Abstract
It is widely accepted that leaf dark respiration is a determining factor for the growth and maintenance of plant tissues and the carbon cycle. However, the underlying effect and mechanism of elevated CO2 concentrations ([CO2]) on dark respiration remain unclear. In this study, tomato plants grown at elevated [CO2] showed consistently higher leaf dark respiratory rate, as compared with ambient control plants. The increased respiratory capacity was driven by a greater abundance of proteins, carbohydrates and transcripts involved in pathways of glycolysis carbohydrate metabolism, the tricarboxylic acid cycle and mitochondrial electron transport energy metabolism. This study provides substantial evidence in support of the concept that leaf dark respiration is increased by elevated [CO2] in tomato plants and suggests that the increased availability of carbohydrates and the increased energy status are involved in the increased rate of dark respiration in response to elevated [CO2].
Similar content being viewed by others
Introduction
The global atmospheric concentration of carbon dioxide ([CO2]) has increased from 280 μmol mol−1 during the pre-industrial period to 388.5 μmol mol−1 in 2010 (Dr. Pieter Tans, NOAA/ESRL, www.esrl.noaa.gov/gmd/ccgg/trends/) and is projected to increase to 700 μmol mol−1 by the end of the 21st century1. In addition, concentrations of the greenhouse gas CO2 in the atmosphere are becoming increasingly elevated, as commercial growers throughout the world apply supplemental CO2 to enhance agricultural yields and crop quality. Many studies have sought to understand the response of plants to elevated [CO2]. Although an increased rate of photosynthesis and a partial closure of stomata in response to elevated [CO2] have been widely observed2,3, there is an apparent lack of consistency with regard to the described effects of plant growth in response to elevated [CO2] on leaf mitochondrial respiration (also referred to as dark respiration, as opposed to photorespiration). For example, studies have shown that dark respiration rates are either diminished, enhanced, or unaffected in plants grown under elevated [CO2]2,4,5,6,7. Because dark respiration is a major determinant in carbon cycles at all levels, including individual cells, whole plants, ecosystems and the global carbon cycle and can return as much as 40–50% of photosynthetically fixed carbon to the atmosphere8, it is critical that we improve our understanding of respiratory responses to rising atmospheric [CO2] and elucidate the mechanisms underlying these responses.
A growing body of evidence suggests that mitochondrial respiration plays an important role in the maintenance of optimal rates of photosynthesis9. Rotenone, the mitochondrial complex I inhibitor, decreased photosynthesis and the cytosolic ATP/ADP ratio in barley protoplasts10. The fact that both energy demand and substrate supply can leverage respiration rates is an indication that plants optimize the allocation of carbon and nutrients to maximize photosynthesis and growth11,12. Increases in photosynthesis and growth under elevated [CO2], therefore, should result in higher respiration rates. However, elevated [CO2] has been typically reported to cause an instantaneous reduction in leaf dark respiration, as measured based on CO2 efflux4,13,14. This decrease is difficult to explain due to the absence of a metabolic step with adequate sensitivity and control. Previous analyses have shown that the measurement of respiratory CO2 efflux is prone to several experimental artifacts7,15. First, respiratory gas exchange fluxes are low and it is difficult to accurately measure small variations in such fluxes. Second, the gas being measured is in a state of alteration, as the concentration gradient of [CO2] between the cuvette enclosing the leaf and the surrounding air is altered when [CO2] is changed. Third, CO2 is easily absorbed, adsorbed and leaked both via chamber seals and via the intercellular air spaces of leaves16,17. Fourth, leaf CO2 efflux is usually measured only at growth CO2 levels18,19, which will result in an over- or under-estimation of the specific respiration; this factor is critical but has not been accounted for in previous reports. In the present study, these artifacts are avoided by using CO2 efflux reciprocal measurements at both ambient and elevated [CO2] with an open gas exchange system and respiratory O2 uptake in normal air with an oxygen electrode. The O2 uptake measurement using the oxygen electrode method has been demonstrated to lack sensitivity with regard to dark respiration to changes in [CO2] from 0–2000 μmol mol−1 for a wide range of species7.
We hypothesized that under elevated CO2 conditions, the availability of photosynthates and energy demand in response to maintenance and growth costs involved in the regulation of respiration at the tissue-cellular level would increase. In this study, the changes in the two aforementioned factors were examined using commercial greenhouse tomato plants (Solanum lycopersicum L.), which were maintained at either 380 μmol mol−1 or 800 μmol mol−1, corresponding to the “ambient CO2” and “elevated CO2” conditions, respectively. The dark respiratory rate was assayed using CO2 efflux reciprocal measurement at both ambient and elevated [CO2] on whole leaves as well as O2 uptake measurements on leaf segments. Transcriptomic, proteomic and biochemical characterizations were conducted to extend the knowledge of the effects of elevated [CO2] on the gene and protein expression profiles of the respiratory components. Our study will therefore help to elucidate the mechanisms underlying the effects of higher atmospheric [CO2] on plant respiration as well as carbon management in the whole plant.
Results
Elevated [CO2] treatment increased both the CO2 efflux of intact leaves and O2 uptake of leaf segments
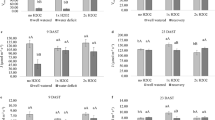
To determine the short- and long-term effects of elevated [CO2] on CO2 efflux, reciprocal measurements at ambient and elevated CO2 concentrations were performed from 0.5 to 18 days after treatment. The results showed that both the treatment and measurement [CO2] affected the daytime dark leaf respiration in the light (RL) and dark (RD) and nighttime dark leaf respiration (RN) (Fig. 1). From the beginning to the end of the experiment, RL, RD and RN were significantly higher when measured at 380 μmol mol−1 CO2 than when measured under the elevated CO2 level of 800 μmol mol−1. Plant growth under elevated [CO2] generally increased the CO2 efflux rate and prior to 12 d, both the RL and RD were significantly higher for the elevated [CO2]-treated plants than for their ambient counterparts when measured at the same CO2 concentration. Thereafter, RL and RD were reduced but still higher in the elevated [CO2]-treated plants; however, no significant quantitative changes between these parameters were observed. By contrast, the CO2 concentration had no consistent effect on the leaf RN under the 380 μmol mol−1 measurement CO2 condition; i.e., although the leaf RN was higher at elevated than at ambient [CO2] before 9 d, it was significantly lower from 12 d to the end of the experiment. Under the 800 μmol mol−1 measurement CO2 condition, the leaf RN was only slightly higher at the early stage of CO2 elevation.
Changes in the rate of CO2 efflux of tomato plants grown at ambient (380 μmol mol−1) or elevated CO2 concentrations (800 μmol mol−1).
The RL (a), RD (b) and RN (c) CO2 efflux rates were measured at different reciprocal CO2 concentrations.  , ambient CO2-treated plants measured under 380 μmol mol−1 CO2;
, ambient CO2-treated plants measured under 380 μmol mol−1 CO2;  , elevated CO2-treated plants measured under 380 μmol mol−1 CO2;
, elevated CO2-treated plants measured under 380 μmol mol−1 CO2;  , ambient CO2-treated plants measured under 800 μmol mol−1 CO2;
, ambient CO2-treated plants measured under 800 μmol mol−1 CO2;  , elevated CO2-treated plants measured under 800 μmol mol−1 CO2.Values are means ± SD (n = 6).
, elevated CO2-treated plants measured under 800 μmol mol−1 CO2.Values are means ± SD (n = 6).
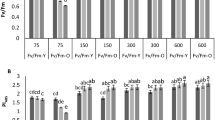
The effects of elevated [CO2] on leaf respiration were further explored by determining the O2 uptake of leaf segments (Fig. 2). We observed that the total, CN-resistant and SHAM-resistant respiration of elevated [CO2]-treated plants were higher than those of ambient-CO2 plants throughout the experimental period. The difference between the ambient- and elevated [CO2] plants was greater for O2 uptake rates than for CO2 efflux rates measured under the same CO2 concentration.
Changes in the O2 uptake rate and cytochrome c oxidase activity of tomato plants grown at ambient (380 μmol mol−1) or elevated CO2 concentrations (800 μmol mol−1).
(a), total respiration; (b), CN-resistant respiration; (c), SHAM-resistant respiration; d, cytochrome c oxidase activity. Values are means ± SD (n = 6).
Similar to the changes in the total and SHAM-resistant respiration, the elevated [CO2] treatment also induced a significant increase in the enzyme activity of the cytochrome pathway. When all measurements were averaged, the COX activity of tomato plants under elevated [CO2] was approximately 1.6-fold higher than that under the ambient CO2 condition (Fig. 2d).
Elevated [CO2] increased carbohydrates accumulation in tomato plants
The leaf carbohydrates determinations showed that the starch, total soluble sugar and sucrose concentrations increased significantly in plants exposed to 800 μmol mol−1 CO2: the concentrations of the three carbohydrates were increased by 90%, 60% and 44%, respectively, at the final harvest (Fig. 3a–c).
Changes in the carbohydrates concentrations and the effects of sucrose addition on O2 uptake rates of tomato plants grown at ambient (380 μmol mol−1) or elevated CO2 concentrations (800 μmol mol−1).
(a–c), Concentrations of starch (a), total soluble sugar (b) and sucrose (c); d, Effects of the addition of sucrose on the rate of O2 uptake. Values are means ± SD (n = 6).
To assess whether the substrate availability affected the respiratory rate under elevated [CO2], the leaf respiration rate was compared in the presence or absence of sucrose as a substrate. As shown in Fig. 3d, sucrose had no clear effects on the respiration of either ambient or elevated [CO2] plants, as the addition of sucrose to the leaf tissue did not significantly increase the respiration rate.
Effects of elevated [CO2] on cellular energy status
The energy status often has an effect on plant respiration. Thus, we investigated whether adenylate concentrations were different between ambient and elevated [CO2] plants. The ATP concentration in plants under elevated [CO2] decreased sharply by the sixth day of treatment (Fig. 4). Thereafter, the ATP concentration increased gradually to the initial level until the end of the experiment. By contrast, [CO2] had no effects on the ADP content at the early stage, although from 6 d to 15 d, the ADP content was slightly elevated under elevated [CO2]. These dissimilar changes in ATP and ADP concentrations resulted in a significantly lower ATP/ADP ratio for elevated [CO2] plants.
Effects of elevated [CO2] on the transcripts and proteomes of mitochondrial respiratory components
The transcriptomic response of fully expanded leaves to elevated [CO2] was analyzed using Affymetrix tomato cDNA microarrays (Fig. 5 and Supplementary Table S1 online). The abundance of 10,209 RNA transcripts encoding metabolically active, regulatory and structural proteins in the tomato was determined. An analysis of variance revealed 1,016 genes with different expression levels (P < 0.05) in elevated [CO2] plants compared to their ambient [CO2] counterparts. We were specifically interested in how elevated [CO2] alters genes related to leaf dark respiration, as these genes represent potential control points of the respiratory rate. We identified a total of 48 transcripts encoding many components of glycolysis, tricarboxylic acid (TCA) cycle and mitochondrial electron transport chain (miETC) genes in response to CO2 enrichment (P < 0.05). Among these transcripts, five were down-regulated and 43 were up-regulated. These up-regulated gene groups included major genes encoding glycolytic fructose-bisphosphate aldolase, glyceraldehyde-3-phosphate dehydrogenase, phosphoglycerate kinase and the TCA cycle components pyruvate dehydrogenase complex E1 alpha subunit, aconitate hydratase 2 and the succinate dehydrogenase complex. Downstream from the TCA cycle, there were increased transcript levels of the NADH dehydrogenase complex, the first committed step in miETC and most of the enzymes in the ATP-coupling, SHAM-resistant cytochrome pathway and CN-resistant respiratory pathway. Elevated [CO2] also led to highly increased transcript levels of COX and alternative oxidase (AOX), which correspond to the terminal oxidases of the cytochrome pathway and CN-resistant pathway, respectively. There was also a marked increase in transcripts of ATP synthase and uncoupling protein (UCP) under elevated [CO2].
Graphical representation of the transcript changes of the respiration metabolism in response to the elevated CO2 concentration (800 μmol mol−1) compared with the ambient CO2 concentration (380 μmol mol−1).
Tomato leaves used for microarray analysis were harvested at 9:00–11:00 am after six days of treatment. Arrows represent metabolic steps. Each colored box represents the statistically significant treatment response (P < 0.05) of a unique transcript encoding an enzyme or protein structure (details on the transcriptional responses are provided in Supplementary Table S1 online). Three independent biological replicates of each treatment were analyzed.
The effect of elevated [CO2] on the leaf protein pattern in tomato plants was also studied using 2-DE and a representative gel image stained by CBB is presented in Fig. 6. A total of 176 spots were detected when we set the threshold as a difference of 5–10-fold (P < 0.05) in expression under ambient versus elevated CO2 conditions. Among these protein spots, 90 exhibited up-regulation and the remaining 86 showed down-regulation under elevated CO2 conditions. We then randomly selected 15 up-regulated and 15 down-regulated spots for MALDI-TOF MS identification. Interestingly, six of these 30 identified proteins were involved in the dark respiration pathway (Table 1). Among these proteins, four (spots 3–6) are glycolysis-related components, one (spot 2) is a TCA cycle component and one (spot 1) is a miETC component. Interestingly, all of these six protein encoding genes were observed to be greatly up-regulated from cDNA microarray data (Supplementary Table S1 online).
Representative Coomassie brilliant blue-stained 2-DE gel of proteins extracted from tomato leaves grown under ambient and elevated CO2 conditions for six days.
Proteins were separated in the first dimension on an IPG strip (pH 4–7) and in the second dimension on a 12% polyacrylamide gel (for details, see “Materials and Methods”). The spots numbered on the gel (1–6) exhibited 5–10-fold differences (P < 0.05) between the two treatments. Three gels from biological repeats for each treatment were used.
Effects of elevated [CO2] on mitochondrial counts
As described above, the mitochondrial respiration and the protein expression profiles among the respiratory components dramatically changed during CO2 enrichment. We further examined the changes in the number of active mitochondria using Rhodamine 123, a fluorescent probe that selectively accumulates in active mitochondria based on the membrane potential20. After staining with Rhodamine 123, active mitochondria usually exhibit a bright green color. As shown in Fig. 7, plant growth at elevated [CO2] resulted in a marked increase in the number of functional mitochondria per cell when compared to the ambient [CO2]-grown plants.
Representative confocal microscopy images of active mitochondria of tomato leaves grown at either ambient (380 μmol mol−1) or elevated (800 μmol mol−1) CO2 concentrations for six days.
Active mitochondria exhibit bright green fluorescence, one representative active mitochondrion is indicated by an arrow in each image.
Discussion
Despite the common acceptance that leaf dark respiration is a determining factor for the growth and maintenance of plant tissues and the carbon cycle, the underlying mechanism and effect of elevated CO2 conditions on dark respiration have not been fully elucidated. It is critical to address this issue to better understanding how plants implement an effective carbon budget under elevated [CO2]. In the present study, the careful measurement of respiration in combination with transcriptomic and proteomic characterization provided evidence that respiratory metabolism is up-regulated when the CO2 concentration is elevated and we argue that the availability of carbohydrates and energy demand are involved in the increase in dark respiration in response to elevated [CO2].
Previous papers have variously concluded that elevated [CO2] will cause plant respiration to increase by as much as 30%, decrease by as much as 60%, or not change2,4,5,15. Although much of the variability is dependent on the species, developmental stages or growth conditions21,22, systematic errors in gas exchange measurement systems have also been reported to account for the over- or under-estimation of the effects of CO2 on leaf dark respiration2,7. In the present study, after methodological errors were eliminated, reciprocal CO2 efflux measurements at ambient and elevated [CO2] indicated that both the treatment and measurement CO2 concentrations have significant effects on leaf respiration and the CO2 efflux rate was evidently higher when measured at 380 μmol mol−1 CO2 than when measured at an elevated [CO2] of 800 μmol mol−1 (Fig. 1). To the best of our knowledge, few previous studies have investigated the leaf CO2 efflux rate at reciprocal CO2 concentrations; measurement only at growth [CO2] conditions18,19 might result in the underestimation and even the inhibition, of leaf dark respiration under elevated CO2 conditions. In addition, although the inconsistent effect of elevated [CO2] on leaf RN was difficult to explain, this finding was in agreement with previous results for Xanthium strumarium18, which showed that leaf RN was from 32% higher to 29% lower at elevated than at ambient [CO2] depending on the time after CO2 treatment. Furthermore, the increase of RD and RL was in accordance with the mitochondrial O2 uptake data obtained using the oxygen electrode system, which was free of CO2 leaks (Figs. 1–2). Plant mitochondria utilize two electron transport pathways: the ATP-coupling SHAN-resistant cytochrome pathway with COX as the terminal oxidase and the CN-resistant respiratory pathway23. The O2 uptake data in the present study showed that both the SHAN-resistant cytochrome pathway and CN-resistant alternative pathway respiration of leaf sections were strongly increased and the increase in the cytochrome pathway was accompanied by a simultaneous elevation of COX activity (Fig. 2). However, previous studies have described direct inhibitory effects of CO2 on mitochondrial COX activity6,24; but the inhibitory effects on COX are not always translated to the tissue level, many of the reported direct effects of CO2 on tissue respiration are much larger than expected based solely on the inhibition of COX4,7,15. These discrepancies may be dependent upon a variety of factors related to the plant's environment, development, treatment duration, species and tissue-specific cues. Furthermore, the increases of leaf dark respiration demonstrated here and in other studies1,5 are supported by the increased abundance of proteins in the glycolysis, TCA cycle and miETC pathway (Fig. 6 and Table 1). In addition, we observed a greater abundance of transcripts for the entire respiratory pathway rather than only a few individual enzymes (Fig. 5 and Supplementary Table S1 online), which provides further evidence supporting the stimulation of respiration at elevated [CO2]. Previous studies conducted on the soybean revealed that exposure to elevated [CO2] increased the abundance of transcripts associated with dark respiration, even though the magnitude of increase of the responses was generally less than 100% when exposed to elevated [CO2], similar to the level observed in the present study5,25.
Although the leaf dark respiration increases in response to exposure to elevated CO2, the precise mechanisms controlling these responses remain largely unknown. Because respiration is important to providing intermediates and energy for biosynthesis, we propose that the increase of carbohydrates availability and increased energy demand are involved in the increased dark respiration in response to elevated [CO2]. There is a general consensus that photosynthesis is enhanced by elevated [CO2]2,26,27. Metabolite analyses showed that leaf carbohydrates status, in general, including starch, total soluble sugar and sucrose, all increased significantly with elevated [CO2] in tomato plants (Fig. 3). The addition of sucrose to leaf segments did not significantly influence the O2 uptake rates (Fig. 3d), further indicating that substrate availability was more than sufficient for leaf respiration under elevated [CO2]. Previous studies argued that the availability of substrates, primarily carbohydrates, determines the longer-term capacity for respiration5,28. Based on the combined transcript and protein profiles as well as the metabolite data (Fig. 3, 5 and 6; Table 1 and Supplementary Table S1 online), we conclude that the elevated [CO2]-induced carbohydrates accumulation from enhanced photosynthesis might be a response to higher substrate availability to increase the flux through glycolysis and the TCA cycle25,29,30. Recent studies31 also reported that the ‘ability’ to respire recently assimilated carbon may contribute to the prevention of carbohydrates build-up and, consequently, to the avoidance of photosynthetic acclimation.
Cellular energy status, expressed as the ratio of ATP to ADP, might also regulate respiration in elevated [CO2]-treated plants32. In the present study, the ATP concentration experienced a transient decrease for the first six days of treatment, whereas the ADP concentration slowly increased under the elevated CO2 condition, which resulted in a lower ATP/ADP ratio in elevated [CO2] plants (Fig. 4). A decrease in the ATP concentration was unexpected because the ATP-coupling cytochrome pathway components were evidently increased under elevated [CO2], including the cytochrome pathway respiration rate, cytochrome pathway-related gene transcripts (complexes III, IV and V) and protein abundance and activity (COX and the putative mitochondrial ATP synthase beta subunit) (Figs. 3, 5 and 6; Tables 1 and Supplementary Table S1 online). Another ATP synthesis process, photosynthetic phosphorylation in the chloroplasts, should also be increased, as reflected by the increased net photosynthesis rate29,30. Thus, the ATP consumption rate, i.e., the energy demand, must have exceeded its production, which resulted in the observed transient lower ATP concentration and lower ATP/ADP ratio in the elevated [CO2] plants. The cellular energy status further exerts control via the allosteric effects of ADP and ATP on respiratory pathway enzymes, such as phosphofructokinase, pyruvate kinase and pyruvate dehydrogenase activities33. It should be noted that the respiratory energy demand of tissues is generally attributed to protein turnover, carbohydrate translocation, sucrose synthesis, ion gradient maintenance and other factors34 and these processes have been reported to increase in response to CO2 elevation35. In agreement with these observations, elevated [CO2]-grown Xanthium strumarium plants also had a higher demand for energy and carbon skeletons than did ambient [CO2]-grown plants18.
In addition to the previously described cytochrome pathway respiration that couples electron transport to the generation of a proton motive force for the synthesis of ATP, there is another respiratory pathway in higher plants, i.e., an alternative pathway that bypasses two sites of H+ transport and has a lower energy efficiency36. This alternative pathway respiration is sensitive to environmental stress and is usually enhanced by the presence of pathogens, chilling, wounding, drought and nutrient stress37,38,39. However, we found that the alternative pathway and gene transcripts (AOX1a, AOX1b, AOX1c and AOX2) were significantly increased under elevated [CO2] (Figs. 3 and 5, Supplementary Table S1 online). Thus, the increases in the alternative pathway activity under elevated [CO2] in the present study and in other studies4 could be interpreted as a reduction in the efficiency of carbohydrate conversion to ATP rather than a stress response. Similarly, cDNA microarray analysis revealed a significant increase in UCP transcripts (Fig. 5 and Supplementary Table S1 online), which also represents a bypass of the cytochrome pathway to catalyze proton conductance across the mitochondrial membrane and dissipation of the mitochondrial proton gradient23. UCP may function in a cooperative and complementary manner with AOX to potentiate a high TCA cycle flux and maintain the carbon balance at the cellular level37,4041.
This higher energy requirement can be met by a more efficient cell metabolism and/or a larger number of energy-producing organelles18, which would be needed to generate more mitochondria per cell, as observed in the leaves of plants grown at elevated [CO2] (Fig. 7). It is generally recognized that the number of mitochondria in a plant cell is typically related to the metabolic activity of the tissue and that cells with higher energy demands have a higher number of mitochondria42,43. The leaves of many species contain greater numbers of mitochondria when grown under elevated [CO2] compared with those grown at ambient [CO2]13. However, other studies have observed that mitochondrial numbers decrease or are not coupled to respiratory responses to elevated [CO2]42,44. These discrepancies might be explained by the varied methodological approaches used in these studies4. In the present study, we used CLSM to assay only metabolically active mitochondria, which may be more coupled to respiration activity relative to the total number of mitochondria, as measured using transmission electron microscopy.
In summary, the data presented here provide substantial evidence that leaf dark respiration is increased by elevated [CO2] in tomato plants; the coordinated response among biochemical, transcriptomic and proteomic results suggests that the increase in carbohydrates availability and energy demand caused the observed changes in the O2 uptake rates. Our study will therefore help to elucidate the mechanisms underlying the effects of higher atmospheric CO2 on plant respiration and thus the carbon management of the whole plant.
Methods
Plant material and growth conditions
Tomato (Solanum lycopersicum L. cv. Hezuo 903) seeds were sown in a medium containing a mixture of soil and perlite (1:3, v:v) in plastic pots (diameter, 10.5 cm; depth, 17.5 cm; one plant per pot) in a controlled environment. The growth conditions were as follows: the photosynthetic photo flux density (PPFD) was 600 μmol m−2 s−1, the photoperiod was 14/10 (h) (day/night), the day/night air temperature was 26/22°C and the relative humidity was 75%. The seedlings (four- to five-leaf stage) were transferred in duplicate to controlled-environment cabinets (Conviron, Winnipeg, Canada), where the atmospheric [CO2] was maintained at either 380 μmol mol−1 or 800 μmol mol−1, corresponding to the “ambient CO2” and “elevated CO2” conditions, respectively. All plants were watered daily to prevent water stress and were fertilized with Hoagland's solution every three days. The leaf samples were collected at different time points as indicated for respiration and metabolite assays. Unless other-wise stated, the transcriptomic, proteomic and mitochondrial count measurements were carried out after six days of treatment.
Measurements of leaf respiration by CO2 efflux
Leaf respiration measured based on CO2 efflux was determined using an open gas exchange system (LI-6400; LI-COR Inc., Lincoln, NE, USA) on the second fully developed leaves. The leaf temperature, relative humidity and leaf-to-air vapor pressure deficit were set at 25°C, 85% and 1–1.3 kPa, respectively. Gas exchange was measured first at the CO2 concentration under which the plants were grown and then at the reciprocal CO2 concentration. Three types of dark respiration were assayed18. Briefly, dark leaf respiration in the light, RL, was estimated by plotting the net CO2 assimilation rate against eight low PPFD levels (100, 80, 60, 40, 30, 20, 10 and 0 μmol m−2 s−1) using the Kok method. Responses of net CO2 assimilation rate versus PPFD were measured on at least four plants, the leaves were initially allowed to equilibrate at 100 μmol m−2 s−1 PPFD and the PPFD was then reduced stepwise to 0 μmol m−2 s−1, allowing 5–6 min for equilibration at each PPFD level. The leaf RD was obtained by averaging the three CO2 efflux rates at zero PPFD for each plant. The leaf RN was measured at the end of the daily dark period from 06:00 to 07:00 h.
Measurement of leaf respiration by O2 uptake and cytochrome c oxidase (COX) activity analysis
The O2 uptake of leaf segments was measured using a Clark-type liquid-phase oxygen electrode (Oxygraph-lab, Hansatech, UK) with some modifications45. Briefly, the plants were kept in the dark for 30 min to avoid any light-enhanced photosynthesis; then, 0.1 g leaf samples were cut into pieces for the measurement of respiration at 25°C in 2 mL of air-saturated 20 mM potassium phosphate buffer (pH 6.8). After a constant rate of oxygen uptake was obtained, potassium cyanide (1 mM) or salicylhydroxamic acid (SHAM, 20 mM) was added for the estimation of cyanide (CN)- or SHAM-resistant respiration, respectively. Similarly, after a constant rate of O2 uptake was attained in the buffer without any reagents, the sucrose-induced leaf respiration was analyzed by adding 110 mM sucrose46.
COX activity (EC1.9.3.1) was measured at 550 nm (25°C) in the presence of 20 μM reduced cytochrome c in 10 mM phosphate buffer (pH 7.2)45.
Metabolites assay
Carbohydrates were extracted from 0.2 g of freeze-dried leaf samples in 50 mL 80% (v/v) ethanol using five extraction steps and carbohydrate concentrations were determined using anthrone47. The supernatant was analyzed for sucrose and soluble sugars and the residue was boiled for 3 h in 10 mL of 2% HCl (v/v) to hydrolyze starch as described by Zhou et al48.
Adenylate metabolites were extracted according to the method of Dordas et al49. with some modifications. Fresh leaf samples (0.2 g) were homogenized with 1.5 mL of 10% perchloric acid and then centrifuged at 14,000 × g for 10 min at 4°C. The supernatant was adjusted to a pH of 7.8 using 5 M KOH and 1 M triethanolamine. The neutralized extract was centrifuged as described above and the supernatant was used for metabolite determinations. ATP concentrations were measured using a luciferin-luciferase system with an ATP bioluminescent assay kit from Sigma. ADP was converted to ATP using pyruvate kinase. The overall luminescence was quantified using a multi-label plate reader (2300 EnSpire; Perkin-Elmer, Turku, Finland).
RNA preparation, microarray hybridization and data analysis
Total RNAs were isolated from the leaves of the tomato plants after six days of treatment under either ambient or elevated CO2 conditions. Three independent biological replicates of each treatment were analyzed. Probe labeling and chip hybridization were performed using a GeneChip® two-cycle cDNA synthesis kit. The tomato genome array (Affymetrix, CA, USA), which is designed specifically to monitor gene expression in the tomato, was used. All procedures for probe preparation, hybridization, washing, staining and scanning of the GeneChip® tomato arrays, as well as data collection, were performed by the Affymetrix custom service (CapitalBio, Beijing, China) according to standard protocols (http://www.affymetrix.com/support/technical/manual/expression-manual.affx). Normalization was performed according to the standard Affymetrix protocol to allow for the comparison of the samples from each set of experiments. The fold changes and P-values of the probe sets were calculated using the limma nested F-test and the P-values for multiple testing were corrected using the false-discovery rate.
Protein extraction and two-dimensional gel electrophoresis (2-DE)
One biological sample was obtained by pooling the leaves from five plants and a total of three biological repeats were analyzed. Briefly, proteins were extracted using a TCA-acetone precipitation step50. The extracted proteins were suspended in buffer containing 7 M urea, 2 M thiourea, 2% CHAPS, 13 mM DTT, 1.25% CA and 1.7% PMSF and were stored at room temperature for 1 h prior to 2-DE. The total soluble protein concentrations were determined according to the method of Bradford51.
Extract containing approximately 1 mg of total proteins was subsequently loaded onto an IPG strip holder with 17 cm linear gradient IPG strips (Bio-Rad, Hercules, CA, USA), pH 4–7 and rehydrated for 24 h at 17°C. Focusing was then performed using a PROTEAN IEF cell (Bio-Rad, Hercules, CA, USA). After the first-dimensional gel electrophoresis step, the gel was removed and equilibrated for 10 min using an equilibration solution (50 mM Tris, pH 8.8, 6 M urea, 30% glycerol, 2% SDS, 0.002% bromophenol blue) containing 1% DTT for 15 min and subsequently with equilibration solution containing 4% iodoacetamide for an additional 15 min. The separation of proteins in the second dimension was performed using a PROTEAN II xi cell (Bio-Rad, Hercules, CA, USA). The temperature of the gel was controlled at 15°C.
Preparative gels were stained with colloidal Coomassie brilliant blue (CBB) G-250. Image analysis was performed using PDQuest Software Version 7.1 according the user's manual. All selected spots were manually excised from the 2-DE gels. In-gel digestion and matrix-assisted laser desorption-ionization time-of-flight mass spectrometry (MALDI-TOF MS) fingerprint spectra acquisition were performed according to Fan et al52. The peptide mass fingerprints were searched against the NCBInr (http://www.ncbi.nlm.nih.gov/) protein databases using MASCOT Version 2.1. Protein identification was based on the combination of a MASCOT score higher than 60 (threshold), maximum peptide coverage and additional experimental pI and Mr of the protein spots on the gels. The protein analysis was repeated twice.
Measurement of mitochondrial counts
Mitochondrial count detection using confocal laser scanning microscopy (CLSM) was performed4. Briefly, leaf sections (0.5 by 0.5 cm) were placed into a loading buffer with 250 mM Rhodamine 123 (Sigma Aldrich) and incubated for 12 min at 37°C in the dark. Excess dye was eliminated by washing the discs three times in 30 mM MES buffer with 0.2 mM CaCl2 at room temperature followed by immediately examination using a CLSM system (Leica TCS SP5; Leica Microsystems, Wetzlar, Germany). The sections were excited with the 488-nm line of an argon laser and dye emissions were recorded using a 505- to 530-nm band-pass filter.
Statistical analysis
At least three independent replicates were conducted for each determination. Data were subjected to analysis of variance and the means were compared using Tukey's test at the 5% level.
References
Aranjuelo, I. et al. Does ear C sink strength contribute to overcoming photosynthetic acclimation of wheat plants exposed to elevated CO2? J. Exp. Bot. 62, 3957–3969 (2011).
Long, S. P., Ainsworth, E. A., Rogers, A. & Ort, D. R. Rising atmospheric carbon dioxide: Plants face the future. Annu. Rev. Plant Biol. 55, 591–628 (2004).
Gifford, R. M. The CO2 fertilising effect - does it occur in the real world? New Phytol. 163, 221–225 (2004).
Gomez-Casanovas, N., Blanc-Betes, E., Gonzàlez-Meler, M. A. & Azcon-Bieto, J. Changes in respiratory mitochondrial machinery and cytochrome and alternative pathway activities in response to energy demand underlie the acclimation of respiration to elevated CO2 in the invasive Opuntia ficus-indica. Plant Physiol. 145, 49–61 (2007).
Leakey, A. D. B. et al. Genomic basis for stimulated respiration by plants growing under elevated carbon dioxide. Proc. Natl. Acad. Sci. U.S.A. 106, 3597–3602 (2009).
Gonzàlez-Meler, M. A., Ribas-Carbó, M., Siedow, J. N. & Drake, B. G. Direct inhibition of plant mitochondrial respiration by elevated CO2 . Plant Physiol. 112, 1349–1355 (1996).
Davey, P. A. et al. Respiratory oxygen uptake is not decreased by an instantaneous elevation of [CO2], but is increased with long-term growth in the field at elevated [CO2]. Plant Physiol. 134, 520–527 (2004).
Amthor, J. S. Terrestrial higher-plant response to increasing atmospheric [CO2] in relation to the global carbon cycle. Global Change Biol. 1, 243–274 (1995).
Fuentes, D. et al. A deficiency in the flavoprotein of Arabidopsis mitochondrial complex II results in elevated photosynthesis and better growth in nitrogen-limiting conditions. Plant Physiol. 157, 1114–1127 (2011).
Igamberdiev, A. U., Hurry, V., Kromer, S. & Gardestrom, P. The role of mitochondrial electron transport during photosynthetic induction. A study with barley (Hordeum vulgare) protoplasts incubated with rotenone and oligomycin. Physiol. Plant. 104, 431–439 (1998).
Thornley, J. H. M. A model to describe the partitioning of photosynthate during vegetative plant growth. Ann. Bot. 36, 419–430 (1972).
Gonzàlez-Meler, M. A., Blanc-Betes, E., Flower, C. E., Ward, J. K. & Gomez-Casanovas, N. Plastic and adaptive responses of plant respiration to changes in atmospheric CO2 concentration. Physiol. Plant. 137, 473–484 (2009).
Griffin, K. L. et al. Plant growth in elevated CO2 alters mitochondrial number and chloroplast fine structure. Proc. Natl. Acad. Sci. U.S.A. 98, 2473–2478 (2001).
Drake, B. G. et al. Does elevated atmospheric CO2 concentration inhibit mitochondrial respiration in green plants? Plant Cell Environ. 22, 649–657 (1999).
Gonzàlez-Meler, M. A. & Siedow, J. N. Direct inhibition of mitochondrial respiratory enzymes by elevated CO2: does it matter at the tissue or whole-plant level? Tree Physiol. 19, 253–259 (1999).
Jahnke, S. & Krewitt, M. Atmospheric CO2 concentration may directly affect leaf respiration measurement in tobacco, but not respiration itself. Plant Cell Environ. 25, 641–651 (2002).
Amthor, J. S., Koch, G. W., Willms, J. R. & Layzell, D. B. Leaf O2 uptake in the dark is independent of coincident CO2 partial pressure. J. Exp. Bot. 52, 2235–2238 (2001).
Wang, X. Z., Lewis, J. D., Tissue, D. T., Seemann, J. R. & Griffin, K. L. Effects of elevated atmospheric CO2 concentration on leaf dark respiration of Xanthium strumarium in light and in darkness. Proc. Natl. Acad. Sci. U.S.A. 98, 2479–2484 (2001).
Beerling, D. J. & Osborne, C. P. Physiological ecology of mesozoic polar forests in a high CO2 environment. Ann. Bot. 89, 329–339 (2002).
Petit, P. X. Flow cytometric analysis of rhodamine-123 fluorescence during modulation of the membrane-potential in plant-mitochondria. Plant Physiol. 98, 279–286 (1992).
Wang, X. Z. & Curtis, P. A meta-analytical test of elevated CO2 effects on plant respiration. Plant Ecol. 161, 251–261 (2002).
Hymus, G. J., Snead, T. G., Johnson, D. P., Hungate, B. A. & Drake, B. G. Acclimation of photosynthesis and respiration to elevated atmospheric CO2 in two Scrub Oaks. Global Change Biol. 8, 317–328 (2002).
Millar, A. H., Whelan, J., Soole, K. L. & Day, D. A. Organization and regulation of mitochondrial respiration in plants. Annu. Rev. Plant Biol. 62, 79–104 (2011).
Bruhn, D., Wiskich, J. T. & Atkin, O. K. Contrasting responses by respiration to elevated CO2 in intact tissue and isolated mitochondria. Func. Plant Biol. 34, 112–117 (2007).
Ainsworth, E. A., Rogers, A., Vodkin, L. O., Walter, A. & Schurr, U. The effects of elevated CO2 concentration on soybean gene expression. An analysis of growing and mature leaves. Plant Physiol. 142, 135–147 (2006).
Xu, Z. J. et al. Effect of free-air atmospheric CO2 enrichment on dark respiration of rice plants (Oryza sativa L.). Agr. Ecosyst. Environ. 115, 105–112 (2006).
Tuba, Z. & Lichtenthaler, H. K. Long-term acclimation of plants to elevated CO2 and its interaction with stresses. Ann. N.Y. Acad. Sci. 1113, 135–146 (2007).
Williams, J. H. H. & Farrar, J. F. Control of barley root respiration. Physiol. Plant. 79, 259–266 (1990).
Jiang, Y. P. et al. Interactive effects of CO2 enrichment and brassinosteroid on CO2 assimilation and photosynthetic electron transport in Cucumis sativus. Environ. Exp. Bot. 75, 98–106 (2012).
Liu, J., Zhang, D., Zhou, G. & Duan, H. Changes in leaf nutrient traits and photosynthesis of four tree species: effects of elevated [CO2], N fertilization and canopy positions. J. Plant Ecol. 5, 376–390 (2012).
Aranjuelo, I. et al. Leaf carbon management in slow-growing plants exposed to elevated CO2 . Global Change Biol. 15, 97–109 (2009).
Klotz, K. L., Finger, F. L. & Anderson, M. D. Respiration in postharvest sugarbeet roots is not limited by respiratory capacity or adenylates. J. Plant Physiol. 165, 1500–1510 (2008).
Farrar, J. F. The respiratory source of CO2 . Plant Cell Environ. 8, 427–438 (1985).
Bouma, T. J. et al. Respiratory energy requirements and rate of protein turnover in vivo determined by the use of an inhibitor of protein synthesis and a probe to assess its effect. Physiol. Plant. 92, 585–594 (1994).
Noguchi, K. & Yoshida, K. Interaction between photosynthesis and respiration in illuminated leaves. Mitochondrion 8, 87–99 (2008).
Moore, A. L. & Siedow, J. N. The regulation and nature of the cyanide-resistant alternative oxidase of plant mitochondria. Biochim. Biophys. Acta 1059, 121–140 (1991).
Shi, K. et al. Flexible change and cooperation between mitochondrial electron transport and cytosolic glycolysis as the basis for chilling tolerance in tomato plants. Planta 237, 589–601 (2013).
Liao, Y. W. K. et al. The reduction of reactive oxygen species formation by mitochondrial alternative respiration in tomato basal defense against TMV infection. Planta 235, 225–238 (2012).
Yoshida, K. et al. Distinct responses of the mitochondrial respiratory chain to long- and short-term high-light environments in Arabidopsis thaliana. Plant Cell Environ. 34, 618–628 (2011).
Watanabe, C. K., Hachiya, T., Terashima, I. & Noguchi, K. The lack of alternative oxidase at low temperature leads to a disruption of the balance in carbon and nitrogen metabolism and to an up-regulation of antioxidant defence systems in Arabidopsis thaliana leaves. Plant Cell Environ. 31, 1190–1202 (2008).
Nunes-Nesi, A., Araujo, W. L. & Fernie, A. R. Targeting mitochondrial metabolism and machinery as a means to enhance photosynthesis. Plant Physiol. 155, 101–107 (2011).
Wang, X. Z., Anderson, O. R. & Griffin, K. L. Chloroplast numbers, mitochondrion numbers and carbon assimilation physiology of Nicotiana sylvestris as affected by CO2 concentration. Environ. Exp. Bot. 51, 21–31 (2004).
Moyes, C. D., Battersby, B. J. & Leary, S. C. Regulation of muscle mitochondrial design. J. Exp. Biol. 201, 299–307 (1998).
Gonzelez-Meller, M. A., Taneva, L. & Trueman, R. J. Plant respiration and elevated atmospheric CO2 concentration: cellular responses and global significance. Ann. Bot. 94, 647–656 (2004).
Millenaar, F. F., Gonzàlez-Meler, M. A., Siedow, J. N., Wagner, A. M. & Lambers, H. Role of sugars and organic acids in regulating the concentration and activity of the alternative oxidase in Poa annua roots. J. Exp. Bot. 53, 1081–1088 (2002).
Noguchi, K., Nakajima, N. & Terashima, I. Acclimation of leaf respiratory properties in Alocasia odora following reciprocal transfers of plants between high- and low-light environments. Plant Cell Environ. 24, 831–839 (2001).
Yemm, E. W. & Willis, A. J. The estimation of carbohydrates in plant extracts by anthrone. Biochem. J. 57, 508–514 (1954).
Zhou, Y. H., Yu, J. Q., Huang, L. F. & Nogues, S. The relationship between CO2 assimilation, photosynthetic electron transport and water-water cycle in chill-exposed cucumber leaves under low light and subsequent recovery. Plant Cell Environ. 27, 1503–1514 (2004).
Dordas, C. et al. Expression of a stress-induced hemoglobin affects NO levels produced by alfalfa root cultures under hypoxic stress. Plant J. 35, 763–770 (2003).
Wang, W. et al. Protein extraction for two-dimensional electrophoresis from olive leaf, a plant tissue containing high levels of interfering compounds. Electrophoresis 24, 2369–2375 (2003).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Fan, P. X., Wang, X. C., Kuang, T. Y. & Li, Y. X. An efficient method for the extraction of chloroplast proteins compatible for 2-DE and MS analysis. Electrophoresis 30, 3024–3033 (2009).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31071832, 31372108) and the National Key Technology R&D Program of China (2013AA102406, 2011BAD12B04).
Author information
Authors and Affiliations
Contributions
K.S. and J.Y. designed the research; X.L., G.Z., B.S., S.Z., Y.Z., Y.L., Y.Z. and X.X. performed the experiments; K.S. and X.L. wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Li, X., Zhang, G., Sun, B. et al. Stimulated Leaf Dark Respiration in Tomato in an Elevated Carbon Dioxide Atmosphere. Sci Rep 3, 3433 (2013). https://doi.org/10.1038/srep03433
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03433
This article is cited by
-
Roles of stomata in gramineous crops growth and biomass production
Cereal Research Communications (2022)
-
Temperature effect on water dynamics in tetramer phosphofructokinase matrix and the super-arrhenius respiration rate
Scientific Reports (2021)
-
Effects of elevated CO2 on resistant and susceptible rice cultivar and its primary host, brown planthopper (BPH), Nilaparvata lugens (Stål)
Scientific Reports (2021)
-
Improving photosynthesis to increase grain yield potential: an analysis of maize hybrids released in different years in China
Photosynthesis Research (2021)
-
Short photoperiod attenuates CO2 fertilization effect on shoot biomass in Arabidopsis thaliana
Physiology and Molecular Biology of Plants (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.