Abstract
Contact structure, a critical driver of infectious disease transmission, is not completely understood and characterized for environmentally transmitted pathogens. In this study, we assessed the effects of temporal and spatial heterogeneity in animal contact structures on the dynamics of environmentally transmitted pathogens. We used real-time animal position data to describe contact between animals and specific environmental areas used for feeding and watering calves. The generated contact structure varied across days and among animals. We integrated animal and environmental heterogeneity into an agent-based simulation model for Escherichia coli O157 environmental transmission in cattle to simulate four different scenarios with different environmental bacteria concentrations at different areas. The simulation results suggest heterogeneity in environmental contact structure among cattle influences pathogen prevalence and exposure associated with each environment. Our findings suggest that interventions that target environmental areas, even relatively small areas, with high bacterial concentration can result in effective mitigation of environmentally transmitted pathogens.
Similar content being viewed by others
Introduction
The transmission of most pathogens involves the passage of the pathogen from an infectious to a susceptible individual through the environment1,2. For environmentally transmitted pathogens, the understanding of pathogen dynamics outside the host and the contact between a host and its associated environment is necessary to fully characterize transmission, as well as to identify effective mitigation strategies. Some environments may have the potential to contribute to a disproportional number of infections; for example, high pathogen survival and growth rates in the environment, or a high contact rate between host and environment can create hotspots for infection3,4,5. For some environmentally transmitted pathogens, such as Shiga-toxigenic Escherichia coli, enteric viruses and multidrug-resistant pathogens, pharmacological interventions are limited, making interventions aimed at blocking pathogen transmission through the environment the cornerstone of their control6. Despite its central role in transmission and disease control, the environment is often poorly represented in infectious disease models used to understand transmission and evaluate control strategies. Models that include environmental reservoirs often represent the environment as a homogenous compartment and the contact rates between the environment and hosts are assumed to be random7,8,9,10.
Pathogen transmission, both direct and indirect, is ultimately determined by the contact structure among hosts and their environments (e.g. number of contacts, duration of each contact and total duration of all contacts in a given time), making the variation in contact structure one of the most important sources of transmission heterogeneity in infectious diseases11,12. Individual animals spend different amounts of time doing different activities in various areas13,14. These heterogeneous behavioral patterns among animals can affect the contact structure (in this study defined as the duration of total contact with specific areas in a pen in 1 day).
However, one of the main limitations in modeling indirect transmission through the environment has been the lack of appropriate empirical data to characterize contact between hosts and their associated environments. Recently, the availability of telemetric technologies has facilitated the collection of empirical data that can be used to characterize the contact structure among hosts and between hosts and their environment. Proximity loggers have been used to describe close social encounters in both humans and animals15,16,17. Animal movement data collected using global positioning and other triangulation and accelerometer systems can be used to both characterize direct contacts and to describe the interactions between animals and their associated environment11,14. These systems provide novel empirical data to describe contact for environmentally transmitted diseases and can be integrated with infectious disease transmission models to provide a more accurate description of pathogen transmission throughout the environment. The overall aim of this study was to assess how spatial and temporal heterogeneity in contact structure between animals and their associated environment are able to influence pathogen transmission through the environment. This study highlights the potential importance of contact structure and temporal-spatial heterogeneity in environmental transmission and potential mitigation strategies.
Results
Spatial and temporal heterogeneity associated with animal movement
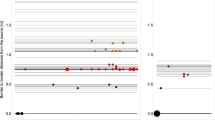
There was clear spatial and temporal heterogeneity associated with animal position and movement during the monitoring period (8 days) for all 3 pens at 10-s resolution. The daily contact durations summarized for the three pens were presented in Table 1 and Figure 1. The animals spent much more time around the hay bunk than around the water and grain bunk and contact with water was limited to only a small amount of time in a day (less than 1%). This pattern is consistent across all 3 pens. Based on the ANOVA results, there was a significant daily difference in animals' contact structure for all 3 areas: grain bunk, water and hay bunk (Fig. 1 panel 1–3; P = 2.6*10−9, 4.8*10−2, 2.2*10−16, respectively). Also, there was a significant difference in total daily contact duration at pen level for the grain bunk, water and hay bunk (P = 3.8*10−5, 1.6*10−8, 2.7*10−15, respectively) and at individual animal level (P = 4.4*10−3, 8.9*10−3, 2.0*10−2, respectively). There was no significant interaction between day and pen (P = 4.0*10−1, 3.9*10−1, 5.1*10−1 for grain bunk, water and hay bunk, respectively). In successive simulations, these sources of heterogeneity were incorporated in the agent-based model.
Simulated prevalence dynamics with spatial heterogeneity
The bacterial concentration was calculated as 106 CFU/m2 (colony forming unit) for water, 105 for both grain bunk and hay bunk and 104 for the remaining pen floor (approximate to the nearest magnitude) for the baseline scenario (see Appendix B in supplemental material for details). So we considered water source as a high bacterial concentration area, grain bunk and hay bunk as medium concentration areas and other pen environment as a low concentration area. We present the sizes of high, medium and low concentration areas; relative percent of the entire pen area size; average duration of contact in a day; and percent of contact in a day in Table 2. The high concentration area (water) contributed only a small amount of area in the pen (1%) and in daily contact (0.8%).
In Figure 2, we showed the time series of prevalence obtained by simulating the agent-based model for different conditions (C1 to C3 as heterogeneous conditions and H1 as homogeneous condition) and summarized the predicted maximum prevalence and its corresponding occurrence day in Table 2. The overall shapes of prevalence dynamics under these conditions were similar; however, peak prevalence differed substantially (Table 3). Condition C2 and C3 represent different potential mitigation approaches. C2 focused on the high concentration area (water) with reduced bacteria concentration only in the water source, while C3 further reduced bacteria concentration in the grain bunk and hay bunk, as well. In C2, we reduced the maximum prevalence by approximately 6%. In C3, a further reduction to 1/10 of the bacteria concentration for the grain bunk and hay bunk (with already lowered bacteria concentration in water as C2) resulted in about a 12.7% reduction of prevalence. Finally, if we lower the bacteria concentration in water to 104 CFU/m2 (which is a homogeneous pen with 104 CFU/m2 bacteria concentration everywhere), we would have less than 70% maximum prevalence. All these conditions (C2, C3 and H1) focus only on controlling bacterial concentration in the water, grain bunk and hay bunk and the results show that targeting these specific areas can be useful to control bacteria colonization and infection in the pen.
Time series of mean prevalence under different conditions.
Solid black line: mean prevalence; dashed red line: 25% and 75% quantiles. Four figures represent condition C1, C2, C3 and H1. Details of these conditions are listed in table 3.
Not only did the simulated infection dynamics and maximum prevalence change under different conditions (with different environmental bacteria concentrations), we also quantified the risk of infection and demonstrated the relative risk (probability of infection) changed substantially at different areas in the pen under these conditions (Table 3). In C1 the water had a high bacteria concentration level (106 CFU/m2) and exposure to the water area for only 10 min (less than 1% of total time in a day) resulted in more than 40% of total new infections. While the calves spent over 85% of their time in the general pen area, the infection probability in the general pen area was less than 40%. When the bacteria concentration was reduced in the water area (in C2, the water area had the same bacteria concentration as did the grain and hay bunks), the importance of the water area as a potential source of infection was substantially decreased and the importance of the general pen area was substantially increased. In C3 where water had a bacteria concentration 10 times higher than in any other area in the pen, the water area became the second-most important source of infection and the most infectious source was the general pen. Contribution to infection from the water area was not as significant as in C1, because in C1 the bacteria concentration was much higher (100 times) than in the general pen area. In a homogeneous setting (as shown in H1), the risk of infection depended only on the total duration that an animal spent around each area; hence, the general pen area served as the primary source of infection. All these results indicate the importance of temporal-spatial heterogeneity in infection dynamics.
Discussion
While contact structure and individual heterogeneity is intensively studied in directly transmitted disease18,19, it is unclear how contact structure among individual animals and various environments influences the transmission dynamics of environmentally transmitted pathogens. The temporal and spatial heterogeneity in our simulations led to some interesting findings. The risk of infection from the environment (i.e. infection or colonization probability) was dependent on both the environmental bacteria concentration and total contact duration. Although the prevalence time series were similar for the simulated scenarios, the heterogeneity in environmental concentration and contact had a large effect in determining the contribution of different environments to new infections. The previous result section illustrates the importance of total contact duration with the high bacteria concentration area. We suggest identifying and quantifying the high-concentration areas (“hotspots” of the pathogen) as one of the most important tasks in clarifying transmission and control options. This can lead to designing more effective control and mitigation strategies (as already shown in the results section when comparing various conditions in simulation). For instance, while it is not feasible to shorten the duration an animal stays near certain areas in the pen, we may be able to reduce environmental contamination with the bacteria (hence reduce probability of infection). Depending on labor and budget, it may be effective to target specific areas (e.g. the “hotspots,” and not entire pen) to reduce prevalence.
In this study, we combined field observation datasets of animal behavior and contact structure with an agent-based modeling framework and quantitatively assessed the temporal-spatial heterogeneity and how it changes infection dynamics within the pen. It is relatively easy and straightforward to adopt the existing agent-based models to study other similarly transmitted pathogens such as Salmonella. Furthermore, environmental transmission pathways occur in many other pathogens, even those assumed to be mostly directly transmitted, such as influenza and pathogens with other transmission routes, such as waterborne and foodborne pathogens20,21,22. Consequently, we can integrate the contact structure to the environment components with these models as well and assess the importance of both environmental transmission and direct transmission pathways. Targeting environmental pathogen sources and pathways is particularly relevant for those pathogens for which pharmacological interventions are limited6.
One of the implicit assumptions in this research is that the contact structure remains the same when the epidemic is established in the population. It is possible that the pathogen might change animal behavior and influence/change the contact structure between the animal and the environment11,15. Nevertheless, for the specific calf and E. coli O157 system in this study, the pathogen behaves as commensal, so it does not cause recognizable symptoms in the animals and as such, it seems unlikely the calves' behavior would change when infected. Future study may consider monitoring the animals' contact structure during the time surrounding inoculation so as to provide more detailed information about behavioral change before and after infection.
In summary, our study is the first of this kind to integrates real-time contact data and an agent-based modeling framework to comprehensively investigate temporal-spatial heterogeneity in the contact structure between animals and their associated environments and relate it to environmental pathogen transmission dynamics. Host behavior and pathogen environmental dynamics can generate heterogeneity in pathogen exposure that has implications for pathogen control.
Methods
Data acquisition and standardization
Animal position data were collected at the research facilities of Kansas State University, Manhattan, Kansas. All the experiments were approved by and complied with animal regulation policy of Kansas State University. Three pens with 21, 21 and 28 calves (approximately 76–78 days of age and average weight of 76.9 ± 11.1 kg at trial initiation), respectively, were monitored. Each pen was approximately 11.9 m × 25.6 m (80 ft × 40 ft) in size (see supplementary figure 1 for a diagram of pen settings). Calves were continuously monitored from August 11, 2011, to August 18, 2011. Each calf had an ultra wide-band tag (Ubisense Series 7000 Compact Tag, Ubisense) attached to its ear. The tags transmit ultra wide-band signals to 7 receivers around the pens; the receivers then transmit to a central computer that logs the 2-dimensional (X-Y coordinates) position data. To save power, the tag did not send data when it did not detect animal movement. The system has been previously validated and used to describe animal behavior as well as quantify time spent by the animals near specific pen locations11,17. All distances between the animal and given areas (e.g. grain, hay, or water) were computed based on 2-dimensional Euclidean distance.
The original position data, which were reported from each tag, were first standardized by aggregating to 10-s resolution (time unit was 10 s). That is, if the tag reported multiple readings of position data (XY coordinates) within the same 10 s, we averaged all these readings for that 10-s period to get a single position (XY coordinates) representing the location of the animal in that period; if the tag had not updated the location information for more than 10 s, we used the last known position data to interpolate the missing readings. After this standardization process, we had 8,640 × 2 data points for each animal in each day (3,600 s/h × 24 h/day/10 s = 8,640 data points/day; 2 columns represent X and Y coordinates). The XY coordinate data for specific pen areas for watering and feeding animals (grain bunk, water source and hay bunk) in each of the 3 pens were also recorded. At each time interval, the animals' position data were compared with position data of grain bunk, water source and hay bunk. If the animal was within approximately 0.3 m (~1 ft) of the threshold outside the boundary of the grain bunk (or water or hay bunk) then we assigned that time interval as a (close) contact to the specific area. The total contact duration in each day around grain bunk, water and hay bunk were computed by adding all the close contact intervals together.
Modeling procedure
Identifying spatial and temporal heterogeneity in the contacts between hosts and specific environments
We quantitatively examined and identified spatial and temporal heterogeneity from the original animal position and contact data described in the data acquisition and preparation section. We expected animals might visit some areas more frequently and with longer duration than other areas in the environment. We specifically investigated whether the contact between calves and 3 specific areas (grain bunk, water source and hay bunk) varied across the observation period (8 days, from August 11 to August 18, 2011) and across the 3 pens by conducting an analysis of variance (ANOVA). The model was specified with equation (1):

In this model, T was the daily contact duration in day i for animal k in the pen j; u was the mean daily contact duration of all counts in the entire period for all animals in all 3 pens; αI was effect of day and was treated as a fixed effect; βj was the effect of pen and treated as random effect; αβij was the random effect of interaction between day and pen; γk|j was the random individual effect nested within each pen and εijk was random error. We ran the ANOVA test for grain bunk, water and hay bunk.
The sources of variation (e.g. at pen, day, and/or individual level) that were found to be significant were incorporated into the model simulations (see next section).
Modeling environmental transmission with spatial and temporal exposure heterogeneity
To evaluate the effects of contact structure heterogeneity on the exposure and transmission of an environmentally transmitted pathogen, we modified our previous agent-based model for the transmission of the zoonotic pathogen E. coli O157 in the cattle reservoir by including spatial and contact heterogeneity in the model. In our previous approach23, the infectious agents shed the bacteria into the surrounding environment and susceptible agents were colonized by contact with the bacteria in the environment. There were 4 epidemiological states for the animal: primary susceptible, primary infectious, secondary susceptible and secondary infectious. After animals recover from the primary infection, they acquire partial immunity. Secondary susceptible individuals require a higher dose of bacteria to become re-infected. If infected again, secondary infectious individuals are assumed to shed lower bacteria concentrations. Given contact between the animal and environment, the daily transmission probability depends on the environmental concentration of E. coli, which was assumed to be homogeneous in our previous study. A more detailed description of the original modeling system is provided23.
In this study, we incorporated spatial and temporal heterogeneity in the environment. The overall pen environment was disaggregated into 4 main areas that differ in the duration of time spent by the animals and in bacterial concentration. These areas are the water source, grain bunk, hay bunk and the remaining pen floor. The bacteria concentrations in these areas were 106 CFU/m2, 105 CFU/m2, 105 CFU/m2 and 104 CFU/m2, respectively26,27,28,29,30. Based on these numbers, we considered water as a high bacteria concentration level area, grain bunk and hay bunk as medium concentration and the rest of the pen floor as low concentration. A detailed derivation of these concentrations can be found in supplementary section 2.
In order to introduce variability in the contact between animals and environment, we applied bootstrapping methods from the observed mean contact duration and its variance (aggregated by all the 21 + 21 + 27 = 69 animals) to generate a time series of daily contact duration for each individual throughout the entire simulation period. As shown later in the results section, there was significant day-to-day variability in animal contact structure; therefore, in each day of the simulation, we sampled one mean contact duration from 8 observations of contact duration (August 11–18 data) and used its associated variance in that day to generate contact durations for all 100 animals. We repeated this process 200 times to construct the 200-day time series. Here, we assumed the contact duration was independent from day to day and was population density independent (population size did not affect contact duration at each area).
Another source of individual variability (besides contact duration) was duration of the infectious period. The animals had a mean primary infectious period of 48 days and standard deviation of 5 days and secondary period mean of 16 days with standard deviation of 3 days, according to the data of oral challenge studies used previously23.
We simulated the duration of time a calf was around the grain bunk, hay bunk, water and the rest of the pen floor in each day for each animal. We determined the probability of infection given contact in each of these 4 areas using a logistic dose-response curve23 and then averaged the probabilities of infection at these areas (weighted to the percent of duration of contact at each area by total time in a day) to get daily effective probability of transmission. We also determined whether a transition from susceptible to infected occurred around the grain bunk, hay bunk, water, or the rest of pen area, by comparing the probability of infection from these areas. In this study, we did not track the shedding amount of pathogen (which is infeasible due to lack of observation data as well as computational burden) and assumed the bacteria concentration reached the environmental carrying capacity so that daily total new shedding of bacteria equaled the total loss and the concentration of bacteria remained constant for each of the 4 areas. Supplementary section 3 describes the overview-design-details protocol for describing agent-based models24. We monitored and aggregated individual animal epidemiological states by day, tracked the prevalence dynamics in the population for the entire simulation period (200 days) and replicated the simulation 100 times.
We evaluated 4 scenarios representing likely environmental concentrations. The baseline scenario (C1) is described in Table 2. Bacteria concentration levels were low, medium and high at 104, 105 and 106 CFU/m2 (as shown in the results section) in the general pen area, grain bunk, hay bunk and water, respectively. We further simulated 3 other scenarios: C2 had bacteria concentration in water reduced from 106 to 105 CFU/m2 while the other concentrations remained the same as C1; C3 had bacteria concentration as 104, 104, 104 and 105 CFU/m2 for the general pen area, grain bunk, hay bunk and water. Besides all these heterogeneous conditions, we also simulated a homogenous condition (H1) where bacteria concentration was 104 CFU/m2 for all areas. In this condition, we did not have to consider any spatial or temporal heterogeneity caused by animal contact pattern.
All the data analysis and simulations were written and run in R 2.1525 on Newton, a high-performance computing server at the University of Tennessee.
References
Breban, R. Role of environmental persistence in pathogen transmission: a mathematical modeling approach. J. Math. Biol. 66, 535–546 (2013).
Li, S., Eisenberg, J., Spicknall, I. & Koopman, J. Dynamics and control of infections transmitted from person to person through the environment. Am. J. Epidemiol. 170, 257–265 (2009).
Van Donkersgoed, J. et al. Concurrence of foodborne bacteria in Alberta feedlots. Can. Vet. J. 50, 166–72 (2009).
Hota, B. Contamination, disinfection and cross-colonization: are hospital surfaces reservoirs for nosocomial infection? Clin. Infect. Dis. 39, 1182–1189 (2004).
Mathiason, C. K. et al. Infectious prions in pre-clinical deer and transmission of chronic wasting disease solely by environmental exposure. PLoS One 4, e5916 (2009).
Boone, S. A. & Gerba, C. P. Significance of formites in the spread of respiratory and enteric viral disease. Appl. Environ. Microbiol. 73, 1687–1696 (2007).
Turner, J., Bowers, R. G., Clancy, D., Behnke, M. C. & Christley, R. M. A network model of E. coli O157 transmission within a typical UK dairy herd: The effect of heterogeneity and clustering on the prevalence of infection. J. Theor. Biol. 254, 45–54 (2008).
Breban, R., Drake, J. M. & Rohani, P. A general multi-strain model with environmental transmission: invasion conditions for the disease-free and endemic states. J. Theor. Biol. 264, 729–736 (2010).
Roche, B. & Rohani, P. Environmental transmission scrambles coexistence patterns of avian influenza viruses. Epidemics 2, 92–98 (2010).
Allen, L. J. et al. Mathematical modeling of viral zoonoses in wildlife. Nat. Resour. Model. 25, 5–51 (2012).
Theurer, M. E. et al. Assessment of behavioral change associated with oral meloxicam administration at time of dehorning in calves using a remote triangulation device and accelerometers. BMC Vet. Res. 8, 48 (2012).
Eames, K. T. D. Modelling disease spread through random and regular contacts in clustered populations. Theor. Popul. Biol. 73, 104–111 (2008).
Isella, L. et al. Close Encounters in a Pediatric Ward: Measuring Face-to-Face Proximity and Mixing Patterns with Wearable Sensors. PLoS One 6, e17144 (2011).
Duncan, A. J., Gunn, G. J., Lewis, F. I., Umstatter, C. & Humphry, R. W. The influence of empirical contact networks on modeling disease in cattle. Epidemics 4, 117–123 (2012).
Theurer, M. E., Amrine, D. E. & White, B. J. Remote noninvasive assessment of pain and health status in cattle. Vet. Clin. N. Am. 29, 59–74 (2013).
Handcock, R. et al. Monitoring animal behaviour and environmental interactions using wireless sensor networks, GPS collars and satellite remote sensing. Sensors 9, 3586–3603 (2009).
White, B. J. et al. Clinical, behavioral and pulmonary changes in calves following inoculation with Mycoplasma bovis. Am. J. Vet. Res. 73, 490–497 (2012).
Woolhouse, M. et al. Heterogeneities in the transmission of infectious agents: implications for the design of control programs. Proc. Natl. Acad. Sci. USA 94, 338–342 (1997).
Woolhouse, M. & Gaunt, E. Ecological origins of novel human pathogens. Crit. Rev. Microbiol. 33, 231–242 (2007).
Eisenberg, J. N. S. et al. Environmental determinants of infectious disease: a framework for tracking causal links and guiding public health research. Environ. Health. Perspect. 115, 1216–1223 (2007).
Winfield, M. D. & Groisman, E. A. Role of nonhost environments in the lifestyles of Salmonella and Escherichia coli. Appl. Environ. Microbiol. 69, 3687–3694 (2003).
Breban, R., Drake, J. M., Stallknecht, D. E. & Rohani, P. The role of environmental transmission in recurrent avian influenza epidemics. PLoS Comp. Biol. 5, e1000346 (2010).
Chen, S., Sanderson, M. & Lanzas, C. Investigating effects of between- and within-host variability on Escherichia coli O157 shedding pattern and transmission. Prev. Vet. Med. 109, 47–57 (2013).
Grimm, V. et al. The ODD protocol: a review and first update. Ecol. Model. 221, 2760–2768 (2010).
R Development Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria (2012).
Parham, P. E. & Ferguson, N. M. Space and contact networks: capturing the locality of disease transmission. J. R. Soc. Interface 3, 483–493 (2006).
Lynn, T. V. et al. The occurrence and replication of Escherichia coli in cattle feeds. J. Dairy Sci. 81, 1102–1108 (1998).
LeJeune, J. T., Besser, T. E. & Hancock, D. D. Cattle water troughs as reservoirs of Escherichia coli O157. Appl. Environ. Microbiol. 67, 3053–3057 (2001).
McGee, P., Bolton, D. J., Sheridan, J. J., Earley, B., Kelly, G. & Leonard, N. Survival of Escherichia coli O157:H7 in farm water: its role as a vector in the transmission of the organism within herds. J. Appl. Microbiol. 93, 706–713 (2002).
Scott, L., McGee, P., Sheridan, J. J., Earley, B. & Leonard, N. A comparison of the survival in feces and water of Escherichia coli O157:H7 grown under laboratory conditions or obtained from cattle feces. J. Food Prot. 69, 6–11 (2006).
Acknowledgements
This work was conducted with partial funding provided at the National Institute for Mathematical and Biological Synthesis, an Institute sponsored by the National Science Foundation, the U.S. Department of Homeland Security and the U.S. Department of Agriculture through NSF Award #EF-0832858, with additional support from The University of Tennessee, Knoxville. The authors thank Misty Bailey from the University of Tennessee for providing editorial comments.
Author information
Authors and Affiliations
Contributions
B.W. and D.A. planned and collected the field data. S.C., C.L. and M.S. participated in model design. S.C. performed the data analysis and simulation. S.C. and C.L. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figures and Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Chen, S., Sanderson, M., White, B. et al. Temporal-spatial heterogeneity in animal-environment contact: Implications for the exposure and transmission of pathogens. Sci Rep 3, 3112 (2013). https://doi.org/10.1038/srep03112
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03112
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.