Abstract
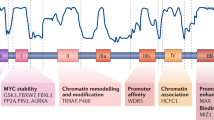
The c-myc protooncogene encodes the Myc transcription factor, a global regulator of fundamental cellular processes. Deregulation of c-myc leads to tumorigenesis and c-myc is an important driver in human cancer. Myc and its dimerization partner Max are bHLH-Zip DNA binding proteins involved in transcriptional regulation of target genes. Non-transcriptional functions have also been attributed to the Myc protein, notably direct interaction with the pre-replicative complex (pre-RC) controlling the initiation of DNA replication. A key component of the pre-RC is the Cdt1 protein, an essential factor in origin licensing. Here we present data suggesting that the CDT1 gene is a transcriptional target of the Myc-Max complex. Expression of the CDT1 gene in v-myc-transformed cells directly correlates with myc expression. Also, human tumor cells with elevated c-myc expression display increased CDT1 expression. Occupation of the CDT1 promoter by Myc-Max is demonstrated by chromatin immunoprecipitation and transactivation by Myc-Max is shown in reporter assays. Ectopic expression of CDT1 leads to cell transformation. Our results provide a possible direct mechanistic link of Myc's canonical function as a transcription factor to DNA replication. Furthermore, we suggest that aberrant transcriptional activation of CDT1 by deregulated myc alleles contributes to the genomic instabilities observed in tumor cells.
Similar content being viewed by others
Introduction
The myc oncogene was originally discovered as the transforming principle in the genome of avian acute leukemia virus MC291, representing a transduced retroviral allele (v-myc) derived from the chicken cellular protooncogene c-myc2,3. The Myc protein product, initially identified as a Gag-Myc hybrid protein specified by MC294, is a transcription factor with strong oncogenic potential2,3,5,6. Myc is a bHLH-Zip protein, forms heterodimers with the Max protein, binds to specific DNA sequence elements (E-boxes) and is the central hub of a global transcriptional regulator network7,8. Myc and Max homologs with conserved basic functions were found in primitive metazoans9 and there is even evidence for a premetazoan evolutionary origin of Myc and Max proteins10. The human Myc transcription factor network controls thousands of genes involved in fundamental cellular processes including growth, proliferation, differentiation, metabolism and apoptosis5,6,11,12,13. The basic function of the Myc-Max transcription factor complex is transcriptional activation of distinct target genes, but Myc has also been implicated in transcriptional repression of specific genes5,6,11. The target genes transcriptionally activated by Myc-Max are related to important pathways of cell growth and metabolism, including protein synthesis, ribosomal biogenesis, glycolysis, mitochondrial function and cell cycle progression5,6,11,14. The genes repressed by Myc are typically involved in cell cycle arrest, cell adhesion and cell-to-cell communication5,6,11, or encode inhibitors of Myc-induced cell transformation15. The discovery of rearrangements and transcriptional deregulation of the human MYC gene in Burkitt's lymphoma was the first indication that the cellular homolog of the retroviral v-myc oncogene is involved in human tumorigenesis16 and it is now established that MYC is one of the crucial drivers in many, if not most human cancers3,5,6,17.
Myc has also been associated with non-transcriptional functions, possibly not even requiring dimerization with Max5,6. An important example is non-transcriptional control of DNA replication by Myc18, providing a possible link to genomic instability typically observed in cells with deregulated Myc expression19,20,21. Genetic instabilities, including changes at the nucleotide level, aneuploidy, chromosome translocations and gene amplification, are a hallmark of many human cancers22,23. It has been proposed that the non-transcriptional control of DNA replication involves direct interaction of the Myc protein with components of the pre-replicative complex (pre-RC)18,24. Eukaryotic DNA replication is tightly regulated both spatially and temporally to ensure correct copying of the entire genome only once in every cell cycle. To prevent rereplication, licensing of specific replication origins in the G1 phase of the cell cycle is achieved by the assembly of the pre-RC onto chromatin, starting with recruitment of the origin recognition complex (ORC), followed by loading of the minichromosome maintenance complex (MCM) mediated by the Cdc6 and Cdt1 proteins and additional replication proteins25,26. The Cdt1 protein, originally identified in yeast and then in insects and vertebrates26,27,28,29, promotes the loading of MCM and is the key factor in the licensing process. In higher eukaryotes, Cdt1 activity is therefore strictly regulated by ubiquitin-dependent degradation and binding of the specific inhibitor geminin to ensure temporal confinement of licensing to the G1 phase30,31. Here we report that the CDT1 gene is a transcriptional target of the Myc-Max complex and that deregulated Myc expression in transformed cells leads to increased expression of the essential DNA replication factor Cdt1. Our results suggest a direct implication of Myc's fundamental function as a transcriptional regulator in genomic instabilities observed in tumor cells.
Results
Activation of CDT1 in myc-transformed cells
Using a conditional cell transformation system in which expression of the MC29 v-myc allele is controlled by doxycycline32, several partial cDNA clones representing candidate myc target genes were isolated by representational difference analysis (RDA), a polymerase chain reaction (PCR)-based subtractive hybridization procedure. One of these clones was of particular interest since it proved to be derived from the gene encoding the DNA replication licensing factor Cdt1, providing a possible link of Myc transcription factor function with DNA replication. The tight correlation of v-myc and CDT1 mRNA levels was demonstrated in the conditional cell transformation systems Q/tMON and Q/tM8 in which v-myc expression is controlled by a doxycycline-dependent or a doxycycline-inhibited transactivator, respectively32. In time course experiments, expression of CDT1 mRNA closely parallels that of the conditional v-myc alleles in both cell systems and every activation/deactivation of the oncogene by addition or removal of the drug, even repeatedly, is precisely reflected in concurrent changes in CDT1 expression (Figure 1a and Supplementary Figure 1). Notably, the CDT1 expression pattern parallels that of the specific myc target WS5 (also called Mmp115 in the chicken genome) encoding a protein related to human melanoma glycoproteins33 and is exactly opposite to the expression pattern of BASP1, a specific target suppressed by Myc and shown to be an inhibitor of Myc's transforming function15. Comparative expression analyses were also performed with normal quail embryo fibroblasts (QEF) or quail cell lines transformed by v-myc (Q8, QEF/MC29, QEF/Rc-Myc), v-jun (VJ, VCD), v-src (R(-)3), or by a chemical carcinogen (QT6). All rapidly growing transformed cells contain elevated levels of CDT1 mRNA, but the highest levels were found in v-myc-transformed cells (Figure 1b). Expression of the myc target WS5 was used as a control. Simultaneous activation of CDT1 and the WS5/Mmp115 control gene was also observed in chicken embryo fibroblasts (CEF) transformed by the MC29 v-myc allele (Figure 1c). Furthermore, expression analysis using the human leukemia cell lines K-562, MOLT-4 and HL-60 and the colon carcinoma line SW-480 revealed a strong correlation of elevated c-myc mRNA levels with CDT1 expression (Figure 1d), indicating that deregulated c-myc, like v-myc, is possibly involved in CDT1 activation. To test if CDT1 activation by v-myc is also detectable at the protein level, a 726-bp fragment of quail CDT1 cDNA (see below) was cloned into prokaryotic expression vector pET-21a and used for the production of a truncated Cdt1 recombinant protein. The purity and identity of recombinant Cdt1(200–441) was verified by mass spectrometry and fragment ion mapping (Supplementary Figure 2) and the protein was used for the generation of a polyclonal antiserum. Using this serum, a translational product with an apparent Mr of 74,000 (p74CDT1) was detected in the Q8 and QEF/MC29 cell lines which are transformed by the p110 Gag-Myc hybrid protein, but not in normal QEF (Figure 1e).
CDT1 activation in cells with elevated v-myc or c-myc expression.
(a) Kinetics of CDT1 activation monitored by Northern analysis using total RNAs from the quail cell line Q/tMON conditionally transformed by a v-myc allele controlled by a doxycycline-dependent transactivator32. Cells were first grown continuously in the presence (+) of doxycycline, at day −6 the drug was removed, re-added at day 0, removed again after 6 days and the cells were incubated for further 6 days. RNAs were isolated before removal or addition of the drug and at the time points indicated. (b) Northern analysis of poly(A)+-selected RNAs (2.5 μg) from normal quail embryo fibroblasts (QEF), or the quail cell lines Q8, QEF/MC29, QEF/Rc-Myc transformed by v-myc, QT6 chemically transformed by methylcholanthrene, VJ and VCD transformed by v-jun, or R(-)3 transformed by v-src. Filters in (a) and (b) were hybridized with 32P-labeled cDNA probes as indicated: quail CDT1, WS5 and GAPDH, chicken BASP1 and MC29 v-myc. Sizes of the mRNAs were: CDT1, 2.5 kb; WS5, 2.8 kb; BASP1, 2.0 kb; v-myc, 1.9 kb; GAPDH, 1.4 kb. (c) Northern analysis of poly(A)+-selected RNAs (5.0 μg) from normal chicken embryo fibroblasts (CEF), or clonal cultures of CEF transformed by MC29 (v-myc). Filters were hybridized with probes as in (a) and (b). Sizes of the mRNAs were: CDT1, 2.5 kb; WS5 (Mmp115), 2.6 kb; GAPDH, 1.4 kb. (d) Northern analysis of poly(A)+-selected RNAs (2.5 μg) from normal human fibroblasts (hFB), or from the human cancer cell lines K-562 (chronic myeloid leukemia), MOLT-4 (T-cell leukemia), HL-60 (acute myeloid leukemia) and SW-480 (colon adenocarcinoma). The filter was hybridized with 32P-labeled cDNA probes specific for the human CDT1, c-myc and β-actin (ACTB) genes. Sizes of the mRNAs were: CDT1, 2.7 kb and 2.0 kb; c-myc, 2.4 kb; ACTB, 2.0 kb. (e) SDS-PAGE (10% wt/vol) analysis of v-Myc (Gag-Myc), c-Myc, Cdt1 and tubulin α (TUBA) proteins using metabolically [35S]methionine-labeled lysates from QEF, or the v-myc transformed quail cell lines Q8 and QEF/MC29. Aliquots (1 × 107 cpm) of the lysates were immunoprecipitated with the antisera indicated. Full-length gels and blots are included in the supplementary information.
Structural organization of the CDT1 gene and protein
A full-length 2,162-bp CDT1 cDNA clone was assembled from overlapping clones obtained by screening a cDNA library from the MC29-transformed quail cell line Q84 with a CDT1 RDA fragment (see above) and from four overlapping 5′ RACE products. The deduced 588-amino acid quail Cdt1 protein has a calculated Mr of 64,690 and an isoelectric point of 9.94. An alignment of the amino acid sequences of the quail Cdt1 protein and the chicken, mouse and human homologs revealed extensive sequence similarities (Supplementary Figure 3), particularly in the binding domains for geminin and the MCM complex34,35. To determine the structural organization of the CDT1 gene, the nucleotide sequence of a 5,677-bp quail genomic DNA segment hybridizing with a CDT1 cDNA probe was determined. The quail CDT1 gene contains 10 exons and displays a similar architecture (Figure 2a) like its chicken ortholog (contig NC_006098). The nucleotide sequences of the quail and chicken CDT1 promoter regions are highly conserved and contain two canonical Myc binding sites (CACGTG) immediately upstream of the transcriptional start sites (Figure 2b). Comparison of the avian CDT1 promoters with those from mouse and human (Figure 2c) showed that the mammalian regulatory regions also contain several canonical and non-canonical E-boxes within 2 kbp upstream of the transcription start sites. A further common feature of the avian and mammalian promoters is the presence of a consensus E2F binding site in close proximity to the transcription start site (Figure 2c).
Structure of the CDT1 gene.
(a) In the schematic diagram of the quail CDT1 gene, the ten exons are depicted as boxes with the coding region shown in black. The structure of a cDNA clone encoding the 588-amino acid Cdt1 protein is shown above. (b) Nucleotide sequence alignment of the quail (q) and chicken (ck) CDT1 5′ regulatory regions (accession nos. KF494239, NC_006098.2). Identical nucleotides are shaded in yellow, the conserved distal (d) and proximal (p) canonical Myc binding sites (E-boxes) are boxed in red, the transcription start (TS) sites in black. The positions of the 5′- and 3′-primers (pr.) used for ChIP and quantitative (qnt.) ChIP are underlined. (c) Schematic diagram of the quail (Coturnix japonica), chicken (Gallus gallus), mouse (Mus muculus) (accession no. NT_078575) and human (Homo sapiens) (accession no. AC092384) CDT1 promoters. Transcription start sites (arrows) and the positions of possible binding sites for Myc (canonical, non-canonical) or E2F are indicated.
Myc binds to the CDT1 promoter
To test if Myc binds to the CDT1 promoter in vivo, chromatin immunoprecipitation (ChIP) analysis was performed using chromatin from normal QEF and from QEF transformed by MC29 encoding the p110 Gag-Myc hybrid protein4, or by the Rc-Myc construct encoding a p52 v-Myc protein not fused to viral structural proteins15. Both QEF/MC29 and QEF/Rc-Myc contain high levels of CDT1 mRNA (cf. Figure 1b). Antisera directed against Myc or Max and NRS as a control and PCR primers flanking the canonical E-boxes in the CDT1 promoter (cf. Figure 2b) or primers flanking the Myc binding sites in the bona fide Myc target WS533 were used (Figure 3a). Strikingly, the principal ChIP pattern obtained with the CDT1 primers is nearly identical to that obtained for the WS5 promoter. In normal QEF, the CDT1 and WS5 promoters are mainly occupied by Max which also forms stable homodimers and by only low amounts of endogenous c-Myc protein. In the v-myc-transformed cells, both promoters are bound by equal amounts of Myc and Max, presumably Myc-Max heterodimers. This was fully confirmed when occupation of the chicken CDT1 promoter was tested in normal CEF and CEF/MC29 (Figure 3b). Again, the ChIP patterns for CDT1 and WS5 were very similar. As an additional control, primers from an unrelated gene region (BASP1) lacking E-boxes were used in the ChIP analysis. Further ChIP analyses included the chemically transformed cell line QT6 and the Q8 cell line transformed by MC29 (Figure 3c). To directly prove that viral Myc protein products occupy the CDT1 promoter in v-myc-transformed cells, an α-Gag serum was also used. The binding pattern of the CDT1 promoter in chemically transformed cells was similar to that in normal QEF. Occupation of the promoter in cells transformed by the p110 Gag-Myc hybrid protein (QEF/MC29, Q8) was demonstrated not only with Myc and Max antisera, but also with the Gag antiserum, whereas ChIP on cells transformed by a v-Myc protein unlinked to viral structural proteins (QEF/Rc-Myc) was positive only with the Myc and Max antisera (Figure 3c). In order to quantify binding of Myc and Max to the CDT1 promoter, real-time quantitative PCR was performed on the ChIP products (Figure 3d). The results are in complete agreement with the qualitative analysis. A strong increase of Myc binding to the CDT1 promoter was observed in the v-myc-transformed cell lines QEF/MC29 and QEF/Rc-Myc compared to normal QEF, whereas Max binding is observed in normal and transformed cells.
In vivo binding of Myc to the quail and chicken CDT1 promoters.
(a) ChIP analysis using chromatin from normal QEFs, or from QEF/MC29 and QEF/Rc-Myc transformed by the v-Myc protein. (b) ChIP using chromatin from normal CEF and from CEF/MC29. (c) ChIP using chromatin from normal QEF, from the chemically transformed quail cell line QT6, or from the v-myc-transformed cell lines QEF/MC29, QEF/Rc-Myc and Q8. The QEF(CEF)/MC29 cells and the Q8 cell line are transformed by the p110 Gag-Myc hybrid protein, whereas the QEF/Rc-Myc cells are transformed by a v-Myc protein not fused to viral structural protein sequences. Antisera directed against Myc, Max, or Gag were used for immunoprecipitation, followed by PCR amplification of DNA from the quail CDT1 (241 bp fragment) or chicken CDT1 (253 bp fragment) regulatory regions both containing two canonical Myc binding sites, or of DNA from the quail WS5 (290 or 273 bp fragment) and chicken WS5 (Mmp115) (293 bp fragment) promoters both containing four Myc binding sites33. Primers specific for a chicken BASP1 exon 2 fragment (197 bp) containing no E-boxes (control) and reactions with normal rabbit serum (NRS) or 1.6% of total chromatin (input) were used as controls. Fragments were analyzed by agarose (1.5% wt/vol) gel electrophoresis. (d) ChIP-quantitative PCR (ChIP-qPCR) using chromatin from normal QEF, or from QEF/MC29 and QEF/Rc-Myc. Immunoprecipitated DNA was analyzed in triplicate by qPCR using primers to amplify a 167-bp segment of the regulatory region from the quail CDT1 promoter containing the two canonical Myc binding sites. Signals were normalized to input chromatin and shown as % input. Full-length gels are included in the supplementary information.
Electrophoretic mobility shift analysis (EMSA) was employed to test in vitro binding of a recombinant Myc-Max heterodimer complex36 to DNA probes derived from the CDT1 promoter region containing either the proximal or the distal E-box (Supplementary Figure 4a). The purified Myc-Max complex (Supplementary Figure 4b) bound to double-stranded DNA containing the CDT1 consensus Myc binding sites with high affinity (Supplementary Figure 4c and d). Application of DNA probes in which the E-boxes were either mutated or deleted (Supplementary Figure 4a) led to significantly reduced binding. As controls, DNA probes containing a consensus E-box (USF E) for Myc binding36 or no E-box (control) and a recombinant Cdt1 protein fragment instead of the Myc-Max complex were used. To quantify the binding affinities, increasing amounts of Myc-Max proteins were added to constant amounts of DNA in EMSA experiments (Supplementary Figure 5a), the ratios of bound to total DNA were determined and the dissociation constants (Kd) for the protein-DNA complexes were calculated (Supplementary Figure 5b). The Kd values for protein-DNA complexes formed by Myc-Max and DNA containing the proximal or distal E-box of the CDT1 promoter were 1.9 and 2.2 nM, respectively, similar to the Kd determined previously for the USF E control probe36. The residual albeit strongly reduced binding of the mutated DNA probes may be due to the nonspecific E-box-independent DNA binding capacity of Myc5,13.
Transcriptional activation of the CDT1 promoter by Myc
To determine whether Myc is indeed involved in transcriptional activation of the CDT1 gene, the quail CDT1 promoter region encompassing nucleotides −217 through +42 (cf. Figure 2) was cloned into a luciferase reporter plasmid to yield pGL3-CDT1. For comparison, the reporter plasmid pGL3-WS5 containing the promoter region of the Myc target WS515,33 was analyzed in parallel. The analysis revealed that the CDT1 promoter was efficiently activated in v-myc-transformed QEF/Rc-Myc or QEF(CEF)/MC29 similar to the WS5 promoter and both promoters showed only basal activity in normal QEF or CEF (Figure 4a). Furthermore, efficient transactivation of the pGL3-CDT1 and pGL3-WS5 promoter constructs was observed in normal QEF or CEF that were transiently co-transfected with a pRc/RSV-derived expression vector (pRc-Myc) encoding the p52 v-Myc protein, as compared to cells co-transfected with the empty pRc vector (Figure 4b). Promoter analysis of pGL3-CDT1 mutants in which the two E-boxes had either been mutated (*) or deleted (Δ) revealed that intact E-boxes are required for efficient transactivation (Figure 4b and Supplementary Figure 6). Expression of ectopic v-Myc and endogenous c-Myc proteins in transfected cells was monitored by immunoblotting. To analyze the contributions of the proximal and distal canonical E-boxes in transactivation of the CDT1 promoter, reporter constructs were created in which either one or both of the binding sites were mutated (Supplementary Figure 7). Co-transfection with pRc-Myc and luciferase analysis revealed that mutation of the proximal E-box was sufficient to abrogate the activation observed for the wild-type promoter, whereas mutation of the distal site allowed residual promoter activation. We note that the mutational analyses of the CDT1 promoter suggest that the transcriptional regulation by the Myc-Max complex is a direct mechanism.
Transcriptional transactivation of the quail CDT1 promoter.
(a) Aliquots (2.0 μg) of the pGL3-Basic vector, or of the pGL3-CDT1 and pGL3-WS5 reporter constructs were transfected together with 1.0 μg of the pSV-β-galactosidase plasmid into equal numbers (8 × 105) of normal QEF or CEF, or of the v-myc-transformed cell lines QEF/Rc-Myc, QEF/MC29 and CEF/MC29. Luciferase activities in relative light units (RLU) normalized to β-galactosidase activities were determined for 10-μl aliquots of cell extracts prepared 24 h after transfection. (b) Aliquots (2.0 μg) of the pGL3-Basic vector, or of the reporter constructs pGL3-CDT1, pGL3-WS5, or pGL3-CDT1*Epd (in which both Myc binding sites have been mutated) were cotransfected with 2.0-μg aliquots of a pRc-derived expression vector encoding the v-Myc protein (pRc-Myc) or of the empty expression vector (pRc), together with the pSV-β-galactosidase plasmid (1.0 μg) into 8 × 105 normal QEF or CEF. Luciferase assays were performed as in (a). For control of protein expression, equal amounts of cell extracts (30 μl for QEF, 60 μl for CEF) were analyzed by SDS-PAGE (10% wt/vol) and v-Myc, c-Myc and tubulin α proteins were detected by immunoblotting. Full-length blots are included in the supplementary information.
Deregulated CDT1 expression induces cell transformation
The strong activation of CDT1 in v-myc-transformed cells prompted us to test if overexpression of CDT1 by itself would induce parameters of the transformed phenotype. Therefore, the entire CDT1 coding sequence was inserted into the replication-competent retroviral pRCAS vector (Figure 5a) and the pRCAS-CDT1 construct was transfected into QEF. For control and comparison, QEF were also transfected with the empty pRCAS vector or with pRCAS-MC2915. Expression of the proteins encoded by the retroviral constructs was verified by immunoprecipitation and SDS-PAGE. In QEF transfected with pRCAS-CDT1 or pRCAS-MC29, the ectopic p74 Cdt1 and p110 Gag-Myc proteins, as well as increased levels of endogenous p74 Cdt1 in cells transformed by pRCAS-MC29 were detected (Figure 5b). Cells transfected with the MC29 construct showed the typical morphology and capacity for anchorage-independent growth of fully v-myc-transformed cells4. QEF transfected with pRCAS-CDT1 were able to grow in semi-solid medium at relatively high numbers, although the colonies were significantly smaller than those induced by transformation with pRCAS-MC29 (Figure 5c). Similar results were obtained when CEF were used (Supplementary Figure 8). CEF were transfected with the empty pRCAS vector, pRCAS-CDT1, pRCAS-MC29, or pRCAS-WS533. The overexpressed p74 Cdt1 protein from RCAS-CDT1-transfected CEF co-migrated with in vitro translated Cdt1 in the SDS-PAGE and the MC29 Gag-Myc hybrid protein and the p118 WS5 protein33 were efficiently expressed from the retroviral vectors (Supplementary Figure 8a). MC29-transfected CEF showed full transformation and WS5 overexpression led to partial cell transformation and colony formation as reported previously33. Ectopic expression of CDT1 led to morphological changes and colony forming capacity in CEF, although the numbers of soft agar colonies were significantly lower both for CDT1 and WS5 compared to those obtained with the MC29 v-myc oncogene (Supplementary Figure 8b).
Transforming potential of ectopically expressed CDT1.
(a) Structure of the retroviral pRCAS vector with a ClaI cloning site. Insertion of adapter DNA carrying a cDNA fragment containing the entire CDT1 coding region yielded plasmid construct pRCAS-CDT1 specifying a replication-competent retrovirus encoding the quail Cdt1 protein. LTR, long terminal repeat; gag, pol, env, essential retroviral genes; bla, β-lactamase. (b) SDS-PAGE (10% wt/vol) analysis of v-Myc (Gag-Myc) and Cdt1 proteins ectopically expressed after transfection of QEF with the replication-defective construct pRCAS-MC29 together with pRCAS as a helper virus construct, with the replication-competent construct pRCAS-CDT1, or with pRCAS as a control. Aliquots (1 × 107 cpm) of lysates from cells metabolically labeled with [35S]methionine were immunoprecipitated with the antisera indicated. (c) Upper panels: phase-contrast micrographs of QEF transfected with vector DNA (pRCAS), or with the retroviral constructs pRCAS-CDT1 or pRCAS-MC29/pRCAS. Lower panels: agar colony formation by cells transfected with pRCAS-CDT1 or pRCAS-MC29/pRCAS. Equal numbers of cells (1 × 105) were seeded in soft agar. pRCAS-transfected QEF were used as controls. Bright-field micrographs were taken after 3 weeks. Numbers of colonies/1000 cells seeded are indicated.
Discussion
Extensive investigations into Myc biology and biochemistry over more than 30 years have revealed that the myc gene is essential for normal cell function, but also one of the major driving forces in human cancer3,6. The physiological role and the oncogenic capacity of the Myc protein are largely based on its principal biochemical function as a regulator of gene expression, exhibiting both transcriptional activating and repressing potential. Elucidation of the cellular pathways affected by Myc therefore requires identification of the relevant target genes. The unambiguous definition of distinct targets and their detailed mechanistic link to Myc's important biological functions is substantially complicated by the vast number of genes apparently controlled by the Myc-Max network. Combinations of microarray screening, methylation marking procedures, global chromatin immunoprecipitation, high throughput sequencing and expression profiling have been used to identify genome-wide binding sites and putative targets for Myc12,13,37,38,39,40,41. Obviously, for the majority of the thousands of putative regulatory targets identified, over 7000 alone in Burkitt lymphoma cells40, detailed analyses of their transcriptional regulation is not available and not all of the binding sites may mediate transcriptional regulation5. Despite the enormous complexity of the Myc-directed transcriptome, previous detailed analyses of the regulation and biochemical function of distinct Myc targets6,11 and the overlapping patterns of the recent high throughput approaches have provided an emerging picture of major cellular processes controlled by Myc5,6. Myc regulates energy metabolism by controlling a large number of genes involved in glycolysis, glutamine metabolism and mitochondrial biogenesis5,6,11,14,42,43. Furthermore, Myc controls genes involved in protein synthesis and ribosome biogenesis, key processes in biomass accumulation prior to cell division5,6. Myc is also involved in cell cycle regulation, induces G1-S progression and directly activates the genes encoding cyclin D2 and CDK46,44,45,46.
In this report, we provide a possible direct molecular link of the canonical transcriptional function of Myc to DNA replication, another fundamental cellular process. We show that deregulated Myc expression leads to transcriptional activation of the CDT1 gene, encoding the key DNA replication licensing factor Cdt1 in eukaryotes. Through late mitosis and G1, replication origins become licensed for DNA replication in the S phase by loading MCM proteins in order to assemble the pre-RC complex25,47. The Cdt1 protein, together with ORC and Cdc6, is responsible for recruitment of the MCM complex. To ensure that chromosomal DNA is replicated only once per cell cycle, origin licensing is restricted to late mitosis and G1. Metazoans prevent licensing during S phase and G2 mainly by downregulation of Cdt1 activity by ubiquitin-dependent proteolysis and activation of the Cdt1 inhibitor geminin30,31,47,48,49. Our results provide a comprehensive molecular definition of the CDT1 gene as a transcriptional target of Myc, shown by correlation of CDT1 expression with conditional or non-conditional Myc expression and by extensive promoter analyses including ChIP, EMSA and reporter assays. Notably, in normal non-transformed cells the CDT1 promoter is mainly occupied by the Max protein, possibly homodimeric or in complex with another E-box binding factor and to a lesser degree by c-Myc. In myc-transformed cells, the ChIP analyses show equal occupation by Myc and Max, as expected for the heterodimeric structure of the Myc-Max complex. It has been postulated that deregulated Myc expression leads to a switch in the regulation of E-box genes that are normally controlled by other E-box transcription factors and that this may be an important mechanism contributing to Myc-induced oncogenesis6. In addition to canonical and non-canonical Myc binding sites, all CDT1 promoter regions compared (cf. Figure 2) contain a binding site for E2F transcription factors near the transcription start site. Based on the genome-wide analysis of Myc binding sites in the human B-cell system, it was found that many of the candidate Myc targets also contain E2F binding sites, suggesting that Myc targets may require additional regulators of gene expression6,12,13. It was indeed reported that human CDT1 expression is regulated by E2F50. Interestingly, based on genome-wide high throughput technologies genes encoding other components of DNA replication licensing, notably MCM proteins, were listed among putative Myc-regulated genes12,13,38,41. In a single study based on a microarray approach, CDT1 was also listed as an initial candidate target, but then, in contrast to our current results, classified as a gene not regulated by Myc and the mechanism of its transcriptional regulation was not analyzed further51. Although direct transcriptional activation of CDT1 by Myc ist the most plausible explanation for the results of several independent experimental approaches as reported here, we would like to point out that indirect mechanisms like changes in CDT1 expression due to altered cell cycle distribution by deregulated Myc or due to activation of E2F by Myc52 cannot be ruled out.
Recent reports have demonstrated that Myc has a non-transcriptional role in the control of DNA replication18,53. Intriguingly, this involves direct interaction of the Myc protein with components of the pre-RC, including MCM proteins and Cdt1. It was shown that overexpression of Myc causes increased replication origin activity and subsequent DNA damage18 and it was proposed that genomic instability observed in cancer cells with high Myc expression is related to deregulation of DNA replication by Myc - pre-RC interactions18,24. Our current results demonstrate that the CDT1 gene encoding the major regulator of DNA replication and important safeguard against rereplication in metazoans is a transcriptional target of Myc, providing a mechanistic link of Myc's canonical function as a transcription factor to DNA replication licensing, in addition to its non-transcriptional function. Addition of Cdt1 to replicated nuclei does indeed stimulate rereplication48 and we (Figure 5 and Supplementary Figure 8) and others54 have shown that deregulated CDT1 exhibits oncogenic potential. Our results suggest that aberrant transcriptional activation of the CDT1 gene by constitutive Myc expression in tumor cells leads to genomic instabilities observed not only in myc-transformed cells19,20,21, but considered as a major hallmark of human cancer22,23. Further analyses are required to assess in detail the putative role of CDT1 as an effector in Myc-induced carcinogenesis.
Methods
Cells and retroviruses
Cell culture, DNA transfection and transformation of quail or chicken embryo fibroblasts (QEF, CEF) were performed as described4,15. The established quail cell lines Q8, QEF/MC29, QEF/Rc-Myc, QT6, VJ, VCD, R(-)3 and the cell lines Q/tM8 and Q/tMON conditionally transformed by v-myc were used4,15,32,33. To generate CEF freshly transformed by the MC29 retroviruses, primary cells were transfected with pRCAS-MC29 as described15. The human cancer cell lines K-562, MOLT-4, HL-60 and SW-480 which are derived from chronic myeloid leukemia cells in blast crisis, acute lymphoblastic leukemia T lymphoblasts, acute myeloid leukemia cells, or colon adenocarcinoma cells, respectively and human immortalized fibroblasts (hFB) were provided by J. Troppmair (Medical University Innsbruck). The retroviral expression vector pRCAS-WS5 has been described33. For construction of pRCAS-CDT1, a 1767-bp cDNA fragment containing the CDT1 coding region was inserted into the ClaI site of the pRCAS vector as described15. DNA transfection and colony assays of transfected cells in 0.3% (wt/vol) agarose were done as described4,33.
DNA cloning and nucleic acid analysis
Molecular cloning, DNA sequencing, Northern analysis and 5′-rapid amplification of cDNA ends (RACE) have been described15,33. Subtractive probe generation and representational difference analysis (RDA) were done as described55 using RNAs isolated from Q/tM832 grown in the absence of doxycycline and from the same cells 48 h after addition of doxycycline for generation of tester and driver cDNAs, respectively. For the isolation of cDNA clones representing the quail CDT1 gene, a Q8-specific cDNA library was screened as described33 using an RDA fragment as a probe. 5′-RACE was applied to obtain a 2,162-bp full-length cDNA sequence. For determination of the CDT1 gene structure, clones were isolated from a quail genomic DNA library as described33. Bioinformatic analyses of nucleotide and amino-acid sequences of the CDT1 gene were performed as described15,33,55. For multiple sequence alignments, the ClustalW algorithm (DS/Gene, Accelrys, San Diego, CA) was used. The nucleotide and deduced amino-acid sequences have been deposited in the GenBank database (accession nos. HQ840726, KF494239). Hybridization probes for Northern analysis included cDNA fragments derived from quail CDT1, WS5, GAPDH and v-myc as described previously33. To obtain DNA probes specific for human CDT1 and β-actin, PCR was performed on cDNAs derived from MOLT-4 or human bronchial epithelial cells (HBEpC), respectively, using gene specific primers. A human c-myc probe specifying a region from c-myc exon 3 was amplified using genomic DNA as a template.
Chromatin immunoprecipitation analyses
ChIP analyses were carried out as described by using sheared extracts derived from formaldehyde-treated normal QEF or CEF and from the transformed cell lines QT6, Q8, QEF/MC29, QEF/Rc-Myc and CEF/MC2915,33. Immunoprecipitations were performed with specific antibodies followed by PCR amplification of fragments from the quail CDT1 (241 bp) or chicken CDT1 (253 bp) regulatory regions, or from the quail WS5 (QEF, QEF/MC29: 290 bp; QEF/Rc-Myc: 273 bp) and chicken WS5/Mmp115 (293 bp) promoters using the primer pairs 5′-GGTAACACAGTACAGCCCTCGC-3′/5-GCCGAAGAAATCGGTGAGGC-3′, 5′-CACTCCCAAACAGAAAGCACG-3′/5′-CCGAAGAAATCGGTGAGGC-3′, 5′-GGTCCCTATATGAACGTGCC-3′/5′-AGAAGGGACCCTCTTTACATAACC-3′, 5′-CCGCAGCACCATCGCTGTGC-3′/5′-GTGCTCCGACTCCGGGAGAGG-3′, 5′-AGACCCTCACGGAGGTCTCAC-3′/5′-CTGCTGTGCCGTGACCAG-3′, respectively. A primer pair specific for a 197-bp chicken BASP1 exon 2 segment (5′-AGGAGCAGCAACTGAGGAAGAG-3′/5′-GTTCTGCTTTCTGGGCTTCTTC-3′), reactions with normal rabbit serum (NRS), or with 1.6% of total chromatin (input) were used as controls. The antisera α-Myc, α-Max and α-Gag have been described33. For ChIP - real-time quantitative PCR (ChIP-qPCR), immunoprecipitated DNAs were analyzed on a Step One Real Time PCR System (Applied Biosystems, Carlsbad, CA) using the primer pair 5′-GGTAACACAGTACAGCCCTCGC-3′/5′-GCGAAACTCGCCGCGCTTGG-3′, amplifying a 167-bp segment from the quail CDT1 promoter adjacent to the transcription start site. Signals were normalized to input chromatin and shown as % input. The raw cycle threshold (Ct) values of the 16.6% input (dilution factor: 6.02) was adjusted to 100% by calculating raw Ct – log26.02. To calculate the % input of the immunoprecipitations, the equation 100 × 2[Ct (adjusted input to 100%) – Ct (IP)] was applied.
Transactivation analysis
The reporter construct pGL3-WS5 (pLUC-WS5) containing the quail WS5 promoter has been described15. To construct pGL3-CDT1, a 1246-bp fragment encompassing nucleotides −1204 to +42 from the quail CDT1 promoter region was first inserted into the BglII and HindIII sites of the pGL3-Basic vector (Promega, Madison, WI) yielding pGL3-CDT1-L. Deletion of a 996-bp SmaI/PstI fragment from this construct yielded pGL3-CDT1 encompassing nucleotides −217 to +42 from the quail CDT1 promoter. To create derivatives of pGL3-CDT1 in which the proximal, the distal, or both canonical E-boxes (5′-CACGTG-3′) were either mutated to 5′-GCCGTG-3′ or deleted, site-directed mutagenesis or overlapping PCR were performed as described15,33. The expression vector pRc-Myc, DNA transfer into cells using the calcium phosphate method and transcriptional transactivation analysis using the luciferase reporter system have been described15.
Electrophoretic mobility shift assay (EMSA)
Protein-DNA binding reactions were performed as described9. Protein-DNA complexes were resolved by native 6% (wt/vol) polyacrylamide gel electrophoresis (PAGE) and radioactive signals were quantified using a PhosphorImager (GE Healthcare, Little Chalfond, UK) as described9. A recombinant Myc/Max protein complex was expressed in Escherichia coli and purified as described36.
Purification of recombinant quail Cdt1(200–441) protein and antiserum preparation
A 726-bp cDNA fragment encoding amino acid residues 200–441 of quail Cdt1 flanked by start/stop codons and cloning sites provided by the PCR primers was amplified and cloned into the NdeI and EcoRI restriction sites of the prokaryotic expression vector pET-21a (Novagen, Darmstadt, Germany), yielding the pET-qCDT1(200–441) expression construct. The plasmid was transformed into Escherichia coli strain BL21 (DE3) CodonPlus-RIL (Stratagene, Santa Clara, CA). Induction of protein expression, cell lysis and protein purification were essentially done as described9. Recombinant Cdt1(200–441) protein was used for polyclonal antiserum preparation as described15.
Mass spectrometry (MS) of Cdt1 recombinant protein
The recombinant Cdt1(200–441) protein was desalted for MS as described9. The final protein concentration of the electrospray ionization (ESI) solution (H2O:MeOH 1:1, 1% vol/vol acetic acid) was 1 μM. ESI MS using a 7 Tesla Fourier transform ion cyclotron resonance (FT-ICR) instrument (Bruker Austria) for top-down experiments using collisionally activated dissociation (CAD) was performed as described9.
Protein analysis
In vitro translation, immunoprecipitation of L-[35S]methionine-labeled proteins, SDS-PAGE and immunoblotting were done as described using antisera directed against Myc and WS515,33, Cdt1 (this report) and tubulin α (Sigma-Aldrich, St. Louis, MO). To construct pET-CDT1 used for in vitro translation of full-length Cdt1 protein, a 1767-bp cDNA fragment encompassing the CDT1 coding region was inserted into pET-21a.
References
Duesberg, P. H., Bister, K. & Vogt, P. K. The RNA of avian acute leukemia virus MC29. Proc. Natl. Acad. Sci. USA 74, 4320–4324 (1977).
Bister, K. & Jansen, H. W. Oncogenes in retroviruses and cells: biochemistry and molecular genetics. Adv. Cancer Res. 47, 99–188 (1986).
Vogt, P. K. Retroviral oncogenes: a historical primer. Nat. Rev. Cancer 12, 639–648 (2012).
Bister, K., Hayman, M. J. & Vogt, P. K. Defectiveness of avian myelocytomatosis virus MC29: Isolation of long-term nonproducer cultures and analysis of virus-specific polypeptide synthesis. Virology 82, 431–448 (1977).
Eilers, M. & Eisenman, R. N. Myc's broad reach. Genes Dev. 22, 2755–2766 (2008).
Dang, C. V. MYC on the path to cancer. Cell 149, 22–35 (2012).
Blackwood, E. M. & Eisenman, R. N. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science 251, 1211–1217 (1991).
Eisenman, R. N. Deconstructing Myc. Genes Dev. 15, 2023–2030 (2001).
Hartl, M. et al. Stem cell-specific activation of an ancestral myc protooncogene with conserved basic functions in the early metazoan Hydra. Proc. Natl. Acad. Sci. USA 107, 4051–4056 (2010).
Young, S. L. et al. Premetazoan ancestry of the Myc-Max network. Mol. Biol. Evol. 28, 2961–2971 (2011).
Dang, C. V. c-Myc target genes involved in cell growth, apoptosis and metabolism. Mol. Cell. Biol. 19, 1–11 (1999).
Li, Z. et al. A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells. Proc. Natl. Acad. Sci. USA 100, 8164–8169 (2003).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl. Acad. Sci. USA 103, 17834–17839 (2006).
Shim, H. et al. c-Myc transactivation of LDH-A: Implications for tumor metabolism and growth. Proc. Natl. Acad. Sci. USA 94, 6658–6663 (1997).
Hartl, M., Nist, A., Khan, M. I., Valovka, T. & Bister, K. Inhibition of Myc-induced cell transformation by brain acid-soluble protein 1 (BASP1). Proc. Natl. Acad. Sci. USA 106, 5604–5609 (2009).
Dalla-Favera, R. et al. Human c-myconc gene is located on the region of chromosome 8 that is translocated in Burkitt lymphoma cells. Proc. Natl. Acad. Sci. USA 79, 7824–7827 (1982).
Nesbit, C. E., Tersak, J. M. & Prochownik, E. V. MYC oncogenes and human neoplastic disease. Oncogene 18, 3004–3016 (1999).
Dominguez-Sola, D. et al. Non-transcriptional control of DNA replication by c-Myc. Nature 448, 445–451 (2007).
Felsher, D. W. & Bishop, J. M. Transient excess of MYC activity can elicit genomic instability and tumorigenesis. Proc. Natl. Acad. Sci. USA 96, 3940–3944 (1999).
Yin, X. Y., Grove, L., Datta, N. S., Long, M. W. & Prochownik, E. V. c-myc overexpression and p53 loss cooperate to promote genomic instability. Oncogene 18, 1177–1184 (1999).
Li, Q. & Dang, C. V. c-Myc overexpression uncouples DNA replication from mitosis. Mol. Cell. Biol. 19, 5339–5351 (1999).
Lengauer, C., Kinzler, K. W. & Vogelstein, B. Genetic instabilities in human cancer. Nature 396, 643–649 (1998).
Prochownik, E. V. & Li, Y. The ever expanding role for c-Myc in promoting genomic instability. Cell Cycle 6, 1024–1029 (2007).
Herold, S., Herkert, B. & Eilers, M. Facilitating replication under stress: an oncogenic function of MYC? Nat. Rev. Cancer 9, 441–444 (2009).
Dutta, A. & Bell, S. P. Initiation of DNA replication in eukaryotic cells. Annu. Rev. Cell. Dev. Biol. 13, 293–332 (1997).
Nishitani, H., Lygerou, Z., Nishimoto, T. & Nurse, P. The Cdt1 protein is required to license DNA for replication in fission yeast. Nature 404, 625–628 (2000).
Hofmann, J. F. X. & Beach, D. cdt1 is an essential target of the Cdc10/Sct1 transcription factor: requirement for DNA replication and inhibition of mitosis. EMBO J. 13, 425–434 (1994).
Maiorano, D., Moreau, J. & Méchali, M. XCDT1 is required for the assembly of pre-replicative complexes in Xenopuslaevis. Nature 404, 622–625 (2000).
Whittaker, A. J., Royzman, I. & Orr-Weaver, T. L. Drosophila Double parked: a conserved, essential replication protein that colocalizes with the origin recognition complex and links DNA replication with mitosis and the down-regulation of S phase transcripts. Genes Dev. 14, 1765–1776 (2000).
Wohlschlegel, J. A. et al. Inhibition of eukaryotic DNA replication by geminin binding to Cdt1. Science 290, 2309–2312 (2000).
Tada, S., Li, A., Maiorano, D., Méchali, M. & Blow, J. J. Repression of origin assembly in metaphase depends on inhibition of RLF-B/Cdt1 by geminin. Nat. Cell Biol. 3, 107–113 (2001).
Oberst, C., Hartl, M., Weiskirchen, R. & Bister, K. Conditional cell transformation by doxycycline-controlled expression of the MC29 v-myc allele. Virology 253, 193–207 (1999).
Reiter, F., Hartl, M., Karagiannidis, A. I. & Bister, K. WS5, a direct target of oncogenic transcription factor Myc, is related to human melanoma glycoprotein genes and has oncogenic potential. Oncogene 26, 1769–1779 (2007).
Lee, C. et al. Structural basis for inhibition of the replication licensing factor Cdt1 by geminin. Nature 430, 913–917 (2004).
De Marco, V. et al. Quaternary structure of the human Cdt1-geminin complex regulates DNA replication licensing. Proc. Natl. Acad. Sci. USA 106, 19807–19812 (2009).
Fieber, W. et al. Structure, function and dynamics of the dimerization and DNA binding domain of oncogenic transcription factor v-Myc. J. Mol. Biol. 307, 1395–1410 (2001).
Menssen, A. & Hermeking, H. Characterization of the c-MYC-regulated transcriptome by SAGE: identification and analysis of c-MYC target genes. Proc. Natl. Acad. Sci. USA 99, 6274–6279 (2002).
Orian, A. et al. Genomic binding by the Drosophila Myc, Max, Mad/Mnt transcription factor network. Genes Dev. 17, 1101–1114 (2003).
Ji, H. et al. Cell-type independent MYC target genes reveal a primordial signature involved in biomass accumulation. PloS ONE 6, e26057 (2011).
Seitz, V. et al. Deep sequencing of MYC DNA-binding sites in Burkitt lymphoma. PloS ONE 6, e26837 (2011).
Perna, D. et al. Genome-wide mapping of Myc binding and gene regulation in serum-stimulated fibroblasts. Oncogene 31, 1695–1709 (2012).
Dang, C. V., Kim, J. W., Ping, G. & Yustein, J. The interplay between MYC and HIF in cancer. Nat. Rev. Cancer 8, 51–56 (2008).
Gao, P. et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458, 762–765 (2009).
Hermeking, H. et al. Identification of CDK4 as a target of c-MYC. Proc. Natl. Acad. Sci. USA 97, 2229–2234 (2000).
Grandori, C., Cowley, S. M., James, L. P. & Eisenman, R. N. The MYC/MAX/MAD network and the transcriptional control of cell behavior. Annu. Rev. Cell. Dev. Biol. 16, 653–699 (2000).
Pelengaris, S., Khan, M. & Evan, G. c-Myc: more than just a matter of life and death. Nat. Rev. Cancer 2, 764–776 (2002).
Blow, J. J. & Dutta, A. Preventing re-replication of chromosomal DNA. Nat. Rev. Mol. Cell. Biol. 6, 476–486 (2005).
Arias, E. E. & Walter, J. C. Replication-dependent destruction of Cdt1 limits DNA replication to a single round per cell cycle in Xenopus egg extracts. Genes Dev. 19, 114–126 (2005).
Ballabeni, A., Zamponi, R., Moore, J. K., Helin, K. & Kirschner, M. W. Geminin deploys multiple mechanisms to regulate Cdt1 before cell division thus ensuring the proper execution of DNA replication. Proc. Natl. Acad. Sci. USA 110, E2848–E2853 (2013).
Yoshida, K. & Inoue, I. Regulation of geminin and Cdt1 expression by E2F transcription factors. Oncogene 23, 3802–3812 (2004).
Watson, J. D., Oster, S. K., Shago, M., Khosravi, F. & Penn, L. Z. Identifying genes regulated in a Myc-dependent manner. J. Biol. Chem. 277, 36921–36930 (2002).
Sears, R., Ohtani, K. & Nevins, J. R. Identification of positively and negatively acting elements regulating expression of the E2F2 gene in response to cell growth signals. Mol. Cell. Biol. 17, 5227–5235 (1997).
Srinivasan, S. V., Dominguez-Sola, D., Wang, L. C., Hyrien, O. & Gautier, J. Cdc45 is a critical effector of Myc-dependent DNA replication stress. Cell Rep. 3, 1629–1639 (2013).
Seo, J. et al. Cdt1 transgenic mice develop lymphoblastic lymphoma in the absence of p53. Oncogene 24, 8176–8186 (2005).
Bader, A. G., Schneider, M. L., Bister, K. & Hartl, M. TOJ3, a target of the v-Jun transcription factor, encodes a protein with transforming activity related to human microspherule protein 1 (MCRS1). Oncogene 20, 7524–7535 (2001).
Acknowledgements
We thank Andrea Schraffl and Kane Puglisi for technical assistance and Sonja Spatzenegger for help with RDA screening. This work was supported by Austrian Science Fund grant P23652 (Klaus Bister).
Author information
Authors and Affiliations
Contributions
M.H. and Klaus Bister designed research, T.V., M.H., M.S., P.R., Kathrin Breuker and T.D.-M. performed experimental work, T.V., M.H., M.S., P.R., Kathrin Breuker, T.D.-M. and Klaus Bister analyzed data, T.V., M.H. and Klaus Bister wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Valovka, T., Schönfeld, M., Raffeiner, P. et al. Transcriptional control of DNA replication licensing by Myc. Sci Rep 3, 3444 (2013). https://doi.org/10.1038/srep03444
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03444
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.