Abstract
Energy-coupling factor (ECF) transporters are responsible for uptake of micronutrients in prokaryotes. The recently reported crystal structure of an ECF transporter RibU provided a foundation for understanding the structure and transport mechanism of ECF transporters. In the present study, molecular dynamics (MD) was carried out to study the conformational changes of the S component RibU upon binding by riboflavin. Our result and analysis revealed a critically important gating mechanism, in which part of loop5 (L5′) (eleven residues, missing in the crystal structure) between TM5 and TM6 is dynamically flexible and serves as a gate. Specifically, the L5′ opens a large cavity accessible to riboflavin from the extracellular space in Apo-RibU and closes the cavity upon riboflavin binding through hydrophobic packing with riboflavin. Thus, L5′is proposed to be the gate for riboflavin binding. In addition, steered molecular dynamics (SMD) simulation is employed to investigate the translocation dynamics of RibU during riboflavin transport. The simulation result does not show evidence that the S component alone can carry out the transport function. Since loop regions are very flexible and therefore could not be resolved by crystallography, their dynamics are hard to predict based on crystal structure alone.
Similar content being viewed by others
Introduction
Membrane proteins play a vital role in the interactions of cells with their surrounding environment and they account for about 25% of proteins in eukaryotic genomes1. Membrane proteins carry out various functions2, such as providing the structural framework that shapes cellular compartments, signaling or transporting molecules, etc.
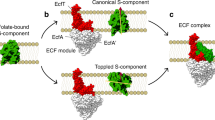
Energy-coupling factor (ECF) transporters are a recently discovered new class of membrane proteins3. They are importers4 of micro-nutrients, such as water-soluble vitamins, metal ions, including Ni2+ and Co+, the amino-acid tryptophan and queuosine, etc. ECF-transporters are composed of four segments5 (Figure 1a), namely, two trans-membrane proteins named S component for substrate binding and T component6 whose function is still unclear and pairs of ATP-binding cassette-containing proteins (A proteins). This arrangement of structure is similar to the better known ATP-binding cassette (ABC7,8) transporters (Figure 1b) which couple two ATP molecules to catalyze the delivery of a great number of molecules across lipid bilayers. The most significant difference between these two types of transporters is that ECF-transporters have no extra-cytoplasmic substrate-binding proteins (BP proteins), which are used to capture various micronutrients in ABC transporters. Similarly, ECF-transporters also utilize the energy released by ATP hydrolysis to transport these micronutrients.
ECF-transporters could be divided into two groups3. In Group I, S components and AT modules are encoded by linked genes, i.e., AT modules are S specific and different S components have different AT modules. The Group II transporters share the same AT modules for various S components. In addition, for ECF-transporters, different S components transport different substrates. For example, if S component is ThiT9, Thiamine (vitamin B1) will be transported and similarly, NiaX10 for Niacin transport (vitamin B3), PanT3 for pantothenate (vitamin B5) and RibU3 pumping riboflavin (vitamin B2) into the cell.
Structural detail of ECF-transporter in atomic level was missing until 2010, when the first crystal structure of an S component (named RibU) in complex with riboflavin (VB2) was reported by Zhang et al11. Recently, two more crystal structures of S components have been reported, ThiT12 with thiamin binding and BioY13 with biotin binding. Although these S components show little sequence similarity, the mentioned three crystal structures of S components have revealed similar fold. That is, all of them have six trans-membranes and two halves, namely, N termini and C termini. RibU, which is the focus of this work, (Figure 2a) is composed of six helices (TM1 to TM6), different from the one previously reported14,15, where TM2 and TM3 are predicted to be one single trans-membrane module. In the complex structure11 the ribityl side chain and polar part of isoallloxazine ring of riboflavin (Figure 2b) is reported to interact with RibU through hydrogen bonds, while non-polar part of riboflavin ring is surrounded by a hydrophobic cage, involving 13 residues of RibU11. In this work11, L1 is proposed to be a gate for riboflavin binding based on static structural analysis of RibU.
(a) The XRD structure of RibU (PDB code: 3P5N11) with missing electron density of I142-L152 (L5′). (b) The structure of riboflavin, composed of a ribityl side chain and an aromatic isoallloxazine ring.
Since the complex lipid bilayer environment makes it difficult to extract the details of protein-ligand interaction dynamics through analysis of crystal structure alone16, it is desirable to perform simulation study to help understand and rationalize the detailed function model of ECF transporters. Molecular Dynamics (MD17,18) simulations can provide much insight and physical mechanism about the structure and dynamics of protein in atomic level. Currently, MD simulation has been widely employed to study membrane proteins19,20,21,22, including interface-associated and trans-membrane peptides, fusion proteins, channel and pore proteins, transporters, ion pumps, ATP-syntheses and G-protein-coupled receptors, etc. Additionally, a recent work23 has performed MD simulation for one of the three S components ThiT12 (PDB ID 3RLB) and has found that substrate-induced conformational changes is important for ThiT binding Thiamin.
It is exciting that two structures24,25 of the quaternary ECF-transporters have been recently determined, namely, EcfS-EcfT-EcfA-EcfA', are all been crystalized. The availability of these two entire or overall crystal structures of ECF-transporter provided the foundation to understand detailed function and working model of ECF-transporter. In this work, we focus on the study of the dynamics of loop5 of S component RibU.
Since electron density of L5′ is missing in the crystal structure, it is proposed that L5′ is quite flexible and may play an important role in the substrate binding and be the focus of our work. MD simulation is employed to study the dynamical structures and functional model of RibU and to address the questions: (1) is there a gating mechanism that controls the binding of riboflavin? (2) If yes, which loop(s) serve as the gate? In addition, we also investigate the translocation mechanism responsible for the transport of riboflavin. In order to find possible answers to the above questions, standard molecule dynamics simulations were performed for both Apo-RibU and Rbf-bound-RibU. Further, molecular mechanics/Poisson Boltzmann (MM-PB) analysis is carried out to study the binding interaction between RibU and two ligands (riboflavin and roseoflaivn). Finally, steered molecule dynamics (SMD26,27) is carried out to investigate the dynamical behaviors of RibU during riboflavin transport.
Results and Discussion
Large conformational change of L5′is observed
A relatively long loop between TM5 and TM6 (the L5 loop) contains 23 amino acids in total. Seven (residues 134–140) of L5 form a short helix (α5′). Eleven amino acids (residues 142–152) of L5, denoted as L5′, are missing in the crystal structure due to randomness of their positions11,. In this study, the missing 3D-structure of L5′is added by the software loopy28,29. Two separate simulations are performed, RibU without VB2 (Apo-RibU) and RibU with binding of VB2 (Rbf-RibU).
The final structures of RibU from two separate simulations (Rbf-RibU and Apo-RibU) are aligned together in Figure 3. The Root Mean Square Deviation (RMSD) between these two conformations is quite large (6.154 Å). The figure shows that most of these two structures are identical and the largest difference comes from the conformation changes of L5′. In Apo-RibU, L5′ (blue color in Figure 3) is almost straight and leaves a large binding pocket accessible for VB2 from the extracellular space, (denoted as open-state of RibU). In Rbf-RibU, L5′ (green color in Figure 3) covers the hydrophobic ring of riboflavin and closes the binding pocket (denoted as closed-state of RibU). The flexibility of L5′ should explain the reason why the electron density of L5′ is missing in the crystal structure of RibU. In contrast, the short helix (α5′) of loop5 sees little changes in open (red color in Figure 3) and close (yellow color in Figure 3) states. Since the short helix is very stable, it is not the focus of this work.
To see the structural stability and dynamics of L5′ during the whole simulation time, the RMSD of L5′ and the entire RibU from Apo-RibU/Rbf-RibU simulations are calculated and plotted in Figure 4. In Apo-RibU (Figure 4a), RMSD of the entire RibU increases to 5 Å and stabilized in the following 60 ns simulation (Figure S6), a similar trend is seen for L5′, indicating large conformational changes for L5′. In contrast, in Rbf-RibU (Figure 4b), the entire RibU (black color) and L5′ (green color) both undergoes relatively smaller conformation changes than that in Apo-RibU, with the largest RMSD of 3.0 Å/2.5 Å. This indicates that riboflavin could stabilize the structure of RibU to some extent. RMSF analysis also shows that L5′ is quite flexible in both Apo-RibU and Rbf-RibU simulations (Figure S7). It is interesting to note that in both Apo-RibU and Rbf-RibU, the major structural fluctuation of RibU comes from the conformational change of L5′. Thus, L5′ is very flexible and may play an important role in the binding process of riboflavin.
L5′ serves as a gate for riboflavin binding
As explained above, the conformation changes of L5′ is smaller in Rbf-RibU than that in Apo-RibU. Figure 5 shows the distance between L5′ and riboflavin (mass centers) as a function of simulation time in Rbf-RibU. The initial distance is 14 Å, followed by a sharp decrease and then leveled off at 10 Å. Clearly, L5′ is closer to riboflavin in Rbf-RibU. Principle Component Analysis (PCA in Figure S8) also shows that L5′ is quite flexible in both Rbf-RibU and Apo-RibU simulations. To help understand the reason why L5′ moves closer to riboflavin, the number of water within 9 Å of riboflavin is plotted as a function of simulation time (Figure 6). The number of water went up and reached 125 in the first of 40 ns and then stabilized around 65 during the following 130 nanoseconds. It is interesting to note that as L5′ moves closer to riboflavin, the number of water begins to decrease. It is thus mostly likely that the closure of L5′ in Rbf-RibU pumps water out of the pocket.
Riboflavin contains a ribityl side chain and an aromatic isoallloxazine ring. Initially, water rushes into the binding pocket to interact with the polar ribityl side chain. But the exposure of the aromatic ring to water is not energy-favorable. There are 7 non-polar residues out of 11 residues in L5′. Energetically, these hydrophobic residues prefer to pack themselves towards the aromatic ring to drive water out of the binding pocket and maintain the hydrophobic environment surrounding riboflavin.
In the current study, the final simulated structure in Apo-RibU is denoted as open-state RibU and the one from Rbf-RibU as closed-state of RibU (Figure 3). Here, an induced-fit model30 is used to describe the RibU-riboflavin binding. The above analysis leads us to propose a gating mechanism as follows (shown in Figure 7). 1) In Apo-RibU, L5′ is straight and opens a large cavity accessible for riboflavin from extracellular space. 2) As riboflavin moves closer to the binding pocket, the RibU gradually changes from the open-state toward the closed-state, during which L5′ covers the hydrophobic ring of riboflavin and prevents RibU from direct contact with the periplasm. In this proposed mechanism, L5′ serves as a gate to control the binding process of riboflavin.
Binding of riboflavin to RibU is hydrophobic
Through the above analysis, it is proposed that hydrophobic effect plays a critical role in binding process of riboflavin. Thus, we performed MM-PB analysis to investigate the detailed interaction energy key for ligand binding. The snapshots extracted from MD simulation are used for MM-PB analysis of binding free energy of two different ligands of RibU, riboflavin and roseoflavin31 (structure shown in supporting information Figure S1). Table 1 shows the computed binding free energy and its energy components.
For riboflavin, electrostatic interaction ΔEes (−17.5 kcal/mol) plus electrostatic salvation energy ΔGPB (39.5 kcal/mol) contribute to binding free energy by about 22.0 kcal/mol. This is quite positive and unfavorable. But, in contrast, van der Waals interactions is −47.8 kcal/mol, which is energetic favorable and makes a substantial contribution to the binding free energy (−25.8 kcal/mol) This shows that rich hydrophobic contacts exist in riboflavin-RibU and we conclude that hydrophobic effect play a critical role in RibU-riboflavin binding.
Also for roseoflavin, the situation is quite similar. Electrostatic interaction negatively contributes to binding free energy by about 26.2 kcal/mol, while Van der Waals interactions contribute −59.3 kcal/mol to the total free energy. Again, hydrophobic effect is important in the process of roseoflavin-RibU binding.
The snapshots used are around the closed-state of RibU with L5′ covering the hydrophobic ring, the hydrophobic contribution of binding free energy mainly comes from the non-polar residues of L5′. Again, the results of MM-PB analysis show that hydrophobic interaction between L5′ and riboflavin is dominant in ligand-RibU binding.
L1 stabilizes the ribityl side chain of riboflavin
The previous work11 proposed that the loop between TM1 and TM2 (L1) serves as a gate upon riboflavin binding. In this work, we specifically study the dynamics of L1 during riboflavin binding. In Rbf-RibU (Figure 4b), the conformation of L1 keeps steady during the entire simulation time of 170 ns (RMSD around 1 Å) and the conformation change is smaller than that in Apo-RibU. The same situation is also seen in the analysis of RMSF (Figure S7). Upon riboflavin binding, L1 sees little conformation changes. In RibU, four hydrogen bonds are present (shown in supporting information Figure S2). Figure 8 shows that three of them are broken and only one remains intact during the simulation. Thus, it is proposed that L1 helps stabilize the ribityl side chain of riboflavin through hydrogen bonds.
Exploring the translocation dynamics of RibU by SMD
It is expected to take a long time for RibU transporting riboflavin across the lipid bilayer and this is beyond the capabilities of present MD simulations. Here, steered molecular dynamics (SMD) simulation is employed to inspect the dynamic behaviors of RibU during transport. The previous work11 proposed a putative path for riboflavin transport. That is, helix I–III moves away from helix IV–VI to form a channel, allowing riboflavin transport from the periplasm to the cytoplasm.
Following this hint, helix I–III is defined as one group and helix IV–VI as another group. Forces are applied on the mass centers of these two groups in the opposite directions along x-axis (Figure S3) for 10 ns simulation time. The initial structure (pink color) and the final SMD simulated structure (lime color) of Rbf-RibU are aligned together and shown in Figure 9. The figure shows that these two defined groups are surely apart from each other and forms a channel under the applied harmonic forces, but it is interesting to find that lipid molecules (green color in Figure 10) go into the channel immediately, block the transport path and prevent riboflavin through the pulled channel. Clearly, it is impossible for riboflavin to reach the cytoplasm by crossing the barrier formed by lipid molecules. Thereafter, riboflavin is not expected to pass through the lipid bilayer in the way proposed by Zhang et al11. A video called smd.mpg (in the supporting information) provides more details about lipid molecules preventing the transport of riboflavin.
Several efforts have been made to attempt to make an isolated cylindrical channel for riboflavin through RibU, such as along the direction that forming 45° with both X-axis and Y-axis, or rotating RibU around Z-axis, but these efforts all failed. The best structure obtained from these simulations is shown in Figure S4. Again and again, lipid molecules entered into the pulled channel and prevented migration of riboflavin to the cytoplasm.
It may conclude that RibU alone is not able to transport riboflavin across the membrane without the help of other ECF-modules. Our observations are consistent with the results from two recently reported XRD structures of S components12,13, which stated that translocation path through S component is impossible. Further, a recent paper (Wang et al., 2013) published the crystal quandary structure of ECF-transporter proposed that RibU is for ligand binding rather than for ligand translation and the translocation path is not formed by RibU, but happens between S component and T component. Based on our analysis, it is proposed that RibU is too small to form a channel for transporting riboflavin and the transport channel should be in the middle of S component and T component, just like the channel ABC32,33 transporters used.
Conclusion
The present work employs MD simulation to study the dynamics of an S component (RibU) of ECF-transporter. Two different simulations of RibU (with/without riboflavin binding) are performed for 170 ns simulation length. Further, SMD is also performed to study the translocation dynamics of RibU for 10 ns simulation time. The followings are major findings from the present work.
-
L5′is a gate controlling riboflavin binding, while L1 stabilizes the ribityl side chain of riboflavin.
-
A gating mechanism is proposed for riboflavin binding controlled by hydrophobic packing between L5′ and riboflavin. MM-PB analysis also provides evidence that hydrophobic effect is dominant in this binding process.
-
Results of SMD study show that RibU alone is not able to transport riboflavin through lipid bilayers without the help of other ECF-modules.
Methods
System setup
The initial structure of RibU for MD simulation comes from its crystal structure in protein data bank (PDB code 3P5N11). As indicated in the published paper, electron density of I142-L152 in loop5 (L5′) is missing and rebuilt in our work by the software loopy28,29. Then two different simulations are performed, namely, riboflavin-binding system (denoted as Rbf-RibU), riboflavin-free system (named Apo-RibU). In Apo-RibU, the starting structure is the crystal structure with riboflavin directly removed from the binding pocket. POPE bilayer is used to wrap up the protein with VMD34. Lipid molecules within 0.6 Å of protein are deleted to avoid lipid overlapped with protein atoms. Then system is solvated by TIP3P water on both sides of the POPE bilayer. Similarly, overlapped water molecules are also deleted to make sure a reasonable simulation system. The final system contains 242 POPE molecules and 14021 water molecules with starting dimension 103 Å × 101 Å × 98 Å and about 76000 atoms in total.
MD simulation
All MD simulations are performed using NAMD35 with CHARMM2736 force field for lipid, protein and TIP3P water molecules. The particle mesh Ewald (PME37) is used to treat the long-range electrostatic interactions with the density of grid points at least 1/Å in all cases. A cutoff of 12 Å is applied to treat van der Waals interactions. Periodic boundary condition (PBC) is imposed on all directions and NPT ensemble is adopted. Langevin dynamics38 is performed to maintain constant temperature with damping coefficient (gamma) of 5/ps and the Nose-Hoover Langevin39 piston method is used to maintain constant pressure with a decay period of 200 fs and an oscillation time of 50 fs. Time-step is 2 fs with SHAKE40 used to constrain all the hydrogen atoms. First, the head group of lipid, water and protein atoms are all fixed to melt the tail of lipid for 1000 energy-minimized steps and 500 ps equilibration at 300 K. Secondly, the head group of lipid, water are released but protein constrained at 2 kcal/mol/Å2 to allow water and lipid pack close to the protein without disturbing the original structure of protein. This procedure is also firstly performed with 1000 energy-minimized steps and then 10 ns equilibration at 300 K. Thirdly, protein are released to allow equilibration of the whole system for 50 ns at 310 K. Finally, production run is performed for 120 ns for Rbf-RibU at 310 K with constant lipid area and the simulation of Apo-RibU is performed for 180 nanoseconds. RMSF analysis is performed with Gromacs41 employing the g_rmsf module. Principle Component Analysis (PCA) is performed through the ProDy42 Interface of the Normal Mode Wizards module in VMD.
MM-PB analysis
In the Molecular Mechanics Poisson-Boltzmann (MM-PB43) analysis, enthalpy of protein-ligand binding could be decomposed into and evaluated as,

Where G is decomposed into contribution from electrostatic (Ees), van der Waals (EvdW) and polar solvation (GPB) term, the binding free energy of a non-covalent association, ΔGbind, can be expressed as,

or

In this study, MD simulations are performed for riboflavin-RibU and roseoflavin-RibU complex, separately. The force parameters used for these two ligands are obtained through the software SwissParam44. One hundred snapshots from the 10 ns equilibrium simulation of these two trajectories at 100 ps intervals are extracted for MM-PB analysis respectively. The PB equation is solved using Delphi45 program with CHARMM27 atomic radii and CHARMM27 charges. Interior dielectric constant is set to 1.0, including protein part and lipid portion. And exterior dielectric constant is defined to 80. A figure about this define is shown in supporting information Figure S5. A perfil of 80 is used. Ees and EvdW are analyzed using the software NAMD2.7.
SMD simulation
By applying an external force, SMD permits to explore long time scale motions and large conformation changes of proteins. SMD has been successfully used to investigate the mechanism of membrane channels and transporter pathways46,47,48. To detect the dynamic behaviors of RibU during transport of riboflavin, SMD is employed to the final equilibrated structure of Rbf-RibU. Based on the putative conduction path proposed by the work of Zhang et al11, helix I–III of RibU are defined as a group and helix IV–VI as another group. The harmonic forces are applied to the mass center of mass of the two groups in the opposite directions along X-axis, separately (shown in the supporting information Figure S3) with constant velocity of 0.01 Å/ps. And the force constant is 0.5 kcal/mol/Å2. The total simulation time is 10 ns before the forces removed from RibU.
In addition, several efforts have been tested (shown in supporting information Table S1). First, the direction of the force, applied to the center of mass of these two groups, has been changed to form an angle with X-axis and be vertical with Z-axis. The angle has varied to 5°, 15°, 45°, respectively. The force constant has been tried for 5 kcal/mol/Å2 and 10 kcal/mol/Å2. That is, there are 6 kinds of simulation in total for this effort. Second, The force is applied only to helix II with other helices constrained. The force constant is 20 kcal/mol/Å2 and the force direction is vertical to Z-axis and at an angle of 45° with X-axis. Third, the force is applied only to helix V with the rest of helices constrained. The force direction is vertical to Z-axis and at an angle of 90° with X-axis. The force constant has been tried for 5 kcal/mol/Å2, 10 kcal/mol/Å2, 20 kcal/mol/Å2 and 50 kcal/mol/Å2. Fourth, the force is applied to riboflavin with the force parallel to Z-axis. The force constant has been tried for 5 kcal/mol/Å2, 10 kcal/mol/Å2 and 100 kcal/mol/Å2. Finally, RibU rotates around an axis parallel the Z-axis and through its center of mass with an angular acceleration −1 rad * ps−2. But all these efforts fail to form a channel for riboflavin transport due to lipid molecules entering into the channel.
References
Wallin, E. & von Heijne, G. Genome-wide analysis of integral membrane proteins from eubacterial, archaean and eukaryotic organisms. Protein Sci. 7, 1029–1038 (1998).
Vinothkumar, K. R. & Henderson, R. Structures of membrane proteins. Q. Rev. Biophys. 43, 65–158 (2010).
Rodionov, D. A. et al. A Novel Class of Modular Transporters for Vitamins in Prokaryotes. J. Bacteriol. 191, 42–51 (2009).
Eitinger, T., Rodionov, D. A., Grote, M. & Schneider, E. Canonical and ECF-type ATP-binding cassette importers in prokaryotes: diversity in modular organization and cellular functions. FEMS Microbiol. Rev. 35, 3–67 (2011).
ter Beek, J., Duurkens, R. H., Erkens, G. B. & Slotboom, D. J. Quaternary Structure and Functional Unit of Energy Coupling Factor (ECF)-type Transporters. J. Biol. Chem. 286, 5471–5475 (2011).
Neubauer, O. et al. Two Essential Arginine Residues in the T Components of Energy-Coupling Factor Transporters. J. Bacteriol. 191, 6482–6488 (2009).
Ivetac, A., Campbell, J. D. & Sansom, M. S. P. Dynamics and function in a bacterial ABC transporter: Simulation studies of the BtuCDF system and its components. Biochemistry (Mosc). 46, 2767–2778 (2007).
Jones, P. M. & George, A. M. Molecular-Dynamics Simulations of the ATP/apo State of a Multidrug ATP-Binding Cassette Transporter Provide a Structural and Mechanistic Basis for the Asymmetric Occluded State. Biophys. J. 100, 3025–3034 (2011).
Erkens, G. B. & Slotboom, D. J. Biochemical Characterization of ThiT from Lactococcus lactis: A Thiamin Transporter with Picomolar Substrate Binding Affinity. Biochemistry (Mosc). 49, 3203–3212 (2010).
Rodionov, D. A. et al. Transcriptional regulation of NAD metabolism in bacteria: genomic reconstruction of NiaR (YrxA) regulon. Nucleic Acids Res. 36, 2032–2046 (2008).
Zhang, P., Wang, J. & Shi, Y. Structure and mechanism of the S component of a bacterial ECF transporter. Nature 468, 717–U148 (2010).
Erkens, G. B. et al. The structural basis of modularity in ECF-type ABC transporters. Nat. Struct. Mol. Biol. 18, 755–U723 (2011).
Berntsson, R. P. A. et al. Structural divergence of paralogous S components from ECF-type ABC transporters. Proc. Natl. Acad. Sci. U. S. A. 109, 13990–13995 (2012).
Burgess, C. M. et al. The riboflavin transporter RibU in Lactococcus lactis: Molecular characterization of gene expression and the transport mechanism. J. Bacteriol. 188, 2752–2760 (2006).
Vogl, C. et al. Characterization of riboflavin (Vitamin B-2) transport proteins from Bacillus subtilis and Corynebacterium glutamicum. J. Bacteriol. 189, 7367–7375 (2007).
Lindahl, E. & Sansom, M. S. P. Membrane proteins: molecular dynamics simulations. Curr. Opin. Struct. Biol. 18, 425–431 (2008).
Gumbart, J., Wang, Y., Aksimentiev, A., Tajkhorshid, E. & Schulten, K. Molecular dynamics simulations of proteins in lipid bilayers. Curr. Opin. Struct. Biol. 15, 423–431 (2005).
Ash, W. L., Zlomislic, M. R., Oloo, E. O. & Tieleman, D. P. Computer simulations of membrane proteins. Biochim. Biophys. Acta, Biomembr. 1666, 158–189 (2004).
Hansson, T., Oostenbrink, C. & van Gunsteren, W. F. Molecular dynamics simulations. Curr. Opin. Struct. Biol. 12, 190–196 (2002).
Karplus, M. & McCammon, J. A. Molecular dynamics simulations of biomolecules. Nat. Struct. Biol. 9, 646–652 (2002).
Woolf, T. B. & Roux, B. Molecular dynamics simulation of the gramicidin channel in a phospholipid bilayer. Proc. Natl. Acad. Sci. U. S. A. 91, 11631–11635 (1994).
Roux, B. & Schulten, K. Computational studies of membrane channels. Structure 12, 1343–1351 (2004).
Majsnerowska, M. et al. Substrate-Induced Conformational Changes in the S-Component ThiT from an Energy Coupling Factor Transporter. Structure 21, 861–867 (2013).
Wang, T. et al. Structure of a bacterial energy-coupling factor transporter. Nature 497, 272–276 (2013).
Xu, K. et al. Crystal structure of a folate energy-coupling factor transporter from Lactobacillus brevis. Nature 497, 268–271 (2013).
Isralewitz, B., Gao, M. & Schulten, K. Steered molecular dynamics and mechanical functions of proteins. Curr. Opin. Struct. Biol. 11, 224–230 (2001).
Hytoenen, V. P. & Vogel, V. How force might activate talin's vinculin binding sites: SMD reveals a structural mechanism. PLoS Comp. Biol. 4, (2008).
Soto, C. S., Fasnacht, M., Zhu, J., Forrest, L. & Honig, B. Loop modeling: Sampling, filtering and scoring. Proteins 70, 834–843 (2008).
Xiang, Z. X., Soto, C. S. & Honig, B. Evaluating conformational free energies: The colony energy and its application to the problem of loop prediction. Proc. Natl. Acad. Sci. U. S. A. 99, 7432–7437 (2002).
Koshland Jr, D. E., Nemethy, G. & Filmer, D. Comparison of experimental binding data and theoretical models in proteins containing subunits. Biochemistry (Mosc). 5, 365–385 (1966).
Duurkens, R. H., Tol, M. B., Geertsma, E. R., Permentier, H. P. & Slotboom, D. J. Flavin binding to the high affinity riboflavin transporter RibU. J. Biol. Chem. 282, 10380–10386 (2007).
Locher, K. P., Lee, A. T. & Rees, D. C. The E-coli BtuCD structure: A framework for ABC transporter architecture and mechanism. Science 296, 1091–1098 (2002).
Oloo, E. O. & Tieleman, D. P. Conformational transitions induced by the binding of MgATP to the vitamin B-12 ATP-binding cassette (ABC) transporter BtuCD. J. Biol. Chem. 279, 45013–45019 (2004).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. Model. 14, 33–38 (1996).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Brooks, B. R. et al. CHARMM: a program for macromolecular energy, minimisation and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N.log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Pastor, R. W., Brooks, B. R. & Szabo, A. An analysis of the accuracy of Langevin and molecular dynamics algorithms. Mol. Phys. 65, 1409–1419 (1988).
Feller, S. E., Yuhong, Z., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Bakan, A., Meireles, L. M. & Bahar, I. ProDy: Protein Dynamics Inferred from Theory and Experiments. Bioinformatics 27, 1575–1577 (2011).
Wang, J. M., Morin, P., Wang, W. & Kollman, P. A. Use of MM-PBSA in reproducing the binding free energies to HIV-1 RT of TIBO derivatives and predicting the binding mode to HIV-1 RT of efavirenz by docking and MM-PBSA. J. Am. Chem. Soc. 123, 5221–5230 (2001).
Zoete, V., Cuendet, M. A., Grosdidier, A. & Michielin, O. SwissParam: A fast force field generation tool for small organic molecules. J. Comput. Chem. 32, 2359–2368 (2011).
Rocchia, W. et al. Rapid grid-based construction of the molecular surface and the use of induced surface charge to calculate reaction field energies: Applications to the molecular systems and geometric objects. J. Comput. Chem. 23, 128–137 (2002).
Jensen, M. O., Park, S., Tajkhorshid, E. & Schulten, K. Energetics of glycerol conduction through aquaglyceroporin GlpF. Proc. Natl. Acad. Sci. U. S. A. 99, 6731–6736 (2002).
Jensen, M. O., Yin, Y., Tajkhorshid, E. & Schulten, K. Sugar transport across lactose permease probed by steered molecular dynamics. Biophys. J. 93, 92–102 (2007).
Liu, X. et al. Mechanics of channel gating of the nicotinic acetylcholine receptor. PLoS Comp. Biol. 4, 0100–0110 (2008).
Acknowledgements
We thank the National Natural Science Foundation of China (Grants No. 21003048, 10974054 and 20933002). C.G.J. is also supported by “the Fundamental Research Funds for the Central Universities” and Open Research Fund of the State Key Laboratory of Precision spectroscopy, East China Normal University. We also thank the Computer Center of ECNU for providing us computational time.
Author information
Authors and Affiliations
Contributions
J.S. performed MD simulation, data analysis and wrote the main manuscript text and prepared all the figures. C.J. and J.Z.H.Z. discussed the results and revised the final draft of the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Info
Supplementary Information
Motion of protein
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Song, J., Ji, C. & Zhang, J. Unveiling the gating mechanism of ECF Transporter RibU. Sci Rep 3, 3566 (2013). https://doi.org/10.1038/srep03566
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03566
This article is cited by
-
Identification of inhibitors targeting the energy-coupling factor (ECF) transporters
Communications Biology (2023)
-
Functional Loop Dynamics of the Streptavidin-Biotin Complex
Scientific Reports (2015)
-
Structures of FolT in substrate-bound and substrate-released conformations reveal a gating mechanism for ECF transporters
Nature Communications (2015)
-
Pyridoxamine is a substrate of the energy-coupling factor transporter HmpT
Cell Discovery (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.