare the associations of gene expression with drug response over 138 drugs administered to a panel of 727 cancer cell lines.
Of these standard collections, C2 provides a significant number of highly enriched gene sets as well as the net highest scoring gene set for many drugs. C2 is a collection composed of sets extracted mainly from biomedical literature and biological databases such as the Kyoto Encyclopedia of Genes and Genomes12 (KEGG) or Reactome13. It is followed in both these categories by C4: a collection of gene sets created by data mining cancer-related microarray data. All other collections perform significantly poorer. We further observe that there is little overlap between the standard collections and that collection performance is predicted by gene count or shared phenotypic characteristics with the phenotype under study. Lastly we showed that a new collection based on co-expressed gene sets extracted from cancer cell lines experiments supplants C2 as the lead collection of gene sets when included in the analysis.
Results
To investigate the impact of a particular collection on GSEA's results, we conducted gene set analyses for 138 drugs tested on 727 cell lines8. The collections tested were the seven standard collections made available through the tool's distributor, together these seven collections are refered to as the Molecular Signatures Database (MSigDB).
There exists no de facto consensus among the community as to which gene set collections are to be used within GSEA. Of the first 32 hits on a PubMed search for GSEA, 9 articles did not specify the source of their gene sets, 7 articles noted a manual curation and omitted the methodology, 14 specified specific particular instances or subsets of the collections within MSigDB and only 1 article specified that the entire MSigDB was used (supplementary file 1). Justifications for the choices made were almost uniformly omitted.
The seven collections are generated using different strategies and thus the number of unique genes accounted for within a collection varies. 12Table 3 shows the number of unique genes contained in each collection.
We compared the number of gene sets in each of the MSigDB collections (Figure 1A, Table 1). With a total of 8761 unique gene sets, the number of gene sets contained in each of the seven collections ranges from 188 (C6) to 3761 (C2). To assess the overlap between these collections we adapted the H index, which is commonly used to estimate a researcher's scientific productivity18, in order to quantify the overlap between two collections of gene sets (see Methods). We used this new overlap index, referred to as the g index, to compute the overlap between each possible pair of gene set collections. Figure 1B represents the resulting g indices as a heatmap. These indices range in value between 0.0106 and 0.0223 (mean = 0.0167, sd = 0.00352.) All scores are available in supplementary file 2. We observed that the highest degree of overlapping is observed between C2, a collection of gene sets curated from biological databases and biomedical literature (Table 1) and C4, a collection composed of gene sets created by mining three large microarray studies (Table 1) with g index of 0.0223, doubling the maximal overlap score of C1. The C4 and the Gene Ontology set C5 share the next highest overlapping index (g index of 0.0212). C1, based on gene position in cytogenetic bands, shows very little overlap with all sets (maximum g index of 0.0118 with the C2 set.) C1 is the only collection based on gene proximity while all other collections attempt to group genes based on phenotypes, pathways or within the classification proposed by Gene Ontology22.
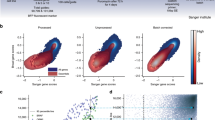
We refer to the distribution of enrichment scores attributed to a collection by GSEA as that collection's enrichment profile. For each drug we performed a gene set enrichment analysis with each individual collection to produce 138 enrichment profiles for each collection. Under the assumption that highly enriched gene sets are more indicative of a collection's value than the overall distribution of its sets, we compared the distributions of normalized enrichment scores with absolute values greater than 2 for each collection (Figure 2A). Within the overall density graph we observed an approximately normal distribution with a mean absolute normalized enrichment score (NES) of around 1.0 for all collections. At the high end of the density curves we note that the C4 collection contains the highest scoring gene sets overall, followed by C2 and C6 (Figure 2A). We also counted the number of enriched gene sets for each drug within each collection (Figure 2B). Overall, GSEA identified significantly more enriched gene sets in the C2 and C4 collections (Table 2).
(A) Density plot representing the distribution of normalized enrichement scores for all drugs in each collection individually. (B) Heatmap of the number of highly enriched gene sets (absolute normalized enrichement score > 2.0, FDR < 25%) for each drug, in each collection. Gene set collections are listed along the bottom of the figure and drugs along the right. Darker hues of blue indicate a greater number of enriched gene sets for a particular drug.
In order to understand the relative effectiveness of each collection in providing highly enriched gene sets in the given context we plotted the fractional contribution of the top scoring sets for the aggregation of all drugs. Each drug was polled for its top scoring gene set (highest absolute NES) and the set's collection of origin was identified. The ratio of sets contributed by a collection to the total of these top scoring sets is that collection's fractional contribution. The number of gene sets polled per drug was incremented from one to fifty and the results are plotted in Figure 3. This procedure permits a competitive analysis of the gene set collections. We observed that the C2 collection is the dominant collection followed by C4 and the remaining collections do remarkably less well with little distinction among them.
(A) Fractional contribution of each collection to the set of top scoring gene sets with n gene sets per drug. n is plotted along the abscise. The ordinance shows the fraction of top gene sets contributed by each collection to the set of top scoring gene sets. As n increases, a higher number of gene sets per drug are assumed to be relevant or significant. Collection C2 is the highest contributor by a large margin, followed by C4, all other collections contribute to a negligible degree. The fractional contribution of C4 peaks before 10 top gene sets per drug, coinciding with C2's low. There is a slight trend downward in C4's contribution afterwards and a lesser trend upwards in the case of C2. (B) Fractional contribution of all Broad's collections plus our data-driven gene set collection, referred to as HGSK.
We introduced a new, data-driven collection to the competitive analysis in order to compare the leading MSigDB collections to an external collection. We also sought to test whether the standard collections offered the highest scoring gene sets for the phenotype under study. This collection was built by computing sets of tightly co-expressed genes in cancer cell lines produced by GlaxoSmithKline and published by Greshock et al.20. We performed a hierarchical clustering analysis to compute the nested structure of co-expression gene sets (Figure 4, see Methods). We repeated the competitive analysis described previously with this new collection of co-expressed gene sets, referred to as HGSK (short for hierarchical GlaxoSmithKline.) As can be seen in Figure 3B the HGSK collection dominates the remainder of the collections mostly at the expense of the C2 collection. However, when a greater number of enriched gene sets are considered (n > 30), C2 contributed more and more gene sets and approached HGSK's contribution. The C4 collection's contribution remains largely unaffected by the inclusion of HGSK. The contributions of other collections remain negligible.
Creation of the HGSK set collection is done by creating a gene-gene distance measure based on the reciprocal of a gene-gene correlation matrix from the expression of tumour cell lines in the GSK data set.
Genes are clustered using traditional hierarchical clustering based on the distance measure. Depth first recursive tree generation is done, iterating over the prior sub-trees of cluster. Sets containing less than 15 genes or more than 500 are discarded.
Discussion
Despite the widespread use of gene set enrichment analyses in biomedical research, the choice of the gene set collection is rarely discussed and its impact on the overall analysis results remains an open question. Here we examine the varied expression profiles yielded by the standard collections when performing gene set enrichment analyses within the specific context of drug response in cancer cell lines. We do this by contrasting the performance of the seven standard collections curated by the Broad Institute. Among these standard collections there is a remarkable variance in the number and strength of association shown in the results. Notably C2 and C4 aggregate significantly more gene sets associated with the phenotypes under study. This is in part unsurprising as collections built around cancer studies may enjoy a positive bias as to gene set association to phenotype given that the phenotypes under study is drug response in cancer cell lines. However a collection creation strategy based on cancer studies is clearly not a predictor of performance on our metrics as is demonstrated by the poor performance of the oncogenic signature collection C6.
To further explore the impact of gene set collection on the GSEA results, we built our own collection, referred to as HGSK, based on hierarchical clustering analysis of co-expressed genes in an independent dataset of cancer cell lines. We then compared the results offered by this collection to simulate an unfiltered data-driven approach to the study of drug response in cancer cell lines. Unsurprisingly, the HGSK collection outperforms the leader among the standard collections. Interestingly, during the competitive analysis HGSK gains come at the expense of C2 (curated primarily from pathway databases) and not C4 (which shares an oncological pedigree with HGSK.) This suggests that the signal provided by the unsupervised clustering algorithm tended towards the identification of genes co-expressed in pathways and not communalities between cancer cultures. However despite its better performance, enriched HGSK gene sets do not lend themselves to immediate biological interpretation, as they are not labeled using prior knowledge. Nonetheless this might be alleviated to a certain degree with third party annotation tools such as the Gene Ontology, which could be used to annotate most HGSK gene sets although not all of them.
A set of results that illustrates the interpretation and association tradeoff particularly well is found within the EGFR/ERBB2-targetting drugs: Erlotinib, Gefitinib, Lapatinib and BIBW2992. The HGSK co-expression based gene set HGSK-547 is attributed a NES over 2.0 in three of these four drugs. STRING-DB23 (Search Tool for the Retrieval of Interacting Genes/Proteins Database) finds the gene set to be significantly enriched in protein-protein interactions (p < 1E-16) and to be enriched in the KEGG pathway Tight Junction (p-value = 4E-5). However little else is known about this set a priori with the exception of the co-expression of its members. On the other hand the standard collections often provide sets that reference literature relevant to the nature and origin of the gene collection. A C2 set JAEGER_METASTASIS_DN is another highly enriched gene set for EGFR targeting drugs, its title is suggestive of biological implications and their source. This second set consists of genes found to be down-regulated in metastases of melanoma in a study geared towards identifying differential expression signatures between primary melanomas and melanoma metastases24. Note that in this case, the gene set is not associated by protein or pathway interactions instead they are revealed by a former study. A second interesting note here is that in this case the C2 set: JAEGER_METASTASIS_DN held a higher aggregate score among a family of drugs (EGFR) than the synthetic HGSK set whereas the co-expression based collection usually provides between 60% to 40% of top scoring gene sets as can be seen from Figure 2.
Results from C2 and HGSK collections concur that chemosensitivity to EGFR/ERBB2 inhibitors is associated with the upregulation of cellular tight junction proteins among including the Claudin family of genes (Claudin-3, 4,7). These proteins assist in maintaining cell polarity and in the recruitment of other signaling proteins and therefore were hypothesized to be involved in tumoregenesis25. Recent work has shown that Claudin-7 inhibits cell migration of human non-small cell lung cancer cells NCI-H1299 via an ERK/MAPK dependent process26. According to Lu and co-workers, the overexpression of Claudin-7 diminished the phosphorylation of ERK1/2 and hence inhibited the aggressiveness of lung cancer through a MAPK/ERK dependent pathway. EGFR is an upstream activator of this pathway and thus it may be that the upregulation of these tight junction protein genes may attenuate cancer invasiveness in the presence of EGFR inhibitors26. Our results suggest that this family of proteins would be an interesting target of further research to elucidate their potential as prognostic biomarkers for patient response to EGFR inhibitors. A recent study showed that Claudin-7 sensitizes lung cancer cells to Cisplatin treatment through a caspase dependent pathway27.
In the CGP study, Garnett et al. identified ERBB2 expression as an indicator of Lapatinib response8. This is supported by the presence of the ERBB2 gene symbol in the top scoring gene set for Lapatinib sensitivity: COLDREN_GEFITINIB_RESISTANCE_DN. This gene set is constructed based on microarray gene expression profiling of Gefitinib testing on non-small cell lung cancer cell lines28.
The results of the GSEA analyses for the MEK1/2 inhibitors were investigated. Selumetinib and PD0325901 are investigational drugs that inhibit the MEK 1 and 2 dual-specificity kinases that upregulate the RAS/RAF/MEK/ERK pathway in MEK-overexpressing tumors. Pathways associated with sensitivity to MEK inhibitors were found to be enriched in genes involved in the innate immune response. For example, a pattern of genes from the Toll-like receptors pathway (TLR2, TLR8, CD86, CD14) is known to activate immune cell responses. Recently a work by Peroval et al, 2013 emphasized the complex role of MAPK signaling pathways in the transcriptional regulation of Toll-like receptors29. It is possible that these receptors would trigger cell death when MEK kinases are degraded.
Thus while GSEA offers interesting results and is valuable in the generation of hypotheses for further investigation, the utility of the standard collections, in the context of drug response in cancer cell lines, varies. In this context, C2 contributes 2 high scoring sets for each submitted by C4. Of further interest is the particularly poor performance of the C5 set which is based on the Gene Ontologies collection and the C6 collection based on oncogenic signatures. The C6 collection was expected to do well given the nature of the cell lines. Both of these fare far worse than a collection based on data mining immunology research. As expected, the HGSK co-expression based gene set collections scores high. This further demonstrates the sensitivity of the GSEA process to the collection used in the analyses. The HGSK collection also highlights the value offered by the annotation of the standard, curated collections. It is important to note that HGSK itself is built from a dataset that closely resembles the dataset being probed. This is done to model a data-driven approach to gene set collection creation, no claims are made about it being a useful collection outside of this context. Our intent here is to show the variation in the results among the collections currently being used by the community. Furthermore only the MSigDB gene set collections are reviewed and solely in the context of drug response in tumour cell lines. In addition, despite its popularity, the gene set analysis method as proposed faces some criticism30.
In conclusion, gene-set association with cancer drug response done by GSEA are sensitive to the gene-set collection used and two gene set collections consistently offer results of a higher significance in the context of drug response in cancer cell lines. Research leveraging GSEA should closely evaluate gene set collection selection criteria. Studies published using the tool should precisely report the nature of the collection used in the analyses.
Methods
The overall analysis design is represented in Figure 5 and the details of each step are described here.
Overall analysis design used in our comparative study.
First we calculated the overlap between each pair of gene set collections. Second we used a large pharmacogenomic dataset (CGP) to rank all the genes with respect to their association to response to each of the 138 drugs. Third we used these rankings together with the gene set collections to run multiple GSEA. Fourth the results are aggregated to compare the most enriched gene sets across collections. The results are then interpreted by taking into account the overlap between collections.
Gene set analysis
The gene set enrichment analysis (GSEA) technique developed by Subramanian and colleagues14 is a widely used method of measuring the association between a set of genes and a phenotype in gene expression profiling data sets. GSEA enables detection of gene sets enriched in genes that are significantly associated with a phenotype of interest. Such enrichment is computed using the Kolmogorov-Smirnov (KS) statistic15. This statistic compares the anticipated random distribution of a set's genes and their actual distribution among a genome-wide list of genes ranked based on their association with the phenotype. The KS statistic is then normalized for gene set size and its significance is adjusted to take into account multiple hypotheses testing. A Java implementation of this method16 is made publicly available by the authors. GSEA requires an a priori gene set collection to be defined. Therefore, alongside the tool, the Broad Institute makes available several gene set collections, referred to as MSigDB17, which is described in the next section.
Gene set collections
Seven collections of gene sets are made available for use with GSEA by the Broad Institute. (http://www.broadinstitute.org/gsea/msigdb/index.jsp) These collections, all together, are referred to as MSigDB17. We downloaded the latest version (4.0) of the collections from the above URL. Table 1 gives a brief description of each collection, summarizing the information available from the website.
Overlap between gene set collections
In order to measure the overlap between collections of gene sets each pair of collections was subjected to a pairwise comparison of gene sets based on the h-index18 and the Jaccard index19 in which the ratio of the cardinality of the intersection of the sets to the cardinality of the union of the sets is calculated. This index, referred to as g′, is calculated using the following formula:

For collections C and D that provide sets C1 through Cm and D1 through Dn respectively, g(C,D) is the largest proportion of the n × m pairings where g′ is greater to or equal to g. We referred to this metric as the gene set overlap index or the g-score.
Co-expression gene set collection based on cancer cell line data
In addition to the Broad's collections of gene sets, we created a new collection based on a fully data-driven analysis of cancer cell lines. This collection of sets was built using gene co-expression data from an independent dataset of 311 cancer cell lines, referred to as the GSK dataset in the literature20. A gene-expression correlation matrix was calculated and a distance matrix was taken as 1 minus the correlation matrix. We then used the resulting distance matrix to generate a hierarchical clustering21 of the cancer cell lines' genes. The clustering was recursively partitioned into all possible sets that respected the hierarchy and were composed of at least 15 and no more than 500 genes in size. Figure 4 summarizes the creation of the HGSK collection. The resulting co-expression gene sets are provided in Supplementary File 4.
Gene ranking based on association with drug response
To compute a genome-wide ranking of genes based on their associations with drug sensitivity, we used the area under the dose response curve (AUC) as a measure of drug sensitivity8 and we assessed the association between gene expression and drug response using a linear regression model controlled for tissue type. For each gene i we fit two linear models, M0 and M1:


where Y denotes the drug sensitivity variable (AUC), G and T denote the expression of gene i and the tissue type respectively and βs are the regression coefficients, i.e., β′0is the intercept, β′tis the regression coefficient for the categorical variable T representing the tissue type and β′i: regression coefficient for the continuous variable G representing the expression of the gene of interest. The strength of gene-drug association is quantified by β′i, above and beyond the relationship between drug sensitivity and tissue type. The variables Y and G are scaled (standard deviation equals to 1) in order to get standardized coefficients from the linear model. Significance of the gene-drug association is estimated by computing the F statistic using the analysis of variance (ANOVA) comparing the two nested models, M0 and M1. All genes are then ranked with respect to their F statistic, that is the significance of the association between their expression and drug sensitivity and the direction of the corresponding association (negative if expression of gene i is association with drug resistance, positive otherwise).
Gene set enrichment analysis
To assess the association of a collection of gene sets with sensitivity to each of the 138 drugs screened in CGP, we used version 2.0.13 of the GSEA tool developed by the Broad Institute. Pre-ranked GSEA requires two input files: a gene set collection (the Broad's collections for instance) and a genome-wide ranking of genes, as described previously. We ran pre-ranked GSEA on each gene set collection to compute enrichment scores for each gene set within the collections. The magnitude of normalized enrichment scores and FDR values were used to evaluate the effectiveness of each collection in identifying candidate gene sets that influence drug response in cancer cell lines.
References
Lockhart, D. J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotech 14, 1675–1680 (1996).
Shi, L. et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat. Biotechnol. 24, 1151–1161 (2006).
Haibe-Kains, B. et al. A Three-Gene Model to Robustly Identify Breast Cancer Molecular Subtypes. JNCI J. Natl. Cancer Inst. 104, 311–325 (2012).
Hung, J.-H., Yang, T.-H., Hu, Z., Weng, Z. & DeLisi, C. Gene set enrichment analysis: performance evaluation and usage guidelines. Brief. Bioinform. 13, 281–291 (2011).
Maciejewski, H. Gene set analysis methods: statistical models and methodological differences. Brief. Bioinform. 10.1093/bib/bbt002 (2013).
Weinstein, J. N. Drug discovery: Cell lines battle cancer. Nature 483, 544–545 (2012).
Shoemaker, R. H. The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 6, 813–823 (2006).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–307 (2012).
Sherman-Baust, C. A., Becker, K. G., Wood Iii, W. H., Zhang, Y. & Morin, P. J. Gene expression and pathway analysis of ovarian cancer cells selected for resistance to cisplatin, paclitaxel, or doxorubicin. J Ovarian Res 4, 21 (2011).
Rusnak, D. W. et al. Assessment of epidermal growth factor receptor (EGFR, ErbB1) and HER2 (ErbB2) protein expression levels and response to lapatinib (Tykerb®, GW572016) in an expanded panel of human normal and tumour cell lines. Cell Prolif. 40, 580–594 (2007).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Croft, D. et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 39, D691–D697 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 102, 15545–15550 (2005).
Sheskin, J. in Handb. Parametr. Nonparametric Stat. Proced. 261–276 (Chapman & Hall, 2011).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J. P. GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23, 3251–3253 (2007).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Hirsch, J. E. An index to quantify an individual's scientific research output. Proc. Natl. Acad. Sci. U. S. A. 102, 16569–16572 (2005).
Levandowsky, M. & Winter, D. Distance between Sets. Nature 234, 34–35 (1971).
Greshock, J. et al. Molecular Target Class Is Predictive of In vitro Response Profile. Cancer Res. 70, 3677–3686 (2010).
R Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2013). at <http://www.R-project.org>.
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Franceschini, A. et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2012).
Jaeger, J. et al. Gene Expression Signatures for Tumor Progression, Tumor Subtype and Tumor Thickness in Laser-Microdissected Melanoma Tissues. Clin. Cancer Res. 13, 806–815 (2007).
Morin, P. J. Claudin proteins in human cancer: promising new targets for diagnosis and therapy. Cancer Res. 65, 9603–9606 (2005).
Lu, Z. et al. Claudin-7 inhibits human lung cancer cell migration and invasion through ERK/MAPK signaling pathway. Exp. Cell Res. 317, 1935–1946 (2011).
Hoggard, J. et al. Claudin-7 increases chemosensitivity to cisplatin through the upregulation of caspase pathway in human NCI-H522 lung cancer cells. Cancer Sci. 104, 611–618 (2013).
Coldren, C. D. Baseline Gene Expression Predicts Sensitivity to Gefitinib in Non-Small Cell Lung Cancer Cell Lines. Mol. Cancer Res. 4, 521–528 (2006).
Peroval, M. Y., Boyd, A. C., Young, J. R. & Smith, A. L. A Critical Role for MAPK Signalling Pathways in the Transcriptional Regulation of Toll Like Receptors. PLoS ONE 8, e51243 (2013).
Tripathi, S., Glazko, G. V. & Emmert-Streib, F. Ensuring the statistical soundness of competitive gene set approaches: gene filtering and genome-scale coverage are essential. Nucleic Acids Res. 41, e82–e82 (2013).
Newman, J. C. & Weiner, A. M. L2L: a simple tool for discovering the hidden significance in microarray expression data. Genome Biol. 6, R81 (2005).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Wingender, E. et al. TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res. 28, 316–319 (2000).
Segal, E., Friedman, N., Koller, D. & Regev, A. A module map showing conditional activity of expression modules in cancer. Nat. Genet. 36, 1090–1098 (2004).
Brentani, H. et al. The generation and utilization of a cancer-oriented representation of the human transcriptome by using expressed sequence tags. Proc. Natl. Acad. Sci. 100, 13418–13423 (2003).
Haibe-Kains, B. El-Hachem, N. Birkbak, N. J. Jin, A. C. Beck, A. H. Aerts, H. J. W. L. & Quackenbush, J. Inconsistency in large pharmacogenomic studies. Nature, 504(7480), 389–393. doi:10.1038/nature12831 (2013).
Author information
Authors and Affiliations
Contributions
A.R.B. and B.H.-K. were responsible for the design and execution of the study, collation of study materials, the microarray analysis of study samples, the collection, assembly and verification of the data, data and statistical analysis and interpretation and final manuscript writing; N.E.-H. assisted with the collation of study materials, the microarray analysis of study samples, data analysis and interpretation and final manuscript writing. A.H.B. and H.J.W.L.A. contributed to the interpretation of the results and participated to the manuscript writing. B.H.K. supervised the study. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Bateman, A., El-Hachem, N., Beck, A. et al. Importance of collection in gene set enrichment analysis of drug response in cancer cell lines. Sci Rep 4, 4092 (2014). https://doi.org/10.1038/srep04092
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04092
This article is cited by
-
Marked intestinal trans-differentiation by autoimmune gastritis along with ectopic pancreatic and pulmonary trans-differentiation
Journal of Gastroenterology (2024)
-
Autoimmune gastritis induces aberrant DNA methylation reflecting its carcinogenic potential
Journal of Gastroenterology (2022)
-
Identification of marker genes and pathways specific to precancerous duodenal adenomas and early stage adenocarcinomas
Journal of Gastroenterology (2019)
-
Data-driven human transcriptomic modules determined by independent component analysis
BMC Bioinformatics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.