Abstract
A model previously developed for the wind-borne spread by midges of bluetongue virus in NW Europe in 2006 is here modified and applied to the spread of Schmallenberg virus in 2011. The model estimates that pregnant animals were infected 113 days before producing malformed young, the commonest symptom of reported infection and explains the spatial and temporal pattern of infection in 70% of the 3,487 affected farms, most of which were infected by midges arriving through downwind movement (62% of explained infections), or a mixture of downwind and random movements (38% of explained infections), during the period of day (1600–2100 h, i.e. dusk) when these insects are known to be most active. The main difference with Bluetongue is the higher rate of spread of SBV, which has important implications for disease control.
Similar content being viewed by others
Introduction
Schmallenberg virus (SBV) (Family Bunyaviridae, genus Orthobunyavirus)1 was detected for the very first time in Europe in 20112. The outbreak of SBV started in the same area (Figure 1) where bluetongue (serotype 8, BTV-8) had emerged five years' previously, in 20063, both diseases causing significant economic losses4,5. As with BTV-86, SBV affects mainly sheep and cattle and occasionally goats, buffaloes, bison, roe, fallow and red deer, elk and moose7,8,9, but not humans10. The clinical features of SBV are mild transient disease in adult cattle characterized by reduced milk production (up to 50%), inappetence, pyrexia and diarrhoea. More importantly, however, SBV can cause severe foetal malformations (of the musculoskeletal and central nervous systems) when pregnant cows, ewes and goats are infected in early to mid-pregnancy, resulting in increased rates of foetal abnormalities and dystocia11,12,13. Similar effects during gestation in sheep, cattle and goats were found in animals vaccinated with BTV-2 and BTV-9 modified live vaccine virus14.
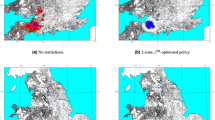
Spatial distribution of SBV from 2011 to 2012 (a single SBV infection in southern Spain is not shown) and of BTV-8 in 2006.
For comparison, the data shown are for the first 121 days of both outbreaks (the full extent of the BTV epizootic in 2006). The inset shows the details of the BTV-8 area. Map created with ArcGIS© software by ESRI.
SBV and BTV-8 are caused by different viruses15 but they are both transmitted by Culicoides midges16, the SBV competence of which has been confirmed in both field and laboratory17 studies that covered the Culicoides obsoletus complex (i.e. C. obsoletus s.s., C. dewulfi, C. scoticus and C. chiopterus18,19,20,21). The virus can also be transmitted transplacentally22, but horizontal (direct animal to animal) transmission is unknown.
Outbreaks of both BTV-8 and SBV began in late summer (July for BTV and August for SBV)3,9. SBV in 2011, however, spread much more rapidly than had Bluetongue in 2006 (Table 1), affecting much larger areas more quickly. From 2011 to date SBV's geographic distribution has steadily increased and now includes virtually all countries of Europe, with only a few exceptions (Lithuania, Portugal and Cyprus)9,22,23,24,25,26,27 (Figure 1 shows the publicly available OIE data).
Much work has been carried out on the possible role of wind in spreading BTV in Europe and elsewhere. The small body size (1–3 mm body length) of midges means that passive flight over land and water bodies for hundreds of kilometres28 is possible under certain conditions of temperature, precipitation, humidity and wind speed29. Using these climatic variables and appropriate entomological parameters, recent research29 on BTV found that infections in the Balearic Islands in 2000 were most likely caused by midges transported from Tunisia and/or Algeria; the UK was infected in 2007 by midges from Belgium; Denmark in 2007 from Germany; Sweden in 2008 from Denmark and/or Germany (as in30); and Norway in 2009 from Denmark.
Once established, the more local spread of BTV, at velocities of between 2 and 5 km/day31,32, also appears to depend upon wind. Of the four possible means of spread of BTV (movement of infected vectors33, or of infected domestic animal hosts34, or of wild ruminants35, or use of infected semen and embryos36) only the carriage of Culicoides by wind fields seems able to explain the direction and extent of the 2006 BTV outbreak in Europe and the importance of wind for BTV spread is now widely accepted29,37. A biologically-informed model (Spatio-temporal Wind Outbreak Trajectories Simulation, SWOTS)31 was applied to the 2006 outbreak in order to extract key variables and parameters associated with the spread of this disease. The results of the BTV-SWOTS analysis suggested that although the majority of farm-to-farm BTV infections occurred through midge movement which was essentially downwind (i.e. passive), or downwind with some random component, more than a third of infections occurred in the upwind direction (Table 2), strongly implying that midges respond to the smell of their hosts by flying upwind to find them. In the BTV outbreak, the modal distance covered in farm-to-farm infections was no more than 1 km31 and upwind infections were associated with shorter average distances between infectious and infected farms than downwind ones.
Given the above conclusions about the spread of BTV by wind-borne midges it seems reasonable to suggest that SBV is spread in the same way but, as far as we are aware, this suggestion has not been quantitatively investigated to date. Risk warnings of possible SBV incursion into the United Kingdom were made by DEFRA (Department of Environment, Food and Rural Affairs of United Kingdom) in 2011 (see for example one of their preliminary outbreak assessment: http://www.defra.gov.uk/animal-diseases/files/poa-schmallenburg-update-120105.pdf). A model for the temperature-dependent spread of SBV suggested that Scotland was at risk of SBV38, a prediction confirmed by reports there of SBV sero-positivity in permanently resident cattle in 201239 and both cattle (including foetal abnormalities) and sheep in 2013 (http://www.defra.gov.uk/ahvla-en/2013/07/01/sbv-updated-testing-results-scotland/).
Here we present a modified version of BTV-SWOTS31 to analyse, understand and simulate the spread of SBV in Europe in 2011–2012 (SBV-SWOTS). The results of BTV-SWOTS and SBV-SWOTS are compared in order to detect and explain similarities and differences between these two diseases, spread by the same insects.
Following previous practice31, we here use the term ‘vector’ only in its mathematical sense and the term ‘midges’ to describe both the insects and their role as carriers of both BTV and SBV. The term ‘midge vector’ therefore refers to a mathematical description of the distance and direction of midge flight.
Results
The largest correlation between infection and midge vectors is 0.62 (p = 0.0098), obtained for 113 days (DA) before birth of a malformed foetus. SBV-SWOTS analysis suggests that infections of farms occur after an average of 6 days of midge flight (DT) from a previously infected farm and that most midge flight occurs during the 1600–2059 h period of each day (TOD) (correlations are highest for this period of the day). The estimated incubation period (IIP + EIP) is 10 days (see Methods for the full definition of these terms). The correlations with individual types of midge movement were: 0.68 for downwind (p = 0.0076), 0.44 for random (p = 0.0036) and 0.17 for upwind (p = 0.0422) movements, indicating a much greater preponderance of downwind movement in SBV than in BTV. The estimated average direction of the midge vectors (of all the infections explained by SBV-SWOTS) was 252 degrees from North (where compass direction North is 0 degrees); i.e. in the mean direction of approximately West-South-West, a quadrant that also contains the average wind direction (Table 2).
Both BTV-SWOTS and SBV-SWOTS were implemented using a high threshold probability (a ratio > = 0.95 of the number of trajectories connecting a particular infectious farm A to a newly infected farm B over the total number of infectious trajectories reaching farm B in the same interval of time), indicating that any particular farm had been infected from one particular previously-infected farm. Farms that did not reach this threshold were considered as unlikely to have been infected by any putative ‘source farm’ within the dataset. This high threshold excluded very few farms in the BTV-SWOTS analysis (8%) strongly indicating that a network of (modelled and quantified) infection routes linked most farms in the dataset. This relatively high certainty of the infection network probably arose because of the relatively short distances (many less than 1 km) between the BTV-infected farms in the landscape. In contrast, the same high probability threshold excludes 30% of SBV-infected farms, probably because these farms were farther apart (the average distance between all infected farms in the dataset is 461 km with standard deviation = 255 km) and so less likely to fall within the plumes of infected midges arising from previously infected farms. Nevertheless, the SBV simulation model successfully connected 70% of all infected farms. This figure rises as the threshold probability is lowered from 0.95 but reaches a maximum of 89% only when the threshold is lowered to 0.13. In other words, even at this low threshold, SBV-SWOTS is still unable to explain 11% of all infections.
Two types of midge movement appeared to be mainly responsible for the spread of SBV: downwind for 62% of explained infections (43% of total infections) and mixed downwind and random for the remaining 38% (27% of total infections; Table 2 and Figure 2). In the latter, the random component accounted for 82% of the distance covered by the midges to arrive at and infect, uninfected farms. There were only weak signs of upwind movement contributing to the spread of SBV (Table 2).
Finally, 55% of the infected farms in the dataset were predicted by the model to be ‘dead ends’ (i.e. infecting no other farm), so that less than half of all farms were involved in infecting other farms.
Discussion
The geographical areas affected by SBV in 2011 overlap many of the areas affected by BTV-8 in 2006 (Figure 1). The cumulative proportions of infected farms over time from the start of each outbreak are similar (Table 1, columns 2 and 3. Kolmogorov-Smirnov test, p = 0,197; Ansari-Bradley test,p = 0.671)40, although BTV-8 case numbers were initially higher, possibly because BTV-8, unlike SBV, was a notifiable disease before the outbreak occurred. The SBV-infected farms were, however, spread over a very much larger area. The rate of increase in total case numbers in both outbreaks diminished after approximately 100 days, probably because of the onset of colder conditions approaching winter (Table 1). Only 10% of both BTV-8 and SBV first year outbreak cases were reported more than 100 days after each outbreak began, continuing at a low level all the way through winter and into the following Spring, indicating an over-wintering mechanism41,42 which is still not fully understood, although adult midges might survive autumn and winter in warm animal barns43,44 and cold-tolerant midges (adult C. obsoletus) have been found in central Italy all year round45. There was no significant change during the post 100-day period in either the average or maximum distance between infected farms.
SWOTS analysis suggests that BTV and SBV are similar in the estimated mean time taken by the midges to move infections from one farm to another (Table 2, DT = 6 and 7 days respectively), the time of day at which midges are flying (DT = 1600–2059 h in both cases, i.e. around about dusk46) and in the estimated combined incubation periods (IIP + EIP = 10 for SBV and IIP of 8 days for BTV). The intrinsic incubation period of SBV on a small sample of cattle was experimentally found to be between 2 and 6 days47 suggesting therefore that the EIP of SBV in midges is between 4 and 8 days.
Whilst the mean values of DT are similar in BTV-SWOTS and SBV-SWOTS (Table 2) the mean distances between infectious and infected farms is very different (Table 2) and are not determined by differences in mean wind speed (for both diseases, mean wind speeds were between 3 and 5 m/s). This conundrum is resolved by looking at the frequency distributions for each of these variables. The modal (as opposed to mean) value of DT for BTV is 1 day and the modal value of the distance between BTV-infected farms is 1 km; for SBV the comparable figures are 6 days and 10 km. In other words, the increase in the modal distance between infected farms was of the order of 1 to 2 km per day of DT in both cases. The proportional frequency distributions of DT for the two diseases are similar beyond values of about seven days; thus the same proportions of midges are flying for these longer periods of time, on average over longer distances, to infect new farms, but very different proportions of midges are causing infections at shorter DTs. We suggest that the main reason for this difference is the very different windows of susceptibility of livestock to the two diseases in question. BTV can cause clinical symptoms in susceptible hosts at more or less any time of year. Whenever a BTV- infected midge encounters a susceptible host, therefore, a resulting infection is possible. Midges are more likely to contact susceptible hosts at closer distances (nearby farms) than farther ones and so the pattern of BTV spread is determined by the proximity and density of farms. SBV, on the other hand, appears to cause the obvious clinical symptom of foetal malformation only if the infectious bite occurs within a restricted window during pregnancy (viraemia of SBV in vertebrate hosts lasts only 5–6 days in adult animals, reviewed by2, while in BT it is much longer48). The average 6 days travelled by the midges to reach another farm (slightly less than a third of the average lifespan of an adult, of about 20 days49) is longer than the feeding interval of midges, the implication being that the midges feed, but do not cause infections, en route, because the animals on which they feed are not susceptible (to foetal abnormalities). Hence whereas the spread of BTV was described as a series of stepping stones, where each stepping stone (a farm where infected midges could transmit infection) was eventually revealed by that farm actually showing infection, the spread of SBV could follow the same stepping stone pattern, but with many stones not revealed by subsequent foetal abnormalities, because fewer hosts were susceptible at the time they were bitten by infected midges. This explanation suggests that more SBV infection is taking place than is eventually revealed by foetal abnormalities and this appears to be the case in the field9. In an extensive survey carried out in Belgium at the beginning of 2012, it was found that all cattle older than 6 months had already been infected by SBV and were therefore probably immune50 to re-infection (for a presently unknown period of time). Similar results were found in sheep51.
This SBV ‘hidden stepping stone’ pattern could explain the lack of upwind movement in SBV infections (Table 2). Upwind flight in response to host odours is only likely to operate over short distances for physically small creatures such as midges52,53; the between-farm distances involved in SBV infections are probably too great for upwind flight to have been important. Nevertheless, in both BTV-8 and SBV the overall direction of midge movement was within the same quadrant as the mean prevailing wind direction, with differences between them of 78 degrees for BTV-8 and 57 degrees for SBV (Table 2). Nuances of midge behaviour that are likely to increase the chance of locally contacting the next host for a blood meal appear to be constrained by the envelope of the prevailing winds at the time of day, dusk, when midges are normally active.
Table 2 shows that SBV infection occurred on average 113 days before the disease case was reported, often as a foetal abnormality in a full-term sheep. Given the average gestation period of sheep (of 150 – 155 days), this indicates that sheep were infected by midges with SBV between 37 to 42 days after conception. This is consistent with what is known for the Akabane virus, in the same genus as SBV and with a similar clinical picture, for which pregnant sheep are vulnerable after 30 days of service/insemination54. Thus if a farm reported an SBV- malformed newborn lamb on the 14th December 2011 (the first case in our dataset), the infectious bite would have occurred on about the 24th August 2011.
Cattle have a longer gestation period, of approximately 280 days, than do sheep. Initially, in an attempt to exclude from SBV-SWOTS analysis the cattle cases (which, it was thought, would affect the DA estimates of sheep) the value of DA was limited to a maximum of 155 days. Nevertheless, 78% of the SBV cases in cattle and 65% of the larger number of cases in sheep were explained by SBV-SWOTS with the calculated value of 113 days for DA. Cattle may be affected by the related Akabane virus (resulting in foetal abnormalities) at any time between about month 3 and month 6 of pregnancy (http://www.dairyaustralia.com.au/Animals-feed-and-environment/Animal-health/Animal-health-fast-facts/Abortion-and-infertility/Akabane-virus.aspx). If SBV is similar in this respect to Akabane then cattle would be susceptible to SBV over a range of DA from 100 to 190 days: more than half of this period would therefore be covered by the SBV-SWOTS allowed maximum value of 155 days.
The 30% of unexplained SBV infections could be due to noise in the dataset, the approximate rather than precise geolocation of the farms, the approximate date of infection, unreported infections/farms, inaccurate (modelled) estimate of wind fields, insects other than midges and host animals other than obvious domestic ones (e.g. wildlife8) being involved in SBV transmission or, finally, to model error. Many of these sources of possible error also apply to the BTV outbreak data, which had only 8% of unexplained infections when analysed by BTV-SWOTS. The errors in the geolocation and time of infection are randomly distributed (Supplementary Information) and hence likely to increase the noise rather than the bias. The number of farms reported early in the outbreak and missing in the OIE dataset do not influence SBV-SWOTS results, since they account for less the 0.1% of the total farms (Supplementary Information). Importantly, our previous BTV-SWOTS analysis showed that up to 50% of all farm cases can be omitted from the dataset without affecting the values of the parameters and variables estimated by SWOTS31 (although the errors tend to be greater). As explained in the Methods, we believe that the SBV outbreak data are sufficiently representative of the SBV outbreak as a whole and that there are relatively minor differences between the ways in which BTV-8 and SBV outbreaks were managed, so that the SWOTS-detected differences between these two diseases are real rather than artefactual, i.e. caused by different biases in the two datasets. We suggest that the difference in the levels of unexplained infections in the BTV and SBV analyses are most likely due to the very much more geographically widespread nature of the SBV outbreak. In the case of BTV a much higher percentage of short-range infections were explained by the BTV-SWOTS model than long range infections (Table 2). Thus SBV, with fewer short range infections, is likely to be more difficult to explain with SBV-SWOTS.
The present results suggest that disease events that happen a long time before their recorded outcome are amenable to the same sort of analysis as was previously applied to BTV with a much shorter time between cause and effect. We stress that all values of DA between 0 and 155 days were explored by the SBV model which, nevertheless, selected a rather precise value of 113 days as the most likely time of infection before the recorded outcome which, in most cases, was a foetal abnormality.
We have no idea of the distribution of farms that were never infected and undoubtedly knowledge of these would help us to work out more precisely how this disease spreads, as would records of the sero-positivity in such farms that, in the absence of obvious clinical outcomes of infection (foetal malformations), were not included in the dataset. Nevertheless, even with the relatively incomplete data available, SBV-SWOTS can mimic accurately the rate of spread of this disease across the network of available farms using only relatively few assumptions about the behaviour of midges in the prevailing wind fields. Of particular interest is the fact that the SBV-SWOTS simulation explains 10 out of 266 UK cases of SBV as being initiated by midges originating from European mainland infections (6 from France and 4 from Belgium), thus tending to confirm the UK Meteorological Office's predictions, before any UK infections were reported, of the possibility of SBV infection from Europe by wind-borne midges on particular days when the wind was blowing in the right direction (http://www.defra.gov.uk/animal-diseases/files/poa-schmallenberg-update-120311.pdf).
To generalise SBV-SWOTS further, data are now required on the distribution of all farms (not just the infected ones), the pregnancy status of the animals on each farm and some idea of the total numbers of midges on each farm42,55,56. Such data would allow SBV-SWOTS to estimate the actual infection risk across the farming landscape. Adding seasonality to the midge numbers would allow this important component to be included to see how the onset of winter conditions limits the seasonal spread of this and other midge-borne diseases. Whilst the great time delay between cause and effect makes the control of SBV very difficult, the new SBV-SWOTS analysis also shows that such control needs to be applied much more quickly and over much wider areas than previously envisaged for BTV, for which the enforced Control Zones had a radius of only 20 km and Protection Zones of 100 km (http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=CELEX:32000L0075:EN:NOT). Midge mobility revealed by the example of SBV and analysed by SBV-SWOTS suggests that Control and Protection Zones need to be considerably bigger, making the control of midge borne diseases that much more difficult.
Methods
Materials
The data used for this analysis were the Schmallenberg cases on farms reported to the World Organisation for Animal Health (OIE) from December 2011 until May 2012. We used the most recent follow-up reports for each country, specifically the 3rd for Belgium, 7th for France, 22nd for Germany, 2nd for Italy, 11th for Luxembourg, 17th for Netherlands, 1st for Spain and the 16th for the United Kingdom (see Supplementary Information). Only the longitude, latitude (GPS co-ordinates) and date of the report of the infection at each farm were used in the analysis.
In total there were 3,487 reported SBV cases in eight countries. This is not the complete dataset because SBV cases in Poland, Sweden and Denmark are either missing or otherwise unavailable (SBV is not notifiable in many countries)2. SBV was declared notifiable in Germany and the Netherlands only from the end of March 2012 but in fact Germany had been contributing SBV case Reports to the OIE since the start of the outbreak and the number of new cases reported there from the end of March shows little change from those reported before this time (thus 2% of all of the 1376 cases from Germany between December 2011 and May 2012 were reported in December 2011, 24% in January 2012, 31% in February, 14% in March, 21% in April and 8% in May). The Netherlands, on the other hand, seems to show a distinct effect of the notification order. Of the total of 345 Dutch cases, 1% were reported in December 2011, 17% in January 2012, 15% in February, 16% in March, 1% April and 50% in May. The Dutch reports in May 2012 (173 cases) will have contributed in only a minor way to the parameter estimates in SBV-SWOTS (which were based on all of the 3,487 case reports) and none of them was explained in the simulation part of the SBV-SWOTS analysis. Notifications for Switzerland (19 cases, reported in July and August 2012), were not included because they occurred outside the period analysed (see above).
In parallel to the OIE database, the European Food Standards Agency (EFSA) also generated a database of SBV cases. The EFSA database contains most of the same information as in the OIE database but includes 7% more infected farms. Unfortunately the EFSA database is not publicly available and suffers the disadvantage of recording infections only to administrative areas (the European Union Nomenclature of Units for Territorial Statistics, NUTS, Level 2, equivalent to English counties or groups of counties) rather than point level, making them unsuitable for present purposes. The EFSA and OIE databases are compared in the Supplementary Information. Both databases undoubtedly suffer from missing data, but there are relatively minor differences between them.
Another difference between the BTV-8 and SBV outbreaks was the imposition of animal movement restrictions in Europe when BTV-8 first appeared, but no restrictions were applied later to SBV. BTV-8, however, spread so quickly that within-country movement restrictions were removed after only two weeks57. Thus differences between the management of the BTV-8 and SBV outbreaks (movement restrictions were never imposed for SBV) are slight and the datasets can be directly compared.
11% of the data used in this analysis were of SBV cases not associated with foetal abnormalities, 62% were recorded as foetal (i.e. full-term) abnormalities in sheep, 35% as foetal abnormalities in cattle and almost 2% as foetal abnormalities in goats (only one case was reported from deer).
Wind data for the analysis were generated by the UK Met Office's numerical weather prediction model, the Unified Model58 and consisted of hourly wind speed and direction estimates at a nominal height of 10 m in a grid of 0.25 degrees resolution covering all of Europe. Before use in the models wind speeds at each grid point were first converted to speeds at a nominal height of 2 m (as in BTV-SWOTS31) and then the corrected wind speeds and their directions were converted to wind vectors (by the triangle law of addition) for five periods within each 24 h cycle; period 1, 0000–0459 h; period 2, 0500–1059 h; period 3, 1100–1559 h; period 4, 1600–2059 h; and period 5, 2100–2359 h as in31 (all times in UTC).
SBV-SWOTS analysis
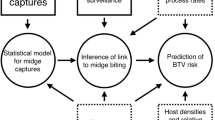
In this study we developed a modified version of the BTV Spatio-temporal Wind Outbreak Trajectories Simulation (BTV-SWOTS)31 to apply to the SBV situation (SBV-SWOTS, Figure 3). In brief, by exploring the correlations between the vectors of infections (between all previously infected farms and all previously uninfected farms that first report an infection in the current interval) and of midge movements (assumed to be related to wind vectors at the infected farms) for each plausible temporal scale, SBV-SWOTS identifies optimal parameter values to describe the SBV outbreak data. Specifically, vectors of infections are the vectors connecting a farm infected at day t to farms infected at day t + DT where DT is the number of days necessary for an infected midge to reach the uninfected farms. Midge movements are vectors obtained by wind vector conversion according to the biologically realistic assumptions of midge flight summarised in Table 3 and may include downwind, upwind, random, downwind and random and upwind and random components (the amount of each is estimated by SBV-SWOTS).
Unlike time-dependent differential models of epidemic spread (where identification of the location and timing of the index case is often crucial), SWOTS adopts more of a ‘mass action’ approach; estimated (averaged) parameter values apply to the entire dataset over the entire course of the outbreak and each disease report makes a more or less equal contribution to the parameter estimates.
The parameters and variables to be optimised by the SBV-SWOTS are the day of infection after fertilisation of the vertebrate host (see below), the number of days of midge travel from farm to farm (days travelled, DT) and the time of day (TOD) associated with the largest correlation between infection and midge vectors (in nature, midges fly mostly at dusk). Once these parameters are optimised, SBV-SWOTS then simulates the outbreak by re-calculating the midge vectors according to the optimised parameters and a stochastic wind field, assuming pre-defined distributions for wind angle and speed centred on their modelled mean values. Sampling from distributions of both wind speed and direction is necessary because the wind can show considerable variation in these variables within a few hours. Wind trajectories were recalculated 1,000 times, with values of speed and direction drawn from their respective mathematical distributions. Hence each infected farm in each interval in SBV-SWOTS is associated with many different possible infected midge trajectories going from it. Some of these trajectories will end up on uninfected farms and eventually infect them. SBV-SWOTS examines the associations between these calculated midge vectors and the calculated (farm to farm) infection vectors for the same time period and identifies the most likely (highest correlation between midge and infection vectors) source and route of infection to each susceptible farm. Because the midge vectors allowed a variety of combinations of up-wind, down-wind and random midge movement, the highest correlations between midge and infection vectors also identified the most likely type of midge movement responsible for infecting one farm from another.
The differences between BTV-8 and SBV required some modifications of BTV-SWOTS for use as SBV-SWOTS. Firstly, the intrinsic incubation period of BTV (IIP, the time between a sheep or cattle host being bitten by an infected midge and that host showing a patent infection) was replaced by a new parameter, the “day of infection before birth of a malformed foetus” (DA) a change required because all the existing literature assumes that SBV infection, resulting in the reported birth abnormalities, occurs at some (relatively early) stage of pregnancy24. We recognise that the real parameter of interest is not DA but the day of infection after fertilisation and conception. Fertilisation of the affected animals, however, occurred at an unrecorded date before the recorded foetal abnormality and hence could not be considered directly in the models. Later, by assuming a fixed gestation period, an estimate of the time of infection after conception can be made. We also later estimated the intrinsic incubation period by calculating the average difference between the time when one farm was shown in the model to be infecting another farm and the time that the infecting farm had itself been first infected. This is a somewhat indirect measure of IIP and will probably also include at least part of the extrinsic incubation period in the midges (EIP, the time taken for the development of a transmissible infection in a newly-infected midge). Hence the incubation periods calculated by SWOTS are likely to be a measure of (EIP + IIP) rather than of EIP or IIP alone. Knowing the IIP from independent studies allows the estimation of EIP from this figure.
A second modification of SBV-SWOTS concerned the wind-fields which, for the BTV-SWOTS analysis, were available (i.e. had been modelled) only at the locations of infected farms. BTV-SWOTS therefore had to interpolate wind speed and direction from these data for any other point of interest. For SBV-SWOTS, the wind speeds and directions were modelled on a regular 0.25 degree grid and SBV-SWOTS simply selected the wind data at the nearest grid point, without further interpolation.
References
Elliott, R. M. et al. Establishment of a reverse genetic system for Schmallenberg virus, a newly emerged orthobunyavirus in Europe. J Gen Virol 94, 851–859 (2012).
Beer, M., Conraths, F. J. & van der Poel, W. H. ‘Schmallenberg virus’--a novel orthobunyavirus emerging in Europe. Epidemiol Infect 141, 1–8 (2013).
Meiswinkel, R. et al. The 2006 outbreak of bluetongue in northern Europe--the entomological perspective. Prev Vet Med 87, 55–63 (2008).
Velthuis, A. G., Saatkamp, H. W., Mourits, M. C., de Koeijer, A. A. & Elbers, A. R. Financial consequences of the Dutch bluetongue serotype 8 epidemics of 2006 and 2007. Prev Vet Med 93, 294–304 (2010).
Martinelle, L., Dal Pozzo, F., Gauthier, B., Kirschvink, N. & Saegerman, C. Field Veterinary Survey on Clinical and Economic Impact of Schmallenberg Virus in Belgium. Transbound Emerg Dis; 10.1111/tbed.12030 (2012).
Falconi, C., Lopez-Olvera, J. R. & Gortazar, C. BTV infection in wild ruminants, with emphasis on red deer: a review. Vet Microbiol 151, 209–219 (2011).
Linden, A. et al. Epizootic spread of Schmallenberg virus among wild cervids, Belgium, Fall 2011. Emerg Infect Dis 18, 2006–2008 (2012).
Larska, M., Krzysiak, M., Smreczak, M., Polak, M. P. & Żmudziński, J. F. First detection of Schmallenberg virus in elk (Alces alces) indicating infection of wildlife in Białowiez·a National Park in Poland. Vet J; 10.1016/j.tvjl.2013.08.013.
EFSA. "Schmallenberg" virus: analysis of the epidemiological data (May 2013). EFSA Technical Report (2013)URL: http://www.efsa.europa.eu/en/supporting/doc/429e.pdf (Accessed 21/10/2013).
Reusken, C. et al. Lack of evidence for zoonotic transmission of Schmallenberg virus. Emerg Infect Dis 18, 1746–1754 (2012).
Herder, V., Wohlsein, P., Peters, M., Hansmann, F. & Baumgartner, W. Salient lesions in domestic ruminants infected with the emerging so-called Schmallenberg virus in Germany. Vet Pathol 49, 588–591 (2012).
Hahn, K. et al. Schmallenberg virus in central nervous system of ruminants. Emerg Infect Dis 19, 154–155 (2012).
van den Brom, R. et al. Epizootic of ovine congenital malformations associated with Schmallenberg virus infection. Tijdschr Diergeneeskd 137, 106–111 (2012).
Savini, G. et al. Bluetongue Serotype 2 and 9 Modified Live Vaccine Viruses as Causative Agents of Abortion in Livestock: A Retrospective Analysis in Italy. Transbound Emerg Dis; 10.1111/tbed.12004 (2012).
Yanase, T. et al. Genetic reassortment between Sathuperi and Shamonda viruses of the genus Orthobunyavirus in nature: implications for their genetic relationship to Schmallenberg virus. Arch Virol 157, 1611–1616 (2012).
Carpenter, S., Wilson, A. & Mellor, P. S. Culicoides and the emergence of bluetongue virus in northern Europe. Trends Microbiol 17, 172–178 (2009).
Schnettler, E. et al. RNA interference targets arbovirus replication in Culicoides cells. J Virol 87, 2441–2454 (2012).
De Regge, N. et al. Detection of Schmallenberg virus in different Culicoides spp. by real-time RT-PCR. Transbound Emerg Dis 59, 471–475 (2012).
Larska, M. et al. First report of Schmallenberg Virus Infection in Cattle and Midges in Poland. Transbound Emerg Dis 60, 97–101 (2013).
Rasmussen, L. D. et al. Culicoids as vectors of Schmallenberg virus. Emerg Infect Dis 18, 1204–1206 (2012).
Elbers, A. R., Meiswinkel, R., van Weezep, E., Sloet van Oldruitenborgh-Oosterbaan, M. M. & Kooi, E. A. Schmallenberg Virus in Culicoides spp. Biting Midges, the Netherlands, 2011. Emerg Infect Dis 19, 106–109 (2013).
Garigliany, M. M., Bayrou, C., Kleijnen, D., Cassart, D. & Desmecht, D. Schmallenberg virus in domestic cattle, Belgium, 2012. Emerg Infect Dis 18, 1512–1514 (2012).
Bradshaw, B. et al. Schmallenberg virus cases identified in Ireland. Vet Rec 171, 540–541 (2012).
Dominguez, M. et al. Preliminary estimate of Schmallenberg virus infection impact in sheep flocks - France. Vet Rec 171, 426, 10.1136/vr.100883 (2012).
Elbers, A. R. et al. Seroprevalence of Schmallenberg virus antibodies among dairy cattle, the Netherlands, winter 2011–2012. Emerg Infect Dis 18, 1065–1071 (2012).
Steinbach, F., Dastjerdi, A., Drew, T., Cook, A. & Davies, I. Continued presentation of cases of Schmallenberg virus in sheep in England. Vet Rec 170, 547 (2012).
News, V. R. Schmallenberg virus antibodies detected in cattle in Wales. Vet Rec 171, 333 (2012).
Gloster, J., Mellor, P. S., Manning, A. J., Webster, H. N. & Hort, M. C. Assessing the risk of windborne spread of bluetongue in the 2006 outbreak of disease in northern Europe. Vet Rec 160, 54–56 (2007).
Burgin, L. E. et al. Investigating Incursions of Bluetongue Virus Using a Model of Long-Distance Culicoides Biting Midge Dispersal. Transbound Emerg Dis 60, 263–272 (2012).
Agren, E. C., Burgin, L., Lewerin, S. S., Gloster, J. & Elvander, M. Possible means of introduction of bluetongue virus serotype 8 (BTV-8) to Sweden in August 2008: comparison of results from two models for atmospheric transport of the Culicoides vector. Vet Rec 167, 484–488 (2010).
Sedda, L. et al. A new algorithm quantifies the roles of wind and midge flight activity in the bluetongue epizootic in northwest Europe. Proceedings. Biological sciences/The Royal Society 279, 2354–2362 (2012).
Pioz, M. et al. Estimating front-wave velocity of infectious diseases: a simple, efficient method applied to bluetongue. Vet Res 42, DOI 10.1186/1297-9716-42-60 (2011).
Napp, S. et al. Assessment of the risk of a bluetongue outbreak in Europe caused by Culicoides midges introduced through intracontinental transport and trade networks. Med Vet Entomol 27, 19–28 (2013).
Turner, J., Bowers, R. G. & Baylis, M. Modelling bluetongue virus transmission between farms using animal and vector movements. Sci Rep 2, 319; 10.1038/Srep00319 (2012).
de Koeijer, A. A. et al. Quantitative analysis of transmission parameters for bluetongue virus serotype 8 in Western Europe in 2006. Vet Res 42, 53; 10.1186/1297-9716-42-53 (2011).
Mayo, C. E. et al. Colostral transmission of bluetongue virus nucleic acid among newborn dairy calves in California. Transbound Emerg Dis 57, 277–281 (2010).
Faes, C. et al. Factors affecting Bluetongue serotype 8 spread in Northern Europe in 2006: The geographical epidemiology. Prev Vet Med 110, 149–158 (2012).
Bessell, P. R. et al. Epidemic potential of an emerging vector borne disease in a marginal environment: Schmallenberg in Scotland. Sci Rep 3, 1178; 10.1038/srep01178 (2013).
Mason, C. et al. SBV in a dairy herd in Scotland. Vet Rec 172, 403–403 (2013).
Sprent, P. & Smeeton, N. C. Applied Nonparametric Statistical Methods, Fourth Edition. (Taylor & Francis, 2010).
Conraths, F. et al. [Bluetongue Disease: An Analysis of the Epidemic in Germany 2006–2009] Arthropods as Vectors of Emerging Diseases (ed. Mehlhorn H.) [103–135] (Springer Berlin Heidelberg, 2012).
Wernike, K. et al. Transmission of Schmallenberg Virus during Winter, Germany. Emerg Infect Dis 19, 1701–1703 (2013).
Baylis, M. et al. Evaluation of housing as a means to protect cattle from Culicoides biting midges, the vectors of bluetongue virus. Med Vet Entomol 24, 38–45 (2010).
Meiswinkel, R. et al. Endophily in Culicoides associated with BTV-infected cattle in the province of Limburg, south-eastern Netherlands, 2006. Prev Vet Med 87, 182–195 (2008).
Liberato, C. D. et al. Biotic and Abiotic Factors Influencing Distribution and Abundance ofCulicoides obsoletusGroup (Diptera: Ceratopogonidae) in Central Italy. Journal of Medical Entomology 47, 313–318 (2010).
Blackwell, A. Diel flight periodicity of the biting midge Culicoides impunctatus and the effects of meteorological conditions. Med Vet Entomol 11, 361–367 (1997).
Hoffmann, B. et al. Novel orthobunyavirus in Cattle, Europe, 2011. Emerg Infect Dis 18, 469–472 (2012).
Bonneau, K. R., DeMaula, C. D., Mullens, B. A. & MacLachlan, N. J. Duration of viraemia infectious to Culicoides sonorensis in bluetongue virus-infected cattle and sheep. Vet Microbiol 88, 115–125 (2002).
Mellor, P. S., Boorman, J. & Baylis, M. Culicoides biting midges: Their role as arbovirus vectors. Annu Rev Entomol 45, 307–340 (2000).
Meroc, E. et al. Large-scale cross-sectional serological survey of schmallenberg virus in belgian cattle at the end of the first vector season. Transbound Emerg Dis 60, 4–8 (2013).
Meroc, E. et al. Distribution of Schmallenberg Virus and Seroprevalence in Belgian Sheep and Goats. Transbound Emerg Dis; 10.1111/tbed.12050 (2013).
Kirkeby, C., Bodker, R., Stockmarr, A., Lind, P. & Heegaard, P. M. H. Quantifying Dispersal of European Culicoides (Diptera: Ceratopogonidae) Vectors between Farms Using a Novel Mark-Release-Recapture Technique. PloS one 8, e61269; 10.1371/journal.pone.0061269 (2013).
Kirkeby, C., Graesboll, K., Stockmarr, A., Christiansen, L. E. & Bodker, R. The range of attraction for light traps catching Culicoides biting midges (Diptera: Ceratopogonidae). Parasite Vector 6, 10.1186/1756-3305-6-67 (2013).
Parsonson, I. M., Dellaporta, A. J. & Snowdon, W. A. Congenital-Abnormalities in Newborn Lambs after Infection of Pregnant Sheep with Akabane Virus. Infect Immun 15, 254–262 (1977).
Kluiters, G. et al. Modelling the spatial distribution of Culicoides biting midges at the local scale. J Appl Ecol 50, 232–242 (2013).
Rigot, T., Drubbel, M. V., Dellecolle, J. C. & Gilbert, M. Farms, pastures and woodlands: the fine-scale distribution of Palearctic Culicoides spp. biting midges along an agro-ecological gradient. Med Vet Entomol 27, 29–38 (2013).
Méroc, E. et al. [Epidemiological analysis of the 2006 bluetongue virus serotype 8 epidemic in north-western Europe–Role of human interventions] Epidemiological analysis of the 2006 bluetongue virus serotype 8 epidemic in north-western Europe (EFSA, Parma, 2007). URL: http://www.efsa.europa.eu/en/efsajournal/doc/34r.pdf (Accessed 21/10/2013).
Davies, T. et al. A new dynamical core for the Met Office's global and regional modelling of the atmosphere. Q J Roy Meteor Soc 131, 1759–1782 (2005).
Acknowledgements
We thank Laura Burgin from UK Met Office for the generation of the wind data and Simon Carpenter, Simon Gubbins and Chris Sanders from the Institute of Animal Health for the useful discussions on the OIE dataset. L.S. and D.J.R. are supported by the European Union grant FP7-261504 EDENext and this paper is catalogued by the EDENext Steering Committee as EDENext188 (http://www.edenext.eu). The contents are the sole responsibility of the authors and do not necessarily reflect the views of the European Commission.
Author information
Authors and Affiliations
Contributions
L.S. and D.J.R. conceived and designed the study and wrote the manuscript. L.S. performed the analysis and prepared all the figures and tables.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Data sources and comparison with EFSA database
Supplementary Information
Modelled Schmallenberg spread in Europe 2011/2012
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sedda, L., Rogers, D. The influence of the wind in the Schmallenberg virus outbreak in Europe. Sci Rep 3, 3361 (2013). https://doi.org/10.1038/srep03361
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03361
This article is cited by
-
Schmallenberg virus: research on viral circulation in Brazil
Brazilian Journal of Microbiology (2022)
-
Coronaviruses in cattle
Tropical Animal Health and Production (2020)
-
Laboratory evaluation of stable isotope labeling of Culicoides (Diptera: Ceratopogonidae) for adult dispersal studies
Parasites & Vectors (2019)
-
Schmallenberg virus: a systematic international literature review (2011-2019) from an Irish perspective
Irish Veterinary Journal (2019)
-
Culicoides species composition and abundance on Irish cattle farms: implications for arboviral disease transmission
Parasites & Vectors (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.