Abstract
Induced pluripotent stem cells (iPSCs) hold much promise in the quest for personalised cell therapies. However, the persistence of founder cell mitochondrial DNA (mtDNA) mutations limits the potential of iPSCs in the development of treatments for mtDNA disease. This problem may be overcome by using oocytes containing healthy mtDNA, to induce somatic cell nuclear reprogramming. However, the extent to which somatic cell mtDNA persists following fusion with human oocytes is unknown. Here we show that human nuclear transfer (NT) embryos contain very low levels of somatic cell mtDNA. In light of a recent report that embryonic stem cells can be derived from human NT embryos, our results highlight the therapeutic potential of NT for mtDNA disease and underscore the importance of using human oocytes to pursue this goal.
Similar content being viewed by others
Introduction
The possibility of generating patient-specific embryonic stem (ES) cells emerged from the seminal discoveries that the oocyte cytoplasm has the capacity to reprogram differentiated cells to an early embryonic state1 and that pluripotency can be harnessed by deriving embryonic stem (ES) cells from the inner cell mass (ICM) of the pre-implantation blastocyst2. Isogenic ES cells have the potential to increase the efficacy of cellular therapies in treating degenerative disease by reducing the risk of immune rejection3.
Fusion of differentiated cells with enucleated oocytes, otherwise known as nuclear transfer (NT), has been reported in a variety of species with variable success rates4. Until recently, there had been only a few isolated reports of blastocyst formation following NT in human oocytes5,6,7 and the only success in generating ES cells from human NT embryos involved leaving the oocyte's nuclear material intact, resulting in the formation of triploid ES cells8. However, it has recently been reported that this limitation can be overcome by using caffeine to prevent premature activation of oocytes during the NT procedure9.
The discovery that ectopic expression of SOX2, OCT4, KLF4 and Myc can induce differentiated cells to revert to an ES cell-like state, resulting in the production of induced pluripotent stem cells (iPSCs)10, raised the possibility of generating patient-specific cell therapies without the need for oocytes. This is an area of intensive research, which holds much promise for the future of isogenic therapies in regenerative medicine. However, the broad spectrum of degenerative diseases associated with mutations in mitochondrial DNA (mtDNA)11 are unlikely to be amenable to iPSC-based therapies due to the persistence of the somatic cell mtDNA mutations.
Mitochondrial DNA mutations cause defects in respiratory chain function resulting in multi-system disease affecting at least 1 in 10,000 adults12. However, pathogenic mutations are more prevalent and are estimated to be present in 1 in 200 babies born13. Pathogenicity is largely determined by the ratio of mutated to non-mutated mtDNA molecules, with a typical minimum critical proportion of 60–90% mtDNA mutation before phenotypic effects are observed11. At present, there are no curative treatments for mtDNA disease and interventions are largely limited to managing symptoms14. While research is underway to develop techniques to prevent transmission of mtDNA mutations by transplanting nuclear DNA between zygotes15 or oocytes16, the development of cellular therapies represents an important step towards treatment of degenerative diseases linked to mtDNA mutations. Given that pathology frequently develops in childhood, the prospect of a lifetime on immunosuppressant drugs makes the development of isogenic therapies for mtDNA disease a particularly pressing goal.
Mitochondrial DNA is transmitted by maternal inheritance17 and oocytes contain orders of magnitude more mtDNA copies than somatic cells18,19. Fusion of a whole somatic cell with an oocyte cytoplasm offers the theoretical possibility of developing NT-derived ES cells in which mutated mtDNA is massively diluted by the mtDNA present in the oocyte. However, the extent to which somatic cell mtDNA persists in NT embryos appears to vary widely between species and between different studies19. For example studies in sheep, including Dolly, indicate that somatic cell mtDNA does not persist following NT20, which raised the possibility that somatic cell mitochondria may be targeted for destruction by a mechanism analogous to that responsible for elimination of sperm mitochondria following fertilisation21. Since then, a number of studies have reported very low levels of donor cell-derived mtDNA following NT in sheep19. By contrast, studies in bovine NT embryos and offspring indicate that somatic cell mtDNA persists to widely varying degrees19. The fate of donor cell mtDNA in human oocytes is unknown. This question is fundamental to the potential of NT in the development of isogenic cell therapies for mtDNA disease.
To address this, we analysed mtDNA in NT-derived human embryos. We found that the mitochondrial biomass of fibroblasts is influenced by the method used to induce arrest in Go/G1. However, following NT, fibroblast mtDNA was undetectable in the majority of embryos and accounted for <2% of mtDNA content in the remainder. Development to the 4-cell stage when using amnion-derived fibroblasts as a nuclear donor was comparable to activated control embryos. However, consistent with the findings of others8,22, we found that the efficiency of development beyond the 8-cell stage is low. Thus, our data indicate that, with the use of optimised techniques for NT as recently published9, isogenic cell therapies for mtDNA disease could be feasible.
Results
Donor cell synchronisation and mitochondrial biomass
We used adult skin fibroblasts or amnion-derived fibroblasts (Fig. 1a) from donors who consented specifically to the use of their cells for NT experiments. While skin fibroblasts have obvious advantages of providing an easily accessible source of donor cells, amnion-derived cells have been found to express some stem cell markers23 and are more amenable to reprogramming during iPSC derivation24 and following NT in other species25. In relation to their therapeutic potential, it would in principle be possible to store amnion of babies born to women who carry mtDNA mutations.
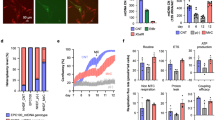
Cell cycle stage and mitochondrial biomass of fibroblasts used for NT.
(a) Brightfield images of human adult skin and amnion-derived fibroblasts in culture. Scale bar = 20 μm. (b) Flow cytometry traces showing enrichment for cells in G0/G1 in cell lines derived from adult skin or amnion. The top panel represents cells maintained at confluency for 4 days. The bottom panel represents cells cultured in 0.5% serum for 3–5 days. Each trace is representative of 3 independent experiments. Refer to Supplementary Figure S1 for an example flow cytometry trace for a population of proliferating amnion fibroblasts. (c) Skin fibroblasts stained with DAPI and MitoTracker Red. Cells maintained at confluency for 4 days (top panel) show increased mitochondrial biomass compared with those grown in 0.5% serum for 3–5 days (bottom panel). Scale bar = 10 μm.
Cell cycle arrest of somatic cells in G0/G1 is normally induced by a prolonged period of confluency and/or serum deprivation before they are used for NT26,27. FACS analysis indicates that both approaches are effective in enriching for G0/G1 cells (Fig. 1b, Supplementary Fig. S1). However, MitoTracker staining indicated that cells maintained at confluency for 4 days had an increased mitochondrial biomass compared with those cultured in serum-deprived medium (Fig. 1c). We therefore used serum deprivation with the aim of minimising somatic cell mtDNA content in NT embryos.
Source of human oocytes
Human oocytes that fail to undergo fertilisation in vitro, although abundant in supply, do not undergo efficient activation and onward development following NT28. We therefore used freshly harvested oocytes donated specifically for this research. The majority of oocytes (n = 239) were obtained from women (n = 26) undergoing IVF treatment who donated up to half of their oocytes, in return for a contribution from research funds towards the cost of their treatment29. A small number of oocytes (n = 26) were also donated by women (n = 9) who had oocytes harvested to reduce the risk of multiple pregnancy following ovarian stimulation and intra-uterine insemination. From a total of 265 oocytes donated for the experiments described here, 224 (85%) were arrested at meiosis II (MII) and were therefore considered suitable for use in NT.
Modification of NT procedures for human oocytes
Our starting point was to replicate the technical procedures used to produce NT-derived rhesus monkey ES cell lines30. Freshly harvested human oocytes were stripped of their cumulus cells and the spindle of those found to be arrested at MII (Fig. 2a) were visualised by polarised light birefringence (Fig. 2b and c) as previously described30. However, we found that a number of modifications were required to adapt NT procedures to human oocytes. Notably laser-assisted thinning of the zona pellucida was required to reduce the risk of oocyte lysis during enucleation. Furthermore, in initial experiments, we found that the electrical pulses (2 × 130 V) used to induce fusion in monkey NT30 resulted in poor rates of fusion with human oocytes. We therefore used inactivated Sendai virus extract, which includes fusogenic proteins of the viral envelope (HVJ-E, Hemagglutinating Virus of Japan – Envelope; Fig. 2d). While HVJ-E was highly effective in promoting successful fusion and oocyte survival, our preliminary experiments indicated that the potential for onward development was poor. Consistent with a recent report9, we found that development was improved by electroporation; the proportion of embryos arresting before the 8-cell stage was reduced by applying two electrical pulses (2 × 70 V) following HVJ-E-mediated fusion.
Enucleation, fusion and activation of human oocytes.
(a) Brightfield image of a human oocyte arrested at MII. Arrow indicates the first polar body (PB). (b) The MII spindle of a human oocyte visualized by polarized light birefringence. Inset shows enlarged MII spindle. (c) Human oocyte enucleated by aspirating a karyoplast containing the MII spindle into a beveled glass pipette using polarized light birefringence. Inset shows enlarged image of the karyoplast containing the MII spindle. (d) Brightfield image of an enucleated human oocyte showing the donor fibroblast bathed in HVJ-E solution positioned under the zona pellucida. Inset shows enlarged image of a fibroblast positioned for fusion. Scale bar = 100 μm. (e) Schematic drawing of the NT procedure. The specific modifications of monkey NT procedures for human oocytes are highlighted in green boxes. (f) Table showing the proportions of human oocytes undergoing successful enucleation, fusion and activation using the modified NT procedures for skin (n = 40) and amnion (n = 34) fibroblasts. Unmanipulated parthenotes (n = 33), activated using ionomycin and 6-DMAP, were included as controls. (g) Images show example cohorts of human parthenotes and NT embryos at 16–18 hr after exposure to ionomycin and 6-DMAP. Scale bar = 100 μm.
We next used these modified enucleation and fusion procedures (Fig. 2e) in a series of experiments to compare the efficiency of NT using adult skin and amnion-derived fibroblasts. The proportion of oocytes surviving enucleation and fusion were 87.5% for adult skin fibroblasts and 76.5% for amnion-derived fibroblasts (Fig. 2f).
To test the efficacy of the chemical activation procedures used to trigger the onset of embryonic development following NT in monkey oocytes30, we performed a series of experiments in which intact oocytes were treated with ionomycin and 6-DMAP (6-Dimethylaminopurine). As 6-DMAP inhibits formation of the second polar body in intact mammalian oocytes31, activation gives rise to diploid parthenotes containing two sets of maternal chromosomes. Activation was confirmed by the presence of pronuclei at 16–18 hours after exposure to ionomycin (Fig. 2g). By these criteria, activation was observed in 90.9% of parthenotes and in 91.4% and 80.8% of NT embryos that survived enucleation and fusion with adult skin and amnion fibroblasts respectively (not significant; Fig. 2f). These data indicate that chemical activation with ionomycin and 6-DMAP is effective in triggering exit from meiosis in human oocytes and that this is not adversely affected by replacement of the maternal genome with that of a fibroblast.
In initial experiments we observed 2–3 pronuclei (Fig. 2g) in a small proportion of NT embryos (9.4% and 14.3% respectively for skin and amnion fibroblasts). This was sometimes, but not always, associated with resorbtion of the first polar body. We therefore modified the enucleation procedure to include removal of the polar body as well as the spindle (Fig. 2e). In addition, multiple small nuclei (Fig. 2g) were observed in a proportion (12.5%) of skin-derived NT embryos and in 6.7% of parthenotes. This is likely due to chromosome scattering before exit from M-phase, which has been previously described following NT32.
To evaluate the effect of NT on embryonic development, we compared the proportions of activated oocytes reaching key developmental milestones (Fig. 3a). Recent evidence from studies in human embryos indicates that a first wave of embryonic transcription can be detected at the 2–4 cell stage and a further major wave occurs at the 6–10 stage33. Embryos incubated with the transcription inhibitor α-amanitin from the 1-cell stage can develop to the 4-cell stage34 or, occasionally, to the 4–8 cell stage when lower concentrations are used22. Whereas those incubated in α-amanitin from 4-cell stage either remain arrested with 4 cells, or progress to the 8-cell stage and develop no further34. These data imply that the oocyte stockpile of transcripts is sufficient to direct the first two rounds of cell division and is consistent with the second wave of transcription being essential for development beyond the 8-cell stage.
Development of human parthenotes and NT embryos.
(a) Schematic drawing showing development of chemically activated human embryos from the pronucleate to the blastocyst stage. Development to the 4–8 cell stage is driven by the oocyte stockpile of maternal transcripts whereas development to later stages requires transcription from the embryonic genome. (b) Images show examples of embryos on days 2–4 following activation of unmanipulated oocytes (parthenotes) and NT embryos derived from either skin or amnion fibroblasts. Scale bar = 100 μm. (c) Proportion of parthenotes and NT embryos developing to each developmental stage during 6 days of culture following activation. (d) Images showing examples of morulae and blastocysts derived from parthenotes and NT embryos as indicated. Scale bar = 50 μm.
To determine the effect of NT on the maternally-directed early cleavage divisions, we compared the proportion of oocytes arresting with ≤ 4 cells following activation alone, or following NT using skin or amnion fibroblasts. We found that the majority (62.5%) of skin fibroblast-derived NT embryos arrested with ≤ 4 cells compared with only 23.8% of those derived from amnion fibroblasts (P < 0.01; Fig. 3b and 3c). The frequency of arrest at the 4-cell stage did not differ between amnion-derived embryos and control parthenotes (Fig. 3c), indicating that the NT procedure per se does not prevent completion of the maternally-directed early cleavage divisions. The high incidence of arrest in skin fibroblast-derived embryos may reflect a response to chromosome segregation errors, which are prevalent in NT embryos32,35, or to DNA damage, which may be increased in adult skin fibroblasts compared with amnion-derived fibroblasts.
Of the NT embryos that progressed beyond the 4-cell stage, the majority arrested during the next round of cell division with ≤ 8 cells (Fig. 3c). Nonetheless, compaction of embryonic cells, which marks the first step in their allocation to the ICM and trophectoderm lineages36, was observed in 6/32 (18.75%) of skin fibroblast-derived embryos (Fig. 3c and 3d), but they did not develop further. Interestingly, all of these were obtained from the same egg donor and compaction occurred when the embryo contained relatively few cells. Blastocyst formation was observed in 1/21 (4.7%) of the amnion-derived NT embryos, compared with 7/30 (23.3%) of parthenotes (Fig. 3e). While the amnion-derived blastocyst had a discernible ICM, the trophectoderm appeared to contain few cells (Fig. 3e). This is consistent with reports of impaired trophoblast formation in mouse NT embryos37. Explantation of the NT-derived ICM in conditions found to support successful derivation of human ES cell lines38, did not result in the formation of a viable ES colony.
Donor cell mtDNA persistence in NT embryos
In the context of mtDNA disease, the therapeutic potential of SCNT depends not only on maximising blastocyst formation, but also on minimising persistence of donor cell mtDNA. According to our current understanding mtDNA does not replicate during the early stages of embryonic development39, however, earlier replication has been reported in some species following NT19. We therefore measured the fibroblast mtDNA content in NT embryos (n = 23) containing ≥ 4 cells obtained from 7 oocyte donors using amnion (n = 14) and skin (n = 9) fibroblasts as donor cells.
The mtDNA genotypes were determined by sequencing the highly variable D-loop from donor fibroblasts and from ovarian follicle cells obtained from the oocyte donors (Fig. 4a). Based on single nucleotide polymorphic variants (Fig. 4b), we determined the levels of fibroblast mtDNA contained in SCNT embryos by last hot cycle PCR restriction fragment length polymorphism (RFLP). Donor cell mtDNA was below the limit of detection (<0.5%) in 60% of NT embryos (Fig. 4c and 4d) and the maximum level was 0.2% (Fig. 4d). Persistence of fibroblast mtDNA was similar between skin and amnion fibroblasts (Fig. 4d).
Persistence of fibroblast mtDNA in NT embryos.
(a) Sequence electropherogram of the mtDNA non-coding control region highlighting the presence of a polymorphism (m.497 C>T) in amnion-derived fibroblast cells. This polymorphism was not detected in oocyte donor mtDNA obtained from granulosa cells aspirated from ovarian follicles during the oocyte retrieval procedure. (b) Schematic illustrating restriction fragment length polymorphism (RFLP) designed using the m.497C>T sequence variant. In the presence of the m.497C>T variant, digestion with the AciI enzyme generates 2 fragments (44 + 115 bp). In the absence of the polymorphism 3 fragments (27, 44 + 88 bp) are created. (c) Radiolabeled restriction fragments were separated by 12% non-denaturing polyacrylamide gel electrophoresis. E1–E7, NT embryos; OD, ovarian granulosa cells from the oocyte donor; F, amnion fibroblasts; and U, uncut labelled PCR product. The percentage level of fibroblast mtDNA detected in each embryo is indicated (E1–E7). In this example, the fibroblast mtDNA content was below the limit of detection (<0.5%) in 4/7 embryos. (d) Graph showing the level of fibroblast mtDNA as a percentage of the entire mtDNA content of NT embryos derived using skin (n = 9) or amnion (n = 14) fibroblasts. Fibroblast mtDNA was below the limit of detection (<0.5%) in 6/9 skin fibroblast-derived NT embryos and in 6/14 amnion-derived NT embryos. (e) Table shows mtDNA copy number measured in individual enucleated oocytes (n = 8) and in single skin (n = 9) amnion (n = 9) fibroblasts. (f) Graph shows hypothetical levels of fibroblast mtDNA in NT embryos based on all possible combinations of enucleated oocytes and skin (n = 72) or amnion (n = 72) fibroblasts in which mtDNA copy number was measured.
To gain insight into whether the non-detectable levels observed in the majority of embryos was due to elimination or dilution of donor cell mtDNA, we measured mtDNA copy number in enucleated oocytes, single skin fibroblasts and single amnion fibroblasts (Fig. 4e). Consistent with previous reports18, enucleated oocyte mtDNA copy number varied by an order of magnitude. Skin and amnion fibroblasts had similar mtDNA copy numbers (1,750 ± 1,071 and 1,517 ± 870 respectively). Assuming that the donor cell mtDNA is neither preferentially replicated nor eliminated, we calculated the expected level of donor cell mtDNA based on all possible combinations of donor cells and enucleated oocytes (n = 144; Fig. 4f). We found that 80% of values were below the limit of detection (<0.5%). This suggests that the non-detectable levels in 60% of NT embryos (Fig. 4d) represent dilution rather than elimination of donor cell mtDNA. Moreover, the range of values fitted closely with the observed levels of donor cell mtDNA in NT embryos (Fig. 4d and 4f). Together, these data indicate that the donor cell mtDNA accounts for a very low proportion of the total mtDNA in NT embryos and suggest that it is neither eliminated nor preferentially replicated following NT.
While the low levels of fibroblast mtDNA present during early development of NT embryos are encouraging, we cannot rule out the possibility of preferential replication at later developmental stages. Because of the limited development of NT embryos to later stages, it was not possible to test this directly. However, low levels of heteroplasmy have been reported in ES cells from embryos generated by chemical activation following transplantation of MII spindles between oocytes16,40. We therefore compared the mtDNA copy numbers of MII spindle-containing karyoplasts and fibroblasts (Fig. 5a). The mtDNA copy number of karyoplasts (n = 21) was highly variable (Fig. 5a), ranging from 484 to 95,242 and generally higher than that of fibroblasts (Fig. 5a). Consistent with this, we found that the levels of heteroplasmy in embryos created following reciprocal transfer of MII spindles between oocytes from different donors (Fig. 5b) exceeded those of fibroblast-derived NT embryos (Fig. 4c, 4d and 5e). In contrast to NT embryos, in which donor cell mtDNA was below the limits of detection (0.5%) in 60% of embryos (Fig. 3d), we found that karyoplast-associated mtDNA was below the limits of detection in only 1/10 embryos derived from MII spindle transfer oocytes (n = 14; Fig. 5e). The levels measured in the remaining 9/10 embryos varied from 1–3% (Fig. 5e). These data together with the finding that ES cells derived following MII spindle transfer contain very low levels of heteroplasmy16,40 argue in favour of the therapeutic potential of NT for mtDNA disease.
mtDNA content of embryos derived following reciprocal transfer of MII spindles between oocytes.
(a) Box plot shows mtDNA copy number in skin (n = 9) and amnion (n = 9) fibroblasts and in MII spindle karyoplasts (n = 21). (b) Schematic drawing showing reciprocal transfer of MII spindles between oocytes. (c) Sequence electropherogram of the mtDNA non-coding control region highlighting the presence of a polymorphism (m.16239C>T) in granulosa cells obtained from one of two oocyte donors (OD1 and OD2). (d) Schematic illustrating RFLP designed using the m.16239C>T sequence variant. In the presence of the m.16239C>T variant, digestion with the MseI enzyme generates 3 fragments (32, 59 + 80 bp). In the absence of the polymorphism 2 fragments (32 + 139 bp) are created. (e) Radiolabeled restriction fragments were separated by 12% non-denaturing polyacrylamide gel electrophoresis. mtDNA was amplified from spindle transfer embryos and ovarian granulosa cells from two oocyte donors. Embryos E1–E5 were generated by artificial activation of enucleated oocytes from OD1 fused with spindle karyoplasts from OD2. Embryos E6–E10 were generated by artificial activation of enucleated oocytes from OD2 fused with karyoplasts from OD1. U, uncut labelled PCR product.
Discussion
Data presented here suggest that fibroblast mtDNA is neither eliminated nor preferentially replicated following NT in human oocytes, but is below the limit of detection in the majority of NT embryos, most likely due to dilution by the high mtDNA copy numbers present in oocytes. Our results, along with more recent technical advances in human NT9, provide proof of principle for the use of NT to develop isogenic ES cell-based therapies for the treatment of degenerative disease associated with mtDNA mutations. Furthermore, we demonstrated that the problem of obtaining human oocytes can be overcome by donation through an “egg-sharing” scheme, which has a high level of acceptance among women undergoing IVF treatment29,41.
Our findings reveal new insights into the importance of donor cell type and method of preparation for realising the therapeutic potential of NT for mtDNA disease. First, we find that mitochondrial biomass is increased in cells maintained at confluency, which is a commonly used approach to induce cell cycle arrest in G1/G0 in preparation for NT26,27. Thus, the level of donor cell mtDNA in NT-derived ES cell lines is likely to be reduced by using serum deprivation rather than confluency to induce arrest of donor cells in G1/G0. While serum deprivation has been reported to induce donor cell apoptosis42,43, serum deprived donor cells were used to generate human NT-derived ES cells in the recent report from Tachibana et al9. Second, we find that the use of adult skin-derived fibroblasts has a detrimental effect on development to the 4-cell stage, whereas the use of amnion-derived fibroblasts does not. This indicates that the maternally-directed phase of human embryonic development is not adversely affected by the NT procedures per se, but that predisposing factors in fibroblasts from adult skin induce arrest during the early embryonic cell divisions. This may be linked to chromosome segregation errors as multiple small nuclei, which are indicative of chromosome scattering, were observed in activated oocytes following transplantation of skin but not amnion fibroblasts. The recent derivation of ES cell lines from NT embryos9 was achieved using foetal dermal fibroblasts which, in common with amniotic fibroblasts, may be more amenable to reprogramming and less prone to chromosome segregation errors. The therapeutic potential of amnion-derived cells could be realised by storing amnion from babies born to affected women.
Our findings indicate that fibroblast mtDNA persists at a very low level due to dilution by the oocyte vast stock of mtDNA. While we cannot rule out the possibilities that donor cell mtDNA content might be affected by embryo viability, or that amplification of donor cell derived mtDNA might occur at later stages, the latter seems unlikely for a number of reasons. First, a number of ES cell lines generated following MII spindle transfer in human oocytes show very low levels of heteroplasmy16,40 and our data indicate that the levels of heteroplasmy are lower following NT using either skin or amnion fibroblasts compared with MII spindle transfer. Furthermore, somatic cell mtDNA was not detected in two triploid ES cell lines generated from combined maternal and somatic cell genomes8.
The question of whether human NT can, in principle, be used to develop cell-based therapies for degenerative disease associated with mtDNA mutations, has not previously been addressed. Together, our data indicate that NT-derived ES cell lines are likely to carry very low levels of mtDNA mutation and are therefore likely to be suitable for therapeutic use. This approach would provide a means of reprogramming somatic cells to treat degenerative disease caused by mtDNA mutations, with a greatly reduced risk of mtDNA mutations persisting in the cells used for transplantation. Help for affected families may also be possible through the development of assisted reproductive techniques involving the transfer of parental chromosomes between fertilised15 or unfertilised16 eggs, to prevent transmission of mtDNA mutations from mother to child.
Methods
Licensing and approvals
The study was approved by the Local Research Ethics Committee and was licensed by the UK Human Fertilisation and Embryology Authority.
Fibroblast culture and preparation for NT
Skin and amniotic fibroblast cell lines were established from tissue obtained with consent from women undergoing caesarean section. Tissue was chopped, treated with collagenase and cultured in DMEM + 10% FBS (Life Technologies). Cells were split using TrypLE Select (Life Technologies) and frozen in 10% DMSO at passages 1–4. To induce arrest in G1/G0 by serum deprivation, cells were thawed and cultured for two days in DMEM + 10% FBS, followed by 3–5 days of serum starvation in 0.5% FBS. Cells arrested in G0/G1 by growing to confluency were cultured for 3–5 days and left for 4 days at confluency. For both treatments cells were trypsinised and washed in G-MOPS medium (Vitrolife) immediately before nuclear transfer.
DNA content analysis of fibroblasts
Following serum starvation for four days, or culture to confluency, cells were dissociated with TrypLE (Invitrogen). Dissociated cells were treated with nuclear extraction buffer (Partec), vortexed and suspended in extraction buffer (Partec) for 15 minutes at room temperature. DNA was stained with DAPI (Partec) for 30 minutes and donor cell DNA content was determined with a FACSCanto flow cytometer and Diva software (BD Biosciences).
Mitochondrial staining
Cells were incubated in culture medium containing 100 nM of MitoTracker Red (Life Technologies) for 30 minutes at 37°C. For brightfield images, fibroblasts were imaged using a Nikon Eclipse TE2000-U inverted microscope, a 20x/0.40 NA Plan Fluor objective and a cooled charge-coupled device (CCD) camera (Oosight, CRi). For immunofluorescence, fibroblasts were imaged using a Nikon Eclipse TE2000-U inverted microscope, a 10x/0.30 NA Plan Fluor objective and a Photometrics CoolSnapHQ interline CCD camera (Roper Scientific). Fibroblasts were illuminated using a Xenon bulb (Osram) and appropriate filter sets (DAPI: Excitation 2Ax UV, Emission 2 Am, Dichroic UV-2A (Nikon). MitoTracker Red: Excitation S580/20x, Emission S630/60m, Dichroic 86006 bs (Chroma Tech)). The filter wheels and focus controllers (Prior Scientific) were driven by MetaMorph (Universal Imaging) imaging software. Images were acquired using MetaMorph and processed using ImageJ software.
Oocyte preparation for nuclear transfer
Oocytes were donated by women undergoing infertility treatment, either as part of an egg sharing programme or following egg collection to prevent multiple pregnancy following ovarian stimulation with exogenous gonadotrophins and intra-uterine insemination. Consenting women shared if ≥6 oocytes were harvested. Women, from whom 5 or fewer oocytes were collected, did not donate eggs but still received subsidised treatment29. Oocytes were collected by ultrasound-guided follicle aspiration. The surrounding cumulus cells were removed using hyaluronidase (HyAse, Vitrolife). MII oocytes were identified by the presence of the first polar body. Oocytes were incubated in G1 Plus medium (Vitrolife) at 37°C and 7% CO2 until nuclear transfer.
Nuclear transfer
Micromanipulations were carried out on a Nikon TE-2000U, equipped with Oosight birefringence imaging system (CRi), Integra Ti micromanipulators and SAS air microinjectors (Research Instruments), Saturn Active thermal laser (Research Instruments). The micromanipulation microscope, dissection microscope and incubators were contained within an enclosed workstation in which the ambient conditions were maintained at 37°C and 7% CO244 (Vitrosafe Ltd). MII oocytes were transferred into the 5 μl droplets of G1 medium (Vitrolife) containing 2.5 μg/ml of cytochalasin B (Sigma) under mineral oil (Vitrolife), in glass-bottomed petri dishes (Willco). For enucleation, the spindle was visualised by polarised light birefringence (Oosight, CRI). Oocytes were positioned on a holding pipette with the spindle at the 4 o'clock position, where the zona pellucida was thinned at a point adjacent to the spindle using a Saturn Active laser (Research Instruments). A bevelled and spiked glass enucleation pipette with inner diameter of 20 μm was pushed through the thinned zona pellucida and the first polar body was removed before the spindle was aspirated into the pipette. The pipette was then slowly withdrawn, resulting in the “budding off” of a spindle karyoplast without breaking the oocyte membrane. A suspension of prepared fibroblasts was placed in a small droplet of culture medium supplemented with 0.5% HSA. In selecting cells for fusion, we avoided abnormally small or large cells, or those showing signs of membrane blebbing. Individual cells were briefly placed in a suspension of HVJ-E, GenomONE-CF Ex (Cosmo Bio, Japan) and were aspirated into the pipette with one to two cell volumes of HVJ-E suspension. The oocyte was positioned on the holding pipette with the polar body at the 9 o'clock position. The zona pellucida was thinned with the laser at the 3 o'clock position and the pipette containing the fibroblast cell was pushed through the remaining zona pellucida. The cell and HVJ-E solution were gently expelled and the pipette withdrawn. Oocyte-somatic cell couplets were then incubated for 1–2 hours prior to activation.
Embryos were imaged using either a Nikon SMZ-1000 stereo microscope on 8× magnification with a Nikon Digital Sight camera, or a Nikon Eclipse TE2000-U inverted microscope, a 40x/0.60 NA PLAN APO objective and a cooled CCD camera (Oosight, CRi).
MII spindle transfer
Enucleation was carried out as described and karyoplasts containing spindles were fused with the enucleated oocytes using HVJ-E as described above.
Activation and culture
NT oocytes were checked for membrane fusion prior to activation and were transferred to 50:50 G1: Fusion medium (0.25 M D-Sorbitol, 100 μM CaOAc, 500 μM MgOAc) and then straight fusion medium. One to three NT oocytes were transferred between the electrodes of a BTX microslide chamber filled with 25 ml of fusion medium. Two 70 V 50 μs electrical pulses were generated by a BTX 830 electrofusion generator (BTX), fused couplets were then transferred to 50:50 G1: Fusion medium and then into G1 culture medium.
Chemical activation was a four-step procedure. NT oocytes were first placed in a solution of G1 with 1 mg/ml HSA, 5 μM ionomycin and 1 mM 6-dimethylaminopurine (6-DMAP) for five minutes, then G1 with 30 mg/ml HSA, 1 mM 6-DMAP for 5 mins, followed by G1 with 10 mg/ml HSA and 1 mM 6-DMAP for 5 hours. Embryos were then carefully washed in G1 with 10 mg/ml HSA and cultured in G1 medium until day 3, when they were transferred to G2 culture medium (Vitrolife) supplemented with 10 mg/ml HSA. Arrested embryos were individually transferred to PBS in a PCR tube and stored at −20°C for mtDNA analysis.
mtDNA Sequencing
The mitochondrial genotypes of fibroblasts and granulosa cells isolated from the ovarian follicular fluid of oocyte donors were identified by sequencing the highly variable D-loop. DNA extraction was performed using the QIAamp® DNA Micro kit (Qiagen, Crawley, UK) as per the manufacturer's instructions. Isolated DNA (1 μl) was amplified in 25 μl reaction volumes using overlapping M13-tailed primers targeted to the mtDNA D-loop (primer sets D1, D2, D3 and D4), as described previously45. PCR product clean-up was carried out with ExoSAP-IT (GE Healthcare, Buckinghamshire, UK). Samples were then sequenced using BigDye Terminator v3.1 Cycle Sequencing chemistry (Applied Biosystems) on an ABI3100 Genetic Analyzer (Applied Biosystems). The resulting sequences were compared to the revised Cambridge Reference Sequence for human mtDNA (GenBank reference NC_012920.1) using SeqScape software (v2.6. Applied Biosystems).
Measurement of fibroblast and spindle karyoplast mtDNA content in NT and ST embryos
The levels of mtDNA carryover from somatic cells and spindle karyoplasts in reconstituted embryos were determined by last hot cycle PCR/restriction fragment length polymorphism (RFLP) analysis. Embryos were first lysed for 2 hours at 55°C in 15 μl of a standard lysis buffer (50 mM Tris-HCl, pH 8.5, 1 mM EDTA, 0.5% Tween-20 and 200 μg/ml proteinase K), then incubated at 95°C for 10 minutes to inactivate the enzyme. Last hot cycle PCR/RFLP assays were designed to target specific polymorphisms that differed between the recipient oocyte and donor fibroblasts or spindle karyoplasts and that resulted in the creation or removal of a restriction enzyme recognition site. The assays were performed as described previously46 with 1 μl embryo lysate and the modifications detailed in Table 1.
MtDNA carryover was calculated as a percentage of total mtDNA present in reconstituted embryos.
Measurement of mtDNA copy number
Amniotic and adult skin fibroblasts arrested in G0/G1 were selected using the same criteria as for NT and frozen individually in PBS in PCR tubes. Enucleated oocytes and spindle karyoplasts were frozen individually in PCR tubes. MtDNA copy number was determined by real-time PCR using established TaqMan probe and PCR primers targeting the MT-ND1 gene of the mitochondrial genome47. Reactions were composed of 1 μl lysate, 300 nM forward and reverse primers, 100 nM fluorogenic probe and 12.5 μl TaqMan Universal MasterMix (Applied Biosystems). Samples were analysed in triplicate. Mitochondrial DNA copy numbers for enucleated oocytes, spindle karyoplasts and fibroblasts were calculated from a standard curve produced from triplicate 10-fold serial dilutions of an MT-ND1 template (nucleotide positions: 3017–4057) of known copy number.
Statistical analysis
Data are means ± s.d. Statistical analysis was performed using a student's t-test. P < 0.05 was considered significant.
References
Gurdon, J. B., Elsdale, T. R. & Fischberg, M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature. 182, 64–65 (1958).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature. 292, 154–156 (1981).
Chidgey, A. P., Layton, D., Trounson, A. & Boyd, R. L. Tolerance strategies for stem-cell-based therapies. Nature. 453, 330–337 (2008).
Rodriguez, R. M. et al. in Stem Cell Transplantation Advances in Experimental Medicine and Biology [Vol. 741] 276–289 (Springer US, 2012).
Stojkovic, M. et al. Derivation of a human blastocyst after heterologous nuclear transfer to donated oocytes. Reprod Biomed Online. 11, 226–231 (2005).
French, A. J. et al. Development of human cloned blastocysts following somatic cell nuclear transfer with adult fibroblasts. Stem Cells. 26, 485–493 (2008).
Li, J. Y. et al. Human Embryos Derived by Somatic Cell Nuclear Transfer Using an Alternative Enucleation Approach. Cloning Stem Cells. 11, 39–50 (2009).
Noggle, S. et al. Human oocytes reprogram somatic cells to a pluripotent state. Nature. 478, 70–75 (2011).
Tachibana, M. et al. Human embryonic stem cells derived by somatic cell nuclear transfer. Cell. 153, 1228–1238 (2013).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 126, 663–676 (2006).
Taylor, R. W. & Turnbull, D. M. Mitochondrial DNA mutations in human disease. Nat. Rev. Genet. 6, 389–402 (2005).
Schaefer, A. M. et al. Prevalence of mitochondrial DNA disease in adults. Ann. Neurol. 63, 35–39 (2008).
Elliott, H. R., Samuels, D. C., Eden, J. A., Relton, C. L. & Chinnery, P. F. Pathogenic mitochondrial DNA mutations are common in the general population. Am. J. Hum. Genet. 83, 254–260 (2008).
Pfeffer, G., Majamaa, K., Turnbull, D. M., Thorburn, D. & Chinnery, P. F. Treatment for mitochondrial disorders. Cochrane Database of Systematic Rev. 4, 10.1002/14651858.CD004426.pub3 (2012).
Craven, L. et al. Pronuclear transfer in human embryos to prevent transmission of mitochondrial DNA disease. Nature. 465, 82–85 (2010).
Tachibana, M. et al. Towards germline gene therapy of inherited mitochondrial diseases. Nature. 493, 627–631 (2013).
Giles, R. E., Blanc, H., Cann, H. M. & Wallace, D. C. Maternal inheritance of human mitochondrial DNA. Proc. Natl. Acad. Sci. U. S. A. 77, 6715–6719 (1980).
Wai, T. et al. The role of mitochondrial DNA copy number in mammalian fertility. Biol. Reprod. 83, 52–62 (2010).
St John, J. C., Facucho-Oliveira, J., Jiang, Y., Kelly, R. & Salah, R. Mitochondrial DNA transmission, replication and inheritance: a journey from the gamete through the embryo and into offspring and embryonic stem cells. Hum. Reprod. Update. 16, 488–509 (2010).
Evans, M. J. et al. Mitochondrial DNA genotypes in nuclear transfer-derived cloned sheep. Nat. Genet. 23, 90–93 (1999).
Al Rawi, S. et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science. 334, 1144–1147 (2011).
Egli, D. et al. Reprogramming within hours following nuclear transfer into mouse but not human zygotes. Nat Commun. 2, 488 (2011).
Miki, T., Lehmann, T., Cai, H., Stolz, D. B. & Strom, S. C. Stem cell characteristics of amniotic epithelial cells. Stem Cells. 23, 1549–1559 (2005).
Easley, C. A. et al. Human Amniotic Epithelial Cells are Reprogrammed More Efficiently by Induced Pluripotency than Adult Fibroblasts. Cell Reprogram. 14, 193–203 (2012).
Okahara-Narita, J., Tsuchiya, H., Takada, T. & Torii, R. Cloned blastocysts produced by nuclear transfer from somatic cells in cynomolgus monkeys (Macaca fascicularis). Primates. 48, 232–240 (2007).
Kasinathan, P., Knott, J. G., Wang, Z., Jerry, D. J. & Robl, J. M. Production of calves from G1 fibroblasts. Nat. Biotechnol. 19, 1176–1178 (2001).
Kishigami, S. et al. Production of cloned mice by somatic cell nuclear transfer. Nat. Protoc. 1, 125–138 (2006).
Hall, V. J. et al. Developmental competence of human in vitro aged oocytes as host cells for nuclear transfer. Hum. Reprod. 22, 52–62 (2007).
Choudhary, M. et al. Egg sharing for research: a successful outcome for patients and researchers. Cell Stem Cell. 10, 239–240 (2012).
Byrne, J. A. et al. Producing primate embryonic stem cells by somatic cell nuclear transfer. Nature. 450, 497–502 (2007).
Szollosi, M. S. et al. Inhibition of protein kinases by 6-dimethylaminopurine accelerates the transition to interphase in activated mouse oocytes. J. Cell Sci. 104 (Pt 3), 861–872 (1993).
Simerly, C. et al. Molecular correlates of primate nuclear transfer failures. Science. 300, 297 (2003).
Vassena, R. et al. Waves of early transcriptional activation and pluripotency program initiation during human preimplantation development. Development. 138, 3699–3709 (2011).
Braude, P., Bolton, V. & Moore, S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature. 332, 459–461 (1988).
Mizutani, E. et al. Abnormal chromosome segregation at early cleavage is a major cause of the full-term developmental failure of mouse clones. Dev. Biol. 364, 56–65 (2012).
Cockburn, K. & Rossant, J. Making the blastocyst: lessons from the mouse. J. Clin. Invest. 120, 995–1003 (2010).
Lin, J. et al. Defects in trophoblast cell lineage account for the impaired in vivo development of cloned embryos generated by somatic nuclear transfer. Cell Stem Cell. 8, 371–375 (2011).
Prathalingam, N. et al. Production and validation of a good manufacturing practice grade human fibroblast line for supporting human embryonic stem cell derivation and culture. Stem Cell. Res. Ther. 3, 12 (2012).
Thundathil, J., Filion, F. & Smith, L. C. Molecular control of mitochondrial function in preimplantation mouse embryos. Mol. Reprod. Dev. 71, 405–413 (2005).
Paull, D. et al. Nuclear genome transfer in human oocytes eliminates mitochondrial DNA variants. Nature. 493, 632–637 (2013).
Haimes, E., Taylor, K. & Turkmendag, I. Eggs, ethics and exploitation? Investigating women's experiences of an egg sharing scheme. Sociol. Health Illn. 34, 1199–1214 (2012).
Dalman, A. et al. Synchronizing cell cycle of goat fibroblasts by serum starvation causes apoptosis. Reprod Domest Anim. 45, e46–53 (2010).
Miranda Mdos, S. et al. Serum-starved apoptotic fibroblasts reduce blastocyst production but enable development to term after SCNT in cattle. Cloning Stem Cells. 11, 565–573 (2009).
Hyslop, L. et al. A novel isolator-based system promotes viability of human embryos during laboratory processing. PLoS One. 7, e31010 (2012).
Tuppen, H. A. et al. The p.M292T NDUFS2 mutation causes complex I-deficient Leigh syndrome in multiple families. Brain. 133, 2952–2963 (2010).
Taylor, R. W. et al. A homoplasmic mitochondrial transfer ribonucleic acid mutation as a cause of maternally inherited hypertrophic cardiomyopathy. J. Am. Coll. Cardiol. 41, 1786–1796 (2003).
He, L. et al. Detection and quantification of mitochondrial DNA deletions in individual cells by real-time PCR. Nucleic Acids Res. 30, e68 (2002).
Acknowledgements
We thank the patients who donated oocytes for use in this research. We also thank the Research Nurses Maria Nesbitt and Linda Burgess and the embryology and clinical team at Newcastle Fertility Centre. We acknowledge help and advice given by Dr Shoukhrat Mitalipov, Oregon Health & Science University. The work was funded by the Medical Research Council (Grant Reference Number G0601157) and the Wellcome Trust (Grant Reference Number 096919/Z/11/Z).
Author information
Authors and Affiliations
Contributions
M.H. and A.P.M. conceived the study. A.P.M. conceived and directed the oocyte donation program. M.H., G.D.G. and L.M.L. D.M.T. and H.A.L.T. designed experiments. G.D.G. and L.M.L. performed the NT experiments. Z.P. refined the fusion procedures. G.D.G. and Q.Z. established fibroblast cultures and Q.Z. and L.M.L. prepared and tested cells for use in NT. L.A.H. coordinated donation of oocytes. N.P. processed embryos for ES cell derivation. H.A.L.T., L.H.N., L.C. and D.M.T. performed the analysis of mtDNA. G.D.G., L.M.L., H.A.L.T., L.A.H., Q.Z. and M.H. analysed the data. M.H., G.D.G., L.M.L. and H.A.L.T. prepared the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figure S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Greggains, G., Lister, L., Tuppen, H. et al. Therapeutic potential of somatic cell nuclear transfer for degenerative disease caused by mitochondrial DNA mutations. Sci Rep 4, 3844 (2014). https://doi.org/10.1038/srep03844
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03844
This article is cited by
-
Double sperm cloning (DSC) is a promising strategy in mammalian genetic engineering and stem cell research
Stem Cell Research & Therapy (2020)
-
Patient-specific neural progenitor cells derived from induced pluripotent stem cells offer a promise of good models for mitochondrial disease
Cell and Tissue Research (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.