Abstract
In order to apply human embryonic stem cells (hESCs) and induced pluripotent stem cells (hiPSCs) to regenerative medicine, the cells should be produced under restricted conditions conforming to GMP guidelines. Since the conventional culture system has some issues that need to be addressed to achieve this goal, we developed a novel culture system. We found that recombinant laminin-511 E8 fragments are useful matrices for maintaining hESCs and hiPSCs when used in combination with a completely xeno-free (Xf) medium, StemFit™. Using this system, hESCs and hiPSCs can be easily and stably passaged by dissociating the cells into single cells for long periods, without any karyotype abnormalities. Human iPSCs could be generated under feeder-free (Ff) and Xf culture systems from human primary fibroblasts and blood cells and they possessed differentiation abilities. These results indicate that hiPSCs can be generated and maintained under this novel Ff and Xf culture system.
Similar content being viewed by others
Introduction
Human embryonic stem cells (hESCs) and induced pluripotent stem cells (hiPSCs) hold promise as tools for regenerative medicine. Recently, several reports have discussed the potential use of stem cells in clinical applications. Geron has initiated treatment of neural disease using neuronal cells derived from hESCs. Advanced Cell Technologies is making efforts to treat eye diseases with ESC-derived cells1. This approach involves the production of retinal pigment epithelium from hESCs that are then transplanted into patients. Regenerative medicine using stem cells, particularly pluripotent stem cells, will certainly advance over the coming years as new discoveries are made.
Researchers usually use feeder cells and serum-containing medium in conventional culture systems for hESCs and hiPSCs2,3. Murine-derived feeder cells are widely used to maintain hESCs and hiPSCs. Human-derived feeder cells are also used for hESC/iPSC culture; however, in some cases, these cells have proven unsuitable for stem cell maintenance4,5. The feeder cell preparation requires significant time and effort. Fetal bovine serum (FBS)-containing medium is normally used for the culture of feeder cells. The reduction or complete removal of serum and animal-derived products is required to satisfy Standard for Biological Ingredients. Moving towards feeder-free culture systems for hESCs and hiPSCs would represent a significant improvement over conventional culture systems.
To address these issues, we sought to develop a novel culture system applicable for human stem cell maintenance and hiPSC derivation. Feeder-free (Ff) and xeno-free (Xf) conditions appear to be acceptable for culturing hESCs and hiPSCs. Various matrices can be used to replace feeder cells, such as Matrigel6,7,8, CELLstart9,10, recombinant proteins11,12,13 and synthetic polymers14,15. Xeno-free media are also available commercially, including TeSR2, NutriStem and Essential E8 medium13, among others. Although we examined most of these materials with respect to whether the hESCs and hiPSCs could be stably and efficiently cultivated in our laboratory, we were unable to identify an efficacious combination of matrix and medium.
It has previously been reported that laminin-511 supports the stable culture of hESCs and hiPSCs11. Recently, a shorter fragment of laminin-511, referred to as the laminin-511 E8 fragment (LN511E8), was also shown to efficiently maintain hESCs and hiPSCs12. Recombinantly expressed LN511E8 (rLN511E8) is isolated more easily and with a greater yield and purity, than full-length laminin-511. For these reasons, we chose rLN511E8 as a matrix for our novel culture system for hESCs and hiPSCs. Next, we examined whether a new xeno-free medium, StemFit™, could be used for our novel culture system with rLN511E8.
Employing these materials, we successfully developed a novel culture system for hESCs and hiPSCs using rLN511E8 and StemFit™ that is easy to use, expandable and reproducible, as clinical-grade hiPSCs must be manufactured according to Standard Operating Procedures (SOPs) in order to meet Cell Processing Center (CPC) standards.
Human ESCs and iPSCs were stably passaged for long periods by dissociating the cells into single cells. Moreover, hiPSCs were successfully established from primary fibroblasts, peripheral blood and cord blood under these conditions using episomal vectors16,17. These Ff-hiPSCs displayed the capacity to differentiate into various types of somatic cells, including all three germ layers. These results indicate that Ff-hiPSCs are suitable for manufacturing in a CPC setting and should prove useful for future research and clinical applications.
Results
Development of a novel culture system for hiPSCs
To develop feeder-free (Ff) and xeno-free (Xf) hiPSC culture conditions, we tested Matrigel, CellStart and the recombinant laminin-511 E8 fragment (rLN511E8) as coating matrices12. H9 hESCs were dissociated into single cells and plated onto the coated culture plates. The hESCs efficiently formed colonies on rLN511E8 but not on the other matrices (Figure S1A). We therefore selected rLN511E8 as the coating matrix for our system. Using rLN511E8, we attempted to cultivate hiPSCs using a variety of commercially available Xf-medium (Figure S1B). TeSR2 did not support the maintenance of hiPSCs (32R118) on rLN511E8. When we used NutriStem, the hiPSCs formed flattened colonies. Although the mixture of TeSR2 and NutriStem supported hESC-like colony formation, the morphology was not good (many granules were detected in cells). Since we were unable to obtain good results, we chose to try StemFit™, a newly developed Xf-medium for hiPSC culture from Ajinomoto Co., Inc. Using StemFit™, we obtained hiPSCs colonies similar to those cultivated on feeder cells2 (Figure S1B).
We examined whether hESCs and hiPSCs, which were previously established and maintained on feeder cells, can be cultivated under the Ff and Xf conditions using rLN511E8 matrix and StemFit™ (Figure 1A). After two or three passages, most of the hESCs and hiPSCs adapted to the Ff and Xf culture conditions. The combination of rLN511E8 and StemFit™ demonstrated efficacy for the hESCs and hiPSC culture.
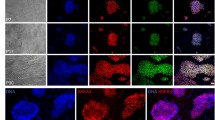
The feeder-free (Ff) and xeno-free (Xf) culture system for hES/iPS cells.
(A) The morphology of hESCs (KhES1) and hiPSCs (32R1, 414C231, 585A131 and 606A131) cultivated on rLN511E8-coated cell culture plates with StemFit™. Scale bar, 100 μm. (B) The growth curves of the hESCs (H9 and KhES1) and hiPSCs (201B7) cultured under Ff and Xf conditions. Each dot represents a passage of cells. (C) The morphology of the indicated passage numbers of 201B7 and H9 cells. The 201B7 and H9 cells at passage numbers p80 and p54, respectively, were immunostained for the indicated pluripotency markers, followed by phase contrast imaging (Ph). Scale bar, 100 μm.
Human iPSCs were then dissociated into single cells and reproducibly plated according to the exact cell number (Figure S2A), an important consideration for standardizing culture conditions and developing a reliable experimental design. The cells cultured on rLN511E8 became confluent within 8–10 days after plating (the average fold change was 132 in each passage (Figure S2B)). The average doubling time was 28.34 hours (Figure S2B). This period was faster than that of hiPSCs cultivated on feeder cells2. Surprisingly, the high cell viability permitted a split ratio of nearly 1:100 (Figure S2B). Frozen stocks were prepared at −80°C using a standard slow-freezing method and were thawed in a 37°C water bath (Figure S2A).
We next examined whether the hES/iPSCs could be stably cultivated over long periods using the new culture conditions. We used H9 hESCs, KhES1 hESCs and 201B7 hiPSCs for this experiment. The cells were stably maintained for 20–30 passages and markers of pluripotency, such as Oct3/4 and TRA-1-60, were still detected (Figures 1B and 1C). Passage number 54 of the 201B7 cells still exhibited the ability to differentiate into all three germ layers in vitro (Figure S2C). We concluded that our method is sufficient and efficient for hESC and hiPSC culture. This culture system has already been tested and showed similar results at several other laboratories in Japan.
Establishment of hiPSCs under Ff and Xf conditions
Although we successfully developed a highly efficient system for the culture of hESCs and hiPSCs using rLN511E8 and StemFit™, hiPSCs intended for clinical application should be generated under similar conditions. First, we attempted to establish hiPSCs from human primary fibroblasts. Skin tissues were collected using biopsies and fibroblasts were generated from the skin tissues. Fibroblasts were established under Xf conditions with medium containing 10% autologous serum. The fibroblasts were electroporated with episomal vectors containing reprogramming factors. Twenty to thirty days after electroporation, hiPSC colonies were observed and selected to establish feeder-free hiPSC (Ff-hiPSC) clones, 987A3 and 987A7 (Figure 2A). The morphology of the fibroblast-derived Ff-hiPSCs was similar to that of 201B7 or H9 cells cultivated on rLN511E8. The loss of episomal vectors was confirmed using a genomic PCR analysis (Figure S3A).
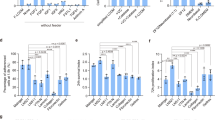
Establishment of iPS cell clones under the feeder-free (Ff) and xeno-free (Xf) culture system.
(A) Human iPSCs established from skin-derived primary fibroblasts (987A3 and 987A7), peripheral blood-derived T-cells (1027B3 and 1027B6), peripheral blood-derived non-T-cells (1120C7 and 1120C12) and cord blood cells (1156D2 and 1156D15) under the Ff and Xf culture system. Photographs were taken between passages p5 and p10. Scale bar, 100 μm. (B) The gene expression levels of pluripotency markers as determined using RT-PCR. “LN” indicates that the cells were cultivated using the Ff and Xf culture system reported herein. “Feeder” indicates cells that were cultivated on murine feeders with non-Xf medium. (C) The growth curve of the feeder-free iPS cells cultured under Ff and Xf culture conditions.
The expression levels of markers of pluripotency were examined using RT-PCR and immunostaining. The fibroblast-derived Ff-hiPSC clones exhibited similar expression levels to those of 201B7 and H9 cells (Figures 2B and S3B). The expression levels of genes related to pluripotency were similar in the cells cultivated on rLN511E8 and feeder cells (Figure 2B). The fibroblast-derived Ff-hiPSCs were stably passaged for long periods (Figure 2C) and had normal karyotypes (Figure S3C).
Ff-hiPSCs were also established from peripheral blood-derived T-cells, non-T-cells and cord blood. The morphology, marker gene expression levels and stability for long-term culture of these cells were similar to those of fibroblast-derived Ff-hiPSCs (Figures 2 and S3D).
The efficiency of Ff-hiPSC generation is summarized in Supplemental Table 1.
Differentiation capacity of Ff-hiPSCs
We examined whether Ff-hiPSCs have the ability to differentiate into several types of somatic cells. First, Ff-hiPSCs were cultivated on rLN511E8 with StemFit™ in the absence of bFGF. The cells efficiently attached to rLN511E8-coated plates and grew, exhibiting spontaneous differentiation. Two weeks after differentiation, we confirmed the expression levels of Sox17, α-smooth muscle actin (SMA) and βIII tubulin by immunostaining (Figure S4). Ff-hiPSCs were able to differentiate into all three germ layers in vitro. Moreover, in the teratoma assays, Ff-hiPSCs differentiated into various tissues of the three germ layers, including gut-like epithelial tissue, cartilage and neural tissue (Figure 3A). Another three Ff-hiPSC clones were also tested for the teratoma assays. These results demonstrated that the Ff-hiPSCs are functionally equivalent to iPSCs derived under feeder conditions2 and have the potential to spontaneously differentiate into all three germ layers both in vitro and in vivo.
The differentiation capacity of the T-cell-derived feeder-free hiPSC clone, 1027B6 (p7–p12).
(A) Hematoxylin and eosin staining of teratomas showing representative derivatives of all three germ layers. Scale bar, 100 μm. (B) Differentiation into mesencephalic dopaminergic neurons under Xf conditions. Photomicrographic images of immunostaining for Tuj1 (green), tyrosine hydroxylase (TH: red), Foxa2 or Nurr1 (blue) and DAPI (white). Scale bar, 100 μm. (C) May-Grünwald-Giemsa staining of differentiated blood cells on day 16 showing hematopoietic precursor cells, myeloid precursor cells, macrophages and erythroblasts. Scale bar, 100 μm. (D) Feeder-free hiPSCs differentiated into insulin-producing cells. After 23 days of culture under the differentiation conditions, the cells were fixed and stained with Hoechst33342 (blue) and anti-insulin antibodies (magenta). Scale bar, 100 μm.
The directed differentiation of hiPSCs has the potential to generate various somatic cells for disease modeling, drug discovery, toxicology, prediction of side effects and eventually, transplantation therapy. Therefore, we next examined whether Ff-hiPSCs could be specifically induced to differentiate towards somatic cells of therapeutic interest.
Parkinson’s disease is characterized by the loss of dopaminergic (DA) neurons; therefore, hiPSC-derived DA neurons may be good sources for cell transplant therapy. Mature and functional DA neurons have been produced through long-term culture of hiPSCs on Matrigel19. We subjected Ff-hiPSCs to the neuronal differentiation protocol with dual SMAD inhibition. Consequently, the Ff-hiPSCs differentiated into DA neurons expressing tyrosine hydroxylase (TH), βIII tubulin (Tuj1), Nurr1 and Foxa2 (Figure 3B). This experiment was performed under Xf conditions. Therefore, these results indicate that DA neurons can be successfully generated under Xf conditions from human tissue samples obtained using Ff-hiPSCs cultured without Matrigel.
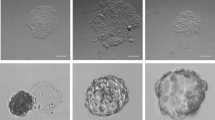
As a second target lineage with therapeutic potential, we chose to differentiate Ff-hiPSC into blood cells. Ff-hiPSCs were cultivated in a low-binding cell culture plate to promote the formation of EB-like spheres, which were sequentially treated with cytokines to invoke blood cell differentiation, as described previously20. Erythroblasts, macrophages and myeloid lineage cells were produced from Ff-hiPSCs, as demonstrated using May-Grunwald-Giemsa staining (Figure 3C).
The third target cells induced were insulin-producing cells. The differentiation of Ff-hiPSCs into insulin-producing cells has been reported previously21 and we made use of a similar protocol. Consequently, insulin-producing cells were generated from Ff-hiPSCs (Figure 3D). These results indicate that Ff-hiPSCs cultured under Xf conditions have the ability to differentiate into specific cells of interest using established in vitro induction protocols, with some minor modifications to maintain the Xf conditions. The efficiency and quality should be examined by the future experiments.
Discussion
We developed a novel efficient culture system for hES/iPSCs without feeder cells. Recombinant LN511E8 strongly supported hESC and hiPSC culture for long periods. StemFit™, a newly developed Xf-medium, was the best medium for hESC and hiPSC culture with rLN511E8. Under this novel culture system, hESCs and hiPSCs were passaged by dissociating them into single cells. Moreover, hESC and hiPSCs could be cryopreserved at −80°C by the slow-freezing method. Ff-hiPSCs showed the ability to differentiate into several somatic cell types, similar to conventional hiPSCs cultured on feeder cells2.
The culture system using rLN511E8 has been reported previously by Miyazaki et al.12. The authors also passaged hES/iPSCs by dissociating into single cells. Despite the basic idea is the same, we could develop more efficient method using StemFit™, achieving significantly better attachment efficiency at 6 hours after plating (Supplementary Table 2). Moreover, we confirmed the requirement of the glutamic acid residue in the C-terminal tail of the laminin γ1 chain, substitution of which with glutamine abolishes the integrin binding activity of LN511E824, in our culture method using an EQ mutant of rLN511E8. The colony formation of 201B7 was not observed on EQ mutant-coated plate (Figure S5).
Recently, the defined and simplified Xf medium, E8, was reported13. The major difference in formulation between E8 and StemFit™ is albumin. StemFit™ includes human albumin (E8 does not include any albumins). Albumin has many biological and physical roles. Thus, this factor may confer difference in performance between the both.
We are planning to build up a bank of hiPSCs for transplantation therapy. The human leukocyte antigen (HLA) is a key factor that mediates the immune-related rejection after transplantation. To minimize the immune system-related rejection, it is necessary to match the HLA type of the donor and recipient. Matching the HLA type is difficult because of the large number of HLA types present in each individual. However, the Japanese population is relatively homogeneous compared to other populations and it has been reported that 50–140 HLA-homozygous cell lines would match 90% of the Japanese population17,22, a HLA-homozygous hiPSC bank would therefore be a helpful resource for therapeutic application in Japan.
Ensuring the quality and safety of hiPSC are important for their clinical application. Manufacturing hiPSCs should be performed in the Cell Processing Center under the GMP guidelines. Our novel hiPSC culture system is comparable to that of standard cell lines, such as 293 cells or HeLa cells, making previously complex steps more routine. Employing this easy to use, reproducible and expandable culture system, a large amount of clinical-grade hiPSC stock can be made at early passage numbers at the same time. Moreover, the procedures needed to establish and maintain Ff-hiPSCs should be minimal and simple in the CPC. In order to apply Ff-hiPSCs for clinical applications, it is necessary to reduce or completely eliminate the use of animal-derived materials. To achieve this, we selected the StemFit™ medium, which does not contain animal-derived materials. This culture system is a promising method for manufacturing clinical-grade hiPSCs. In addition, it is necessary to use defined culture system for the source of iPS cells, such as fibroblasts or blood cells13,23.
Ff-hiPSCs were generated from skin-derived fibroblasts and blood-derived cells. Although the efficiency of Ff-hiPSC generation was lower than that of iPSCs established on feeder cells, we were able to obtain an adequate number of Ff-hiPSC clones. The cord blood cells appeared to be more potent sources for Ff-hiPSC generation. Almost all Ff-hiPSCs have differentiation potential and can differentiate into several kinds of somatic-type cells. These findings indicate that the Ff-hiPSCs established under Ff and Xf conditions from several types of somatic cells are similar to the hiPSCs established using the conventional system with feeders.
In conclusion, the present results indicate that hiPSCs with equivalent growth and differentiation potential to feeder-grown hiPSCs can be generated from various human tissue samples under Ff and Xf conditions. Since this method is easy to use, expandable and reproducible, it should prove valuable for generating clinical-grade hiPSCs intended for drug screening and therapeutics, or even basic research applications.
Methods
Production of laminin-511 E8
Recombinant LN511E8 were expressed in 293-F cells (Life Technologies) with 6 × His, HA and FLAG tags at the N-termini of the α5E8, β1E8 and γ1E8 chains, respectively and were purified using affinity chromatography with Ni-NTA resin and anti-FLAG antibody agarose, as described previously24. rLN511E8 was also produced in cGMP-banked Freedom CHO-S cells (Life Technologies) with an N-terminal 6 × His tag on the α5E8 chain (iMatrix-511™, Nippi, Inc.). The cGMP-compatible rLN511E8 was purified using Ni-NTA affinity chromatography and ion-exchange chromatography. cGMP-compatible rLN511E8 was manufactured in a controlled clean room (Nippi and Yokohama Biopharmaceutical Research and Development Center).
Components of StemFit™
In this study, we used newly developed medium manufactured by Ajinomoto Co., Inc., Japan. StemFit™ contains twenty-one amino acids (L-Alanine, L-Arginine, L-Asparagine, L-Aspartic Acid, L-Cysteine, L-Cystine, L-Glutamic Acid, L-Glutamine, Glycine, L-Histidine, L-lsoleucine, L-Leucine, L-Lysine, L-Methionine, L-Phenylalanine, L-Proline, L-Serine, L-Threonine, L-Tryptophan, L-Tyrosine and L-Valine), ten vitamins (L-Ascorbic Acid, Cobalamin, Biotin, Folic Acid, I-inositol, Niacinamide, D-Calcium Pantothenate, Pyridoxine Hydrochloride, Riboflavin and Thiamine Hydrochloride), five trace minerals (Cupric Sulfate, Ferric Sulfate, Ferric Nitrate, Zinc Sulfate and Sodium selenite) and growth factors, including bFGF. All ingredients are highly defined or purified materials and none are derived from non-human animal sources. Large-scale manufacturing of StemFit™ has been achieved in well-validated and -qualified factories in Japan.
Feeder-free culture system for hESCs and hiPSCs
The hESCs/iPSCs were cultivated on rLN511E8-coated (0.5 μg/cm2) cell culture plates with StemFit™ and passaged via dissociation into single cells using TrypLE Select (Life Technologies). Although this approach is similar to a previously published method12, we separately developed a single cell culture system for hESCs (H93 and KhES125) and hiPSCs (201B72 and 32R118). Before plating the cells, cell culture dishes or plates were coated with rLN511E8 (0.5 μg/cm2). The rLN511E8 solution was diluted with PBS(−) and the diluents were transferred to cell culture dishes or plates. The dishes or plates were incubated for one hour in a CO2 incubator (37°C). The hESCs/iPSCs were dissociated into single cells by treatment with 0.5× TrypLE Select (1× TrypLE Select diluted 1:1 with 0.5 mM EDTA/PBS(−)) for four minutes at 37°C. Since the cells remained attached to the rLN511E8-coated plates, we aspirated the 0.5× TrypLE Select and carefully washed the cells with PBS(−). The cells were scraped under the StemFit™ and dissociated into single cells by pipetting the cells 10 times. The number of dissociated single cells was counted using a Countess Automated Cell Counter (Life Technologies). We typically plated 1.3 × 104 live cells into the rLN511E8-coated wells of a 6-well cell culture plate ( = 1.35 × 103 live cells/cm2). A Rock inhibitor (Y-27632, final concentration 10 μM) was used only at the time of plating26. The next day, the medium was changed to fresh StemFit™ without Y-27632. The medium was changed every other day. Eight to ten days after plating, the cells reached 80–90% confluency and were ready for the next passage. Frozen stocks of Ff-hiPSC were similarly prepared as single cells and frozen in STEM-CELLBANKER (1 × 106 live cells/mL) (Nippon Zenyaku Kogyo) at −80°C using a standard slow-freezing method.
Establishment of Ff-hiPSCs from fibroblasts and blood cells
Human primary fibroblasts were derived from biopsied skin tissue samples. The fibroblasts were established and expanded with DMEM containing 10% autologous human serum. Using these fibroblasts, iPS cells were generated as described previously17. Briefly, following electroporation of reprogramming factors with episomal vectors using the Neon system (Life Technologies), the cells were plated on a non-coated cell culture plate. iPS cells were induced by changing the medium to StemFit™. Twenty to thirty days after plating, iPS cell colonies were observed.
Blood cell-derived iPS cells were generated as described previously16. Briefly, mononuclear cells were prepared from peripheral blood using the Ficoll-Paque PREMIUM (GE Healthcare) separation method. The cells were electroporated with episomal vectors using a Nucleofector 4D system (with P3 Primary Cell Kit, Lonza) and plated on rLN511E8-coated cell culture plates. The iPSCs were induced by changing the medium to StemFit™. Twenty to thirty days after plating, iPS cell colonies were observed. A similar method was used to generate Ff-hiPSCs from human cord blood (provided by the RIKEN Bioresource Center Cell Bank). We generated several clones of Ff-hiPSCs from each experiment.
The experimental protocols dealing with human subjects were approved by the institutional review board at our institute (Kyoto University Graduate School and Faculty of Medicine, Ethics Committee). Written informed consent was provided by each donor.
Genomic PCR analysis of the integration of the episomal vectors
Purified genomic DNA was used as a template to detect the remaining episomal vectors in the hiPSCs using a PCR analysis. The episomal vectors were detected using the primer set for EP4. The EP4 primer set was constructed in the EBNA1 region. The Fbx15 gene was amplified with a primer set for Fbx as an internal control. The episomal vector, pCXLE-EGFP, was used as a positive control (“Plasmid”). The primer sets are listed in Supplementary Table 3.
Antibodies used for immunostaining
The antibodies used in this study included anti-Oct3/4 (BD, 611202), -Nanog (R&D, AF1997), -SSEA4 (BD, 560073), -TRA-1-60 (BD, 560071), -Sox17 (R&D, AF1924), -α-smooth muscle actin (SMA) (Dako, M0851), -α-1-fetoprotein (AFP) (Dako, N1501) and -βIII tubulin (Millipore, MAB1637) antibodies.
RNA isolation and reverse transcription
Total RNA purification and RT-PCR were performed as described previously2,27,28,29. The expressions of Nanog, Sox2, Oct3/4, Lin28, L-Myc, c-Myc, Klf4 and G3PDH were detected using the primer sets listed in Supplementary Table 3.
In vitro differentiation
To examine whether Ff-hiPSCs spontaneously differentiate into the three germ layers, the Ff-hiPSCs were cultivated with StemFit™ lacking bFGF (StemFit™-bFGF) for two weeks on rLN511E8-coated cell culture plates. The differentiated cells were immunostained with the indicated antibodies.
Directed differentiation into dopaminergic neurons
Dopaminergic neurons were induced from Ff-hiPSCs under xeno-free conditions. The dissociated Ff-hiPSCs were plated on rLN511E8-coated plates at high density (7.5 × 105 cells/cm2; Day 0). Glasgow minimum essential medium (GMEM, Life Technologies) was supplemented with 8% Xeno-free knockout serum replacement (Life Technologies), sodium pyruvate (SIGMA), 2-mercapto ethanol (Wako), MEM NEAA (Life Technologies), 500 nM of A-83-01 (Activin inhibitor, Wako), 100 nM of LDN193189 (BMP inhibitor, STEMGENT) and 10 μM Y-27632 (Wako). To induce neural cells with a ventral mesencephalic phenotype, 2 μM Purmorphamine (Wako) and 100 ng/mL of FGF-8 (Wako) were added starting the next day. Starting on day 3, 3 μM CHIR99021 (GSK3β inhibitor, Wako) was also added. On day 12, the first passage was performed with TrypLE CTS (Life Technologies) and the neural cells were replated on a fresh rLN511E8 surface at the same density as on day 0. The basal medium was switched to Neurobasal medium supplemented with Xeno-free B27 (Life Technologies), 200 μM of ascorbic acid (SIGMA), 400 μM of dbc AMP (SIGMA), 2 ng/mL of GDNF (R&D), 20 ng/mL of BDNF (R&D) and 10 μM Y-27632. Throughout the neural induction process, the medium was changed every day. On days 12, 20 and 28, the culture was passaged onto a new rLN511E8 surface. Y-27632 was added to the medium on the day of passage. Immunostaining was performed using anti-TH (Chemicon, AB152), -Foxa2 (Santa Cruz, SC6554) and –Nurr1 (kindly provided by KAN Research Institute) antibodies.
Directed differentiation into blood cells
Ff-hiPSCs were differentiated into blood cells as described previously30 with some key modifications. Briefly, Ff-hiPSCs were cultivated in Ultra Low Attachment 6-well cell culture plates (Corning) with StemFit™ to prepare EB-like spheres. The spheres were subsequently used for blood cell differentiation according to the established protocol20.
Directed differentiation into insulin-producing cells
Ff-hiPSCs were differentiated into insulin-producing cells as described previously21. Briefly, human Ff-hiPSCs were dissociated and plated on rLN511E8 and cultured with the reported factors required for pancreatic differentiation. The dissociated cells were treated with 100 ng/ml of activin A and 3 μM CHIR99021 for 24 hours and then were treated with 100 ng/ml of activin A and 1 μM CHIR99021 for four days. Next, the cells were cultured with 1 μM dorsomorphin (Calbiochem, San Diego, CA), 2 μM retinoic acid (Sigma, St. Louis, MO) and 10 μM SB431542 (Sigma) for six days. Subsequently, the cells were cultured with 10 μM forskolin, (Wako), 10 μM dexamethasone (Wako), 5 μM Alk5 inhibitor II (Calbiochem) and 10 mM nicotinamide (StemCell Technologies, Vancouver, BC) for 12 days. Following the culture, the cells were fixed and stained with Hoechst33342 (Life Technologies) and antibodies against insulin (Dako).
References
Schwartz, S. D. et al. Embryonic stem cell trials for macular degeneration: a preliminary report. Lancet 379, 713–720 (2012).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Takahashi, K., Narita, M., Yokura, M., Ichisaka, T. & Yamanaka, S. Human induced pluripotent stem cells on autologous feeders. PLoS One 4, e8067 (2009).
Rodriguez-Piza, I. et al. Reprogramming of human fibroblasts to induced pluripotent stem cells under xeno-free conditions. Stem Cells 28, 36–44 (2010).
Xu, C. et al. Feeder-free growth of undifferentiated human embryonic stem cells. Nat Biotechnol 19, 971–974 (2001).
Stover, A. E. & Schwartz, P. H. Adaptation of human pluripotent stem cells to feeder-free conditions in chemically defined medium with enzymatic single-cell passaging. Methods Mol Biol 767, 137–146 (2011).
Totonchi, M. et al. Feeder- and serum-free establishment and expansion of human induced pluripotent stem cells. Int J Dev Biol 54, 877–886 (2010).
Ausubel, L. J., Lopez, P. M. & Couture, L. A. GMP scale-up and banking of pluripotent stem cells for cellular therapy applications. Methods Mol Biol 767, 147–159 (2011).
Bergstrom, R., Strom, S., Holm, F., Feki, A. & Hovatta, O. Xeno-free culture of human pluripotent stem cells. Methods Mol Biol 767, 125–136 (2011).
Rodin, S. et al. Long-term self-renewal of human pluripotent stem cells on human recombinant laminin-511. Nat Biotechnol 28, 611–615 (2010).
Miyazaki, T. et al. Laminin E8 fragments support efficient adhesion and expansion of dissociated human pluripotent stem cells. Nat Commun 3, 1236 (2012).
Chen, G. et al. Chemically defined conditions for human iPSC derivation and culture. Nat Methods 8, 424–429 (2011).
Lu, H. F. et al. A 3D microfibrous scaffold for long-term human pluripotent stem cell self-renewal under chemically defined conditions. Biomaterials 33, 2419–2430 (2012).
Mei, Y. et al. Combinatorial development of biomaterials for clonal growth of human pluripotent stem cells. Nat Mater 9, 768–778 (2010).
Okita, K. et al. An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells 31, 458–466 (2013).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat Methods 8, 409–412 (2011).
Nakagawa, M., Takizawa, N., Narita, M., Ichisaka, T. & Yamanaka, S. Promotion of direct reprogramming by transformation-deficient Myc. Proc Natl Acad Sci U S A 107, 14152–14157 (2010).
Doi, D. et al. Prolonged maturation culture favors a reduction in the tumorigenicity and the dopaminergic function of human ESC-derived neural cells in a primate model of Parkinson's disease. Stem Cells 30, 935–945 (2012).
Grigoriadis, A. E. et al. Directed differentiation of hematopoietic precursors and functional osteoclasts from human ES and iPS cells. Blood 115, 2769–2776 (2010).
Kunisada, Y., Tsubooka-Yamazoe, N., Shoji, M. & Hosoya, M. Small molecules induce efficient differentiation into insulin-producing cells from human induced pluripotent stem cells. Stem Cell Res 8, 274–284 (2012).
Nakatsuji, N., Nakajima, F. & Tokunaga, K. HLA-haplotype banking and iPS cells. Nat Biotechnol 26, 739–740 (2008).
Saha, K. et al. Surface-engineered substrates for improved human pluripotent stem cell culture under fully defined conditions. Proc Natl Acad Sci U S A 108, 18714–18719 (2011).
Ido, H. et al. The requirement of the glutamic acid residue at the third position from the carboxyl termini of the laminin gamma chains in integrin binding by laminins. J Biol Chem 282, 11144–11154 (2007).
Suemori, H. et al. Efficient establishment of human embryonic stem cell lines and long-term maintenance with stable karyotype by enzymatic bulk passage. Biochem Biophys Res Commun 345, 926–932 (2006).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat Biotechnol 25, 681–686 (2007).
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germ-line competent induced pluripotent stem cells. Nature 448, 313–317 (2007).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Nakagawa, M. et al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat Biotechnol 26, 101–106 (2008).
Grigoriadis, A. E. et al. Directed differentiation of hematopoietic precursors and functional osteoclasts from human ES and iPS cells. Blood 115, 2769–2776 (2010).
Kajiwara, M. et al. Donor-dependent variations in hepatic differentiation from human-induced pluripotent stem cells. Proc Natl Acad Sci U S A 109, 12538–12543 (2012).
Acknowledgements
We would like to thank Junko Toga and Emiko Yagi for producing the research-grade rLN511E8 (Osaka Univ.), Drs. Satoru Okamoto and Tsuyoshi Kobayoshi for developing and producing the StemFit™ (Ajinomoto), Drs. Shunji Hattori, Takuji Yamamoto and Kazumasa Fujita for producing rLN511E8 (Nippi), Yokohama BioResearch & Supply (YBRS) for manufacturing cGMP-compatible rLN511E8, Drs. Tokiko Ohkame and Yusuke Ohkame for performing the karyotype analysis (CiRA), Drs. Takashi Aoi (Kobe University) and Takafumi Kimura (CiRA) for providing valuable advice on this research and Dr. Knut Woltjen for valuable help in preparing the manuscript (CiRA). The hESC clones H9 and KhES1 were kindly provided from the WiCell Research Institute (Dr. James A. Thomson) and the Institute for Frontier Medical Sciences (Kyoto University), respectively. This research was supported in part by a grant from the Leading Project of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), a grant from the Funding Program for World-Leading Innovative Research and Development on Science and Technology (FIRST Program) of Japan Society for the Promotion of Science, Grants-in-Aid for Scientific Research of the Japan Society for the Promotion of Science and MEXT (to S.Y.), New Energy and Industrial Technology Development Organization (NEDO) of Japan (to K.S.) and the Coordination, Support and Training Program for Translational Research of the Ministry of Education Culture, Sports, Science and Technology of Japan (to K.S.).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: M. Nakagawa, S.S. and S.Y. Performed the experiments: M. Nakagawa, S.S., Y.T., N.T., T.I., K.A., A.M., J.T., T.T., K.O., M. Nishizawa, Y.Y. and K.S. Analyzed the data: M. Nakagawa, S.S. and Y.T. Contributed reagents/materials/analysis tools: M. Nakagawa, S.S., Y.T. and K.S. Wrote the paper: M. Nakagawa.
Ethics declarations
Competing interests
The authors declare no competing financial interests. S.Y. is a member without salary of the scientific advisory boards of iPierian, iPS Academia Japan, Megakaryon Corporation and HEALIOS K.K. Japan.
Electronic supplementary material
Supplementary Information
Fano resonance in anodic aluminum oxide based photonic crystals
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Nakagawa, M., Taniguchi, Y., Senda, S. et al. A novel efficient feeder-free culture system for the derivation of human induced pluripotent stem cells. Sci Rep 4, 3594 (2014). https://doi.org/10.1038/srep03594
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03594
This article is cited by
-
Pre-clinical evaluation of the efficacy and safety of human induced pluripotent stem cell-derived cardiomyocyte patch
Stem Cell Research & Therapy (2024)
-
Mutant α-synuclein causes death of human cortical neurons via ERK1/2 and JNK activation
Molecular Brain (2024)
-
A human induced pluripotent stem cell model from a patient with hereditary cerebral small vessel disease carrying a heterozygous R302Q mutation in HTRA1
Inflammation and Regeneration (2023)
-
An in vitro culture platform for studying the effect of collective cell migration on spatial self-organization within induced pluripotent stem cell colonies
Journal of Biological Engineering (2023)
-
CDK8/19 inhibition plays an important role in pancreatic β-cell induction from human iPSCs
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.